ISSN: 0973-7510
E-ISSN: 2581-690X
The production of weak organic acids by microorganisms has been attributed as the prime reason for the solubilization of insoluble phosphates under both in vitro and soil conditions. Literature seems to be heavily biased towards gluconic acid production by microbes and its subsequent release into the environment as the key factor responsible for phosphate solubilization. This has found credibility since gluconic acid being a product of the Kreb’s cycle is often detected in large quantities in the culture media, when assayed under in vitro conditions. In the present work, the organic acid profiles of four elite phosphate solubilising isolates were determined in the presence of different insoluble sources of phosphates, under in vitro buffered culture conditions by HPLC (High-Performance Liquid Chromatography). While most previous studies did not use a buffered culture media for elucidating the organic acid profile of phosphate solubilizing bacterial isolates, we used a buffered media for estimation of the organic acid profiles. The results revealed that apart from gluconic acid, malic acid is produced in significant levels by phosphate solubilizing bacterial isolates, and there seems to be a differential pattern of production of these two organic acids by the isolates in the presence of different insoluble phosphate sources.
Phosphate Solubilising Bacteria, Organic Acids, Malic acid, Gluconic acid
Phosphorus is an essential nutrient for plant growth and reproduction. Phosphorus is available in soil primarily in organic or inorganic forms, but most forms are not available for plant uptake due to its tendency to form complexes with cations such as Calcium (Ca) in alkaline soils and Aluminum (Al) and Iron(Fe) in acidic soils. Several studies have advocated the use of biological formulations to overcome P deficiency in soil. Many rhizospheric microorganisms have been reported to solubilize mineral phosphates by different mechanisms. The most commonly encountered phosphate solubilizing bacterial genera are Pseudomonas, Bacillus, Rhizobium and Enterobacter. Their ability to solubilize phosphates has been mainly attributed to their organic acid secretion potential1, 2.
One of the most well-known mechanisms of phosphate solubilization is the release of weak organic acids by phosphate solubilizing microorganisms, which solubilize the elemental-phosphatic complexes thereby releasing the solubilized phosphorus into the culture media. Most often different divalent and trivalent organic anions such as malate, citrate and oxalate are produced by the microbes and are implicated to play an important role in the solubilization of phosphates 3. This observation has been supported by several studies on the organic acid production profile under in vitro culture conditions 4-7. Under in vitro culture conditions, the production of organic acids by bacterial strains is largely influenced either by the growth medium or by the presence of insoluble phosphate sources. Gluconic acid has been reported as the main organic acid secreted by many phosphate solubilizing microorganisms under laboratory conditions and is well known for its chelation abilities 8,9. Gluconic acid is produced by the direct extra-cellular oxidation of glucose catalysed by enzyme glucose dehydrogenase coupled with the co-factor pyrroloquinoline quinone (PQQ) 10,11. The oxidation of gluconic acid gives rise to the production of 2-ketogluconic acid, a substantial carboxylic acid that can chelate calcium ions and dissolve hydroxyapatites12.. Though there have been reports of other organic acids viz., oxalic acid, citric acid, acetic acid, malic acid, succinic acid, tartaric acid, propanoic acid, lactic acid, fumaric acid, pyruvic acid etc9,13-18 being produced by phosphate solubilizing bacterial strains, gluconic acid has gained much importance due to its ability to solubilise phosphates under in vitro conditions 12,16,18-20.
Arvind & Gulati13 were the first to decipher the organic acid profile of phosphate solubilizing Pseudomonads using multiple insoluble substrates. But this study was carried out in a non-buffered medium.They presented a detailed description of organic acids produced by different strains of Pseudomonas that solubilised Udaipur rock phosphate (URP), Mussorie rock phosphate (MRP) and North Carolina rock phosphate (NCRP) under non- buffered in vitro conditions and concluded that gluconic acid and 2-ketogluconic acid are the major organic acids responsible for phosphate solubilization under in vitro conditions. A positive correlation between the levels of phosphate solubilisation and the quantity of gluconic acid produced by Enterobacter cloacae EB 27, Serratia marcescens EB 67, Serratia sp. EB 75, Pseudomonas sp. CDB 35 and Pseudomonas sp. BWB 2 was established by Hameeda et al21. The bacterium Azospirillum has been reported to release gluconic acid at high concentrations under in vitro conditions which was implicated in the solubilization of phosphates14.
While several studies in the past have focussed on utility of gluconic acid in phosphate solubilization, they seem to have missed the significant levels of another important organic acid viz., malic acid in the solubilization of phosphates. The present study highlights the extra cellular release of malic acid in significant quantities by strains of three different bacterial genera viz., Pseudomonas, Bacillus and Paraburkholderia species under in vitro conditions. The isolates were carefully chosen from a collection of phosphate solubilizing microbes in order to represent the three major soil genera that play a vital role in phosphate solubilization in the soil. Malic acid has been reported as a crucial metal chelator that is released majorly by fungal species such as Aspergillus niger while solubilizing rock phosphate under in vitro conditions20. There are very few reports about the utility of malic acid production in significant levels by bacterial species and its utility in phosphate solubilisation, nor is its mechanism of phosphate solubilization has been clearly defined. Also concentrations of malic acid equivalent to or at par to gluconic acid have never been reported. This study mainly focuses organic acid production profiles of elite bacterial isolates in a buffered medium under in vitro conditions in the presence of three different insoluble substrates viz., tricalcium phosphate (TCP), Aluminium phosphate Al-P and Ferrous phosphate Fe-P.
Bacterial strains and determination of the phosphate solubilization potential of the isolates
Four efficient phosphate solubilizing bacterial isolates were included in the present study viz. Pseudomonas sp (Ban-4) isolated from Banana (Musa sp.) rhizosphere, Pseudomonas sp (DIN-5) isolated from Sapota (Manilkara zapota) rhizosphere , Bacillus sp (NE-4) isolated from Rice (Oryza sativa) and Paraburkholderia tropica (P-31) isolated from Pomegranate (Punica granatum) rhizosphere.These strains were characterized based on their phenotypic characters and 16S rRNA gene sequencing. The bacterial strains were maintained at -70°C in 50 % gycerol stocks.
The phosphate solubilization potential of the elite bacterial strains was studied using three insoluble phosphate sources viz., Tricalcium phosphate ( TCP), Aluminium phosphate Al-P, Iron phosphate Fe-P in a buffered medium. The media contained 100 mM glucose, 25 µM Magnesium sulphate (MgSO4, 10 mM Ammonium chloride (NH4Cl) and the following micronutrients (mg/l) viz., Ferrous sulphate (FeSO4)-3.5; Zinc sulphate (ZnSO4)-0.16; Copper sulphate (CuSO4)–-0.08; Boric acid (H3BO3)- 0.5; Calcium chloride (CaCl2) – 0.03; Manganese sulphate (MnSO4) – 0.4 and it was buffered with 100 mMTris Hydrochloride (HCl) (pH 8.0)22-24. The elite bacterial strains were grown in 100 ml minimal buffered broth supplemented with either 0.5% tricalcium phosphate (TCP) or Aluminium phosphate (Al-P) or Ferrous phosphate (Fe-P) separately and incubated at 30°C for 7 days under shaking conditions. On completion of the incubation period the culture was centrifuged at 10,000 rpm for 10 min and the pH of the culture supernatant was recorded. Subsequently the soluble P released by the bacterial isolates was determined quantitively by the spectrophotometric method25. All the studies were conducted in triplicate.
Identification and quantification of the organic acid profile of the promising isolates under using high performance liquid chromatography (HPLC)
The cell free culture supernatant obtained mentioned above was filter sterilized using a 0.22 µm nylon filter. The detection and quantification of organic acids was done using a HPLC (Shimadzu, Japan), equipped with photodiode array detector (SPD-M2OA) and Synergi 4µm RP-C18 column (Phenomenex, USA, 250X4.6mm). The mobile phase consisted of 1mM Sulphuric acid (H2SO4) + 8mM Sodium sulphate (Na2SO4) (1:1v/v) containing 0.1% orthophosphoric acid at 0.5ml/min flow rate. The organic acids present in the culture filtrate were quantified at 210 nm using standards of different organic acids viz., tartaric acid, formic acid, malic acid, malonic acid, lactic acid, citric acid, succinic acid, propionic acid, gluconic acid, oxalic acid, Keto-D glutarate. The elutates were detected and quantified with reference to the peak areas obtained by using the respective standards.
Phosphate solubilizing potential of the bacterial strains
Among four strains Paraburkholderia tropica P-31 released the highest levels of soluble P (50.0 μg/ml) when TCP was used as insoluble source of phosphorus and drop in pH was observed from 7.0 to 4.4. The isolate Bacillus sp. NE-4 released the highest levels of soluble P (19.0 μg/ml) when Fe-P was used as insoluble source of phosphorus and a drop in pH was observed from 7 to 5.0. Pseudomonas sp DIN-5, released the highest levels of soluble P i.e. 21.9 (μg/ml) when Al-P was used as insoluble source of phosphorus in the medium but a concomitant drop in pH was not observed (Table -1). It should be noted here that solubility of Calcium phosphate (Ca- P) increases exponentially with decreasing pH whereas solubility of Iron phosphate (Fe-P) decreases with the decrease in the pH in the range of 4.5 to 3.5 and solubility of Aluminum Phosphate ( Al-P) is lowest in the pH range of 5.5-4.522-24. The results of the present study are therefore in accordance with the observations of Merbach23 and Henry24. Microorganisms have been known to deploy different mechanisms for the dissolution of different phosphate complexes. Acidification of the medium is proposed as a major mechanism for the dissolution of the Ca-P complex, this mechanism results in the acidification of the medium, due to the release of protons. Alternatively metal complexing and metal reduction are the possible mechanisms of dissolution of Al and Fe phosphates. By forming metal complexes the organic acids reduce metals from their variable oxidation states to lower oxidation states thereby resulting in the release of soluble P forms26.
Table (1):
Quantitative estimation of the P solubilising abilities of the elite phosphate solubilising bacterial isolates under in vitro conditions.
| Isolate | TCP | Fe-P | Al-P | |||
|---|---|---|---|---|---|---|
| pH |
Soluble P (ppm) |
pH | Soluble P (ppm) |
pH | Soluble P (ppm) |
|
| Pseudomonas sp. (DIN-5) | 4.6 | 46.4 | 7.3 | 6.0 | 7.4 | 21.9 |
| Paraburkholderia tropica (P-31) | 4.4 | 50.0 | 7.3 | 6.7 | 7.3 | 8.2 |
| Bacillus sp.(NE-4) | 5.2 | 20.3 | 5.0 | 19.0 | 5.0 | 6.3 |
| Pseudomonas sp. (BAN-4) | 4.6 | 35.6 | 5.2 | 1.0 | 4.8 | 7.3 |
There are many reports of PSB’s which solubilize TCP but do not dissolve Fe-P and Al-P complexes and are therefore of not much utility under dynamic field conditions27,28 because of two major reasons, viz., a contrasting range of soil pH levels and is the strong buffering capacity of soils 29. The experimental framework of the present study therefore took cognizance of the above factors. Though, buffering of media drastically lowers the phosphate solubilizing capacity of microbes the elite phosphate solubilisers are likely to sustain their performance under buffered conditions30.
Table (2):
Organic acid profile of the elite isolates under in vitro conditions.
| Bacterial Isolates | Organic Acid Concentration ( μg/ml) | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Insoluble P sources | Tartaric acid | Formic acid | Malic acid | Malonic acid | Lactic acid | Citric acid | Succinic acid | Propionic acid | Gluconic acid | Oxalic acid | Keto D Gluterate | |
| Pseudomonas sp DIN-5 |
TCP | 20.2 | 19.9 | 1135.38 | 865.6 | ND | 83.8 | 213.9 | ND | 541.38 | ND | ND |
| Fe-P | 18.9 | 16.6 | 751.78 | 91.3 | 92.5 | 91.4 | 652.7 | ND | 1125.43 | ND | ND | |
| Al-P | ND | 86.3 | 1025.65 | ND | ND | 62.9 | 262.8 | ND | 742.33 | ND | 36.9 | |
| Bacillus sp NE-4 |
TCP | 42.8 | 20.5 | 342.48 | 92.2 | ND | 76.2 | 552.7 | ND | 1124.35 | ND | 52.7 |
| Fe-P | 35.7 | 9.3 | 752.28 | ND | 85.3 | ND | 747.9 | ND | 926.50 | ND | ND | |
| Al-P | 23.8 | 8.177 | 1739.03 | 423.2 | ND | 89.4 | 572.3 | ND | 292.13 | 5.5 | 141.5 | |
| Pseudomonas sp Ban-4 |
TCP | 38.9 | 41.1 | 1124.90 | 193.6 | 84.4 | ND | 553.6 | ND | 312.13 | ND | 92.6 |
| Fe-P | 23.9 | 41.3 | 865.33 | 711.4 | 65.3 | 76.6 | 785.5 | ND | 625.95 | 2.65 | 23.6 | |
| Al-P | 52.6 | 15.4 | 928.78 | 212.5 | 91.5 | 235.6 | 411.2 | ND | 618.83 | ND | 21.1 | |
| Paraburkholderia tropica P-31 |
TCP | 37.9 | 58.4 | 752.28 | 547.9 | ND | 52.9 | 411.6 | ND | 784.28 | 5.3 | 37.9 |
| Fe-P | 41.6 | 19.8 | 1110.48 | 835.8 | ND | 21.7 | 112.7 | ND | 0.53 | 5.6 | 41.6 | |
| Al-P | 96.2 | 23.1 | 1408.10 | ND | 22.6 | 131.4 | 841.4 | ND | 465.85 | ND | 96.2 | |
ND- Not detected
Deciphering the organic acid profile of the phosphate solubilizing bacterial isolates under in vitro conditions by High Performance Liquid Chromatography (HPLC)
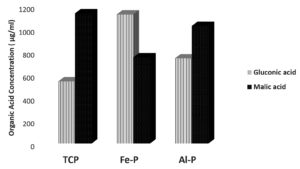
When the organic acid produced by the isolates were quantified under in vitro conditions, the concentration of organic acids viz., gluconic acid and malic acid were found to be higher compared to other acids (Table 2). Apart from and gluconic acids, succinic acid was the other predominant organic acid produced across substrates and isolates. Closer scrutiny of the data revealed a differential pattern of organic acid release by the isolates in the presence of different insolube sources of phosphates. The extracellular concentrations of malic acid were higher when Pseudomonas sp. (DIN-5) was cultured in the presence of TCP and Al-P whereas gluconic acid was detected in higher concentrations when Fe-P was used as a an insoluble substrate of P. Conversely in the case of Bacillus sp. (NE-4) the gluconic acid concentrations were higher in the presence of TCP and Fe-P while extracellular malic acid levels were higher when Al-P was used as an insoluble substrate of P. The strain Pseudomonas sp. (BAN-4) released highest concentrations of malic acid compared to gluconic acid across insoluble substrates of P, while Paraburkholderia tropica (P-31) released highest levels of malic acid when Fe-P and Al-P were used as insoluble substrates of P, while the highest concentrations of gluconic acid was recorded when TCP was used as an insoluble source of P. It could be concluded that the highest levels of malic acid were detected in eight of the twelve scenarios under which the organic acid profiles were estimated (Fig. 1 a, b, c, d).
Fig. 1(a). Relative concentrations of malic and gluconic acids produced by the phospate solubilzing Pseudomonas sp. DIN-5 under in vitro conditions in the presence of different insoluble phosphate sources
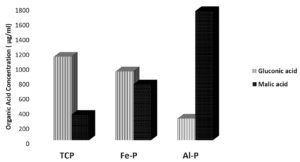
Fig. 1 (b). Relative concentrations of malic and gluconic acids produced by the phospate solubilzing Bacillus sp. NE-4under in vitro conditions in the presence of different insoluble phosphate sources
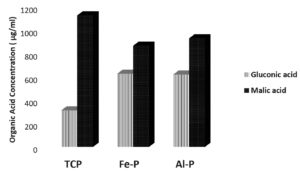
Fig. 1 (c). Relative concentrations of malic and gluconic acids produced by the phospate solubilzing Pseudomonas sp. BAN-4 under in vitro conditions in the presence of different insoluble phosphate sources
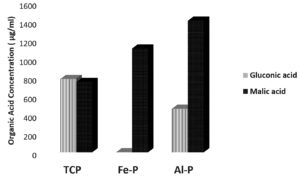
Fig. 1 (d). Relative concentrations of malic and gluconic acids produced by the phospate solubilzing Paraburkholderia tropica. P-31 under in vitro conditions in the presence of different insoluble phosphate sources
The earlier studies on the in vitro organic acid production by phosphate solubilizing bacterial isolates have reported that gluconic acid is produced by several bacterial genera in significant quantities while malic acid is produced by phosphate solubilizing fungal genera like Aspergillus1,20,31-35. The novelty of this study rests in the fact that it has been proved conclusively that bacterial genera apart from Pseudomonas produce malic acid in higher or equivalent quantities to that of gluconic acid across insoluble phosphate sources in a buffered media.
In general malic acid has applications in the liquor, food, synthetic and pharmaceutical industries33 but its agricultural usage is not well known. L-malic acid is known to be produced majorly by species of Aspergillus, Schizophyllum34-36 , Penicillium, Aureobasidium1,30,34, an engineered strain of E.coli35, species of Penicillium, Aureobasidium36-40, and Bacillus subtilis35,41. The literature suggests that the most probable mechanism of malic acid production is the increase in utilisation of glucose via carboxylation of pyruvate, followed by reduction of oxaloacetate36. Saccharomyces cerevisiae isolated from Sake mash was reported to produce very high levels of malic acid by downregulating the production of thymine genes (THI4) and upregulation of stress genes (HSP12)37. Similarly among bacteria a mutant strain of Thermobifida fusca muC accumulated malic acid on a medium containing cellulose where phosphoenolpyruvate is known to get converted to oxaloacetate in the presence of phosphoenol carboxylase and further oxaloacetate being reduced to malate in the presence of malate dehydrogenase39-41. Microorganisms utilising the reductive pathway are known to secrete maximum concentrations of malic acid qualitatively and quantitatively 42-44. Fungal strains of Aspergillus, Ustilago trichophora and Aureobasidium pullulans are known to secrete good quantities of malic acid from biofuel related coproducts and biomass42 . Alternatively, other methods have been mentioned in literature for the increased synthesis of malic acid like in case of Saccharomyces cerevisiae whose cells contained overexpressed gene for fumarate hydratase catalysed the conversion of fumarate to malate43.
Several enzymes are known to play a role in metabolising malic acid. The NADP-malic enzyme (NADP-ME), has been known to produce the essential components of energy and for the biosynthesis of defense components. These defense compounds are implicated in the malate metabolism in plant defense44. Malic enzymes have also been found in Sinorhizobium meliloti45,46, Bacillus stearothermophilus,47 Streptococcus bovis48, Corynebacterium glutamicum49, Lactococcus lactis50 and B. subtilis51,52. The list is exhaustive when it comes industrial production of malic acid and its utility but from the agriculture point of view, we have seen two main studies, one is based on metal chelation and other is known for eliciting plant defences, yet another study focusses on demonstrating the role of malic acid released from the roots of Arabidopsis (Arabidopsis thaliana) which attracts the useful rhizobacterium Bacillus subtilus FB17 and assisted in biofilm formation53. Thus far, no study demonstrated the possible pathway of malic acid production by Gram negative bacteria which is good enough to bring about the solubilization of phoshates. While gluconic acid is known as one of the key organic acids released by several species of Pseudomonas54 and many Gram negative and Gram positive bacterial genera and fungal genera for phosphate solublization55-60. The role of malic acid in phosphate solubilization has not been well documented. This study opens up the exploration of the utility and role of malic acid is the solubilization of phosphates, under in vitro conditions.
This study has shown that under in vitro buffered condiitons apart from gluconic acid, malic acid is also produced in significant levels by the Gram negative phosphate solubilizing bacterial isolates belonging to various genera. There seems to be a differential pattern of the production of theese two major organic acids across isolates and insoluble sources of phosphates. But nevertheless in eight of the twelve scenarios explored in this study, malic acid concentrations were higher than the gluconic acid concentrations, thereby conslusively establishing the role of malic acid in phospahte solubilization. Previous studies have not determined the possible pathways of malic acid production by Gram negative bacteria, which is likely to influence phosphate solubilization. Threfore future work has to be initiated in this direction to explore the pathways of malic acid production in Gram negative bacteria and to further elucidate its role in bacterial phosphate solubilization.
ACKNOWLEDGMENTS
The authors would like to acknowledge Dr. Maryada Sharma, for her valuable assistance in HPLC analysis.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This study was financially supported by the Department of Science and Technology, Government of India under WOS-A scheme (Grant number SR/WOS-A/LS-1080/2014).
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analysed during this study are included in the manuscript.
- Rodrı́guez H, Fraga R. Phosphate solubilizing bacteria and their role in plant growth promotion. Biotechnol Adv. 1999;17(4-5):319-39.
Crossref - Whitelaw MA. Growth promotion of plants inoculated with phosphate solubilizing fungi. Edited by Donald L. Sparks. Adv Agron, Academic press. 1999;69: 99-151.
Crossref - Pohlman AA, Mc Coll JG. Kinetics of metal dissolution from forest soils by soluble organic acids 1. J Environ Qual. 1986;15 (1):86-92.
Crossref - Sperber JI. Solution of apatite by soil microorganisms producing organic acids. Aust J Agric Res. 1958;9 (6) :782-7.
Crossref - Banik S, Dey BK. Available phosphate content of an alluvial soil as influenced by inoculation of some isolated phosphate-solubilizing micro-organisms. Plant Soil. 1982; 69(3):353-64.
Crossref - Goldstein AH. Bacterial solubilization of mineral phosphates: historical perspective and future prospects. Am J Alternative Agr. 1986;1:51-7.
Crossref - Cunningham JE, Kuiack C. Production of citric and oxalic acids and solubilization of calcium phosphate by Penicillium bilaii. Appl. Environ Microbiol. 1992; 58 (5):1451-8.
Crossref - Goldstein AH. Recent progress in understanding the molecular genetics and biochemistry of calcium phosphate solubilization by Gram negative bacteria. Biol Agric Hortic. 1995; 12(2):185-93.
Crossref - Chen YP, Rekha PD, Arun AB, Shen FT, Lai WA, Young CC. Phosphate solubilizing bacteria from subtropical soil and their tricalcium phosphate solubilizing abilities. Appl Soil Ecol. 2006;34(1):33-41.
Crossref - Kim CH, Han SH, Kim KY, et al. Cloning and expression of pyrroloquinoline quinone (PQQ) genes from a phosphate-solubilizing bacterium Enterobacter intermedium. Curr Microbiol. 2003; 47 (6) :457-61.
Crossref - Whiting PH, Midgley M, Dawes EA. The role of glucose limitation in the regulation of the transport of glucose, gluconate and 2-oxogluconate, and of glucose metabolism in Pseudomonas aeruginosa. Microbiology. 1976; 92 (2) :304-10.
Crossref - Moghimi A, Tate ME, Oades JM. Characterization of rhizosphere products especially 2-ketogluconic acid. Soil BiolBiochem. 1978;10(4):283-7.
Crossref - Vyas P, Gulati A. Organic acid production in vitro and plant growth promotion in maize under controlled environment by phosphate-solubilizing fluorescent Pseudomonas. BMC Microbiol. 2009;9:174.
Crossref - Rodriguez H, Gonzalez T, Goire I, Bashan Y. Gluconic acid production and phosphate solubilization by the plant growth-promoting bacterium Azospirillum spp. Naturwissenschaften. 2004 ;91(11):552-5.
Crossref - Buch A, Archana G, Kumar GN. Heterologous expression of phosphoenolpyruvate carboxylase enhances the phosphate solubilizing ability of fluorescent Pseudomonads by altering the glucose catabolism to improve biomass yield. Bioresour Technol. 2010;101(2):679-87.
Crossref - Gulati A, Sharma N, Vyas P, et al. Organic acid production and plant growth promotion as a function of phosphate solubilization by Acinetobacter rhizosphaerae strain BIHB 723 isolated from the cold deserts of the trans-Himalayas. Arch Microbiol. 2010;192(11):975-83.
Crossref - Perez E, Sulbaran M, Ball MM, Yarzabal LA. Isolation and characterization of mineral phosphate-solubilizing bacteria naturally colonizing a limonitic crust in the south-eastern Venezuelan region. Soil Biol Biochem. 2007 ;39(11):2905-14.
Crossref - de Werra P, Péchy-Tarr M, Keel C, Maurhofer M. Role of gluconic acid production in the regulation of biocontrol traits of Pseudomonas fluorescens CHA0. Appl Environ Microbiol. 2009; 75 (12) :4162-74.
Crossref - Ogut M, Er F, Kandemir N. Phosphate solubilization potentials of soil Acinetobacter strains. Biol Fertil Soils. 2010;46(7):707-15.
Crossref - Akintokun AK, Akande GA, Akintokun PO, Popoola TO, Babalola AO. Solubilization of insoluble phosphate by organic acid-producing fungi isolated from Nigerian soil. Int J Soil Sci. 2007;2:301-7.
Crossref - Hameeda B, Reddy YH, Rupela OP, Kumar GN, Reddy G. Effect of carbon substrates on rock phosphate solubilization by bacteria from composts and macrofauna. Curr Microbiol. 2006;53(4):298-302.
Crossref - Merbach W, Fankem H, Deubel A. Influence of rhizosphere bacteria of African oil palm (Elaeis guineensis) on calcium, iron, and aluminum phosphate in vitro mobilization. In International symposium “Root Research and Applications 2009 Sep (pp. 2-4).https://d1wqtxts1xzle7.cloudfront.net/42005465/Influence_of_rhizosphere_bacteria_of_Afr20160203-5553-ybtr5j.pdf?1454555825=&response-content-disposition=inline%3B+filename%3DInfluence_of_rhizosphere_bacteria_of_Afr.pdf&Expires=1622146098&Signature=QRfGIqO
- Merbach W, Deubel A, Gransee A, Ruppel S, Klamroth AK. Phosphorus solubilization in the rhizosphere and its possible importance to determine phosphate plant availability in soil. A review with main emphasis on German results. Arch Agron Soil Sci. 2010 ;56(2):119-38.
Crossref - Henri F, Laurette NN, Annette D, et al. Solubilization of inorganic phosphates and plant growth promotion by strains of Pseudomonas fluorescens isolated from acidic soils of Cameroon. Afr J Microbiol Res. 2008;2(7):171-8.
Crossref - Murphy JA, Riley JP. A modified single solution method for the determination of phosphate in natural waters. Anal Chim Acta. 1962; 27:31-6.
Crossref - Bashan Y, Kamnev AA, de-Bashan LE. Tricalcium phosphate is inappropriate as a universal selection factor for isolating and testing phosphate-solubilizing bacteria that enhance plant growth: a proposal for an alternative procedure. Biol Fertil Soils. 2013; 49(4):465-79.
Crossref - Gyaneshwar P, Kumar GN, Parekh LJ, Poole PS. Role of soil microorganisms in improving P nutrition of plants. Plant Soil. 2002;245(1):83-93.
Crossref - Rengel Z, Marschner P. Nutrient availability and management in the rhizosphere: exploiting genotypic differences. New Phytol. 2005;168(2):305-12.
Crossref - Ae N, Arihara J, Okada K. Phosphorus response of chickpea and evaluation of phosphorus availability in Indian Alfisols and Vertisols. Phosphorus nutrition of grain legumes in the semi-arid tropics (Johansen, C., Lee, KK, and Sahrawat, KL, eds.). Patancheru. 1991; 502(324):33-41.
- Gyaneshwar P, Kumar GN, Parekh LJ. Effect of buffering on the phosphate-solubilizing ability of microorganisms. World J Microbiol Biotechnol. 1998; 14(5):669-73.
Crossref - Satyaprakash M, Nikitha T, Reddi EU, Sadhana B, Vani SS. Phosphorous and phosphate solubilising bacteria and their role in plant nutrition. Int J Curr Microbiol Appl Sci. 2017;6(4):2133-44.
Crossref - Walpola BC, Yoon MH. Prospectus of phosphate solubilizing microorganisms and phosphorus availability in agricultural soils: A review. Afr J Microbiol Res. 2012;27;6(37):6600-5.
Crossref - Mehrvarz S, Chaichi MR, Alikhani HA. Effect of phosphate solubilizing microorganisms and phosphorus chemical fertilizer on forage and grain quality of barely (Hordeum vulgare L.). Agric Environ Sci. 2008;3(6):822-828.
- Kumar A, Kumar A, Patel H. Role of microbes in phosphorus availability and acquisition by plants. Int J Curr Microbiol Appl Sci. 2018;7(5):1344-7.
Crossref - Gaind S. Phosphate dissolving fungi: mechanism and application in alleviation of salt stress in wheat. Microbiol Res. 2016;193:94-102.
Crossref - Zelle RM, de Hulster E, van Winden WA, et al. Malic acid production by Saccharomyces cerevisiae: engineering of pyruvate carboxylation, oxaloacetate reduction, and malate export. Appl Environ Microbiol. 2008 1;74(9):2766-77.
Crossref - Takahiro OBA, Suenaga H, Nakayama S, et al. Properties of a high malic acid-producing strains of Saccharomyces cerevisiae isolated from sake mash. Biosci Biotechnol Biochem. 2011;75(10):2025-2029.
Crossref - Deng WW, SasamotoH, AshiharaH. Effect of caffeine on the expression pattern of water- soluble proteins in rice (Oryza sativa) seedlings. Nat Prod Commun. 2015; 10(5):733-736.
Crossref - Chi Z, Wang ZP, Wang GY, Khan I, Chi ZM. Microbial biosynthesis and secretion of l-malic acid and its applications. Crit Rev Biotechnol. 2016; 36:99-107.
Crossref - Brown SH, Bashkirova L, Berka R, et al. Metabolic engineering of Aspergillus oryzae NRRL 3488 for increased production of L-malic acid. Appl Microbiol Biotechnol. 2013;97:8903-12.
Crossref - Zhang H, Cai J, Dong J, et al. High-level production of poly (β-L-malic acid) with a new isolated Aureobasidium pullulans strain. Appl Microbiol Biotechnol. 2011;92:295-303.
Crossref - West TP. Microbial production of malic acid from biofuel-related coproducts and biomass. Fermentation. 2017;3:14.
Crossref - Neufeld RJ, Peleg Y, Rokem JS, Pines O, Goldberg I. L-Malic acid formation by immobilized Saccharomyces cerevisiae amplified for fumarase. Enzyme Microb Technol. 1991;13:991-6.
Crossref - Casati P, Drincovich MF, Edwards GE, Andreo CS. Malate metabolism by NADP-malic enzyme in plant defense. Photosynth Res. 1999; 61:99-105.
Crossref - Mitsch MJ, Cowie A, Finan TM. Malic enzyme cofactor and domain requirements for symbiotic N2 fixation by Sinorhizobium meliloti. J Bacteriol. 2007; 189:160-8.
Crossref - Voegele RT, Mitsch MJ, Finan TM. Characterization of two members of a novel malic enzyme class. Biochim Biophys Acta Protein Struct Molec Enzym. 1999;1432:275-85.
Crossref - Kobayashi K, Doi S, Negoro S, Urabe I, Okada H. Structure and properties of malic enzyme from Bacillus stearothermophilus. J Biol Chem. 1989;264:3200-5.
Crossref - Kawai S, Suzuki H, Yamamoto K, Inui M, Yukawa H, Kumagai H. Purification and characterization of a malic enzyme from the ruminal bacterium Streptococcus bovis ATCC 15352 and cloning and sequencing of its gene. Appl Environ Microbiol. 1996 ;62:2692-700.
Crossref - Gourdon P, Baucher MF, Lindley ND, Guyonvarch A. Cloning of the malic enzyme gene from Corynebacterium glutamicum and role of the enzyme in lactate metabolism. Appl Environ Microbiol. 2000;66:2981-7.
Crossref - Martín MG, Sender PD, Peirú S, De Mendoza D, Magni C. Acid-inducible transcription of the operon encoding the citrate lyase complex of Lactococcus lactis biovar diacetylactis CRL264. J Bacteriol. 2004;186:5649-60.
Crossref - Doan T, Servant P, Tojo S, et al. The Bacillus subtilis ywkA gene encodes a malic enzyme and its transcription is activated by the YufL/YufM two-component system in response to malate. Microbiology. 2003 ;149(9):2331-43.
Crossref - Lerondel G, Doan T, Zamboni N, Sauer U, Aymerich S. YtsJ has the major physiological role of the four paralogous malic enzyme isoforms in Bacillus subtilis.
J Bacteriol. 2006 ;188:4727-36.
Crossref - Rudrappa T, Czymmek KJ, Paré PW, Bais HP. Root-secreted malic acid recruits beneficial soil bacteria. Plant Physiol. 2008;148:1547-56.
Crossref - Goldstein AH, Liu ST. Molecular cloning and regulation of a mineral phosphate solubilizing gene from Erwinia herbicola. Biotechnol. 1987; 5(1) :72-4.
Crossref - Babu-Khan S, Yeo TC, Martin WL, Duron MR, Rogers RD, Goldstein AH. Cloning of a mineral phosphate-solubilizing gene from Pseudomonas cepacia. Appl Environ Microbiol. 1995; 61(3) :972-8.
Crossref - Kim KY, Jordan D, McDonald GA. Effect of phosphate-solubilizing bacteria and vesicular-arbuscular mycorrhizae on tomato growth and soil microbial activity. Biol Fert Soils. 1997; 26(2):79-87.
Crossref - Kim KY, Jordan D, Krishnan HB. Expression of genes from Rahnella aquatilis that are necessary for mineral phosphate solubilization in Escherichia coli. FEMS Microbiol Lett. 1998; 159 (1) :121-7.
Crossref - Krishnaraj PU, Goldstein AH. Cloning of a Serratia marcescens DNA fragment that induces quinoprotein glucose dehydrogenase-mediated gluconic acid production in Escherichia coli in the presence of stationary phase Serratia marcescens. FEMS Microbiol Lett. 2001; 205 (2) :215-20.
Crossref - Fasim F, Ahmed N, Parsons R, Gadd GM. Solubilization of zinc salts by a bacterium isolated from the air environment of a tannery. FEMS Microbiol Lett. 2002; 213(1) :1-6.
Crossref - Bashan Y, Kamnev AA, de-Bashan LE. A proposal for isolating and testing phosphate-solubilizing bacteria that enhance plant growth. Biol Fert Soils. 2013;49:1-2.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.