ISSN: 0973-7510
E-ISSN: 2581-690X
Candida species are major opportunistic fungal causative agents for significant infections. While C. albicans is considered the most widespread and virulent species, the clinical importance of non-albicans Candida species has substantially risen. This study investigates Palestinian clinical isolates of Candida species by identifying their species type, genotype, mating type, and biofilm formation capability, as well as detecting virulence factors using polymerase chain reaction (PCR). One hundred clinical Candida isolates were obtained from four medical governmental hospitals in Central and Northern Palestine. The PCR technique was used to identify the Candida species and the ABC genotype for the 25S rDNA. The mating type was also determined, besides the detection of hyphal wall protein 1, agglutinin-like sequence genes, phospholipases, and secreted aspartic proteases. Genetic heterogeneity among C. albicans strains was determined by RAPD-PCR typing. C. albicans (53%) and C. tropicalis (15%) are highly represented species. Genotype A was the most prevalent among C. albicans isolates (27%). Notably, the Agglutinin-like sequence genes are present in 59% of the isolates. In contrast, 30% of the isolates have either the SAP9 or/and SAP10 genes. Therefore, the results of RAPD-PCR typing revealed that 14 fragments were produced using the OPI 06 primer with a 100% polymorphism. The 24 C. albicans isolates were clustered into four groups on a 50% cut-off similarity. C. tropicalis was the most frequently isolated non-albicans Candida species. Biofilm formation and the presence of SAP9 and/or SAP10 genes were mainly observed in C. albicans strains, which exhibited low genetic variability.
Agglutinin-like Sequence, Candida Species, Genotyping, Mating Type, Multiplex PCR, RAPD, Virulence Factors
Candidiasis represents a major opportunistic infection, particularly affecting immunocompromised patients and hospitalized children under four years, especially those suffering from respiratory illnesses.1 Part of the Candida genus, most notably C. albicans, normally inhabit human mucosal surfaces as harmless commensals but have increasingly emerged as significant invasive pathogens. This trend is largely associated with the growing population of immunocompromised individuals and widespread antibiotic use.2 The virulence of Candida species arises from multiple factors such as biofilm development, toxin secretion, and the activity of hydrolytic enzymes which collectively enhance tissue invasion and impair host immune responses.3
Nowadays, non-albicans Candida species have been associated with severe illnesses characterized by high morbidity and mortality rates.4 Likewise, vulvovaginitis (VV) is considered one of the most commonly encountered women’s health issues, which can be caused by either microbial or non-microbial sources.5 Most infections are attributed to C. albicans and non-albicans Candida (NAC) species, including C. glabrata, C. parapsilosis, and C. tropicalis,6 which mostly colonize the vaginal and rectal regions, with similar symptoms and complications.7 Furthermore, it has been shown that the NAC species are involved in candidiasis with higher prevalence and clinical impact in comparison to C. albicans candidiasis.8 One of the most prevalent NAC species is C. tropicalis, which is considered an opportunistic type that mainly affects immunocompromised patients, causing critical systemic diseases.9
Candidiasis does not arise from a single cause; rather, its development reflects a convergence of three interrelated domains: the intrinsic characteristics of the fungus, the systemic and local status of the host, and modifications within the microenvironment. Among the Candida species, C. albicans remains the most frequently recovered isolate, constituting roughly 80% of clinical isolates. However, species such as C. tropicalis, C. glabrata, C. parapsilosis, C. krusei, and C. dubliniensis are by no means uncommon in human infections. What makes candidiasis particularly distinctive in the landscape of infectious diseases is that the organism resides harmlessly in a significant proportion of healthy individuals, only manifesting as disease when the equilibrium between host defenses and fungal proliferation is disturbed.10 The organism’s capacity for dimorphic switching between its blastospore and hyphal forms alongside its ability to form biofilms, elaborate proteolytic enzymes, and express surface adhesins, underpins its pathogenic potential when conditions become permissive.11 It is the deterioration of local and systemic host defenses, rather than any sudden surge in fungal virulence, that most consistently precipitates disease.12 At the immunological level, Th17-mediated responses driven by interleukin-17 appear central to mucosal containment of Candida, and disruption of this axis correlates strongly with susceptibility to both superficial and invasive forms of infection.13
The range of host-modifying factors that predispose individuals to candidiasis is considerably broad, spanning endocrine, immunological, iatrogenic, nutritional, and local domains. Metabolic and endocrine disturbances most notably diabetes mellitus, but also pregnancy, renal insufficiency, and hyperthyroidism create a systemic milieu unfavorable to effective antifungal defense.10 Patients whose immunity is compromised, whether through cytotoxic chemotherapy, organ transplantation, HIV infection, hematological malignancies, or aplastic anemia, face a substantially elevated risk, as the fungus readily exploits the resulting gaps in both innate and adaptive immunity.10,12 The therapeutic use of broad-spectrum antibiotics and corticosteroids systemic or inhaled has long been recognized as an iatrogenic risk, the former by dismantling the competitive microbial flora, and the latter by attenuating mucosal immunity.14 Nutritional deficiencies involving iron, folate, and certain vitamins, as well as physiological vulnerability at the extremes of age infancy, where immune maturity is incomplete, and older adulthood, where immune senescence
prevails further compound individual susceptibility.10
The protective role of mucosal secretions is underscored by the observation that conditions causing reduced salivary flow and mucosal dryness, irrespective of their etiology, consistently predispose to Candida overgrowth by depleting key antifungal proteins from mucosal surfaces.11 At the level of the microenvironment, ill-fitting prostheses and poorly maintained appliances offer an ideal niche for biofilm formation, while habitual cigarette smoking and regular alcohol consumption erode mucosal integrity and dampen local immune surveillance, thereby acting as potent environmental co-factors in the pathogenesis of candidiasis.10,14
Therefore, various factors associated with Candida spp. invasion and host immune response escaping, including: cell membrane anatomy, morphological conversion, biofilm formation, and hydrolytic enzymes synthesis.15 Moreover, the expression of these virulence factors is highly common among C. albicans, including biofilm formation, morphology conversion, and filamentous growth. However, NAC species have recently been classified as one of the pathogenic species, causing several complicated candidiasis infections.4
However, Candida infections can be diagnosed either by conventional methods, including serological and microbiological methods with low sensitivity, or with precise and sensitive techniques, including antigen detection methods and molecular approaches. Several types of candidiasis have certain diagnostic challenges due to their undefined manifestations, like visceral candidiasis.16 Additionally, standard identification methods include KOH preparation, germ tube testing, chlamydospore formation, and sugar fermentation and assimilation assays. On Candida differential agar (CDA), C. albicans produces green colonies, C. krusei pink to purple, C. tropicalis dark blue, and C. glabrata cream to white colonies. In contrast, Congo red agar (CRA) is used to assess biofilm formation.17 Besides the use of conventional methods for Candida species identification using biochemical tests and CHROM agar, Recent approaches have shifted toward molecular techniques, including DNA amplification combined with the random amplified polymorphic DNA (RAPD) method, alongside commercial DNA extraction kits and various RAPD primers, e.g., OPA02, OPA03, and OPA08, in addition to band analysis software (BioNumerics Applied). RAPD-PCR enables the detection of polymorphisms and the clustering of Candida strains, highlighting its high resolution as a robust tool for strain- and species-level discrimination.18 Numerous therapeutic studies have explored emerging candidemia to examine the interrelation between the genetic profiles of certain C. albicans strains and their antifungal susceptibility.19
For decades, the identification of Candida species in clinical microbiology laboratories has historically depended on a set of well-established conventional techniques, chief among them the germ tube test, chromogenic agar platforms, and carbohydrate assimilation profiles. These methods have endured largely because of their accessibility and low operational cost, particularly in resource-limited settings. Yet their shortcomings, long recognized by specialists, have grown harder to dismiss as the clinical stakes of accurate fungal identification have risen. One of the most consequential limitations is the time required conventional culture-based workflows frequently demand up to 72 hours before a reliable species-level identification can be rendered, a delay that carries real implications for patients with rapidly progressing infection. Equally problematic is the extent to which accurate identification depends on the individual skill and experience of the analyst, introducing a degree of subjectivity that undermines reproducibility across laboratories and settings.20 The difficulty is compounded when clinically significant yet phenotypically indistinct species enter the picture. C. glabrata, C. parapsilosis, and C. dubliniensis, for instance, share overlapping biochemical signatures that render reliable differentiation through conventional means genuinely unreliable, yet each carries a distinct virulence profile and antifungal susceptibility pattern with direct bearing on treatment decisions.21,22 Molecular methods have reshaped what is possible in clinical mycology, not merely by accelerating species identification but by offering a depth of biological insight that conventional approaches simply cannot provide. PCR-based platforms, including multiplex PCR, High-Resolution Melting Analysis, and Multi-Locus Sequence Typing, have demonstrated both the sensitivity and specificity needed for direct species identification from clinical specimens, often delivering results within hours rather than days.23 Among available molecular targets, sequencing of the internal transcribed spacer (ITS) regions of ribosomal DNA has emerged as the reference standard, capable of resolving species boundaries that phenotypic methods routinely blur.20 The contribution of molecular characterization, however, extends well beyond identification alone. Multiplex PCR assays that specifically target virulence-associated genes, including those encoding hyphal wall protein (HWP1), agglutinin-like sequence proteins (ALS), and secreted aspartyl proteases (SAPs), have made it possible to draw meaningful connections between genetic architecture and clinical behavior, connections that are invisible to conventional diagnostics.22 At the level of antifungal resistance, whole-genome and RNA sequencing approaches have enabled investigators to pinpoint the precise genetic alterations notably mutations in ERG11, FKS1, and FKS2 that confer resistance to azoles and echinocandins, mechanisms that would otherwise escape detection through phenotypic susceptibility testing alone.23,24 The cumulative evidence from these advances strongly supports the integration of molecular methods into routine clinical mycology as an essential complement to, and in many contexts a replacement for, conventional identification approaches.
The body of knowledge surrounding Candida infections in Palestine remains strikingly limited when measured against the vast scope of mycological research published worldwide. The first and, for some time, only molecular study conducted in the region was that of Ali-Shtayeh et al., who genotyped 151 C. albicans isolates from patients with vulvovaginal and cutaneous candidiasis using PCR targeting the transposable group I intron of the 25S rDNA gene. Their findings identified genotype C as the most frequently encountered type, while genotype A isolates were found to be more resistant to fluconazole, flucytosine, and amphotericin B relative to the other genotypes an observation that pointed, provocatively, to a possible association between genotype and resistance phenotype. Significant as this contribution was, it remained bounded by its own design: the study addressed a single species in two specific clinical contexts, and the wider Candida species landscape across the full range of clinical specimen types in Palestine was left entirely unexplored.19 Nearly a decade later, Baniodeh, Hanaa et al. provided what constitutes the first broad-scope prevalence and antifungal susceptibility profile of Candida infections across Palestinian hospital settings, documenting eleven species among clinical isolates and drawing attention to the increasing dominance of non-albicans Candida species and their frequently reduced susceptibility to first-line antifungal agents. However, species identification throughout that study relied entirely on phenotypic methods, specifically CHROMagar and the VITEK 2 Compact system, with no molecular component applied at any stage.25 Read alongside the Ali-Shtayeh et al. study, what emerges is a portrait of a regional literature in which molecular inquiry has barely scratched the surface: one study, a decade old, examining a single species across two infection types, and another offering phenotypic data without any genomic depth. To date, no investigation has attempted a comprehensive molecular characterization of Candida isolates spanning diverse clinical specimen types in Palestine, nor has any study examined virulence gene carriage or resistance-associated genetic mutations among strains circulating within the local healthcare environment. This is not a trivial gap. Antifungal resistance profiles are well-established to fluctuate across geographic regions and institutional settings, and without region-specific molecular data, clinicians are left to extrapolate from foreign surveillance figures that may bear little resemblance to local realities.20,23 The present study was conceived precisely to fill this void undertaking, for the first time in Palestine, a comprehensive molecular characterization of Candida species recovered from clinical specimens across varied anatomical sites, with the intent of building a locally grounded evidence base capable of supporting improved clinical decision-making, strengthening national surveillance capacity, and contributing meaningfully to the mycological literature of the broader Eastern Mediterranean region.
Ethical approval and informed consent
This study was approved by the Ethics Committee, An-Najah National University and informed consent was obtained from all participants in the study before the collection of samples and data acquisition.
Study period
Clinical isolates were consecutively collected from multiple medical centers in Central and Northern Palestine between October 2024 and January 2025.
Sample collection
In total, 100 clinical isolates of Candida spp. were obtained from four governmental hospitals in Palestine, including Palestine Medical Complex, An-Najah National University Hospital, Rafidia Surgical Hospital, and Al-Watani Hospital. The source of specimens which were obtained from patients with Candida infections in this study includes; urine (45%), high vaginal swabs HVS (19%), wound (13%), sputum (9%), trap-collected sputum (TRAP) (8%), body fluids (3%), blood (2%) and nail scraping (1%) (Table 1). Candida isolates were included in this study if they were collected from clinically confirmed cases of candidiasis, as diagnosed by treating physicians, obtained from patients admitted to or attending hospital laboratories. Isolates were eligible for inclusion if they had been previously subcultured on standard culture media, subjected to germ tube testing, and identified using the Vitek 2 automated identification system, with species-level identification available for a subset of isolates processed in laboratories equipped with this system. In cases where mixed-species cultures were encountered, each species was considered eligible for inclusion provided it had been successfully subcultured onto separate culture plates and confirmed as a distinct isolate. All included isolates were subsequently subcultured onto Sabouraud Dextrose Agar (SDA) to obtain fresh, viable colonies suitable for downstream DNA extraction and molecular analysis. Isolates were excluded from the study if they represented duplicate samples from the same patient, defined as repeated isolates of the same Candida species obtained from the same individual at different time points, to avoid redundancy and ensure the independence of isolates. Isolates that could not be successfully subcultured, failed to yield adequate growth on SDA, or lacked sufficient clinical and microbiological data for classification, or those with inadequate DNA yield after extraction were also excluded.
Table (1):
Distribution of 100 Candida isolates according to the health center, specimen source, and patient gender
| Health Center | Specimen Source | Gender | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Urine | High Vaginal Swab | Wound | TRAP | Sputum | Fluid | Blood | Nail scraping | Total | Male | Female | |
| Palestine Medical Complex | 20 (40%) | 18 (36 %) |
7 (14%) | 0 (0%) | 1 (2%) | 1 (2%) | 2 (4%) | 1 (2%) | 50 (50%) | 17 (34%) | 33 (66%) |
| An Najah National University | 14 (38.9%) | 1 (2.8%) | 5 (13.9%) | 7 (19.4%) | 7 (19.4%) | 2 (5.6%) | 0 (0%) | 0 (0%) | 36 (36%) | 22 (61.1%) | 14 (38.9%) |
| Rafidia Hospital | 2 (40%) | 0 (0%) | 1 (20%) | 2 (40%) | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | 5 (5%) | 5 (100%) | 0 (0%) |
| AL-Watani Hospital | 9 (100%) | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | 9 (9%) | 4 (44.4%) | 5 (55.6%) |
| Total | 45 (45%) | 19 (19%) | 13 (13%) | 9 (9%) | 8 (8%) | 3 (3%) | 2 (2%) | 1 (1%) | 100 (100%) | 48 (48%) | 52 (52%) |
Genomic DNA extraction of Candida spp.
The collected isolates were subcultured on SDA, and genomic DNA was extracted from the clinical isolates following a previously described method,26 with minor modifications. A few Candida colonies were removed from an overnight SDA agar plate, mixed with 120 µl of Tris-EDTA (10 mM Tris-HCl, 1 mM EDTA [pH 8]) buffer. Boil water was used for the mixture for 10 min, then immediately kept on ice for 3 min. After that, it was heated in a household microwave oven at high power for 2 min (Universal P70B17L-K4, 230 V, 50 Hz, 700 W; Universal Appliances Ltd., Israel), then incubated in ice again for 3 min. The centrifugation step was performed at 14,000 × g for 5 min using a MiniSpin plus centrifuge (Eppendorf, Hamburg, Germany). The DNA in a collected supernatant was extracted with 40 µl of chloroform and centrifuged at 14,000 × g for 5 min.
The extracted DNA was evaluated for its quantity, quality and purity through A260/A280 ratio assessment and using a micro-volume nanodrop spectrophotometer (Genova Nano; Jenway, UK), and the genomic DNA was stored at -20 °C for subsequent molecular analyses.
Identification of Candida spp. using the PCR technique
Identification of Candida spp. was performed using multiplex polymerase chain reaction (PCR) as previously described,27 with certain modifications. This method amplifies two regions of Candida DNA, including the ITS1-5.8S-ITS2 region, using a combination of two yeast-specific universal primers (UNI1 and UNI2) and six species-specific primers (Invitrogen, Thermo Fisher Scientific, Carlsbad, CA, USA) in a single PCR reaction for each of the following species: C. albicans, C. krusei, C. tropicalis, C. parapsilosis, C. dubliniensis, C. glabrata, and C. lusitaniae (Table 2). Each PCR reaction (25 µl total) contained 12.5 µl DreamTaq™ PCR Master Mix (2X; Thermo Scientific), 0.3 µM of each primer, 3 µl (60-80 ng) of DNA template, and nuclease-free water to reach the final volume. PCR was conducted in a thermal cycler (Mastercycler personal, Eppendorf, Germany) under the following conditions: initial denaturation at 95 °C for 3 min, followed by 30 cycles of 94 °C for 1 min (denaturation), 50 °C for 30 sec (annealing), and 72 °C for 1 min (extension), with a final extension at 72 °C for 5 min. PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100-bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from
C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template.
Table (2):
The sequence of PCR primers used in this study to identify Candida species, and the specific fragment size for each species
| Candida spp. | Primer sequence 5→3 | (bp) length | Ref. |
|---|---|---|---|
| – | UNI1 5′-GTCAAACTTGGTCATTTA-3′ UNI2 5′-TTCTTTTCCTCCGCTTATTG-3′ |
– | 27 |
| C. albicans | Calb 5′-AGCTGCCGCCAGAGGTCTAA-3′ | 446 | |
| C. krusei | Ckru 5′-CTGGCCGAGCGAACTAGACT-3′ | 169 | |
| C. tropicalis | Ctro 5′-GATTTGCTTAATTGCCCCAC-3′ | 507 | |
| C. parapsilosis | Cpar 5′-GTCAACCGATTATTTAATAG-3′ | 370 | |
| C. dubliniensis | Cdub 5′-CTCAAACCCCTAGGGTTTGG-3′ | 217 | |
| C. glabrata | Cgla 5′-TTGTCTGAGCTCGGAGAGAG-3′ | 839 | |
| C. lusitaniae | Clus 5′-TTCGGAGCAACGCCTAACCG-3′ | 329 |
ABC genotyping
ABC genotyping was carried out for all Candida isolates depending on the 25S rDNA sequence using CA-INT-F and CA-INT-R primers (Invitrogen-Thermo Fisher Scientific, Carlsbad, CA, USA) (Table 3). The mixture of PCR consisted of 25 μl total volume including; 12.5 µl DreamTaq™ PCR Master Mix (2X; Thermo Scientific), 0.3 µM of each primer, 3 µl (60-80 ng) of DNA template, and nuclease-free water to reach the final volume. The PCR conditions were as follows: incubation for 3 min at 94 °C before 30 cycles; 94 °C for 1 min, 65 °C for 1 min, 72 °C for 2.5 min, and a final step of 5 min at 72 °C.19 PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100 bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template.
Table (3):
The sequence of primers used in the current study to identify Candida genotype, mating type, biofilm formation, and Phospholipases genes, the length of PCR product in base pairs (bp), annealing temperature, and pool
| Identification | Target gene | Primer sequence 5′→3′ | Fragment length in bp | Primer mix | Annealing temp. | Ref. |
|---|---|---|---|---|---|---|
| ABC genotyping | 25S rDNA | CA–INT–F ATA AGG GAA GTC GGC AAA ATA GAT CCG TAA CA–INT–R 5′CCT TGG CTG TGG TTT CGC TAG ATA GTA GAT |
A (450) | I | 65 °C | 19 |
| B (840) | ||||||
| C (450 and 840) | ||||||
| Mating type | MTLa1 | MTLa1-F TAA GAA TGA AGA CAA CGA GG MTLa1-R CGT GTT TTT CTG CTA TCA ATT CC |
535 | II | 57 °C | 28 |
| MTLα1 | MTLα1-F TAC ATT CTG GTC GCG ATG CTC MTLα1-R GTA ATC CAA AGC CTC GCA TAA |
423 | ||||
| Biofilm-related genes | HWP1 | HWP1-F ATGACTCCAGCTGGTTC HWP1-R ATGACTCCAGCTGGTTC |
503 | III | 55 °C | 29 |
| ALS1
|
ALS1-F ACCAGAAGAAACAGCAGGTG ALS1-R GACTAGTGAACCAACAAATACCAG |
319 | ||||
| ALS3 | ALS3-F CCAAGTGTTCCAACAACTGAA ALS3-R GAACCGGTTGTTGCTATGGT |
185 | ||||
| Phospholipase genes | PLB1 | PLB1-F CCT ATT GCC AAA CAA GCA TTG TC PLB1-R CCA AGC TAC TGA TTT CAC CTG CTC C |
179 | III | 53 ºC | 30 |
| PLB2 | PLB2-F GTG GGA TCT TGC AGA GTT CAA GC PLB2-R CTC AAA GCT CTC CCA TAG ACA TCT G |
270 |
bp: base pair
Mating type
The detection of the mating type locus a1 and α1 for each isolate was performed by using the mating type locus a1 (MTLa1) and α1 (MTLα1) primers (Invitrogen-Thermo Fisher Scientific, Carlsbad, CA, USA) (Table 3). The PCR mixture was prepared as 12.5 µl DreamTaq™ PCR Master Mix (2X; Thermo Scientific), 0.3 µM of each primer, 3 µl (60-80 ng) of DNA template, and nuclease-free water to reach the final volume (25 μl total volume). The PCR amplification was performed under the following conditions: 94 °C for 3 min, 30 cycles; 1 min at 94 °C, 45 sec at 57 °C, 1 min at 72 °C, and a final step at 72 °C for 5 min.28 PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100-bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template.
Biofilm-related genes detection
The previously extracted DNA was used for amplification of biofilm-related genes using Hyphal wall protein 1 (HWP1), Agglutinin-like sequence 1 (ALS1), and Agglutinin-like sequence 3 (ALS3) primers (Invitrogen-Thermo Fisher Scientific, Carlsbad, CA, USA) (Table 3). The PCR mixture contained of 12.5 µl DreamTaq™ PCR Master Mix (2X; Thermo Scientific), 0.3 µM of each primer, 3 µl (60-80 ng) of DNA template, and nuclease-free water to reach the final volume (25 μl total volume). The amplification process was carried out through the following PCR conditions: 94 °C for 5 min, 32 cycles; 94 °C for 1 min, 55 °C for 1 min, 72 °C for 1 min, and a final step of 10 min at 72 °C.29 PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100-bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template.
Detection of Candida Phospholipases PLB1 and PLB2 genes
The extracted DNA was used for amplification of Phospholipases genes using PLB1 and PLB2 primers (Invitrogen- Thermo Fisher Scientific, Carlsbad, CA, USA) (Table 3). The PCR mixture for the detection of PLB1 and PLB2 genes. The PCR mixture contained 12.5 µl DreamTaq™ PCR Master Mix (2X; Thermo Scientific), 0.3 µM of each primer, 3 µl (60-80 ng) of DNA template, and nuclease-free water to reach the final volume (25 μl total volume). The PCR conditions were including; 94 °C for 3 min; 30 cycles; 94 °C for 30 sec, 53 °C for 30 sec, 72 °C for 30 sec, and a final step of 72 °C for 10 min.30 PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100 bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template.
Detection of Secreted Aspartyl Proteinases
Secreted Aspartyl Proteinases SAP1, 3, 4, and 7 genes
The amplification of SAP1, 3, 4, and 7 genes was performed using SAP1, SAP3, SAP4, and SAP7 primers (Invitrogen- Thermo Fisher Scientific, Carlsbad, CA, USA) (Table 4). The PCR mixture was prepared with 25 µl total, containing 12.5 µl DreamTaq™ PCR Master Mix (2X; Thermo Scientific), 0.3 µM of each primer, 3 µl (60-80 ng) of DNA template, and nuclease-free water to reach the final volume and amplification was performed under the following conditions: initial denaturation at 94 °C for 3 min, 30 cycles of 94 °C for 30 sec, 46 °C for 30 sec, and 72 °C for 30 sec, followed by a final extension at 72 °C for 10 min.30 PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100 bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template.
Table (4):
The sequence of primers used in the current study to identify Candida Secreted Aspartyl Proteinases SAP1, SAP3, SAP4, and SAP7 genes, and the length of PCR product in base pairs (bp), and annealing temperature
| Identification | Target gene | Primer sequence 5′→3′ | Fragment length in bp | Annealing temperature | Ref. |
|---|---|---|---|---|---|
| Aspartyl Proteinase genes | SAP1 | SAP1-F TCA ATC AAT TTA CTC TTC CAT TTC TAA CA SAP1-R CCA GTA GCA TTA ACA GGA GTT TTA ATG ACA |
161 | 46 ºC | 30 |
| SAP3 | SAP3-F CCT TCT CTA AAA TTA TGG ATT GGA AC SAP3-R TTG ATT TCA CCT TGG GGA CCA GTA ACA TTT |
231 | 46 ºC | ||
| SAP4 | SAP4-F TTA TTT TTA GAT ATT GAG CCC ACA GAA A SAP4-R GCC AGT GTC AAC AAT AAC GCT AAG TT |
171 | 46 ºC | ||
| SAP7 | SAP7-F GAA ATG CAA AGA GTA TTA GAG TTA TTA C SAP7-R GAA TGA TTT GGT TTA CAT CAT CTT CAA CTG |
196 | 46 ºC |
bp: base pair
Secreted Aspartyl Proteinases SAP2, 5, 6, and 8 genes
The extracted DNA was used to amplify the SAP2, SAP5, SAP6, and SAP8 genes with their corresponding primers (Invitrogen, Thermo Fisher Scientific, Carlsbad, CA, USA) (Table 5). PCR mixtures were prepared as 25 µl total volume of 12.5 µl DreamTaq™ PCR Master Mix (2X; Thermo Scientific), 0.3 µM of each primer, 3 µl (60-80 ng) of DNA template, and nuclease-free water to reach the final volume. Amplification was performed under the following conditions: initial denaturation at 94 °C for 3 min, followed by 30 cycles of 94 °C for 30 sec, 52 °C for 30 sec, and 72 °C for 30 sec, with a final extension at 72 °C for 10 min.30 PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100 bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template.
Table (5):
The sequence of primers used in the current study to identify Candida Secreted Aspartyl Proteinases SAP2, SAP5, SAP6, and SAP8 genes, and the length of PCR product in base pairs (bp), and annealing temperature
| Identification | Target gene | Primer sequence 5′→3′ | Fragment length in bp | Annealing temperature | Ref. |
|---|---|---|---|---|---|
| Aspartyl Proteinases genes | SAP2 | SAP2-F AAC AAC AAC CCA CTA GAC ATC ACC SAP2-R TGA CCA TTA GTA ACT GGG AAT GCT TTA GGA |
178 | 52 ºC | 30 |
| SAP5 | SAP5-F AGA ATT TCC CGT CGA TGA GAC TGG T SAP5-R CAA ATT TTG GGA AGT GCG GGA AGA |
277 | 52 ºC | ||
| SAP6 | SAP6-F CCC GTT TTG AAA TTA AAT ATG CTG ATG G SAP6-R GTC GTA AGG AGT TCT GGT AGC TTC G |
187 | 52 ºC | ||
| SAP8 | SAP8-F GCC GTT GGT GCC AAA TGG AAT AGT TA SAP8-R ATT TGA CTT GAG CCA ACA GAA TGG T |
256 | 52 ºC |
bp: base pair
Secreted Aspartyl Proteinases SAP9 and SAP10 genes
The extracted DNA was used to amplify the SAP9 and SAP10 genes using their respective primers (Invitrogen, Thermo Fisher Scientific, Carlsbad, CA, USA) (Table 6). PCR mixtures were prepared as 12.5 µl DreamTaq™ PCR Master Mix (2X; Thermo Scientific), 0.3 µM of each primer, 3 µl (60-80 ng) of DNA template, and nuclease-free water to reach the final volume (25 μl total volume). Amplification was carried out under the following thermal conditions: initial denaturation at 95 °C for 5 min, followed by 40 cycles of 94 °C for 4 sec, 59 °C for 10 sec, and 72 °C for 30 sec, with a final extension at 72 °C for 3 min.26 PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100 bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template.
Table (6):
The sequence of primers used in the current study to identify Candida Secreted Aspartyl Proteinases SAP9 and SAP10 genes, the length of the PCR product in base pairs (bp), and the annealing temperature
| Identification | Target gene | Primer sequence 5′→3′ | Fragment length in bp | Primer mix | Annealing temperature | Ref. |
|---|---|---|---|---|---|---|
| Aspartyl Proteinases genes | SAP9 | SAP9-F ATT TAC TCC ACA GTT TAT ATC ACT GAA GGT SAP9-R CCA CCA GAA CCA CCC TCA GTT |
80 | III | 59 °C | 26 |
| SAP10 | SAP10-F CCC GGT ATC CAA TAG AAT CGA A SAP10-R TCA GTG AAT GTG ACG AAT TTG AAG A |
80 |
bp: base pair
Randomly Amplified Polymorphic DNA (RAPD)-PCR typing
RAPD-PCR was carried out using the OPI_06 primer sequence 5′-AAGGCGGCAG-3′ (Invitrogen-Thermo Fisher Scientific, Carlsbad, CA, USA) as previously described,31 with minor modifications. The PCR mixture was conducted in a final volume of 25 µl containing 12.5 µl of PCR premix (DreamTaqTM PCR Master Mix (2X), Thermo Scientific), 0.8 µM of the primer, and 3 µl (60-80 ng) of C. albicans DNA template. Additionally, the master mix was modified by adjusting the concentrations of dNTPs to 0.4 mM, MgCl‚ to 3 mM, and Taq DNA polymerase to 1.5 U per reaction. The amplification process was carried out using the following thermal conditions: 94 °C for 3 min, followed by 40 cycles of 1 min at 94 °C, 1 min at 36 °C, and 2 min at 72 °C, with a final step at 72 °C for 5 min. PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel in 1X TAE buffer at 100 V for 50 min. Gels were stained with 0.5 µg/ml ethidium bromide, and fragment sizes were determined using a 100 bp ladder (GeneDireX). Positive control (PC) reactions contained genomic DNA from C. albicans ATCC 90028, while negative controls (NC) included all reagents except the DNA template. DNA banding patterns from gel images were scored using the binary method, recording the presence and absence of bands as 1 and 0, respectively. The resulting binary matrix was analyzed using the Ward linkage method in SPSS Statistics version 27 (IBM).
Statistical analysis
All statistical analyses were performed using SPSS version 27. The chi-square test was applied to conduct a correlation between patient gender and specimen source in relation to Candida species. A P-value of <0.05 was considered statistically significant.
Candida spp. identification
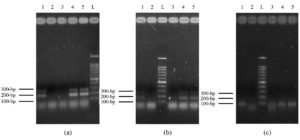
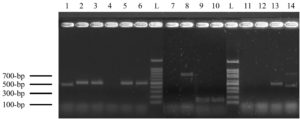
Most isolates showed a band size of 446-bp for C. albicans (n = 53, 53%), followed by C. tropicalis of 507 bp (n = 15, 15%), C. krusei of 169 bp (n = 13, 13%), and C. glabrata of 839 bp (n = 5, 5%). Some specimens have two species, including C. krusei and C. glabrata (n = 3, 3%), C. albicans and C. glabrata (n = 1, 1%), and the remaining 10 isolates (10%) have not shown any bands (Figure 1). There was no statistical association between patients’ gender and specimen source among Candida spp. (χ² (88) = 79.072, P = 0.741).
Figure 1. The polymerase chain reaction (PCR) products of Candida species-specific primers and yeast-specific universal primers for ITS sequence. Lane L: 100-bp ladder, lane-1 represents a band of 446-bp for C. albicans. Lane 2, 3, 5, 6, and 13 bands of 507 for C. tropicalis, lane 8 band of C. glabrata (839-bp), lane 9 and 10 represent C. krusei (band of 169-bp), and lane 14 represents two bands of 446 and 839-bp for C. albicans and C. glabrata
ABC genotyping
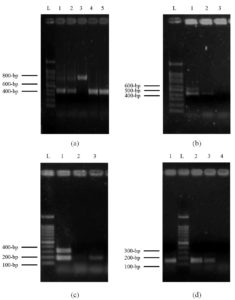
The isolates of C. albicans were categorized into three genotypes; the most frequent were genotype A (32%), which was observed by gel electrophoresis as a 450-bp band size, followed by genotype B of 840 bp (30%), and 13% of the isolates showed dual bands of 450 and 840-bp, which represented genotype C. However, the remaining 25% of the isolates didn’t show any bands with respect to genotyping (Figure 2a).
Mating typing
The mating capability was found among C. albicans isolates with 28% of MTLα homozygosity (423-bp) and 8% as MTL heterozygosity of two band sizes of 535-bp of MTLα and 423-bp of MTLα. In contrast, the remaining (64%) didn’t show any bands. However, 7% of C. tropicalis isolates showed MTLα homozygosity, 20% were MTL heterozygosity, and the remaining (73%) didn’t show any bands (Figure 2b).
Biofilm formation
It was found that 15% of C. albicans isolates have the ALS3 gene (185-bp), 72% showed two bands of 185-bp and 319-bp for ALS3 and ALS1, and 13% showed neither ALS1 nor ALS3. For C. tropicalis, 27% of the isolates have ALS3, 40% have both ALS1 and ALS3, and 33% didn’t show any bands. Whereas, for C. krusei, 23% of the isolates have ALS3, 54% have both ALS1 and ALS3, and 23% have no bands. Finally, for C. glabrata, 20% of the isolates have ALS3, 20% have ALS1 and ALS3, and 60% showed neither ALS1 nor ALS3 genes (Figure 2c). Whereas no isolates were found with the HWP1 gene.
Candida phospholipases
It was found that 64% of C. albicans isolates have a 179-bp band size of the PLB1 gene, and the remaining 36% didn’t show any bands; however, among NAC species, the PLB1 gene was found in 62% of C. krusei, 40% of C. glabrata, and 27% of C. tropicalis isolates. Whereas, PLB2 gene was not detected in all isolates (Figure 2d and Table 7). The Phospholipases and Secreted Aspartic Proteinases patterns among C. albicans and NAC species are shown in Table 7.
Table (7):
The patterns of Phospholipase genes and Secreted Aspartic Proteinases genes among C. albicans and NAC species
Pattern |
C. albicans (n) |
NAC |
(n) |
|---|---|---|---|
PLB1, SAP5, SAP6, SAP3, SAP7, SAP9, SAP10 |
1 |
C. tropicalis |
1 |
PLB1, SAP1, SAP3, SAP9, SAP10 |
1 |
Not found |
0 |
PLB1, SAP2, SAP1 |
0 |
C. krusei |
1 |
PLB1, SAP2, SAP1, SAP9, SAP10 |
0 |
C. krusei |
1 |
PLB1, SAP2, SAP3, SAP7, SAP9, SAP10 |
1 |
Not found |
0 |
PLB1, SAP2, SAP8, SAP1, SAP3, SAP9, SAP10 |
2 |
C. tropicalis |
1 |
PLB1, SAP2, SAP8, SAP3, SAP4, SAP9, SAP10 |
2 |
Not found |
0 |
PLB1, SAP2, SAP8, SAP3, SAP7 |
1 |
Not found |
0 |
PLB1, SAP2, SAP8, SAP3, SAP7, SAP9, SAP10 |
1 |
Not found |
0 |
PLB1, SAP3, SAP7 |
1 |
Not found |
0 |
PLB1, SAP4 |
0 |
C. krusei |
1 |
PLB1, SAP5, SAP6 |
2 |
C. krusei |
1 |
PLB1, SAP5, SAP6, SAP3, SAP7, SAP9, SAP10 |
1 |
Not found |
0 |
PLB1, SAP5, SAP6, SAP1, SAP3 |
1 |
Not found |
0 |
PLB1, SAP5, SAP6, SAP1, SAP3, SAP9, SAP10 |
1 |
Not found |
0 |
PLB1, SAP5, SAP6, SAP3, SAP4 |
1 |
Not found |
0 |
PLB1, SAP5, SAP6, SAP3, SAP4, SAP9, SAP10 |
3 |
Not found |
0 |
PLB1, SAP5, SAP6, SAP3, SAP7 |
1 |
Not found |
0 |
PLB1, SAP5, SAP6, SAP9, SAP10 |
1 |
Not found |
0 |
PLB1, SAP6 |
1 |
Not found |
0 |
PLB1, SAP6, SAP1, SAP9, SAP10 |
0 |
C. krusei |
2 |
PLB1, SAP6, SAP4, SAP9, SAP10 |
0 |
C. krusei |
1 |
PLB1, SAP6, SAP7 |
0 |
C. krusei |
1 |
PLB1, SAP6, SAP9, SAP10 |
1 |
C. krusei |
1 |
PLB1, SAP6, SAP9, SAP10, SAP4 |
0 |
C. glabrata |
1 |
PLB1, SAP9, SAP10 |
1 |
C. krusei, C. glabrata, C. tropicalis |
1, 1, 2 |
SAP3, SAP7, SAP9, SAP10 |
0 |
C. glabrata |
1 |
SAP5, SAP6 |
1 |
Not found |
0 |
SAP5, SAP6, SAP9, SAP10 |
0 |
C. krusei |
1 |
SAP9 or/and SAP10 |
13 |
C. krusei, C. glabrata, C. tropicalis |
3, 1, 5 |
Total |
30 (57%) |
27 (81%) |
NAC, non-albicans Candida
Figure 2. Multiplex PCR profile of Candida genotypes, Mating types, Biofilm formation genes and Phospholipases genes. Lane L: 100-bp ladder. (a) represent the Candida genotyping. Lanes 1 and 2 represent genotype C (two bands of 450 and 840-bp), lane 3 represents genotype B (840-bp), and lanes 4 and 5 represent genotype A (450-bp) of C. albicans isolates. (b) Mating typing. Lane 1 represents heterozygous MTL of two bands of MTLα and MTLα (535 and 423-bp), lane 2 represents homozygous MTLα as one band of 423-bp, and lane 3 shows no band for isolates. (c) Biofilm formation. Lane 1 represents two bands of ALS3 and ALS1 genes (185 and 319-bp), lane 2 shows no bands, and lane 3 shows one band of ALS3 (185-bp). (d) Phospholipases genes. Lane L: 100-bp ladder, lanes 1, 2, and 3 represent bands of the PLB1 gene (179-bp), whereas lane 4 shows no bands
Secreted aspartic proteinases
The findings of this study revealed that the frequency of SAP1, SAP3, SAP4 and SAP7 genes was as follows: C. albicans (n = 4/53; 7.5%, 26/53; 49%, n = 6/53; 11.3% and n = 15/53; 28.3%), C. tropicalis (n = 1/15; 6.7%, 2/15; 13.3%, n = 0/15; 0% and n = 1/15; 6.7% ), C. krusei (n = 2/13; 15.4%, n = 0/13; 0%, n = 2/13; 15.4% and n = 1/13; 7.7%), C. glabrata (n = 0/5;0%, n = 1/5; 20%, n = 1/5; 20% and n = 1/5; 20% ), respectively. Whereas, the frequency of SAP2, SAP5, SAP6 and SAP8 genes was as follows: C. albicans (n = 10/53; 18.9%, 18/53; 34%, n = 20/53; 37.7% and n = 8/53; 15.1%), C. tropicalis (n = 1/15; 6.7%, 1/15; 6.7%, n = 1/15; 6.7% and n = 1/15; 6.7% ), C. krusei (n = 1/13;7.7%, n = 2/13;15.4%, n = 6/13; 46% and n = 0/13; 0% ), C. glabrata (n = 0/5;0%, n = 0/5; 0%, n = 1/5; 20% and n = 0/5; 0% ), respectively. However, it was revealed that the most frequent genes were SAP9 or/and SAP10, which were found among Candida species including: C. albicans (n = 35/53; 66%), C. tropicalis (n = 9/15; 60%), C. krusei (n = 8/13; 61.5%), C. glabrata (n = 4/5; 80% ), respectively. According to virulence factor patterns, the frequency of SAP9 or/and SAP10 among isolates that were incorporated with other virulence factors like PLB1, SAP5, SAP6, SAP3, SAP7, SAP9 or/and SAP10 was as follows: 25% of C. albicans, 33% of C. tropicalis, 23% of C. krusei, and 20% of C. glabrata. (Figure 3 and Table 7).
Figure 3. Multiplex PCR profile of Candida species Secreted Aspartic Proteinases genes. Lane L: 100-bp ladder. (a) Lane 1 represents SAP2 and SAP8 (bands of 178 and 256-bp), lanes 4 and 5 represent the bands of SAP6 and SAP5 (187 and 277-bp). However, lanes 2 and 3 don’t show any bands. (b) Lane 1 represents SAP7 and SAP3 (196 and 231-bp), lanes 4 and 5 represent SAP4 and SAP3 (171 and 231-bp), and lanes 2 and 3 do not show any bands. (c) Lanes 1,4, and 5 represent bands of SAP9 or SAP10 (90-bp), whereas lanes 2 and 3 show no bands
Hierarchical Clustering of C. albicans using Ward linkage according to virulence genes
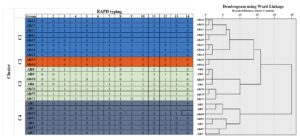
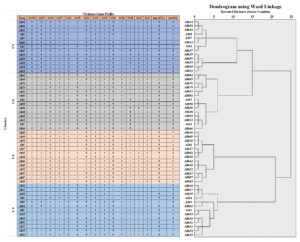
The dendrogram obtained by Ward’s linkage for the 53 C. albicans isolates recovered from Palestine concerning the virulence genes (including MTLa1, MTLα1, ALS1, ALS3, PLB1, PLB2, and SAPs), which were clustered into four groups: C1, C2, C3, and C4 at a cut-off similarity of 50%. However, isolates in clusters C1, C2, C3, and C4 were grouped at a cut-off similarity of about 90%. Cluster C1, C2, and C4 could be subdivided into four sub-clusters (Figure 4).
Figure 4. Dendrogram showed the clusters of virulence factors of 53 isolates of C. albicans recovered from patients with candidiasis in the Northern and Central Palestine, based on the Ward’s linkage method analysis
RAPD-PCR typing
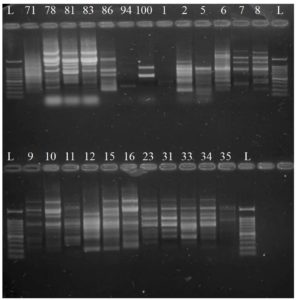
Regarding the RAPD-PCR results, the primer OPI 06 produced 14 fragments. The length of the fragments ranges from 200 to 2000-bp. All of the bands are polymorphic, with a 100% percentage. Data are illustrated in Figure 5.
Figure 5. RAPD-PCR product profiles using the primer OPI_06. Lane L = ladder 100-bp. The 24 labeled Lanes are for the RAPD-PCR product of C. albicans DNA of the recovered isolates in Palestine
The dendrogram obtained by Ward’s linkage for the RAPD-PCR product of 24 C. albicans isolates recovered from Palestine were clustered into four groups (C1, C2, C3, and C4), depending on a 50% cut-off similarity. Therefore, isolates in cluster C1 were grouped at a cut-off similarity of 80%, whereas in C3 and C4 clusters, isolates were grouped at a cut-off similarity of 90%. Cluster C3 and C4 could be subdivided into three and five sub-clusters, respectively. Data are shown in Figure 6.
In the present study, Candida species were investigated through molecular characterization, and the virulence attributes from clinical specimens in Palestine. C. albicans was the most prevalent (53%), then C. tropicalis (16%). These results are consistent with a study on patients with chronic kidney disease, which reported C. albicans (51.1%), C. glabrata (25%), and C. tropicalis (14.8%),32 as well as the findings of Zarrinfar et al.,2 who detected C. albicans (52%), C. tropicalis (24%), C. glabrata (14.7%), C. krusei (5.3%), C. parapsilosis (1.3%), C. kefyr (1.3%), and C. guilliermondii (1.3%) in bronchoalveolar lavage specimens from patients with pulmonary disorders. These observations align with previous studies highlighting the medical relevance of non-albicans Candida species, alongside the severity of infections caused by C. albicans.33 The pathogenic capacity of C. albicans is attributed to numerous virulence factors, including strong adhesion ability, dimorphic transition, and the production of hydrolytic enzymes that facilitate tissue invasion.34 Co-infection is one of the causes that explains the mixed-species infections. Furthermore, it was suggested that the absence of bands for some isolates was caused by the presence of mutations in the gene sequence or other Candida spp.33
The findings of the three genotypes (A, B, and C) are consistent with a previous study, which reported genotype A as the most prevalent (52.3%), followed by genotype B (31.8%) and genotype C (15.9%),35 verifying the evolutionary and pathogenic capabilities of C. albicans with HWP (80.0%), ALS (56.0%), and SAP genes (56.0%).33 Moreover, epidemiological tracking and species-to-species diversity were highly determined through genotyping studies. Whereas, findings of C. albicans without any genotype-specific bands may be due to gene mutations or non-typeable. The huge number of C. albicans strains for which the nucleotide sequence of the whole genome is available in GenBank may be useful to improve the genotype assignment method. The current approach was never designed to detect genotypes other than A, B, and C; therefore, it appears suitable to modify the method to overcome these obstacles.
Genotype A emerged as the predominant C. albicans type among clinical isolates in Palestine, a pattern first brought to light by Ali-Shtayeh et al., is not merely a descriptive curiosity but a finding that takes on considerably greater weight when read against what is now understood about the biological behavior of this genotype. The classification itself rests on the presence or absence of a self-splicing transposable group I intron within the 25S rDNA gene, genotype A is characterized by the absence of this intron, yielding a PCR amplicon of approximately 450 bp, whereas genotypes B and C carry intron sequences of differing lengths producing amplicons of 840 bp and both 450 and 840 bp respectively; a structural distinction that has been consistently associated with differences in clinical behavior and invasive potential across diverse geographic settings.36,37
A study examining clinical blood and vaginal C. albicans isolates confirmed that genotype A was the predominant genotype among bloodstream isolates, accounting for 60% of invasive blood-derived specimens, further reinforcing its established association with deeper and more systemic forms of infection.36 This finding has not remained geographically isolated. Data from southwest China, for instance, showed genotype A accounting for 86.7% of isolates from vulvovaginal candidiasis patients, with genotypes B and C appearing more evenly distributed across other clinical sources, a distribution that mirrors, in broad outline, what Ali-Shtayeh et al.19 and the observations in the Palestinian cohort.38 At the mechanistic level, the greater pathogenicity attributed to genotype A isolates has been linked to their more pronounced expression of proteinase and phospholipase activity, enzymatic properties that equip the organism to adhere to and degrade host tissue proteins with unusual efficiency.19 The resistance profile of this genotype adds another layer of clinical concern; its documented association with reduced susceptibility to fluconazole and flucytosine is particularly troubling in the Palestinian setting, where fluconazole continues to function as the antifungal agent of first resort across most clinical contexts.19,25
Compounding these virulence characteristics is C. albicans’ well-characterized ability to transition between yeast and hyphal morphologies, a plasticity that drives tissue invasion, undermines immune clearance, and creates pathways through which superficial mucosal infections may escalate toward deep-seated systemic disease.39 What the evidence collectively suggests, then, is that the clinical burden of candidiasis in Palestine may be skewed toward more severe and treatment-resistant disease than aggregate incidence figures alone would indicate, a possibility that has yet to be rigorously tested in prospective clinical work and that represents an important priority for future investigation.
Across studies conducted in Iran, Turkey, Burkina Faso, and China, genotype A has consistently emerged as the dominant clinical C. albicans genotype, a pattern stable enough across specimen types and settings to suggest that this lineage has established itself as the evolutionarily favored form in most human infection contexts.37,40 Yet relative proportions of genotypes B and C, and the specific resistance associations each carries, shift considerably from one setting to another in ways that resist straightforward biological explanation, implying that local ecological, therapeutic, and host-related pressures are reshaping genotype distributions in ways that global patterns cannot fully anticipate.38
Placing the Palestinian data within an immediately regional frame, that of neighboring Arab countries and the broader Middle East, is an exercise that the literature renders difficult, because virulence gene profiles from these populations remain incompletely reported, leaving a comparative baseline that is far thinner than the clinical situation warrants. What the molecular literature does establish with reasonable confidence is that virulence gene profiling carries genuine predictive value for antifungal susceptibility. The detection of HWP1 genes through multiplex PCR, for instance, has been associated with significantly elevated rates of resistance to flucytosine, while the carriage of SAP genes has been tied to enhanced biofilm organization and diminished susceptibility across multiple antifungal classes.22
These associations are not merely academic; they point toward a model of antifungal stewardship in which prescribing decisions are informed by the molecular profile of the infecting isolate rather than derived from blanket empirical protocols, an approach that holds particular promise in settings like Palestine, where surveillance infrastructure is still developing and antifungal resources must be deployed judiciously,25,41 against this backdrop, several limitations of the present study deserve Candida acknowledgement. The geographic scope of specimen collection, confined to a limited number of sites within the West Bank, restricts the extent to which the findings can be generalized to the full diversity of Candida strains circulating across the Palestinian population. The cross-sectional design, by its nature, captures only a moment in time and cannot speak to how genotype distributions or resistance profiles may be shifting over months or years. And while the molecular methods employed here represent a meaningful advance over phenotypic identification alone, the absence of whole-genome sequencing means that resistance-conferring point mutations in genes such as ERG11, FKS1, and FKS2, mutations that may drive phenotypic resistance without leaving any detectable signature in virulence gene assays, fall outside the current analysis can detect.41 These boundaries do not undercut the study’s contribution, but they do define the territory within which its conclusions can reasonably be held.
Among the 53 C. albicans isolates, 28% of the isolates were determined as homozygous MTLα, and 8% were heterozygous at the MTL locus. These results align with a previous study, where the presence of MTL heterozygosity among C. albicans isolates was associated with their mating capability.42 However, several virulence properties, such as biofilm formation, adaptive patterns against antifungal agents, and activation of white-to-opaque switch mating pathways, all of which were linked to homozygous MTLα isolates.43 It was investigated that C. albicans exhibits less mating flexibility compared to C. tropicalis isolates.44 This confirms their potential in sexual reproduction as they participate in antifungal resistance under pressure.45
The biofilm results align with several studies, specifically those involving C. albicans isolates with high biofilm capacity, as the presence of these two genes is strongly associated with host tissue adhesion and biofilm formation. In the study of Roudbarmohammadi et al.,46 forty women with recurrent vulvovaginal candidiasis in Iran have ALS1 (75.8%) and ALS3 (77.7%), 74% of the isolates tested positive for both ALS1 and ALS3 mRNA by RT-PCR. This confirmed their role in the colonization of Candida spp. to epithelial cells of the host and medical devices like catheters in a conserved manner among NAC species, with some structural alterations for C. tropicalis.47 In our study, the results indicated that 15%, 27%, 23% and 20% and 100% of C. albicans, C. tropicalis, C. krusei, and C. glabrata were positive for ALS3, respectively. In contrast, a previous study reported that the ALS3 gene was detected in 94% of C. albicans isolates from patients admitted to intensive care units (ICUs) using PCR assays, as well as in 55% of C. glabrata and 13% of C. tropicalis isolates.48
Although hyphal wall proteins contribute to biofilm maturation in C. albicans, the HWP1 gene has not been found in any of the isolates, which conflicts with a previous study that found the HWP1 (77%) is highly conjugated with other virulence genes, e.g., PLB1 (52%).49 All of these findings may show hyphal morphogenesis of lower prevalence, which may be due to amplification procedure limitations or primer mismatch.
Additionally, other virulence genes were examined among Candida isolates, including phospholipase (PLB) and secreted aspartyl proteinase (SAP) genes. Using the PLB1 primer, 64% of the C. albicans isolates were found to carry the corresponding gene, which contributes significantly to tissue invasion and evasion of the host immunity.32 Consequently, SAP9 or SAP10 were found to be highly prevalent, confirming the importance of these two proteinases in cell wall integrity and systemic candidiasis.33 The study’s findings contradicted the results of a previous study regarding the PLB2 gene, which was detected in all strains alongside PLB1, with a high prevalence of SAP1 (94.8%).34 These combinations of SAP genes have been found to increase the species’ proteolytic activity and tissue invasion. In this study, the results for SAP4 among C. albicans, C. glabrata, and C. krusei were 11.3%, 20% and 15.3%, respectively. Which were counteracted by the previous study results,48 which showed 88%, 0%, and 12% for C. albicans, C. krusei, and C. glabrata, respectively.
The study results, related to RAPD-PCR products, demonstrated that the number of fragments produced by the RAPD-PCR primer OPI 06 was 14. The length of the fragments ranged from approximately 200-2000 bp with a 100% polymorphism. These results were matched with a previous study,31 which showed a polymorphism percentage of 96% using the OPI 06 primer. The explanation behind the resulting DNA polymorphisms involves rearrangements or mutations within the priming sites. In addition, the presence of several specimen body sources or intrusive environments of C. albicans isolates. Therefore, the dendrogram results for C. albicans isolates concerning the virulence genes (including MTLa1, MTLα1, ALS1, ALS3, PLB1, PLB2, and SAPs) showed that isolates in clusters C1, C2, C3, and C4 were grouped at a cut-off similarity of 90%. Whereas, RAPD-PCR results showed that isolates in cluster C1 were grouped at a cut-off similarity of 80%, whereas among the C3 and C4 clusters, isolates were grouped at a cut-off similarity of 90%. These observations imply that isolates of Palestinian C. albicans showed low genetic variation in comparison with the results of a previously published study,50 which showed that C. albicans isolates have higher genetic variation with thirteen various groups with respect to their RAPD pattern.
Several research directions emerge naturally from the findings reported here, and among them, the question of SAP9 and SAP10 gene activity stands out as perhaps the most clinically consequential thread yet to be properly pulled in the Palestinian context. Both genes belong to the secreted aspartyl proteinase (SAP) family, a ten-member repertoire that C. albicans deploys to interact with, degrade, and colonize host tissue, but SAP9 and SAP10 occupy a functionally distinctive position within this family: rather than being secreted into the extracellular environment, the proteins they encode are anchored to the fungal cell surface via glycosylphosphatidylinositol (GPI) linkages. This membrane-tethered configuration means that their roles extend beyond proteolysis to encompass cell wall integrity, epithelial adhesion, and structural biofilm organization functions that position them closer to the interface between the fungus and its host than most other SAP family members. The pathological weight of these genes became most visible in deletion studies, where strains lacking SAP9 and SAP10 showed markedly attenuated virulence in epithelial infection models, with substantially reduced tissue invasion and cellular damage; when gene function was restored, full pathogenicity returned, a clean experimental demonstration that these two genes are not incidental players but active drivers of mucosal disease.51 Their clinical relevance is further sharpened by evidence that SAP9 expression rises in biofilms formed by bloodstream isolates and climbs further still when the organism is exposed to fluconazole and caspofungin, an upregulation pattern that is difficult to interpret as anything other than an adaptive molecular response to antifungal pressure, suggesting that SAP9 activity may track treatment resistance in real time rather than simply reflecting constitutive virulence.52
Supporting this interpretation, isolates concurrently carrying both SAP9 and SAP10 have been disproportionately represented among strong biofilm producers and multidrug-resistant strains, with some series finding both genes present in up to 66.7% of the most resistant clinical isolates.53 Taken together, these observations make a persuasive case for treating SAP9 and SAP10 detection as clinically actionable as markers capable of flagging isolates at high risk of forming treatment-refractory biofilms, causing invasive disease, and failing standard antifungal regimens and for their systematic inclusion in future molecular surveillance protocols across Palestine and the broader Eastern Mediterranean region.
Looking further ahead, the gap most urgently in need of filling is a longitudinal, multi-center molecular surveillance program spanning Palestinian hospitals one designed to generate a nationally representative genomic repository of Candida clinical isolates that could support ongoing tracking of genotype shifts, emerging resistance profiles, and evolving virulence gene distributions over time. Such a resource would supply the comparative regional anchor that the current literature conspicuously lacks and would lay a credible scientific foundation for evidence-based antifungal policy in a setting where that foundation does not yet exist. The path toward meaningfully improved candidiasis outcomes in Palestine runs, in the view of this study, through precisely this kind of integrated molecular surveillance, one that moves beyond species-level phenotyping to encompass genotyping, virulence profiling, and genomic resistance characterization as a unified diagnostic and epidemiological enterprise.
This study provides evidence of the significance of molecular analysis for accurately identifying Candida spp. obtained in Palestine. In addition to determining their genotypic level, mating type, and the virulence of the isolated Candida spp. using PCR. It was shown that the most common isolates were belonged to C. albicans with a high prevalence of genotype A, with the presence of several virulence factors, mostly the Agglutinin-like sequence genes and SAP9 and SPA10. Whereas, the resulting DNA fragments through RAPD-PCR and the clustering of the isolates using Ward’s linkage methods showed their level of polymorphism and genetic variability. These findings emphasized the importance of molecular surveillance for species identification, genotypic and virulence determination, as well as genetic heterogeneity among C. albicans strains. This underscores the need for further investigations to curb the increasing pathogenicity of C. albicans and other Candida species in clinical settings. The knowledge of the Candida species prevalence and their virulence factors highlights the pathogenesis understanding and the need for further antifungal agents to suppress the virulence factors for the pathogenic Candida species.
ACKNOWLEDGMENTS
The authors extend their gratitude to the Microbiology Laboratories of An-Najah National University, Nablus, Palestine, the Palestinian Ministry of Health, and An-Najah National University Hospital for their technical assistance and collaboration during this work.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
Both authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This research was supported by An-Najah National University, Nablus, Palestine, through the Graduate Studies Ph.D. Research Fund.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the Ethics Committee, An-Najah National University, Nablus, Palestine.
INFORMED CONSENT
Written informed consent was obtained from the participants before enrolling in the study.
- Kashefi E, Seyedi SJ, Zarrinfar H, Fata A, Mehrad-Majd H, Najafzadeh MJ. Molecular Identification of Candida Species in Bronchoalveolar Lavage Specimens of Hospitalized Children with Pulmonary Disorders. Journal of Babol University of Medical Sciences. 2021;23(1)
- Zarrinfar H, Kaboli S, Dolatabadi S, Mohammadi R. Rapid detection of Candida species in bronchoalveolar lavage fluid from patients with pulmonary symptoms. Braz J Microbiol. 2016;47(1):172-176.
Crossref - Talapko J, Juzbasic M, Matijevic T, et al. Candida albicans-The Virulence Factors and Clinical Manifestations of Infection. J Fungi. 2021;7(2):79.
Crossref - Frias-De-Leon MG, Hernandez-Castro R, Conde-Cuevas E, et al. Candida glabrata Antifungal Resistance and Virulence Factors, a Perfect Pathogenic Combination. Pharmaceutics. 2021;13(10):1529.
Crossref - Goje O, Munoz JL. Vulvovaginitis: Find the cause to treat it. Cleve Clin J Med. 2017;84(3):215-224.
Crossref - Acosta-Mosquera Y, Tapia JC, Armas-Gonzalez R, Caceres-Valdiviezo MJ, Fernandez-Cadena JC, Andrade-Molina D. Prevalence and Species Distribution of Candida Clinical Isolates in a Tertiary Care Hospital in Ecuador Tested from January 2019 to February 2020. J Fungi. 2024;10(5):304.
Crossref - Macias-Paz IU, Perez-Hernandez S, Tavera-Tapia A, Luna-Arias JP, Guerra-Cardenas JE, Reyna-Beltran E. Candida albicans the main opportunistic pathogenic fungus in humans. Revista Argentina de Microbiologia. 2023;55(2):189-198.
Crossref - Amanloo S, Zanjani M, Serajian S, Ahmadi F, Kakavand F. Genotyping of candida albicans isolates obtained from vulvovaginal candidiasis patients in Zanjan, Iran, based on ABC and RPS typing systems. Curr Med Mycol. 2022;8(4):9-14.
Crossref - Munhoz-Alves N, Mimura LAN, Viero RM, et al. Candida tropicalis Systemic Infection Redirects Leukocyte Infiltration to the Kidneys Attenuating Encephalomyelitis. J Fungi. 2021;7(9):757.
Crossref - Saeed S, Hasan S, Kuldeep, Parmar SS. Conventional and Recent Diagnostic Aids in oral Candidal Infections: A Brief overview. Biomed Pharm J. 2017;10(1):419-426.
- Vila T, Sultan AS, Montelongo-Jauregui D, Jabra-Rizk MA. Oral candidiasis: A disease of opportunity. J Fungi. 2020;6(1):15.
Crossref - Yang L, Cheng T, Shao J. Perspective on receptor-associated immune response to Candida albicans single and mixed infections: Implications for therapeutics in oropharyngeal candidiasis. Med Mycol. 2023;61(8):myad077.
Crossref - Launder D, Dillon JT, Wuescher LM, et al. Immunity to pathogenic mucosal C. albicans infections mediated by oral megakaryocytes activated by IL-17 and candidalysin. Mucosal Immunol. 2024;17(2):182-200.
Crossref - Erfaninejad M, Mahmoudabadi AZ, Maraghi E, Hashemzadeh M, Fatahinia M. Epidemiology, prevalence, and associated factors of oral candidiasis in HIV patients from southwest Iran in post-highly active antiretroviral therapy era. Front Microbiol. 2022;13:983348.
Crossref - Sobel JD. Vulvovaginal candidosis. Lancet (London, England). 2007;369(9577):1961-1971.
Crossref - Montagna MT, Caggiano G, Borghi E, Morace G. The role of the laboratory in the diagnosis of invasive candidiasis. Drugs. 2009;69(Suppl 1):59-63.
Crossref - Rachel R, Anuradha M, Leela KV. Biofilm Formation and Antifungal Susceptibility Profile of Candida Species Responsible for Vulvovaginal Candidiasis in Pregnant and Non-Pregnant Women visiting a Tertiary Care Hospital in Southern India. J Pure Appl Microbiol. 2024;18(1):381-388.
Crossref - Madhavan P, Chong PP, Farida J, et al. Antimicrobial susceptibilities and random amplified polymorphic DNA-PCR fingerprint characterization of Candida glabrata, Candida parapsilosis and Candida rugosa from two major hospitals in Kuala Lumpur, Malaysia. Trop Biomed. 2019;36(1):183-193.
- Ali-Shtayeh M, Jamous R, Alothman N, et al. Genotyping and antifungal susceptibility of Candida albicans strains from patients with vulvovaginal and cutaneous candidiasis in Palestine. Afr J Microbiol Res. 2015;9(13):952-959.
Crossref - Evren E, Gocmen JS, Istar EH, et al. Medically important Candida spp. identification: an era beyond traditional methods. Turk J Med Sci. 2022;52(3):834-840.
Crossref - Czajka KM, Venkataraman K, Brabant-Kirwan D, et al. Molecular mechanisms associated with antifungal resistance in pathogenic Candida species. Cells. 2023;12(22):2655.
Crossref - Dawoud AM, Saied SA, Torayah MM, Ramadan AE, Elaskary SA. Antifungal susceptibility and virulence determinants profile of candida species isolated from patients with candidemia. Sci Rep. 2024;14(1):11597.
Crossref - Garcia-Salazar E, Acosta-Altamirano G, Betancourt-Cisneros P, et al. Detection and molecular identification of eight Candida species in clinical samples by simplex PCR. Microorganisms. 2022;10(2):374.
Crossref - Schikora-Tamarit MA, Gabaldon T. Recent gene selection and drug resistance underscore clinical adaptation across Candida species. Nat Microbiol. 2024;9(1):284-307.
Crossref - Baniodeh H, Abu-Helu R, Abulihya M, et al. The first prevalence and antifungal susceptibility profile of Candida infections in Palestine, 2022. BMC Infect Dis. 2024;24(1):1142.
Crossref - Ahmed N, Rizk DE, El Said M, Barwa RM, Elsokary MA, Hassan RH. Molecular identification and virulence factors determination in Candida species isolated from egyptian patients. Am J Microbiol Res. 2019;7(4):108-117.
Crossref - M El-Ganiny A, E Yossef N, A Kamel H. Prevalence and antifungal drug resistance of nosocomial Candida species isolated from two university hospitals in Egypt. Curr Med Mycol. 2021;7(1):31-37.
Crossref - Jafarian H, Gharaghani M, Asnafi AA, Hardani AK, Zarei-Mahmoudabadi A. Phenotype, genotype, and mating type determination in oral Candida albicans isolates from pediatric patients with neutropenia. J Clin Lab Anal. 2022;36(9):e24664.
Crossref - Mohammadi F, Hemmat N, Bajalan Z, Javadi A. Analysis of Biofilm Related Genes and Antifungal Susceptibility Pattern of Vaginal Candida albicans and Non Candida albicans Species. BioMed Res Int. 2021;2021(1):5598907.
Crossref - Bassyouni RH, Wegdan AA, Abdelmoneim A, Said W, AboElnaga F. Phospholipase and Aspartyl Proteinase Activities of Candida Species Causing Vulvovaginal Candidiasis in Patients with Type 2 Diabetes Mellitus. J Microbiol Biotechnol. 2015;25(10):1734-1741.
Crossref - Al-Tekreeti AR, Al-Halbosiy MM, Dheeb BI, Hashim AJ, Al-Zuhairi AFH, Mohammad FI. Molecular identification of clinical Candida isolates by simple and randomly amplified polymorphic DNA-PCR. Arab J Sci Eng. 2018;43(1):163-170.
Crossref - de la Rosa-Garcia E, Olalde-Hernandez M, Irigoyen-Camacho M, Mondragon-Padilla A, Mendoza-Juache A, Sanchez-Vargas L. Antifungal susceptibility of oral isolates of Candida species from chronic kidney disease patients on chronic dialysis. J Mycol Med. 2020;30(3):101009.
Crossref - Makled AF, Ali SAM, Labeeb AZ, et al. Characterization of Candida species isolated from clinical specimens: insights into virulence traits, antifungal resistance and molecular profiles. BMC Microbiol. 2024;24(1):388.
Crossref - Monroy-Pérez E, Paniagua-Contreras GL, Rodríguez-Purata P, et al. High Virulence and Antifungal Resistance in Clinical Strains of Candida albicans. Can J Infect Dis Med Microbiol. 2016;2016:5930489.
Crossref - Mashaly GE, Zeid MS. Candida albicans Genotyping and Relationship of Virulence Factors with Fluconazole Tolerance in Infected Pediatric Patients. Infect Drug Resist. 2022;15:2035-2043.
Crossref - Mahmoodi M, Nouraei H, Nasr R, Zomorodian K, Khodadadi H, Pakshir K. Phenotypes characterization and ABC genotypes distribution of clinical Candida albicans isolates. J Clin Lab Anal. 2023;37(7):e24888.
Crossref - Gharaghani M, Shabanzadeh M, Jafarian H, Zarei Mahmoudabadi A. ABC typing and extracellular enzyme production of Candida albicans isolated from Candida vulvovaginitis. J Clin Lab Anal. 2022;36(1):e24117.
Crossref - Wang M, Cao Y, Xia M, et al. Virulence and antifungal susceptibility of microsatellite genotypes of Candida albicans from superficial and deep locations. Yeast. 2019;36(5):363-373.
Crossref - Naglik JR, Gaffen SL, Hube B. Candidalysin: discovery and function in Candida albicans infections. Curr Opin Microbiol. 2019;52:100-109.
Crossref - Sawadogo PM, Zida A, Soulama I, et al. Genotype analysis of clinical Candida albicans isolates using PCRs targeting 25S rDNA and ALT repeat sequences of the RPS and antifungal susceptibility in Ouagadougou (Burkina Faso). Infect Drug Resist. 2019;12:3859-3866.
Crossref - Frias-De-Leon MG, Hernandez-Castro R, Vite-Garin T, et al. Antifungal Resistance in Candida auris: Molecular Determinants. Antibiotics. 2020;9(9):568.
Crossref - Mba IE, Nweze EI, Eze EA, Anyaegbunam ZKG. Genome plasticity in Candida albicans: A cutting-edge strategy for evolution, adaptation, and survival. Infect Genet Evol. 2022;99:105256.
Crossref - Xie J, Tao L, Nobile CJ, et al. White-opaque switching in natural MTL a/α isolates of Candida albicans: evolutionary implications for roles in host adaptation, pathogenesis, and sex. PLoS Biol. 2013;11(3):e1001525.
Crossref - Porman AM, Hirakawa MP, Jones SK, Wang N, Bennett RJ. MTL–independent phenotypic switching in Candida tropicalis and a dual role for Wor1 in regulating switching and filamentation. PLoS Genet. 2013;9(3):e1003369.
Crossref - Popp C, Ramírez-Zavala B, Schwanfelder S, Krüger I, Morschhäuser J. Evolution of Fluconazole-Resistant Candida albicans Strains by Drug-Induced Mating Competence and Parasexual Recombination. mBio. 2019;10(1):e02740-18.
Crossref - Roudbarmohammadi S, Roudbary M, Bakhshi B, Katiraee F, Mohammadi R, Falahati M. ALS1 and ALS3 gene expression and biofilm formation in Candida albicans isolated from vulvovaginal candidiasis. Adv Biomed Res. 2016;5(1):105.
Crossref - Oh SH, Isenhower A, Rodriguez-Bobadilla R, et al. Pursuing Advances in DNA Sequencing Technology to Solve a Complex Genomic Jigsaw Puzzle: The Agglutinin-Like Sequence (ALS) Genes of Candida tropicalis. Front Microbiol. 2020;11:594531.
Crossref - Ardehali SH, Azimi T, Fallah F, et al. Molecular detection of ALS1, ALS3, HWP1 and SAP4 genes in Candida Genus isolated from hospitalized patients in Intensive Care Unit, Tehran, Iran. Cell Mol Biol (Noisy-le-grand). 2019;65(4):15-22.
- Shrief R, Zaki MES, El-Sehsah EM, Ghaleb S, Mofreh M. Study of antifungal susceptibility, virulence genes and biofilm formation in Candida albicans. Open Microbiol J. 2019;13(1):241-248.
Crossref - Skora M, Rosam K, Namysl M, et al. Candida albicans Horizontal Transmission in COVID-19 Patients Hospitalized in Intensive Care Unit. J Fungi. 2024;10(12):864.
Crossref - Schild L, Heyken A, de Groot PW, et al. Proteolytic cleavage of covalently linked cell wall proteins by Candida albicans Sap9 and Sap10. Eukaryotic cell. 2011;10(1):98-109.
Crossref - Joo MY, Shin JH, Jang HC, et al. Expression of SAP5 and SAP9 in Candida albicans biofilms: comparison of bloodstream isolates with isolates from other sources. Med Mycol. 2013;51(8):892-6.
Crossref - Kadry AA, El-Ganiny AM, El-Baz AM. Relationship between Sap prevalence and biofilm formation among resistant clinical isolates of Candida albicans. Afr Health Sci. 2018;18(4):1166.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.