ISSN: 0973-7510
E-ISSN: 2581-690X
The Arabian Sea environment harbors numerous microorganisms that have developed unique metabolic abilities to ensure their survival in hostile habitats. The bacterial pigments are considered to be important metabolic product which is useful for bacteria and may exert specific biological properties including antibacterial activity. Moreover, there is an imperative need for exploring new antibacterial agents since; there are drug resistance issues with the existing antibiotics. Therefore, the present study was aimed to assess antibacterial activity of pigments extracted from marine bacteria of Arabian Sea water samples and to characterize the potent antibacterial pigment producing isolate. The water samples were obtained from Tithal (Valsad), Diu, Daman and Dandi beaches of India. Total 9 distinct pigmented bacteria were isolated. The pigments extracted from bacterial isolates were assessed for the antibacterial, antioxidant and anticancer activities. Four pigment producing isolates (NP5, NP6, NP8 & NP9) among the nine isolates showed good antibacterial activity against the different bacterial cultures. Among these four isolates, NP9 showed maximum antibacterial activity against all the test cultures. The pigment obtained from NP5 isolate exhibited higher antioxidant property as compared to NP6, NP8 and NP9. However, none of the pigments obtained from NP5, NP6 and NP9 exhibited anti-cancerous activity on MCF-7 cell line. The molecular identification by16srDNA sequencing revealed that NP9 belongs to Candidatus chryseobacterium massiliae (MK213063). Chryseobacterium has been known to produce yellow pigment; however, for the first time, the study suggests that Candidatus chryseobacterium massiliae strain produces red pigment with potent antibacterial activity. However, its antibacterial activity must be tested with more number of pathogens and study in disease animal model is needed to confirm the results of the study.
Arabian sea, Bacteria pigments, antibacterial, antioxidant and anticancer activities.
The Arabian Sea is an attractive resource for investigations due to its affluent biodiversity. However, investigation for marine microbial metabolites is intricate due to its aloofness and many marine microorganisms cannot be cultured.1 Majority of these marine bacteria are pigmented2 and these pigments are produced for various reasons. For example, Cyanobacteria produce phycobilin pigments to carry out photosynthesis3 and certain bacteria produce pigments which can prevent the growth of other bacteria. The marine bacterial pigments exert significant in vitro and/or in vivo biological activities. The pigment producing bacterial strains include Spirillum rubrum (purple pigment), Chromobacterium Violacein (violet pigment), Staphylococcus aureus (golden pigment), Proteus vulgaris (brown pigment), Pseudomonas aeruginosa (blue green pigment) and Micrococcus roseus (red pigment).
The antimicrobial activities of pigments could be used in development of antimicrobial drugs and may have various industrial, pharmaceutical and medical applications4. Pigmented secondary metabolites have been reported to possess potential clinical applications in treatment of human diseases4. The bacterial pigments demonstrated significant anti-inflammatory, anti-malarial, anti-cancerous, immunosuppressive as well as antimicrobial activities5. The pigment producing bacteria can be isolated and grown on various growth mediums6.
Given the fact that diverse marine microbial communities are important source of novel antimicrobial agents7, the present study was aimed to investigate antibacterial activity of pigments extracted from marine bacteria of Arabian Sea water samples and characterization of potent antibacterial pigment producing isolate. In addition to antibacterial activity, the study also investigated the antioxidant and anti-cancerous activities of the marine bacterial pigments.
Collection of Samples
The Arabian Sea water samples were collected from Tithal (Valsad), Diu, Daman and Dandi beaches located on the coast of Arabian Sea of India. The details of sampling conditions during collection of seawater samples are described in Table 1. The water samples (approximately 20 ml) were collected in sterilized bottles as mentioned in Scoop method and stored at low temperature until the isolation of the bacteria was carried out (within 48 hours)8.
Table (1):
Sampling conditions during collection of Arabian Sea water samples.
Sr. No. |
Characteristics |
Tithal BeachwaterSample |
Diu BeachwaterSample |
Daman BeachwaterSample |
Dandi BeachwaterSample |
|---|---|---|---|---|---|
1. |
Temperature(oC) |
29 oC |
27 oC |
34 oC |
27 oC |
2. |
Turbidity |
Clear |
Turbid |
Turbid |
Clear |
3. |
Weather |
Humid |
Hot & cloudy |
Hot & cloudy |
Cloudy |
4. |
Human Activity |
Medium |
Low |
Low |
Medium |
5. |
Depth (Feet) |
1.5 |
1.5 |
1.5 |
1.5 |
6. |
Distance From Shore (Feet) |
23 |
20 |
20 |
24 |
Isolation of pigment-producing bacteria
The Arabian Sea water samples were diluted serially and seeded onto nutrient agar medium plates, followed by incubation at 37°C for 24-48 hours. Plates with discrete pigmented colonies were considered and morphological characteristics of colonies were recorded. Total 9 morphologically dissimilar discrete bacterial colonies were pulled out and streaked on nutrient agar medium plate to obtain pure cultures. The isolated pure cultures were further maintained on nutrient agar slants at 4°C.
Evaluation of pigment-production at different pH and temperatures
Isolated colonies were streaked on nutrient agar plates with different pH (pH 5, 6, 7 & 8) and incubated at different temperatures ranging from 28°C to 37°C for 24 hrs. Further, the plates were observed for the growth and pigmentation of the isolates.
Extraction of pigments
Bacterial culture grown in nutrient broth with 2% glycerol (pH 7.2) was used for extraction of pigment. The pigments were extracted by using the method described by Asker & Ohta (1999)9. One ml of the standard inoculum was added into fifty ml broth in two fifty ml Erlenmeyer flask. This was incubated for 24 hrs in water bath with shaking at 37°C. After incubation, the broth was centrifuged at 10000 rpm for 10 min. and supernatant (colorless) was discarded, followed by resuspension of pelleted cells in distilled water for lysing the cells. Extraction of pigment from suspended cell pellet (kept in water bath at 60°C for 20 min.) was carried out using methanol with repeated centrifugation (10,000 rpm for 10 min.) until cell debris turned colourless.
The supernatants containing the diffused pigment were filtered through membrane filter (0.22μm pore diameter) and filtrates were collected in sterilized screw cap-tubes and dried. The visible absorption spectra of pigments were analyzed by UV-Visible spectrophotometer at 400-600nm (between the wavelength of 350-750 nm) as described by Krishna et al. (2008)10.
Determination of Antibacterial Activity of the Pigments
The following microorganisms were procured from National Collection of Industrial Microorganisms (NCIM), Pune, India: Bacillus cereus (KR078401; NCIM no. 2155), Escherichia coli (KR109284; NCIM 2065), Vibrio cholera (NCIM no. 5316; ATCC 15748), Bacillus subtilis (ATCC 9372; NCIM 2921), Staphylococcus aureus (KR078391; NCIM 5345) and Bacillus megaterium (NCIM no. 2034; NRRL B 1372(1951). The stock cultures were maintained on nutrient agar at 37°C, then sub-cultured in nutrient broth at 37°C, prior to each test.
The antibacterial activity of the different extracted pigments was carried out by agar-cup diffusion method11. The above mentioned test microorganisms were incubated in nutrient broth at 37°C for 24 hrs. The 0.1 ml of culture was inoculated onto nutrient agar plates and 8.0 mm wells were created with the help of sterile cup-borer. To each well different concentration of the pigments were added followed by incubation of plates (in upright position) at 37°C for 24 hrs. Methanol served as negative control. The experiment was done in triplicates and after 24 hrs antibacterial activity was determined by measuring the zone of inhibition in millimetre.
The minimum inhibitory concentration (MIC) values of the pigments (NP5, NP6, NP8 & NP9) were determined using macro-dilution method, against gram positive and gram negative bacterial pathogens: Escherichia coli, Bacillus cereus, Vibrio cholerae, Bacillus subtilis, Staphylococcus aureus and Bacillus megaterium. Different dilutions of the extracted pigments were prepared in methanol. Bacterial suspension of the test organisms were prepared in Mueller-Hinton broth (sterilized). One ml of the dilution was added to each sterilized screw cap tube containing 1ml of compound (pigment) suitably diluted in the sterilized broth medium to make final volume of two ml. The culture medium devoid of sample (pigment) and other without microorganism served as negative controls. Incubation of all tubes was carried out at 37°C for 24 hrs; followed by analysis at 600nm.
Evaluation of Antioxidant Activity of Pigments
Antioxidant activity of bacterial pigments obtained from NP5, NP6, NP8 & NP9 was determined by DPPH (1,1-diphenyl-picrylhydrazyl) assay, in which 0.05 mM solution of DPPH (in methanol) was used. Two ml of DPPH solution was added to the equal volume of pigment solution (at concentration of 150mg/ml; prepared in methanol). The methanol served as control. The optical density (O.D.) was measured after 60 min (at room temperature) at 517 nm using ascorbic acid as standard. The higher DPPH free radical scavenging activity was correlated with lower absorbance of the reaction mixture12. The following formula was used to calculate percentage inhibition of DPPH, suggestive of percentage of scavenging activity of pigment on DPPH radicals:
Inhibition of DPPH % = (Ao – As) x 100 / Ao
Where, ‘Ao’ refers to Absorption of control; ‘As’ refers to Absorption of tested pigment
Cytotoxicity and anti-cancer activity assay of pigments
The MTT [3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide] assay was used to assess cell viability13. In vitro cytotoxicity effects of different pigments were studied using MCF-7 Breast cancer cell line. Briefly, 96 wells cell culture plate 1×104 Cells were seeded with 200µl of DMEM medium with 10% Bovine Fetal Serum followed byincubation of plate at 37°C to get confluent monolayer. Once 80% confluency was arrived, cells were treated with different concentration of pigments NP5 (orange), NP6 (yellow) and NP9 (red) (100 to 1000 µg/ml) respectively and untreated cells were kept as control. Incubation of plate was carried out at 37°C for 24 hrs, followed by cytotoxicity assessment by MTT assay. The MTT was added to all the treatment wells including control and incubation of plate was carried out at 37°C for 4 hrs. Further, the formazan crystals were solubilized by addition of DMSO for 20 min. The cell viability values were recorded in ELISA reader at 570 nm. The cell viability percentage was calculated by following formula:
Cell viability Percentage (%) = (A570 of the test) / (A570 of the control) × 100
Molecular identification
The selected pigment producing isolate was subjected to genomic DNA isolation and 16srDNA PCR was performed. The forward primer: 5’ AGAGTTTGATCCTGGCTCAG3’ and reverse primer: 5’AAGGAGGTGATCCAGCCGCA3’ were used. The PCR products were then sent for 16srDNA sequencing. The DNA sequences were BLAST from the existence microbial DNA database and Phylogenetic trees were evaluated.
Pigment producing isolates
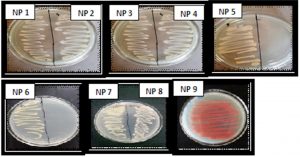
Total 50 morphologically distinct bacterial colonies were obtained from Arabian Sea water samples. The conditions of sampling are shown in Table 1. Of these 50 bacterial colonies, 9 were found to be pigmented. Out of 9 pigmented colonies, 2 were yellow pigmented, 6 were with orange pigment and 1 colony was red in color (Fig. 1). The isolates namely NP5, NP6, NP8 and NP9 were found to be more prominent pigment producers as compared to other isolates and hence they were further considered (Fig. 1). The other isolates such as NP1, NP2, NP3, NP4 and NP7 showed very less or negligible amount of pigment production.
Fig. 1. Isolation of pigment producing bacteria: The bacterial colonies were isolated from different dilutions of marine water samples. Numerous colonies were obtained out of which total 9 pigment producing colonies were selected from higher dilution plates and designated as pigment producing isolates.
Effect of pH on pigment production
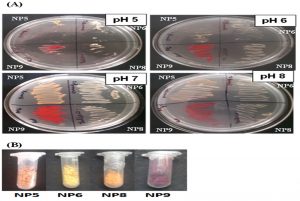
Since, the alteration in pH affects production of pigments; the selected bacterial isolates (NP5, NP6, NP8 & NP9) were streaked on media containing different pH (5.0, 6.0, 7.0 and 8.0). Though, the organisms were capable to grow at different pH; the pigment production capacity and bacterial growth was found to be optimum at pH 7 as compared to other pH (Fig. 2). Hence, the pH 7.0 was considered further for growth of bacteria in broth media and for extraction of pigments.
Fig. 2. (A) Pigment producing bacterial isolates at different pH (5.0, 6.0, 7.0 & 8.0): The pH 7.0 was found to be optimum for bacterial growth as well as pigment production. (B) Extracted Pigments from NP5 (Orange), NP6 (Yellow), NP8 (Orange) and NP9 (Red) isolates after drying: The dried form of pigments were weighed 0.122g, 0.576g, 0.323g & 0.231g per 150ml of culture respectively.
Bacterial Pigment Extraction
Pigments were extracted using solvents such as acetone, ethyl acetate, chloroform and methanol. However, the pigments were not successfully extracted using chloroform, ethyl acetate and acetone but methanol was found to extract the different pigments efficiently from the respective bacterial cells. The pigments were successfully extracted from all the four isolates (NP5, NP6, NP8 & NP9) in the dried form and weighed 0.122g, 0.576g ,0.323g & 0.231g per 150ml of culture broth respectively. (Fig. 2B).
Antibacterial activity of pigments
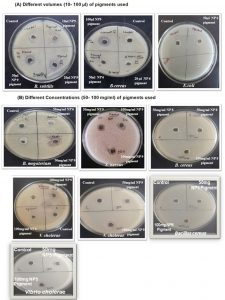
Further, the extracted pigments were subjected to assessment of antibacterial activity against different bacterial pathogens (Table 2). Out of the 9 pigmented bacterial isolates, 4 pigment producing isolates namely NP5, NP6, NP8 and NP9 showed inhibitory activity towards the test pathogens as suggested by zone of inhibition (Fig. 3). Among these four isolates, NP9 showed maximum antibacterial activity against all the test microorganisms (Fig. 3).
Fig. 3. Antibacterial activity of extracted pigments against different test bacteria:
(A) Different pigments such as NP5(Orange), NP 6(Yellow), NP 8(Orange) and NP 9(Red) in different volumes were tested against different pathogens which shows inhibitory activity with increase in amount of pigment used against B. subtilis, B. cereus and E.coli.
(B) At 50mg/ml concentration pigments showed minimum inhibitory activity against B. subtilis, B. cereus B. megaterium, S. aureus and V. cholerae but at 100mg/ml they showed larger zone of inhibition. NP 9(Red) pigment showed potent inhibitory activity as compared to NP 5(Orange), NP6 (Yellow) and NP8 (Orange) pigments.
Table (2):
Antibacterial activity of bacterial pigments against different test bacteria.
| Pigments | Test organisms | ||||
|---|---|---|---|---|---|
| Diameter of zone of inhibition (mm) | |||||
| V. cholerae | B. subtilis | B. megaterium | S. aureus | B. cereus | |
| Pigment Concentration: 50mg/ml | |||||
| NP1 | No zone | No zone | No zone | No zone | No zone |
| NP2 | No zone | No zone | No zone | No zone | No zone |
| NP3 | No zone | No zone | No zone | No zone | No zone |
| NP4 | No zone | No zone | No zone | No zone | No zone |
| NP5 | 1 mm | No zone | No zone | No zone | No zone |
| NP6 | No zone | No zone | No zone | No zone | No zone |
| NP7 | No zone | No zone | No zone | No zone | No zone |
| NP8 | 4.0 | 3.0 | No zone | No zone | No zone |
| NP9 | 6.0 | 5.0 | 5.0 | 4.0 | 6.0 |
| Pigment Concentration: 100mg/ml | |||||
| NP1 | No zone | No zone | No zone | No zone | No zone |
| NP2 | No zone | No zone | No zone | No zone | No zone |
| NP3 | No zone | No zone | No zone | No zone | No zone |
| NP4 | No zone | No zone | No zone | No zone | No zone |
| NP5 | 2 mm | No zone | No zone | No zone | No zone |
| NP6 | No zone | No zone | No zone | No zone | 1.0 |
| NP7 | No zone | No zone | No zone | No zone | No zone |
| NP8 | 6.0 | 5.0 | No zone | No zone | 5.0 |
| NP9 | 8.0 | 6.0 | 7.0 | 6.0 | 8.0 |
Table (3):
Minimum inhibitory concentration (MIC) of bacterial pigments against different test microorganisms.
Isolate & Pigment |
Test microorganisms (Accession No.) |
Concentration of Pigment (mg/ml) |
MIC (mg/ml) |
|---|---|---|---|
NP5 (Orange) |
Bacillus megaterium (NCIM no. 2034; NRRL B 1372(1951) |
200 |
50 |
NP6 (Yellow) |
Bacillus cereus (KR078401; ATCC 6630; NCIM no. 2155) |
300 |
75 |
NP8 (Orange) |
Bacillus cereus (KR078401; ATCC 6630; NCIM no. 2155) |
250 |
62.5 |
NP 9 (Red) |
Vibrio cholerae (NCIM no. 5316; ATCC 15748 ) |
350 |
43.75 |
MIC of pigments
The minimum inhibitory concentrations (MIC) of pigments obtained from NP5, NP6, NP8 and NP9 isolates against different test microorganisms have been shown in Table 3. The pigments from other remaining isolates were not tested for MIC, as they did not show antibacterial activity against any of the pathogens (Table 2). The results suggest that NP9 pigment (red pigment) exhibited effective MIC of 43.75 mg/ml against Vibrio cholerae. While NP8 pigment (yellow pigment) exhibited effective MIC of 63.5 mg/ml against Bacillus cereus as compared to NP6 pigment. The NP5 pigment (orange pigment) exhibited MIC of 50.0 mg/ml against Vibrio cholerae.
Antioxidant activity of pigments
The antioxidant activity of pigments of isolates namely, NP5, NP6, NP8 and NP9 along with ascorbic acid as reference standard was determined by DPPH assay (Suppl. Fig. S1). The antioxidant activity of the pigments from NP5, NP6, NP8 and NP9 (at concentration of 150mg/ml) was found to be 36.6%, 22.22%, 55.55% and 44.44% of respectively. These results suggest that NP8 pigment exhibited higher antioxidant property as compared to NP5, NP6 and NP9.
Cytotoxicity activity assay of pigments on MCF7 Cells
The breast cancer cells (MCF-7) were treated with different doses (100-1000μg) of bacterial pigments: NP5 (Orange), NP6 (yellow) and NP9 (red), respectively (Suppl. Fig. S2) for 24 hrs. There was no significant decrease in viability (Table S3) of the cells was observed in pigment treated group as compared to untreated groups with different doses of all the three pigments.
Cultural characteristics and Molecular identification of isolates
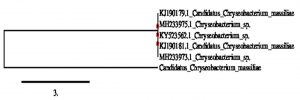
The cultural characteristics were studied for all selected nine isolates (Tables S1). The isolates namely, NP1, NP2 NP3, NP4, NP5, NP6, NP7 and NP8 were revealed as gram positive cocci, whereas isolate NP9 was found to be gram negative rod. The biochemical properties of the isolates are presented in Table S2. Molecular identification of selected isolate NP9, which showed better antibacterial, antioxidant property with novel pigment production, was carried out by 16srDNA sequencing. The results revealed NP9 isolate as Candidatus chryseobacterium massiliae. The sequence submission of 16srDNA was done in GenBank-NCBI with accession number MK210172. The phylogenetic analysis of the NP9 isolate is shown in Fig. 4.
Marine bacteria have potential for producing unique metabolites that make their endurance in hostile environmental conditions such as high salinity, pressure and temperatures14. The marine bioactive pigments of bacteria have been reported, such as prodiginines (red), carotenes (orange), violacein (violet), phenazine Compounds (blue, green, purple, yellow, red & brown), quinones (yellow to red) and tambjamines (yellow & Purple)5. Previously, several investigations of marine bacteria have been carried out for their antibiotic activity. Pyocyanin and pyorubrin pigments produced by Pseudomonas have been reported to exhibit antibacterial activity.15 Similarly, himalomycin A, B, and fridamycin D pigments produced by Streptomycete sp. B6921 showed antibacterial activity16. The tambjamines and tryptanthrin produced by Pseudoalteromonas tunicate and Flexibacteria respectively found to exhibit antibiotic properties17,18.
Since, there are several investigations going on for searching novel antibiotics and drug resistance is also increasing. Therefore, these bacterial pigments could be a good alternative against drug resistant bacteria. Recently, a yellow pigmented coral-associated bacterium exhibiting anti-Bacterial activity against multidrug resistant (MDR) organism has been reported19. The present study isolated pigmented bacteria from Arabian Sea water samples from different coastal regions in India and extracted the different pigments to analyze their antibacterial, antioxidant and anti-cancerous properties. The marine bacteria isolated in the present study were found to produce yellow, red & orange pigments. The isolates namely, NP5, NP6, NP8 and NP9 were found to have antibacterial activity as well as good antioxidant property. The NP9 red pigment was found to be more effective as antibacterial agent as compared to others based on its zone of inhibition and MIC values against different bacteria; suggesting that it was effective in inhibiting the growth of bacterial pathogens at lower concentrations as well. Earlier, one research group reported that 69.4% of the colonies were chromogenic out of thousands of colonies obtained from marine water and mud samples20. In particular, the study found yellow (31.3%), orange (15.2%), brown (9.9%) and red/pink (5.4%)20. Similarly, 60 novel marine bacterial species including yellow (19), brown (5), orange (4) and red (1) were reported21.
In addition to anti-bacterial activity, the marine bacterial pigments have also been reported to exhibit antioxidant activity due to antioxidant compounds such as polyphenolic compounds22. In the present study, the marine bacterial pigments also exhibited good antioxidant activity. In particular, the NP8 (orange) pigment exhibits higher antioxidant property as compared to that of NP6 (yellow pigment), NP5 (orange pigment) and NP9 (red pigment). Ealrier, prodigiosin pigment was shown to contain good study antioxidant activity23. The anti-oxidant property of the bacterial pigments renders them a useful agent as natural colorants in foods so as to increase the shelf life as well.
One of the bioactive properties of the bacterial pigments is their anti-cancerous activity. The bacterial pigments can induce apoptosis and arrest the cell cycle. Earlier, marine bacterial pigments such as undecylprodigiosin produced by Streptomyces rubber24, violacein produced by Pseudoalteromonas sp.25, and chinikomycin A & B, Manumycin A produced by Streptomycete sp.26 have been reported to exhibit anti-cancerous activity. The current study also investigated the anti-cancer potential of the three pigments: orange (NP5), yellow (NP6) and red (NP9). However, none of these three pigments showed inhibitory effect on MCF-7 cells as compared to compared to the untreated control. In addition to antibacterial property of the extracted pigments, red and purple pigments are also known to have colorfastness property and hence, these pigments can be used as colorant for fabrics in textile industry as a safe and eco-friendly colorant to replace harmful synthetic dye27.
The molecular characterization of NP9 by 16srDNA sequencing revealed the isolate as Candidatus chryseobacterium massiliae. In contrast to study by Ahmad et al. (2012)3, in which Chryseobacterium was shown to produce yellow- orange pigment, the present study reports for the first time that Candidatus chryseobacterium massiliae produced red pigment with potent antibacterial activity. The bacterium identified in the present study may be a novel strain of Chryseobacterium massiliae capable of producing red pigment.
Overall, the present study found that the four isolates namely NP5, NP6, NP8 and NP9 obtained from Arabian Sea Water sample produce antibacterial pigments with good antioxidant activity. Among these isolates NP9 was characterized as Candidatus chryseobacterium massiliae (MK213063), which exceptionally produces red pigment and possess higher and diverse anti-bacterial activity among the other isolates and may be employed in various pharmaceutical applications. However, antibacterial activity of these pigments must be confirmed with other bacterial pathogens as well. In addition, further characterization of red pigment of Candidatus chryseobacterium massiliae and disease animal model studies are needed for its pharmaceutical application.
Additional file: Additional Table S1- S3. Additional Figure S1 and S2.
ACKNOWLEDGMENTS
We are thankful to Uka Tarsadia University, Bardoli, Gujarat, India for providing necessary research facilities to conduct the study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
MD designed the experiments; NP & SJ performed the experiments; NP & MD analyzed data, and NP, MD, SJ, RB wrote and edited the manuscript. All authors reviewed the data and the manuscript.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript and/or the Supplementary Files.
- Hugenholtz P & Pace NR. Identifying microbial diversity in the natural environment: a molecular phylogenetic approach. Trends Biotechnol., 1996; 14(6): 190-197.
Crossref - Zobell CE. Marine Microbiology. ChronicaBotanica Co., Waltham, Mass, 1946; pp1-240.
- Ahmad WA, Ahmad WYW, Zakaria ZA & Yusof NZ. Application of bacterial pigments as colorant. In Application of Bacterial Pigments as Colorant. Springer, Berlin, Heidelberg, 2012; pp 57-74.
Crossref - Numan M, Bashir S, Mumtaz R, Tayyab S, Rehman NU, Khan AL, Shinwari ZK & Al-Harrasi A. Therapeutic applications of bacterial pigments: a review of current status and future opportunities. 3 Biotech., 2018; 8(4): 207.
Crossref - Soliev AB, Hosokawa K and Enomoto K. Bioactive Pigments from Marine Bacteria: Applications and Physiological Roles. Evidence-Based Complementary Alternat. Med., 2011; 2011; Article ID 670349, 17 pages.
Crossref - Joshi VK, Attri D, Bala A & Bhushan S. Microbial pigments. Ind J Biotech., 2003; 2: 362-369.
- Giri AV, Anandkumar N, Muthukumaran G & Pennathur G. A novel medium for the enhanced cell growth and production of prodigiosin from Serratia marcescens isolated from soil. BMC Microbiol., 2004; 4(1): 11.
Crossref - Balraj J, Pannerselvam K & Jayaraman A. Isolation of pigmented marine bacteria Exiguobacterium sp. from peninsular region of India and a study on biological activity of purified pigment. Int. J Sci. Tech. Res., 2014; 3(3): 375-384.
- Asker D & Ohta Y. Production of canthaxanthin by extremely halophilic bacteria. J Biosci. Bioengin., 1999; 88(6): 617-621.
Crossref - Krishna JG, Basheer SM, Beena PS & Chandrasekaran M. Marine Bacteria As Source Of Pigment For Application As Dye In Textile Industry. Proc. Internatl. Conf. Biodiv. Conserv. Mgt., 2008; 743-44.
- Grammer A. Antibiotic sensitivity and assay test. Microbiol. Methods, 1976; 235.
- Vora A, Londhe V & Pandita N. Herbosomes enhance the in vivo antioxidant activity and bioavailability of punicalagins from standardized pomegranate extract. J Functional Foods, 2015; 12: 540-548.
Crossref - Mosmann T. Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol. Methods, 1983; 65(1-2): 55-63.
Crossref - Fenical W & Jensen PR. Marine microorganisms: a new biomedical resource. In Pharmaceutical and Bioactive Natural Products. Springer, Boston, MA, 1993; pp 419-457.
Crossref - Saha S, Thavasi R & Jayalakshmi S. Phenazine pigments from Pseudomonas aeruginosa and their application as antibacterial agent and food colourants. Res. J Microbiol., 2008; 3(3): 122–128.
Crossref - Maskey RP, Helmke E & Laatsch H. Himalomycin A and B: isolation and structure elucidation of new fridamycin type antibiotics from a marine Streptomyces isolate. J Antibiotics, 2003; 56(11): 942-949.
Crossref - Franks A, Haywood P, Holmstrom C, Egan S, Kjelleberg S & Kumar N. Isolation and structure elucidation of a novel yellow pigment from the marine bacterium Pseudoalteromonas tunicate. Molecules, 2005; 10(10): 1286-1291.
Crossref - Wagner-Dobler I, Beil W, Lang S, Meiners M & Laatsch H. Integrated approach to explore the potential of marine microorganisms for the production of bioactive metabolites. Adv. Biochem. Eng./Biotechnol., 2002; 74: 207-238.
Crossref - Sibero MT, Bachtiarini TU, Trianto A, Lupita AH, Sari DP, Igarashi Y, Harunari E, Sharma AR, Radjasa OK & Sabdono A. Characterization of a yellow pigmented coral-associated bacterium exhibiting anti-Bacterial Activity Against Multidrug Resistant (MDR) Organism. The Egypt. J Aquatic Res., 2019; 45(1): 81-87.
Crossref - Zobell CE & Feltham CB. Preliminary studies on the distribution and characteristics of marine bacteria. Bull. Scripps Inst. Oceanog. Tech. Ser., 1934; 3: 279-296.
- Zobell CE and Upham HC. A list of marine bacteria including descriptions of sixty new species. Bull. Scripps Inst. Oceanogr., 1944; 5: 239-292.
- Bravo L. Polyphenols: Chemistry, Dietary Sources, Metabolism, and Nutritional Significance. Nutri. Rev., 1998; 56(11): 317-333.
Crossref - Panesar R, Kaur S & Panesar PS. Production of microbial pigments utilizing agro-industrial waste: a review. Curr. Opinion Food Sci., 2015; 1: 70-76.
Crossref - Gerber NN. Prodigiosin-like pigments. CRC Crit. Rev. Microbiol., 1975; 3(4): 469-485.
Crossref - Yada S, Wang Y, Zou Y et al. Isolation and characterization of two groups of novel marine bacteria producing violacein. Marine Biotechnol., 2008; 10(2): 128-132.
Crossref - Li F, Maskey RP, Qin S et al. Chinikomycin A and B: isolation, structure elucidation and biological activity of novel antibiotics from a marine Streptomyces sp. isolate M045. J Nat. Prod., 2005; 68 (3): 349-353.
Crossref - Mumtaz R, Bashir S, Numan M, Shinwari ZK, Ali M. Pigments from Soil Bacteria and Their Therapeutic Properties: A Mini Review. Curr Microbiol. 2019; 76(6): 783-790.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.