ISSN: 0973-7510
E-ISSN: 2581-690X
The volatile organic compounds (VOCs) produced by eight plant growth-promoting bacterial strains originating from surface-sterilized root nodules of cowpea and garden pea were examined using headspace-solid phase microextraction-gas chromatography–mass spectrometry (HS-SPME-GC-MS). One hundred and fifty-nine VOCs were detected in the control and bacterial inoculated flasks. Among the VOCs produced by the bacterial isolates, esters (14.46%), ketones (13.83%), alcohols (12.57%) and hydrocarbons (9.4%) were the most prevalent, while other classes of VOCs were detected at lower frequencies. While 33.13% of the compounds were exclusive to cowpea nodule isolates, 38.50% of the compounds were exclusive to garden pea nodule isolates. The compounds that were common between both groups accounted for 28.40% of the total compounds detected. The major VOCs with plant growth promotion potential produced by Enterobacter sp. CPH64, Enterobacter sp. CPK42, Chryseobacterium sp. CPM11, and Stenotrophomonas sp. CPH62, originating from cowpea root nodules were palmitic acid, dimethyl silanediol, hexanol 2-ethyl, 2-methyl 1- butanol and 2-tridecanone, while Enterobacter sp. GP44, Enterobacter sp. GP71, Enterobacter sp. GP84, and Bacillus sp. GP102, from garden pea root nodules, produced the VOCs, viz. 3-methyl-1-butanol, 2-methyl-1-butanol, 2,3,5-trimethyl pyrazine, dimethyl silanediol, 2-tridecanone, butane 1-methoxy 3-methyl, hexanol 2-ethyl- and palmitic acid. These compounds have been previously shown to promote plant growth, improve plant defence, leaf chlorophyll content, enhance fruit ripening and mitigate stress effects. This is an early report on the VOC profile of bacterial endophytes isolated from root nodules of two vegetable legumes, viz. cowpea (tropical) and garden pea (temperate).
Vegetable Legumes, Root Nodule-associated Bacteria, Volatile Organic Compounds: HS-SPME-GC-MS; Plant Growth Promotion
Vegetable legumes are a diverse and nutritionally rich group of plants that are crucial for the human diet and sustainable agriculture due to their nitrogen-fixing abilities. This category includes peas, beans, lentils, chickpeas, and soybeans. Cowpea (Vigna unguiculata) and garden pea (Pisum sativum) are two vital food legumes. In 2019, global cowpea production reached 8.9 Mt1 while garden pea cultivation covered 2.18 Mha, yielding 21.77 Mt at a productivity rate of 9.99 t ha-1.2 It was previously believed that only rhizobial strains could colonize the interiors of root nodules. However, later studies have indicated that diverse bacteria belonging to different genera, viz. Agrobacterium, Arthrobacter, Acinetobacter, Bacillus, Bosea, Enterobacter, Micromonospora, Mycobacterium, Paenibacillus, Pseudomonas and Stenotrophomonas, are also associated with legume root nodules and play important roles in promoting the growth and development of their hosts.3,4 But research on their functional metabolites remains limited. Most studies on bacterial volatile organic compounds have focused on rhizosphere-associated strains or well-characterized PGPR such as Bacillus and Pseudomonas,5-7 while the VOC profiles of root nodule bacteria of cowpea and gardenpea nodules remains to be deciphered, despite increasing evidence that nodule-associated non-rhizobial bacteria play important roles in plant growth and stress adaptation. This study therefore addresses a clear gap by providing the first VOC profile of bacterial endophytes isolated from root nodules of two vegetable legumes, viz. cowpea (tropical) and garden pea (temperate).
Rhizospheric microbes produce a wide range of volatile organic and inorganic chemical compounds that play a definite role in signalling among bacterial communities, promoting host growth and inducing tolerance to biotic and abiotic stresses.8,9 Inorganic compounds include carbon dioxide (CO2), carbon monoxide (CO), nitric oxide (NO), hydrogen sulfide (H2S), ammonia (NH3) and hydrogen cyanide (HCN), while volatile organic compounds (VOCs) include a range of terpenes, aromatic compounds, hydrocarbons, ketones, alcohols, aldehydes, acids, and metabolites containing sulfur and nitrogen.6,10 The plant growth-promoting properties of VOCs emitted by rhizospheric bacteria have been established previously.11 VOCs are known to stimulate photosynthesis, root growth and the uptake of specific nutrients such as iron and sulfur.12-15 They are also known to directly inhibit the growth of fungal and bacterial phytopathogens, thereby protecting their hosts.16,17
Gas chromatography–mass spectrometry (GC-MS) is particularly well suited for the analysis of VOCs produced by bacterial strains under various conditions due to its sensitivity, selectivity and ability to analyse a wide range of compounds. Prior to their detection by GC-MS, bacterial volatile organic compounds (VOCs) can be efficiently extracted and concentrated by headspace (HS)-solid-phase microextraction (SPME).18-20 The data generated by GC-MS not only aids in compound identification but also facilitates their quantitative analysis. This study was carried out to profile the VOCs produced by eight plant growth-promoting bacterial isolates obtained from the root nodules of vegetable legumes, viz. cowpea and garden pea, by HS-SPME-GC-MS and to elucidate their putative plant growth-promoting roles from previously published literature.
Bacterial strains
The bacterial strains used in this study were originally isolated from the surface-sterilized root nodules of cowpea and garden pea cultivated in different parts of the states of Andhra Pradesh and Karnataka, India, and selected for further study based on their superior plant growth-promoting abilities, viz. ammonia production, phosphate solubilization and growth hormone production.21 The elite bacterial isolates, viz. Chryseobacterium sp. CPM11, Enterobacter sp. CPH64, Enterobacter sp. CPK42, and Stenotrophomonas sp. CPH62 from cowpea root nodules, and Enterobacter sp. GP44, Enterobacter sp. GP71, Enterobacter sp. GP84, and Bacillus sp. GP102 from garden pea root nodules were subjected to volatile profiling by HS-SPME-GC-MS.
Sample preparation and headspace-solid-phase micro-extraction
Individual bacterial isolates were inoculated in nutrient broth in 250 ml Erlenmeyer flasks with a headspace of 60%. The flasks were tightly sealed with plastic film wrap to prevent any loss of volatile compounds and incubated at 30 °C for 48 hrs. An uninoculated control was included to determine the volatile emissions from the sterile media. After incubation, volatile compounds were trapped with a fused-silica SPME fibre coated with a mixed stationary phase of polydimethylsiloxane (PDMS), divinylbenzene (DVB), and carboxen (CAR) on a stainless-steel holder. The DVB/CAR/PDMS fibre (50/30 µm; highly cross-linked) was pre-conditioned at 300 °C for 1 hour.20 The fibre was inserted into the flask through a small opening in plastic film and exposed to the headspace for 2 hours, with gentle stirring. After adsorption, the fibre was transferred to the GC-MS injector for thermal desorption and subsequent analysis of the volatile compounds.
GC-MS analysis of volatile organic compounds
The GC-MS analysis was carried out using a Shimadzu Nexis GC-2030 system connected to a Shimadzu TQ 8040 NX mass detector. After sampling, the SPME fibre was left in the injector during the entire run. The VOCs were separated using a fused-silica capillary column (SH-I-5Sil MS; 30 m × 0.25 mm, 0.25 µm film thickness). The injector was set at 255 °C, and samples were injected in split mode (1:5) for 0.2 min. The detector was set at 270 °C. The oven program started at 40 °C for 3 min, increased by 3 °C/min to 160 °C (held for 2 min), and then increased by 5 °C/min to 230 °C, where it was held for 1 min. The MS was operated in electron-ionization mode with helium as the carrier gas at 1.0 mL/min. The ion source, transfer line, and injector were kept at 230 °C, 240 °C, and 255 °C, respectively, and spectra were recorded in the 50-400 amu range. Total VOC production was estimated by adding all GC peak areas and expressing each compound as a percentage of this total. The VOCs were identified by matching retention indices and mass spectra with the FFNSC and NIST-2020 libraries. The relative abundance of each compound was adjusted by subtracting the peak areas of the uninoculated control.
Statistical analysis
The data was statistically analysed using Operational Statistics (OPSTAT) software. The metabolome data were analysed using MetaboAnalyst 6.0.22 Partial least squares–discriminant analysis (PLS-DA) was performed using MetaboAnalyst (https://www.metaboanalyst.ca), a comprehensive web-based platform, to differentiate the overall volatile profile patterns among bacterial isolates from cowpea nodules (CPH62, CPH64, CPK42, CPM11) and their control group, as well as bacterial isolates from garden pea nodules (GP44, GP71, GP84, GP102) and their respective control group. Prior to analysis, the dataset was formatted to include volatile names and sample names. Additionally, heatmaps were created using normalized data in MetaboAnalyst to visualize the relative abundance of key volatiles. These heatmaps display the major volatiles emitted by each bacterial isolate in a matrix format, highlighting patterns and group-specific differences. A Venn diagram was prepared to represent the common VOC’s between the root nodule associated bacterial groups using the “Venny 2.1.0” tool.23
Partial Least Squares-Discriminant Analysis of the VOCs produced by the bacterial isolates
A total of one hundred and fifty nine VOCs were detected from the uninoculated control and the eight bacterial strains by HS-SPME-GC-MS. Among the VOCs produced by the bacterial isolates, esters (14.46%), ketones (13.83%), alcohols (12.57%) and hydrocarbons (9.4%) were the most prevalent, while other classes of VOCs were detected at lower frequencies. While 33.13% of the compounds were exclusive to cowpea nodule isolates, 38.50% of the compounds were exclusive to garden pea nodule isolates. The compounds that were common between both groups accounted for 28.40% of the total compounds detected. The VOCs with potential plant growth promotion activity are listed in Table. The data matrix of the VOCs produced by the cowpea and garden pea isolates was analysed separately by PLS-DA to identify the variation and grouping trends of the VOCs with respect to the uninoculated control. This distinguished the overall differences in metabolic profiles between the groups.
Table:
Volatile organic compounds with potential plant growth promoting effects produced by the root nodule-associated bacterial strains
| Volatile Organic Compound | Relative Abundance (%) | Produced by group | Functional | References |
|---|---|---|---|---|
| 3-methyl-1 butanol | 19.59 | Enterobacter sp. CPH64, | Alcohol | 24-28 |
| 11.43 | Enterobacter sp. GP44, | |||
| 2.83 | Chryseobacterium sp. CPM11, | |||
| 15.03 | Enterobacter sp.CPK42, | |||
| 23.91 | Enterobacter sp. GP84 | |||
| 2,3,5-Trimethyl pyrazine | 1.28 | Bacillus sp. GP102 | Pyrazine | 17,29-31 |
| 0.58 | Enterobacter sp. GP71, | |||
| 0.8 | Enterobacter sp. CPH64 | |||
| Dimethyl silanediol | 5.23 | Bacillus sp. GP102, | Alcohol | 32 |
| 1.23 | Chryseobacterium sp. CPM11, | |||
| 0.41 | Enterobacter sp. CPH64, | |||
| 2.04 | Stenotrophomonas sp. CPH62 | |||
| 0.74 | Enterobacter sp.CPK42, | |||
| 0.9 | Enterobacter sp. GP44, | |||
| 0.77 | Enterobacter sp. GP84 | |||
| 0.98 | Enterobacter sp. GP71 | |||
| 2-Tridecanone | 6.74 | Enterobacter sp. CPH64, | Ketone | 33-39 |
| 2.02 | Enterobacter sp. GP44, | |||
| 3.17 | Enterobacter sp.CPK42, | |||
| 1.76 | Enterobacter sp. GP84 | |||
| 0.39 | Chryseobacterium sp. CPM11 | |||
| Butane 1- methoxy 3- methyl | 23.88 | Enterobacter sp. GP44, | Methoxy | 5,12,40 |
| 31.94 | Enterobacter sp. GP84 | |||
| Hexanol 2-ethyl- | 12.24 | Enterobacter sp. GP71, | Alcohol | 41,42 |
| 3.23 | Bacillus sp. GP102, | |||
| 1.71 | Enterobacter sp. GP44, | |||
| 1.18 | Chryseobacterium sp. CPM11, | |||
| 6.02 | Stenotrophomonas sp. CPH62, | |||
| 2.97 | Enterobacter sp. CPK42 | |||
| Palmitic acid | 2.22 | Bacillus sp. GP102 | Fatty acids | 43 |
| 0.39 | Enterobacter sp. CPH64, | |||
| 0.46 | Enterobacter sp. CPK42 | |||
| 0.49 | Stenotrophomonas sp. CPH62 | |||
| 0.92 | Enterobacter sp. GP71, | |||
| 0.34 | Enterobacter sp. GP84, | |||
| 2-methyl-1- butanol | 7.08 | Enterobacter sp. CPH64, | Alcohol | 5,44-47 |
| 4.76 | Enterobacter sp. GP44, | |||
| 3.24 | Enterobactersp.CPK42 | |||
| 0.89 | Chryseobacterium sp. CPM11, |
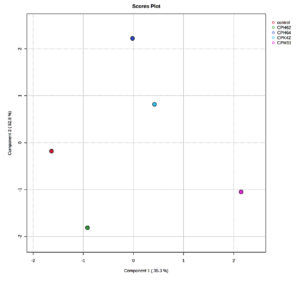
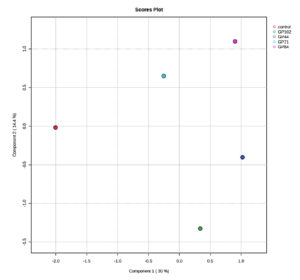
The score plots illustrated clear differences in the volatile compounds produced by the different bacterial groups and the control. The volatile compounds from bacterial isolates of cowpea (Figure 1a) and garden pea (Figure 1b) nodules exhibited distinct separation from each other and from the control, indicating differential expression patterns across the isolates and the control. In the score plot of cowpea nodule isolates, component 1 and component 2 accounted for 35.3% and 52.8%, respectively, of the variation, while in the score plot of garden pea nodules, component 1 and component 2 accounted for 30% and 14.4%, respectively, of the variation.
Figure 1a. Partial Least Squares-Discriminant Analysis (PLS-DA) of the VOCs detected in the uninoculated control and cowpea nodule isolates
Figure 1b. Partial Least Squares-Discriminant Analysis (PLS-DA) of the VOCs detected in the uninoculated control and garden pea root nodule isolates
Venn diagram analysis and hierarchical clustering of VOCs produced by the bacterial isolates
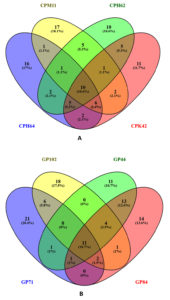
The Venn diagram analysis (Figure 2) was employed to examine the uniqueness and commonalities among the VOCs produced by the cowpea and garden pea isolates. Among the cowpea isolates, Chryseobacterium sp. CPM11 produced seventeen distinct compounds and Enterobacter sp. CPH64 produced sixteen distinct compounds, Stenotrophomonas sp. CPH62 produced ten distinct compounds and Enterobacter sp. CPK42 produced eleven unique compounds. Chryseobacterium sp. CPM11 and Stenotrophomonas sp. CPH62 produced five VOCs that were common amongst them. Two common VOCs were produced by Enterobacter sp. CPH64 and Stenotrophomonas sp. CPH62, while five common VOC’s were detected in Stenotrophomonas sp. CPH62 and Enterobacter sp. CPK42. The collective analysis of the cowpea isolates revealed a shared set of 10 compounds among them. This suggests shared metabolic pathways among these bacterial isolates. The investigation of volatile organic compounds produced by garden pea nodule isolates revealed distinctive chemical profiles for each isolate. Bacillus sp. GP102 featured an exclusive repertoire of 18 compounds, Enterobacter sp. GP84 produced 14, Enterobacter sp. GP71 produced twenty-one unique compounds, and Enterobacter sp. GP44 produced 11 distinctive elements. Commonalities were observed between the Enterobacter sp. GP71 and Bacillus sp. GP102 isolates, which share six common compounds. Intriguingly, no common compounds were identified between Bacillus sp. GP102 and Enterobacter sp. GP44, but one common compound was found between Enterobacter sp. GP71 and Enterobacter sp. GP44. A substantial overlap of eleven common compounds was observed among the four isolates, emphasizing shared metabolic pathways across the isolates.
Figure 2. Venn diagram displaying the number of volatile organic compounds coincident between the bacterial strains isolated from (A) cowpea and (B) garden pea root nodules
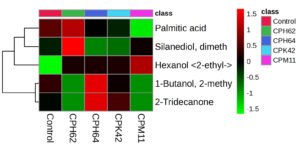
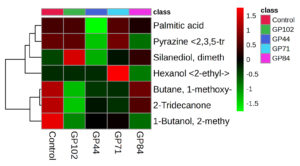
The differential expression of bacterial volatile compounds with plant growth promotion abilities was visualized via hierarchical clustering with heatmaps. The volatile compounds were differentially accumulated across isolates obtained from cowpea, garden pea nodule isolates and the control (Figures 3 and 4). Higher expression levels of volatiles are denoted by red on the heatmap, whereas lower expression levels are represented by green. In summary, the detailed examination of volatile compounds provided valuable insights into the distinctive characteristics and potential functional roles of each bacterial isolate, shedding light on their unique chemical signatures.
Figure 3. Heatmap of the VOCs with putative plant growth promotion abilities produced by bacterial isolates obtained from cowpea nodules. The columns represent the bacterial isolates, and the rows represent the different metabolites identified. The colour codes indicate the abundance of each VOC
Root nodules harbour several non-rhizobial bacteria. Twenty strains were isolated from the root nodules of Phaseolus vulgaris L., which include six different genera, viz.: Enterobacter, Stenotrophomonas, Chryseobacterium, Arthrobacter, Pseudomonas and Rhizobium.48 Members of the family Enterobacteriaceae were the most abundant among the nonrhizobial genera, in cowpea nodules along with Stenotrophomonas and Chryseobacterium.49
In the present study, HS-SPME-GC-MS analysis revealed the production of volatile compounds, such as 3-methyl-1-butanol, 2-methyl-1-butanol, 2,3,5-trimethyl pyrazine, dimethyl silanediol, 2- tridecanone, butane 1-methoxy 3-methyl, hexanol 2-ethyl- and palmitic acid, which were previously shown to have plant growth-promoting activities, plant defence, improved chlorophyll content, fruit ripening and stress mitigation activities.50,51 To analyse these compounds, the Partial Least Squares Discriminant Analysis (PLS-DA) which is a multivariate statistical technique in metabolomics is increasingly being used to analyse volatile profiles of Plant Growth-Promoting Bacteria (PGPB)52-54 was employed. By integrating VOC peak intensities with supervised classification, PLS-DA effectively differentiates bacterial strains based on their characteristic volatile signatures.55 Recent studies have profiled VOC blends from several PGPR, such as Pseudomonas koreensis, P. fluorescens, Lysinibacillus sphaericus, and Paenibacillus alvei, using HS-SPME-GC-MS coupled with PLS-DA.52 This approach has revealed species-specific and strain-specific VOC patterns, identified novel compounds and certain VOCs as potential biomarkers for PGPR classification and functional prediction.52 The PLS-DA has also proven to be effective in distinguishing bacterial VOCs from control samples.55
Alcoholic VOCs such as 3-methyl-1-butanol have been widely recognized for their plant growth-promoting (PGP) properties. The VOC 3-methyl-1-butanol has been shown to enhance the growth of Medicago sativa and Arabidopsis thaliana under iron-limited conditions.24 It has been reported that bacterial strains isolated from agave and cactus roots emitted 3-methyl-1-butanol, which positively influenced the growth and maturation of several Agave species.25 The 3-methyl-1-butanol produced by rhizospheric bacteria was reported to increase superoxide dismutase (SOD) activity, elevate chlorophyll levels and promote the growth of maize and wheat.26 It has also been identified as a strong inducer of plant growth and systemic resistance,5 regulate lateral root formation via jasmonic acid signalling without activating auxin-responsive pathways, thereby enhancing avocado growth and productivity.27 Further confirmation of the PGP potential of 3-methyl-1-butanol by Ayuso Calles et al.28 showed that the VOCs emitted by Rhizobium sp. GPTR29 increased plant development under controlled, salt-stress, and greenhouse conditions.
The alcoholic VOC 2-methyl-1-butanol has been well documented for its plant growth-promoting and antimicrobial properties by Ayuso Calles et al.28 who identified 2-methyl-1-butanol as one of the major compounds associated with growth promotion and the induction of systemic resistance in plants. Several 1-butanol derivatives, including 3-methyl-1-butanol, 2-methyl-1-butanol, and 1-butanol-3-methyl acetate, have been reported to inhibit the mycelial growth of Pythium ultimum, Rhizoctonia solani, and Sclerotium sclerotiorum44,45 demonstrated that VOCs emitted by Fusarium oxysporum, including 3-methyl-1-butanol and 2-methyl-1-butanol, effectively suppressed mycelial growth and reduced the microsclerotia viability of fungal pathogens. In addition, the pathogenic fungus Ceratocystis fimbriata has been shown to be inhibited by Pseudomonas chlororaphis subsp. aureofaciens, which emits both 3-methyl-1-butanol and 2-methyl-1-butanol. It has been reported that VOCs such as 2-methyl-1-butanol, 1-decene, and 2-heptylfuran produced by Trichoderma significantly increased the fresh weight and chlorophyll content of Arabidopsis thaliana.47
The alcoholic VOC 2-ethyl-1-hexanol has been previously reported. The VOC from Bacillus sp. JC03, which emitted relatively high levels of 2-ethyl-1-hexanol, tetrahydrofuran-3-ol, and 2-heptanone, all of which contributed to enhanced plant growth.41 Further demonstrated that 2-ethyl-1-hexanol exhibited strong antifungal activity against Rhizoctonia solani, indicating its potential as a biological control agent for managing rice sheath blight42 confirmed that 2-ethyl-1-hexanol promoted growth in Arabidopsis thaliana.41 VOCs such as dimethyl silanediol produced by Pseudomonas pseudoalcaligenes can alleviate drought stress in maize by modulating the plants defence system.32
The ketonic compound 2-tridecanone, widely regarded as a plant growth-promoting VOC, has been well characterized in earlier studies, several VOCs emitted by Bacillus sp. BCT9—namely 3-hydroxy-2-butanone, 2,3-butanediol, 2-nonanone, 2-undecanone, 2-tridecanone, and 2-pentadecanon which stimulated seed germination and significantly influenced the growth of Lactuca sativa during both germination and seedling stages. High concentrations of 2-tridecanone in leaf tissues have been associated with plant resistance, producing antibiosis and antixenosis effects against spider mites and other arthropods,33,35-37 It has been reported that 2-tridecanone plays a key role in plant resistance mechanisms against the South American tomato pinworm.39
Pyrazine compounds, such as 2,3,5-trimethylpyrazine, were also detected in the present study. It has been reported that 2,3,5-trimethylpyrazine emitted by Bacillus strain HA and Bacillus strain A8a modulated root development, increased the fresh weight of Arabidopsis thaliana, and inhibited the growth of Fusarium solani, and soil-borne oomycetes. Antimicrobial VOCs produced by Bacillus spp., including 2-decanone, 2,3,5-trimethylpyrazine, and 2-methylbutanoic acid, have demonstrated antagonistic activity against major phytopathogens such as Alternaria alternata, Fusarium solani, Fusarium kuroshium, and Phytophthora cinnamomi.17,30,31 Butane-1-methoxy-3-methyl produced by B. subtilis GB03 and B. amyloliquefaciens IN937a promoted growth and induced systemic acquired resistance in A. thaliana.5 The plant growth ability of acidic compounds such as palmitic acid was proven by Xiong et al.,43 who showed that root exudates from the halophyte Limonium sinense help in the colonization by the Bacillus flexus KLBMP 4941, which promotes host plant growth under salt stress due to the presence of organic acids such as palmitic acid, in the root exudates. Palmitic acid also had positive effects on the growth, motility, chemotaxis, and root colonization by the bacterium.
In the past, much emphasis has been greatly placed on bacterial phytohormones for their role in plant growth promotion, but recent trends have shown that other compounds of bacterial origin such as VOCs, also play a major role in plant growth promotion and disease resistance. Given the relatively limited knowledge on the VOCs produced by vegetable root nodule associated bacteria, there is a pressing need for extensive documentation and further research to decipher their roles. Such an exercise would pave the way for discovering novel aspects of nodule associated bacterial metabolism in relation to plant growth promotion and microbial interactions.
ACKNOWLEDGMENTS
The authors acknowledge the ICAR – Indian Institute of Horticultural Research, Bengaluru, for the laboratory and analytical facilities.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
GS conceived the study. ASB executed the study. AVG, GS and DK supervised the study. ANL and KSS carried out the HS-SPME-GC‒MS analysis. SK carried out the statistical analysis. ASB wrote the manuscript. GS revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies on human participants or animals performed by any of the authors.
- Food and Agriculture Organization of the United Nations. FAOSTAT database. Published 2019. Accessed October 01, 2024. https://www.fao.org/faostat
- Food and Agriculture Organization of the United Nations. FAOSTAT database. Published 2021. Accessed October 01, 2024. https://www.fao.org/faostat
- Velázquez E, Martínez-Hidalgo P, Carro L, et al. Nodular endophytes: an untapped diversity. In: Beneficial Plant–Microbial Interactions: Ecology and Applications. CRC Press;2013:215-235.
- De Meyer SE, De Beuf K, Vekeman B, Willems A. A large diversity of nonrhizobial endophytes found in legume root nodules in Flanders (Belgium). Soil Biol Biochem. 2015;83:1-11.
Crossref - Farag MA, Ryu CM, Summer LW, Pare PW. GC-MS SPME profiling of rhizobacterial volatiles reveals prospective inducers of growth promotion and induced systemic resistance in plants. Phytochemistry. 2006;67(20):2262-2268.
Crossref - Audrain B, Farag MA, Ryu CM, Ghigo JM. Role of bacterial volatile compounds in bacterial biology. FEMS Microbiol Rev. 2015;39(2):222-233.
Crossref - Sharifi R, Ryu CM. Sniffing bacterial volatile compounds for healthier plants. Curr Opin Plant Biol. 2018;44:88-97.
Crossref - Garbeva P, Weisskopf L. Airborne medicine: bacterial volatiles and their influence on plant health. New Phytol. 2020;226(1):32-43.
Crossref - Thomas G, Withall D, Birkett M. Harnessing microbial volatiles to replace pesticides and fertilizers. Microb Biotechnol. 2020;13(5): 1366-1376.
Crossref - Lemfack MC, Gohlke BO, Toguem SMT, Preissner S, Piechulla B, Preissner R. mVOC 2.0: a database of microbial volatiles. Nucleic Acids Res. 2018;46(D1): D1261-D1265.
Crossref - Blom D, Fabbri C, Connor EC, et al. Production of plant growth-modulating volatiles is widespread among rhizosphere bacteria and strongly depends on culture conditions. Environ Microbiol. 2011;13(11): 3047-3058.
Crossref - Ryu CM, Farag MA, Hu CH, et al. Bacterial volatiles promote growth in Arabidopsis. Proc Natl Acad Sci U S A. 2003;100(8):4927-4932.
Crossref - Zhang H, Xie X, Kim MS, et al. Soil bacteria augment Arabidopsis photosynthesis by decreasing glucose sensing and abscisic acid levels in planta. Plant J. 2008;56(2): 264-273.
Crossref - Zhang H, Sun Y, Xie X, Kim M, Dowd SE, Paré PW. A soil bacterium regulates plant acquisition of iron via deficiency-inducible mechanisms. Plant J. 2009;58(4):568-577.
Crossref - Meldau DG, Meldau S, Hoang LH, Underberg S, Wunsche H, Baldwin IT. Dimethyl disulfide produced by the naturally associated bacterium Bacillus sp B55 promotes Nicotiana attenuata growth by enhancing sulfur nutrition. Plant Cell. 2013;25(7):2731-2747.
Crossref - Vespermann A, Kai M, Piechulla B. Rhizobacterial volatiles affect the growth of fungi and Arabidopsis thaliana. Appl Environ Microbiol. 2007;73(17): 5639-5641.
Crossref - Asari S, Matzén S, Petersen MA, Bejai S, Meijer J. Multiple effects of Bacillus amyloliquefaciens volatile compounds. FEMS Microbiol Ecol. 2016;92.
Crossref - Zechman JM, Aldinger S, Labows JN Jr. Characterization of pathogenic bacteria by automated headspace concentration–gas chromatography. J Chromatogr B. 1986;377:49-57.
Crossref - Labows JN, McGinley KJ, Webster GF, Leyden JJ. Headspace analysis of volatile metabolites of Pseudomonas aeruginosa by gas chromatography–mass spectrometry. J Clin Microbiol. 1980;12(4):521-526.
Crossref - de V. Facundo HV, Garruti DS, Cordenunsi BR, Lajolo FM. Isolation of volatile compounds in banana by HS-SPME. Int J Biosci Biochem Bioinform. 2013;3(2):110-115.
Crossref - Sarath Babu A, Selvakumar G, Gopal AV, Kalaivanan D. Isolation and determination of plant growth promotion traits of non-nodulating root nodule-associated bacteria in vegetable legumes. Andhra Agric J. 2023;70(4):479-485.
- Pang Z, Chong J, Zhou G, et al. MetaboAnalyst 5.0: narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 2021;49(W1): W388-W396.
Crossref - Oliveros JC. Venny: an interactive tool for comparing lists with Venn’s diagrams. Published 2015. Accessed 2024. https://bioinfogp.cnb.csic.es/tools/venny/
- Kong WL, Wang YH, Wu XQ. Enhanced Iron uptake in plants by volatile emissions of Rahnella aquatilis JZ-GX1. Front Plant Sci. 2021;12:704000.
Crossref - Camarena-Pozos DA, Flores-Núñez VM, López MG, López-Bucio J, Partida-Martínez LP. Smells from the desert: Microbial volatiles that affect plant growth and development of native and non-native plant species. Plant Cell Environ. 2018;42(4):1368-1380.
Crossref - Li CY, Zhang JJ, Zhao CJ, et al. Effects of 3-methyl-1-butanol on seed germination and seedling growth of maize and wheat. Bull Bot Res. 2018;38(5):785-789.
Crossref - Gamboa-Becerra R, Desgarennes D, Molina-Torres J, et al. Plant growth-promoting and non-promoting rhizobacteria from avocado trees differentially emit volatiles that influence growth of Arabidopsis thaliana. PROTOPLASMA. 2021;259(4):835-854.
Crossref - Ayuso-Calles M, Flores-Félix JD, Amaro F, et al. Effect of Rhizobium mechanisms in improving tolerance to saline stress in lettuce plants. Chem Biol Technol Agric. 2023;10(1):89.
Crossref - Cortazar-Murillo EM, Méndez-Bravo A, Monribot-Villanueva JL, et al. Biocontrol and plant growth promoting traits of two avocado rhizobacteria orchestrated by diffusible and volatile compounds. Front Microbiol. 2023;14:1152597.
Crossref - Méndez-Bravo A, Cortazar-Murillo EM, Guevara-Avendaño E, et al. Plant growth-promoting rhizobacteria associated with avocado display antagonistic activity against Phytophthora cinnamomi through volatile emissions. PLoS One. 2018;13(3): e0194665.
Crossref - Wang D, Li Y, Yuan Y, et al. Identification of non-volatile and volatile organic compounds produced by Bacillus siamensis LZ88 and their antifungal activity against Alternaria alternata. Biol Control. 2022;169:104901.
Crossref - Yasmin H, Rashid U, Hassan MN, et al. Volatile organic compounds produced by Pseudomonas pseudoalcaligenes alleviated drought stress by modulating defense system in maize (Zea mays L.). Physiologia Plantarum. 2020;172(2):896-911.
Crossref - Williams WG, Kennedy GG, Yamamoto RT, Thacker JD, Bordner J. 2-Tridecanone: a naturally occurring insecticide from the wild tomato Lycopersicon hirsutum f. glabratum. Science. 1980;207(4433): 888-889.
Crossref - Fincheira P, Parra L, Mutis A, Parada M, Quiroz A. Volatiles emitted by Bacillus sp. BCT9 act as growth modulating agents on Lactuca sativa seedlings. Microbiol Res. 2017;203:47-56.
Crossref - Fery RL, Kennedy GG. Genetic analysis of 2-tridecanone concentration, leaf trichome characteristics, and tobacco hornworm resistance in tomato. J Amer Soc Hort Sci. 1987;112(5):886-891.
- Weston PA, Johnson DA, Burton HT, Snyder JC. Trichome secretion composition, trichome densities, and spider mite resistance of ten accessions of Lycopersicon hirsutum. J Am Soc Hortic Sci. 1989;114(3):492-498.
- Eigenbrode SD, Trumble JT. Antibiosis to beet armyworm (Spodoptera exigua) in Lycopersicon Accessions. HortScience. 1993;28(9):932-934.
Crossref - Eigenbrode SD, Trumble JT, Jones RA. Resistance to beet armyworm, hemipterans, and Liriomyza spp. in Lycopersicon accessions. J Am Soc Hortic Sci. 1993;118(4):525-530.
Crossref - Maluf WR, Barbosa LV, Santa-Cecília LVC. 2-Tridecanone-mediated mechanisms of resistance to the South American tomato pinworm Scrobipalpuloides absoluta (Meyrick, 1917) (Lepidoptera-Gelechiidae) in Lycopersicon spp. Euphytica. 1997;93(2):189-194.
Crossref - Fiddaman PJ, Rossall S. Effect of substrate on the production of antifungal volatiles from Bacillus subtilis. J Appl Bacteriol. 1994;76:395-405.
Crossref - Jiang CH, Xie YS, Zhu K, et al. Volatile organic compounds emitted by Bacillus sp. JC03 promote plant growth through the action of auxin and strigolactone. Plant Growth Regul. 2019;87:317-328.
Crossref - Wang E, Liu X, Si Z, et al. Volatile organic compounds from rice rhizosphere bacteria inhibit growth of Rhizoctonia solani. Agriculture. 2021;11(4):368.
Crossref - Xiong YW, Li XW, Wang TT, et al. Root exudates-driven rhizosphere recruitment of Bacillus flexus KLBMP 4941 and its growth-promoting effect on Limonium sinense under salt stress. Ecotoxicol Environ Saf. 2020;194: 110374.
Crossref - Fialho MB, Ferreira LFR, Monteiro RTR, Pascholati SF. Antimicrobial volatile organic compounds affect morphogenesis-related enzymes in Guignardia citricarpa, causal agent of citrus black spot. Biocontrol Sci Technol. 2011;21(7):797-807.
Crossref - Strobel GA, Dirkse E, Sears J, Markworth C. Volatile antimicrobials from Muscodor albus, a novel endophytic fungus. Microbiology. 2001;147(11): 2943-2950.
Crossref - Mulero-Aparicio A, Cernava T, Turrà D, et al. The role of volatile organic compounds and rhizosphere competence in mode of action of the nonpathogenic Fusarium oxysporum FO12 toward Verticillium wilt. Front Microbiol. 2019;10:1808.
Crossref - Lee S, Behringer G, Hung R, Bennett J. Effects of fungal volatile organic compounds on Arabidopsis thaliana growth and gene expression. Fungal Ecol. 2019;37:1-9.
Crossref - Sánchez AC, Gutiérrez RT, Santana RC, et al. Effects of co-inoculation of native Rhizobium and Pseudomonas strains on growth parameters and yield of two contrasting Phaseolus vulgaris L. genotypes under Cuban soil conditions. Eur J Soil Biol. 2014;62:105-112.
Crossref - Leite J, Fischer D, Rouws LFM, et al. Cowpea nodules harbor nonrhizobial bacterial communities that are shaped by soil type rather than plant genotype. Front Plant Sci. 2017;7:2064.
Crossref - Wang K, Lin Z, Dou J, Jiang M, Shen N, Feng J. Identification and surveys of plant growth VOCs from Biocontrol Bacteria Paenibacillus peoriae GXUN15128. Microbiol Spectr. 2023;11(3):e0434622.
Crossref - Chandrasekaran M, Paramasivan M, Sahayarayan JJ. Microbial volatile organic compounds: an alternative for chemical fertilizers in sustainable agriculture development. Microorganisms. 2023;11(1):42.
Crossref - Mhlongo MI, Piater LA, Dubery IA. Profiling of volatile organic compounds from four plant growth-promoting rhizobacteria by SPME–GC–MS: a metabolomics study. Metabolites. 2022;12(8): 763.
Crossref - Márquez DGP, Munaro D, Uarrota VG. Metabolomics as a tool to study volatile organic compounds produced by plant growth-promoting rhizobacteria. In: Sayyed, R.Z., Uarrota, V.G. (eds) Secondary Metabolites and Volatiles of PGPR in Plant-Growth Promotion. Springer, Cham. 2022;21-44.
Crossref - Azizan KA, Baharum SN, Ressom HW, Noor NM. GC-MS analysis and PLS-DA validation of the trimethyl silyl-derivatization techniques. Am J Appl Sci. 2012;9(7): 1124-1136.
Crossref - Kate A, Tiwari S, Gujar JP, et al. Spotting of volatile signatures through GC-MS analysis of bacterial and fungal infections in stored potatoes (Solanum tuberosum L.). Foods. 2023;12(10):2083.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.