ISSN: 0973-7510
E-ISSN: 2581-690X
Multivariate statistical models were utilized to identify the interaction between the inhibitory activity and the molecular properties of the different antimycotics against Microsporum canis. Information on the inhibitory potency against M. canis and molecular properties of antifungal agents were obtained from literature. The relationship between the inhibitory potency and the molecular properties of the different antimycotics against M. canis was established using multiple linear regression analysis (MLRA) and principal component analysis (PCA). Three major descriptors: topological polar surface area, molecular weight, and rotatable bond count of the antimycotics were identified to confer inhibitory action against M. canis using MLRA (r2=0.8968, p<0.0001) and PCA (95.86% total contribution rate). Both MLRA and PCA as statistical approaches demonstrate their potential as tools in computational structure design and for possible synthesis of next generation antimycotics as more effective treatments of fungal infections.
dermatophytes, fungal infections, molecular descriptors, multivariate data analysis, regression equation.
Superficial fungal infections, which are commonly caused by dermatophytes including Microsporum canis, are treated with topical antimycotics, but resistance to these topical agents was observed in cases of tinea unguium and tinea capitis which necessitate the use of systemic antifungal drugs.1-3 Molecular properties such as topological polar surface area, heavy atom count, hydrogen bond acceptor and donor counts, rotatable bond count, complexity, lipophilicity, and molecular weight were important descriptors in assessing the inhibitory effect of drugs. Since statistical modelling is a very important tool in predicting biological activities of compounds4, antifungal drugs against M. canis were statistically evaluated using multiple linear regression analysis (MLRA) and principal component analysis (PCA). In drug design, establishing the relationship between the activity and the properties of a compound is of utmost importance.5 The generated relationship will likewise unveil the relevant molecular properties of the existing antifungal drugs that would account for their inhibitory action against M. canis. Findings of the investigation would offer additional insights relevant to the synthesis of new derivatives of antimycotics as more effective treatments of dermatophytoses.
Inhibitory Potency of Antifungal Drugs
The biological activity of 16 different antifungal drugs against M. canis was obtained from literature.6 Six of these agents which are systemic drugs include voriconazole, terbinafine, ketoconazole, griseofulvine, fluconazole, and itraconazole. The 10 topical agents were amorolfine, butenafine, clotrimazole, econazole, miconazole, naftifine, tioconazole, tolciclate, tolnaftate, and undecylenic acid.
Molecular Descriptors of Antifungal Drugs
Information on the molecular descriptors of the 16 drugs (Table 1) were retrieved from NCBI (National Center for Biotechnology Information) databases7-22, and these include topological polar surface area (tpsa), logarithm of molecular weight (logmw), heavy atom count (hac), lipophilicity (xlogp), logarithm of complexity (logcomp), hydrogen bond acceptor count (hbac), rotatable bond count (rbc), and hydrogen bond donor count (hbdc).
Multiple Linear Regression Analysis and Principal Component Analysis
For the model construction, quantitative relationship between the inhibitory activity of antimycotics and their molecular properties was established using MLRA. This multivariate statistical approach is a computational method frequently utilized in drug design owing to its simplicity and reproducibility.23 In variable selection, backward elimination was employed in the construction of the regression model. The adequacy of the obtained regression model was evaluated using the computed r2, squared correlation coefficient24. In addition, bivariate correlation studies examined the presence of outliers, linearity, normality, independence, homoskedasticity, and multicollinearity among the significant molecular predictors.
To assess the similarity among the antimycotics and to identify possible correlations in the different drug properties, principal component analysis was employed. Using this multivariate statistical approach, information regarding the data set is summarized into principal components where each component contains uncorrelated linear combinations of variables having the maximum variance.25 Among correlated variables, linear transformation was performed and these transformed variables were then sorted in order of decreasing variance. In the present study, principal components with loading eigenvalues greater than 1.0 were included. An absolute value correlation coefficient of at least 0.39 between the index and its principal component retains the molecular property within the given component. Data analyses were performed using STATA® V12.0 at 0.05 significance level.
The computed loading values of the three principal components have contribution rates of 50.30%, 32.55%, and 13.01% (Table 2). Complexity, heavy atom and hydrogen bond acceptor counts, molecular weight, and topological polar surface area were in the 1st principal component. Lipophilicity and hydrogen bond donor count were in the 2nd component, while rotatable bond count was in the 3rd principal component. When PCA technique was used in assessing the inhibitory effect of the 16 antimycotics, comprehensive scores (range: 6.62-34.93) were obtained (Table 3). Relatively higher comprehensive scores were observed in systemic antifungal drugs compared with topical antimycotics.
Table (1):
Descriptors of the different antimycotics.
| Antimycotic | Properties | Reference | |||||||
|---|---|---|---|---|---|---|---|---|---|
| tpsa | logmw | hac | xlogp | logcomp | hbac | hbdc | rbc | ||
| Amorolfine | 12.5 | 2.502 | 23 | 5.7 | 2.526 | 2 | 0 | 6 | 7 |
| Butenafine | 3.2 | 2.502 | 24 | 6.3 | 2.573 | 1 | 0 | 5 | 8 |
| Clotrimazole | 17.8 | 2.538 | 25 | 5 | 2.598 | 1 | 0 | 4 | 9 |
| Econazole | 27 | 2.582 | 24 | 5.3 | 2.579 | 2 | 0 | 6 | 10 |
| Fluconazole | 81.6 | 2.486 | 22 | 0.4 | 2.554 | 7 | 1 | 5 | 11 |
| Griseofulvin | 71.1 | 2.547 | 24 | 2.2 | 2.76 | 6 | 0 | 3 | 12 |
| Itraconazole | 101 | 2.849 | 49 | 5.7 | 3.049 | 9 | 0 | 11 | 13 |
| Ketoconazole | 69.1 | 2.725 | 36 | 4.3 | 2.866 | 6 | 0 | 7 | 14 |
| Miconazole | 27 | 2.619 | 25 | 5.3 | 2.62 | 2 | 0 | 6 | 15 |
| Naftifine | 3.2 | 2.458 | 22 | 5.1 | 2.534 | 1 | 0 | 5 | 16 |
| Terbinafine | 3.2 | 2.465 | 22 | 5.6 | 2.631 | 1 | 0 | 5 | 17 |
| Tioconazole | 55.3 | 2.588 | 23 | 5.3 | 2.579 | 3 | 0 | 6 | 18 |
| Tolciclate | 44.6 | 2.51 | 23 | 5.5 | 2.656 | 2 | 0 | 3 | 19 |
| Tolnaftate | 44.6 | 2.488 | 22 | 5.5 | 2.587 | 2 | 0 | 3 | 20 |
| Undecylenic acid | 37.3 | 2.265 | 13 | 3.9 | 2.149 | 2 | 1 | 9 | 21 |
| Voriconazole | 76.7 | 2.543 | 25 | 1.5 | 2.651 | 8 | 1 | 5 | 22 |
Table (2):
Principal component loading values and eigenvectors.
Antimycotic Property |
1stComponent |
2ndComponent |
3rdComponent |
|---|---|---|---|
logmw |
0.4470 |
-0.2337 |
-0.0616 |
xlogp |
-0.0860 |
-0.5646 |
0.2922 |
logcomp |
0.4369 |
-0.2103 |
-0.2980 |
hbac |
0.4218 |
0.3192 |
-0.0534 |
hbdc |
-0.0401 |
0.5843 |
0.2351 |
rbc |
0.2284 |
0.0147 |
0.8643 |
tpsa |
0.3966 |
0.3152 |
-0.0775 |
hac |
0.4615 |
-0.1988 |
0.1043 |
Proportion |
0.5030 |
0.3255 |
0.1301 |
Cumulative |
0.5030 |
0.8285 |
0.9586 |
Eigenvalue |
4.0242 |
2.6041 |
1.0408 |
Table (3):
Comprehensive scores of the antimycotics using principal component analysis.
| Antimycotic | Comprehensive evaluation | Rank | |||
|---|---|---|---|---|---|
| 1stComponent | 2ndComponent | 3rdComponent | Comprehensive Score | ||
| Itraconazole | 69.07 | -3.22 | 9.51 | 34.93 | 1 |
| Fluconazole | 47.70 | 0.36 | 4.32 | 24.67 | 2 |
| Ketoconazole | 49.02 | -2.43 | 6.05 | 24.65 | 3 |
| Voriconazole | 47.63 | -0.26 | 4.32 | 24.43 | 4 |
| Griseofulvin | 44.15 | -1.24 | 2.59 | 22.14 | 5 |
| Tioconazole | 36.10 | -2.99 | 5.19 | 17.86 | 6 |
| Tolciclate | 31.43 | -3.11 | 2.59 | 15.14 | 7 |
| Tolnaftate | 30.93 | -3.11 | 2.59 | 14.88 | 8 |
| Miconazole | 25.40 | -2.99 | 5.19 | 12.48 | 9 |
| Undecylenic acid | 23.59 | -1.62 | 7.78 | 12.35 | 10 |
| Econazole | 24.91 | -2.99 | 5.19 | 12.23 | 11 |
| Clotrimazole | 21.29 | -2.82 | 3.46 | 10.24 | 12 |
| Amorolfine | 18.64 | -3.22 | 5.19 | 9.00 | 13 |
| Butenafine | 15.01 | -3.56 | 4.32 | 6.95 | 14 |
| Naftifine | 14.05 | -2.88 | 4.32 | 6.69 | 15 |
| Terbinafine | 14.10 | -3.16 | 4.32 | 6.62 | 16 |
Statistical evaluation of the molecular descriptors of the antifungal drugs showed that molecular weight exhibited a positive correlation with heavy atom count, hydrogen bond acceptor count, and topological polar surface area (Table 4). Topological polar surface area and heavy atom count were directly correlated with hydrogen bond acceptor count. Among the antifungal agents, positive correlation between the numbers of heavy atom and rotatable bond exists. Complexity of the antimycotics was positively correlated with hydrogen bond acceptor and heavy atom counts, topological polar surface area, and molecular weight. However, lipophilicity of the antimycotics was negatively correlated with hydrogen bond donor count, hydrogen bond acceptor count, and topological polar surface area.
Table (4):
Correlational analysis on the molecular descriptors of antimycotics.
logmw |
xlogp |
logcomp |
rbc |
hbac |
hbdc |
tpsa |
hac |
|
|---|---|---|---|---|---|---|---|---|
logmw |
1.000 |
0.1561 |
0.9165** |
0.3466 |
0.5547* |
-0.4352 |
0.5127* |
0.9323** |
xlogp |
0.1561 |
1.000 |
0.0621 |
0.1431 |
-0.6285** |
-0.7601** |
-0.5768* |
0.1609 |
hbdc |
-0.4352 |
-0.7601** |
-0.4498 |
0.1797 |
0.4013 |
1.000 |
0.3674 |
-0.3298 |
hbac |
0.5547* |
-0.6285** |
0.5891* |
0.3531 |
1.000 |
0.4013 |
0.9294** |
0.6122* |
rbc |
0.3466 |
0.1431 |
0.1261 |
1.000 |
0.3531 |
0.1797 |
0.2996 |
0.5033* |
tpsa |
0.5127* |
-0.5768* |
0.5190* |
0.2996 |
0.9294** |
0.3674 |
1.000 |
0.5368 |
hac |
0.9323** |
0.1609 |
0.9007** |
0.5033* |
0.6122* |
-0.3298 |
0.5368 |
1.000 |
logcomp |
0.9165** |
0.0621 |
1.000 |
0.1261 |
0.5891* |
-0.4498 |
0.5190* |
0.9007** |
**1%, *5% level of significance
When preliminary analysis using stepwise backward elimination of MLRA was employed, heavy atom count, complexity, hydrogen bond acceptor and donor counts, and lipophilicity were removed from the full model. Topological polar surface area, molecular weight, and rotatable bond count were retained as significant independent variables accounting in the variation on the biological activity of these antimycotics against M. canis. For every independent variable included in the regression model, a minimum of five samples is required.24,26 Since there were only 16 available antimycotics in this study, the constructed regression equation included at most three molecular descriptors. The generated regression model (Eq. 1) has identified three predictors and is adequate (r2=0.8968, p=0.0000) in explaining the potency values (MIC, minimum inhibitory concentration) of the antimycotics against M. canis. The three descriptors were tpsa (topological polar surface area, p=0.005), logmw (molecular weight, p=0.000), and rbc (rotatable bond count, p=0.000), with rotatable bond count and molecular weight accounting to 79.65% of the fluctuations in the response variable (MIC). The negative coefficient of logmw suggests that increasing the molecular weight lowers MIC, thus improving the antimycotic potency. The positive coefficients for tpsa and rbc indicate that MIC increases as topological polar surface area and rotatable bond count increase.
MIC = 301.33(±34.55) – 130.68(±14.31) logmw + 4.99(±0.76) rbc + 0.19(±0.06)tpsa …(Eq.1)
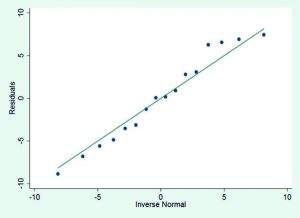
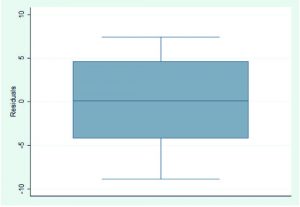
After identifying the significant molecular descriptors, various assumptions of MLRA were examined. A large F-value, F(3,12)=34.74 (p=0.0000), would indicate that the constructed regression function satisfied the requirement for linearity. Independence of the error terms was likewise met (z=-0.52, p=0.60). The linear plot obtained between the inverse normal values and the residuals (Fig. 1) suggested normally distributed error terms (p=0.482), while test on homoskedasticity identified constant variance (p=0.936). The generated boxplot (Fig. 2) showed nonexistence of an outlier. Multicollinearity among significant molecular descriptors was not present since variance inflation factor (VIF) values were all less than 10 (logmw, 1.44; tpsa, 1.39; rbc, 1.16).
The effects of other molecular descriptors obtained from the statistical assessments as regard with the antifungal activity of the drugs were correlated with the included significant properties in the regression equation (Table 4). Apparently, positive correlations exist between molecular weight and hydrogen bond acceptor count (r=0.5547, p<0.05), heavy atom count (r=0.9323, p<0.01), and complexity (r=0.9165, p<0.01). Moreover, rotatable bond count is directly correlated with heavy atom count (r=0.5033, p<0.05) while topological polar surface area is inversely related with lipophilicity (r=-0.5768, p<0.05), but directly related with hydrogen bond acceptor count (r=0.9294, p<0.01) and complexity (r=0.5190, p<0.05).
Molecular and structural properties of compounds were assessed based on their hydrogen bond acceptor and donor counts, molecular weight, complexity, rotatable bond and heavy atom counts, lipophilicity, and topological polar surface area. The present study identified molecular weight, rotatable bond count, and topological polar surface area as significant properties of antifungal drugs which confer inhibitory action against dermatophytoses particularly caused by Microsporum canis. In drug design, r2 is a useful determinant of model validity24 and a value of at least 0.6 is considered satisfactory27. In the present study, the regression equation is adequate to capture the functional relationship between the inhibitory activity and molecular properties of the antimycotics against M. canis.
Evaluation of the various descriptors of the antimycotics against M. canis revealed that molecular weight exhibited a positive correlation with heavy atom count, complexity, topological polar surface area, and hydrogen bond acceptor count. Molecular weight is a very important property especially in small molecule drug discovery28,29 because it affects several pharmacologic events30. Molecular weight of the antimycotics was directly correlated with molecular complexity and these molecular descriptors were identified as significant properties included in the first principal component. Molecular complexity is positively correlated with heavy atom and hydrogen bond acceptor counts, and accounts for various properties such as carbon hybridization, chirality, and aromaticity. Aromaticity is directly related with lipophilicity and albumin affinity.31 Aromatic molecules with more than three rings had poor compound developability and were associated with increased toxicity risks.32 In the selection of lead compounds, moderately complex structures are preferred because of their binding specificity33 and are associated with pharmacologic properties34 and biological activity35. Furthermore, topological polar surface area has been identified as one of the significant molecular descriptors in the regression equation and in the second principal component. This molecular property, defined as the total polar atom surfaces, has correlation with drug transport property, permeability, and absorption.36 Complexity and hydrogen bond acceptor count are directly related with topological polar surface area. Topological polar surface area and rotatable bond count influence molecular complexity and compound bioavailability.37
There are five classes of available antimycotics intended as treatment for fungal infections: allylamines, azoles, echinocandins, nucleoside analogs, and polyenes.38 In the present study, antimycotics were classified as either a systemic drug or a topical agent. The use of these antifungal drugs depends on the sensitivity profile of species, anatomical site it was found, and the type of infection.39 Echinocandins such as anidulafungin, caspofungin, and micafungin are considered as a new class of drugs exhibiting fungicidal effects.40 The mechanism of action of echinocandins prevents glucan synthesis41 which renders inability of fungal cell wall to withstand osmotic stress42. On the other hand, thiocarbamates and allylamines, such as naftifine and terbinafine are antifungals which rupture fungal cell membrane and promote accumulation of squalene which inhibits the synthesis of ergosterol.41 Terbinafine is the most potent systemic drug against dermatophytoses.6 In addition, allylamines also restrict sterol derivatives production.39 Another class of antimycotics are polyenes, which include amphotericin B, which are responsible in the leakage of intracellular constituents including magnesium, sugars, and potassium which disrupt the fungal membrane structure.43 In spite of its effectivity, amphotericin B has high toxicity restricting its use compared to azoles.
Finally, azoles are semi-synthetic and synthetic compounds with broad spectrum of activity and are classified either as triazoles (itraconazole, fluconazole, and voriconazole) or imidazoles (clotrimazole, econazole, ketoconazole, and miconazole).39,44 Triazoles are intended for systemic and superficial mycoses.45 Despite structural differences between triazoles and imidazoles, imidazoles have similar mechanism of action as triazoles.46 Imidazoles, except for ketoconazole, are used in treating superficial infections.44 In the present study, azoles intended as systemic drugs against dermatophytoses were classified to have higher comprehensive scores using hierarchical weighted principal component analysis when compared with azoles as topical agents. Azoles affect fungal cell growth and proliferation41, but there are limitations on the utility of these antifungal drugs because of hepatotoxicity and resistance47. In general, mucosal or superficial mycoses can usually be successfully treated by imidazole topical agents48 although triazoles have a broader application in the therapeutic management of both invasive and superficial mycoses49.
Computational approach was employed in classifying antimycotics and in establishing functional relationship between their inhibitory activity and molecular properties. Topological polar surface area, molecular weight, and rotatable bond count were identified as significant molecular descriptors of these antifungal drugs conferring their inhibitory action against dermatophytoses caused by Microsporum canis. Increase in the molecular weight and decrease in the rotatable bond count and topological polar surface area of the antimycotics will render better potency.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct, and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets analyzed in the study are included in the manuscript and presented as tables.
- Roberts DT. Onychomycosis: current treatment and future challenges. Br. J. Dermatol., 1999; 141(Suppl. 56): 1-4.
Crossref - Del Rosso JQ. Current management of onychomycosis and dermatomycoses. Curr. Infect. Dis. Rep., 2000; 2: 438-445.
Crossref - Gupta AK, Adam P, Dlova N, Lynde CW, Hofstader S, Morar N, Aboobaker J, Summerbell RC. Therapeutic options for the treatment of tinea capitis caused by Trichophyton species: griseofulvin versus the new oral antifungal agents, terbinafine, itraconazole, and fluconazole. Pediatr. Dermatol., 2001; 18: 433-438.
Crossref - Sumalapao DEP, Janairo JIB, Gloriani NG. Dipole moment, solvation energy, and ovality account for the variations in the biological activity of HIV-1 reverse transcriptase inhibitor fragments. Annu. Res. Rev. Biol., 2018; 22: 1-8.
Crossref - Warr WA. 2011. Some trends in chem(o)informatics. In: Chemoinformatics and Computational Chemical Biology, Bajorath J. Ed., Humana Press, New York. 1-38.
Crossref - Favre B, Hofbauer B, Hildering K, Ryder NS. Comparison of in vitro activities of 17 antifungal drugs against a panel of 20 dermatophytes by using a microdilution assay. J. Clin. Microbiol., 2003; 41(10): 4817-4819.
Crossref - National Center for Biotechnology Information. 2019a. PubChem Database. Amorolfine, CID=54260, https://pubchem.ncbi.nlm.nih.gov/compound/Amorolfine (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019b. PubChem Database. Butenafine, CID=2484, https://pubchem.ncbi.nlm.nih.gov/compound/Butenafine (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019c. PubChem Database. Clotrimazole, CID=2812, https://pubchem.ncbi.nlm.nih.gov/compound/Clotrimazole (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019d. PubChem Database. Econazole, CID=3198, https://pubchem.ncbi.nlm.nih.gov/compound/Econazole (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019e. PubChem Database. Fluconazole, CID=3365, https://pubchem.ncbi.nlm.nih.gov/compound/Fluconazole (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019f. PubChem Database. Griseofulvin, CID=441140, https://pubchem.ncbi.nlm.nih.gov/compound/Griseofulvin (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019g. PubChem Database. Itraconazole, CID=3793, https://pubchem.ncbi.nlm.nih.gov/compound/Itraconazole (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019h. PubChem Database. Ketoconazole, CID=47576, https://pubchem.ncbi.nlm.nih.gov/compound/Ketoconazole (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019i. PubChem Database. Miconazole, CID=4189, https://pubchem.ncbi.nlm.nih.gov/compound/Miconazole (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019j. PubChem Database. Naftifine, CID=47641, https://pubchem.ncbi.nlm.nih.gov/compound/Naftifine (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019k. PubChem Database. Terbinafine, CID=1549008, https://pubchem.ncbi.nlm.nih.gov/compound/Terbinafine (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019l. PubChem Database. Tioconazole, CID=5482, https://pubchem.ncbi.nlm.nih.gov/compound/Tioconazole (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019m. PubChem Database. Tolciclate, CID=5506, https://pubchem.ncbi.nlm.nih.gov/compound/Tolciclate (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019n. PubChem Database. Tolnaftate, CID=5510, https://pubchem.ncbi.nlm.nih.gov/compound/Tolnaftate (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019o. PubChem Database. Undecylenic acid, CID=5634, https://pubchem.ncbi.nlm.nih.gov/compound/Undecylenic-acid (accessed on Dec. 12, 2019)
- National Center for Biotechnology Information. 2019p. PubChem Database. Voriconazole, CID=71616, https://pubchem.ncbi.nlm.nih.gov/compound/Voriconazole (accessed on Dec. 12, 2019)
- Snedecor GW, Cochran WG. 1967. Statistical methods. Oxford and IBH. New Delhi, India.
- Leach AR, Gillet VJ. 2007. Computational models. In: An introduction to cheminformatics, revised edition, Springer, The Netherlands. 75-97.
Crossref - Hastie T, Tibshirani R, Friedman J. 2009. Principal components. In: The elements of statistical learning. 2nd edition. Springer, 534-541.
Crossref - Tabachnick BG, Fidell LS. 2007. Multiple regression. In: Using multivariate statistics. 5th edition, London: Pearson/Allyn & Bacon. 117-194.
- Golbraikh A, Tropsha A. Beware of q2. J. Mol. Graph. Model., 2002; 20: 269-276.
Crossref - Meanwell NA. Improving drug candidates by design: a focus on physicochemical properties as a means of improving compound disposition and safety. Chem. Res. Toxicol., 2011; 24(9): 1420-1456.
Crossref - Leeson PD. Molecular inflation, attrition and the rule of five. Adv. Drug Deliv. Rev., 2016; 101: 22-33.
Crossref - Gleeson MP. Generation of a set of simple, interpretable ADMET rules of thumb. J. Med. Chem., 2008; 51: 817-834.
Crossref - Ritchie TJ, Macdonald SJ. The impact of aromatic ring count on compound developability – are too many aromatic rings a liability in drug design? Drug Discov. Today, 2009; 14(21-22): 1011-1020.
Crossref - Lovering F, Bikker J, Humblet C. Escape from flatland: increasing saturation as an approach to improving clinical success. J. Med. Chem., 2009; 52: 6752-6756.
Crossref - Hann MM, Leach AR, Harper G. Molecular complexity and its impact on the probability of finding leads for drug discovery. J. Chem. Inf. Comput. Sci., 2001; 41: 856-864.
Crossref - Lovering F. Escape from Flatland 2: complexity and promiscuity. Med. Chem. Comm., 2013; 515-519.
Crossref - Hann MM, Oprea TI. Pursuing the lead likeness concept in pharmaceutical research. Curr. Opin. Chem. Biol., 2004; 8: 225-263.
Crossref - Ertl P, Rohde B, Selzer P. Fast calculation of molecular polar surface area as a sum of fragment based contributions and its application to the prediction of drug transport properties. J. Med. Chem., 2000; 43: 3714-3717.
Crossref - Veber DF, Johnson SR, Cheng HY, Smith BR, Ward KW, Kopple KD. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem., 2002; 45(12): 2615-2623.
Crossref - Trevor AJ, Katzung BG, Kruidering-Hall MM, Masters SB. 2013. Chapter 48: Antifungal agents, Katzung & Trevor’s Pharmacology: Examination & board review (10th edition).
- De Oliveira Santos GC, Vasconcelos CC, Lopes AJO, de Sousa Cartagenes M, Do S, Filho AKDB, do Nascimento FRF, de Andrade Monteiro C. Candida infections and therapeutic strategies: mechanisms of action for traditional and alternative agents. Front. Microbiol., 2018; 9:1351
Crossref - Nett JE, Andes DR. Antifungal agents: spectrum of activity, pharmacology, and clinical indications. Infect. Dis. Clin. North Am., 2016; 30(1): 51-83.
Crossref - Lewis, R.E. Current concepts in antifungal pharmacology. Mayo Clin. Proc., 2011; 86(8): 805-817.
Crossref - Chen SC, Sorrell TC. Antifungal agents. Med. J. Aust., 2007; 187(7): 404-409.
Crossref - Mesa-Arango AC, Scorzoni L, Zaragoza O. It only takes one to do many jobs: Amphotericin B as antifungal and immunomodulatory drug. Front. Microbiol., 2012; 3: 286.
Crossref - Gavarkar PS, Adnaik RS, Mohite SK. An overview of azole antifungals. Int. J. Pharm. Sci. Res., 2013; 4(11): 4083-4089.
- Sheehan D, Hitchcock C, Sibley C. Current and emerging azole antifungal agents. Clin. Microbiol. Rev., 1999; 12(1): 40-79.
Crossref - Sud I, Feingold D. Mechanisms of actions of the antimycotic imidazoles. J. Invest. Dermatol., 1981; 76: 438-441.
Crossref - Carrillo-Munoz AJ, Giusiano G, Ezkurra PA, Quindos G. Antifungal agents: mode of action in yeast cells. Rev. Esp. Quimioter., 2006; 19(2): 130-139.
- National Health and Medical Research Council. 1999. A guide to the development, implementation and evaluation of clinical practice guidelines. Canberra: NHMRC.
- Boucher HW, Groll AH, Chiou C, Walsh TJ. Newer systemic antifungal agents. Drugs, 2004; 64: 1997-2020.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.