ISSN: 0973-7510
E-ISSN: 2581-690X
Diseases of the respiratory system are a common cause of antibiotic prescription in Iraq and worldwide. Technology has been recently used for its diagnosis, such as the Film Array Respiratory Panel. This study aims to identify the correlation between the diagnosis and treatment of respiratory tract infections with the result of polymerase chain reaction (PCR) for respiratory viruses. A descriptive, cross-sectional, retrospective study included 134 patients treated at Alkharama Hospital and the Private Hospital in Baghdad, Iraq, in the period from January 2020 to March 2020 For all cases, the results of the panel and the treatment received by the patients were analysed. 58% received antibiotic treatment upon admission, 13% combined treatment (antibiotic + antiviral), 27% received symptomatic treatment, and 2% were treated with the first-instance antiviral. After the result, 38% continued with antibiotics, 30% with antibiotics and antivirals, 13.8% with antivirals and 18.2% with symptomatic treatment. Despite the worldwide alarm over antimicrobial resistance, patients continue to be treated with antibiotics due to a situation that is influenced by several factors.
Respiratory viral infections, PR FilmArray, antibiotics, antimicrobial resistance
Diseases of the respiratory system represent one of the leading causes of medical care in the world. They are common ailments of all ages, whose aetiology varies according to age, environmental circumstances, climate, healthcare setting, and underlying diseases.1,2 Most respiratory infections only affect the upper respiratory tract and can be considered mild, benign, and self-limited (common cold, rhinitis, and pharyngotonsillitis). In Iraq, acute upper respiratory infection is considered the leading cause of illness and a primary reason for seeking medical attention. Respiratory viruses are the main cause in up to 70–90% of cases. Rhinovirus has been documented in adulthood, followed by influenza A and B viruses, coronavirus, and adenovirus; the most common in children are respiratory syncytial virus (RSV), parainfluenza virus 1,2,3, influenza A and B virus, adenovirus and rhinovirus. In a smaller proportion, between 15 and 30% of cases in children and between 5 and 20% in adults, the aetiology is bacterial: Streptococcus pyogenes, Streptococcus pneumoniae, Mycoplasma pneumoniae, Neisseria meningitides, and Neisseria gonorrhoeae.3,4
However, only 5% Of respiratory diseases it is estimated that may involve the lower and middle respiratory tract (bronchitis, bronchiolitis, and pneumonia); they are potentially more severe and, most times, require hospital admission.5 The complications of these diseases in adulthood are related to comorbidities in the patient and senescence.2,6,7 Acute lower respiratory tract infections and other chronic lung diseases are considered among the most common causes of severe illness and death worldwide.8
In previous years, identifying the aetiology of the virus was methodologically difficult, which is why few hospitals followed this approach. In recent years, techniques for diagnosing respiratory tract infections have advanced, currently using the detection of nucleic acids of the virus, with tests such as polymerase chain reaction (PCR), with a sensitivity of 95–100% and 99–100% specificity.9 Because it is common for more than one virus to be involved in these infections, it was necessary to design multiplex PCR methods in which different viruses can be simultaneously identified.10 Other options are PCR coupled to optical enzyme immunoassay, which are systems for detecting amplification products using probes immobilised on a different chemical nature surface.1
The current use of antibiotics, antivirals, and other antimicrobials are increasingly alarming. However, it is known that antimicrobial resistance is a phenomenon that appears naturally over time. This process is accelerated by the inappropriate use of antimicrobials, as well as their excessive prescription. The use of antibiotics is considered unnecessary or inappropriate in up to half of the patients.11-13
It is hoped that when the doctors have studies to identify the possible aetiology of the respiratory infection being treated, they will make more rational use of antibiotics. This study evaluated therapeutic behaviour in a private hospital when a positive result was given in identifying respiratory viruses. The most frequent viral agents and the respiratory pathologies in those who underwent the test are described.
A descriptive, cross-sectional, and retrospective study was conducted. Patients of all ages who had had positive respiratory virus results in a multiplex PCR reaction test carried out in the period from January 2020 to March 2020 were selected from Al-karama hospital – Baghdad, Iraq.
All patients included in the study underwent a respiratory virus PCR test during their hospital stay. The Film Array Respiratory Panel® was used (RP), a qualitative test for the simultaneous detection and identification of multiple nucleic acids of viruses and bacteria in the nasopharyngeal aspirate. A minimum sample volume of 0.3 ml (300 ml) was taken from each patient’s respiratory tract. It is a test that simultaneously identifies the following viruses and bacteria: adenovirus, coronavirus 229E, coronavirus HKU1, coronavirus NL63, coronavirus OC43, influenza A virus (with subtyping for hemagglutinin genes H1, H1-2009 and H3) and influenza B, human metapneumovirus, parainfluenza virus 1, parainfluenza 2, parainfluenza 3, and parainfluenza 4, respiratory syncytial virus, rhinovirus or enterovirus, Bordetella pertussis, Chlamydophila pneumoniae and Mycoplasma pneumoniae.
The sample was introduced into the FilmArray RP cartridge, which was then placed in the FilmArray Module; a test report was automatically generated at the end of the analysis. The entire process takes about an hour. The test has a sensitivity and specificity of 95 and 99%, respectively.14
In each positive case, the following were analysed: the clinical, radiological diagnosis established by the treating physician, the prescribed treatment, the behaviour regarding the treatment that was followed when the test result was obtained, the antimicrobial prescription time and the conditions at the patient’s discharge.
The initial sample consisted of 172 patients of all ages, of which 38 were excluded due to not having complete medical records.
The final sample was 134 patients, of which 49.2% were men and 50.8% were women. Patients of all ages were included: 56% were infants, 16% preschool, 2% schoolchildren, 4% adolescents, and 22% adults.
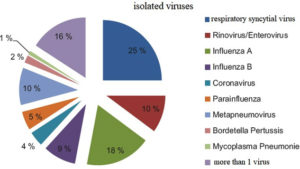
From the results of the viral panel, RSV was identified in 25% of the samples, followed by influenza A (18%), rhinovirus or enterovirus (10%), metapneumovirus (10%), influenza B (9%), parainfluenza (5%), coronavirus (4%), B. pertussis (2%), M. pneumoniae (1%) and in 16% of the samples more than one virus was isolated (Fig. 1).
Fig. 1. Percentage distribution of viruses identified by a polymerase chain reaction in nasopharyngeal samples
Radiological clinical diagnoses in all patients were pneumonia in 42%, 10% with bronchiolitis, 17% with influenza, 8% with bronchitis, asthmatic attacks in 2%, rhinopharyngitis in 2%, 19% with other diagnoses.
Of all the patients, 58% received only antibiotic treatment upon admission, 13% received combined treatment (antibiotic + antiviral), 27% received symptomatic treatment, and 2% were treated first with an antiviral as shown in ( Table 1).
Table (1):
Illustrate medicine received by the patient upon admission to the hospital.
Medication |
Number of patients |
percentage |
|---|---|---|
received only antibiotic |
78 |
58% |
antibiotic + antiviral |
17 |
13% |
symptomatic treatment |
37 |
27% |
Antiviral |
3 |
2% |
Total number of patients |
134 |
Of the patients who initially received antibiotics (71%), on obtaining the result of the positive viral panel, only 10% discontinued this treatment; 2.3% continued only with the antiviral that was administered after their admission. In 1.5%, the antibiotic was suspended and an antiviral was added, and in 6.2%, the antibiotic was suspended, and they were treated symptomatically. A total of 22% added antiviral to the antibiotic treatment they already had and 8% continued with combined treatment; 31% continued with antibiotics only. 12% of the patients continued only with symptomatic treatment, 7% added an antibiotic to their symptomatic treatment after the respiratory panel’s positive result, and 10% added antiviral to the symptomatic treatment.
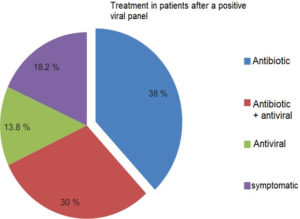
After the positive result, 38% received antibiotics as definitive treatment, 30% were treated with combined therapy (antibiotic + antiviral), 13.8% were treated with antiviral alone, and 18.2% symptomatically (Fig. 2).
Of the 68% of patients who received antibiotics empirically or as definitive treatment, 41% received more than one antibiotic during their hospital stay. As monotherapy or associated with another antibiotic, 45% received a cephalosporin, 17% macrolides, 11% quinolones, and 5% others as shown in (table 2). In 39% of the patients, diagnoses or laboratory studies (such as a culture of bronchial secretion) were found that justified the use of antibiotics.
Table (2):
Illustrate the antibiotic category prescribed to the patient.
Type of antibiotic |
Patient (percentage to total patient) |
Mostly used antibiotic |
|---|---|---|
Cephalosporin |
60(45%) |
Ceftriaxone |
Macrolides |
23(17%) |
azithromycin |
Quinolones |
15(11%) |
Levofloxacin |
Other antibiotic |
7(5%) |
Metronidazole |
The days of hospital stay in patients who received antibiotic treatment despite obtaining a positive viral panel (38%) were reported between 5 to 6 days and 30% who were treated with antibiotics and antivirals. In the 13.8% who received antiviral treatment, the stay ranged from 3 to 4 days. In those who received symptomatic treatment, their hospital stays ranged from 4 to 5 days. There were two deaths (1.5%), one patient required transfer to a high speciality unit, and 131 did not present complications and were discharged due to improvement.
Several studies show a high prevalence of viral agents as the cause of acute respiratory processes in adult and paediatric patients, leading to hospitalisation.15,16
Knowing the aetiology and clinical picture of respiratory diseases, supported by current methods for their diagnosis, should facilitate decision-making on the appropriate treatment for each patient. When the clinical picture adequately correlates with the laboratory findings, the doctor has the elements to support his therapeutic decision.6
This study evaluated how much a positive result of the respiratory panel by PCR influences the treatment that the patients received.
At a global level, reducing the use of antibiotics by having a positive result in a viral panel is variable. In a study in Sweden of patients identified with a virus, 21% were definitively treated with antibiotics.10 In the present study, 71% of the patients received an antibiotic upon admission, and 68% received it as definitive treatment, even though they subsequently had a positive result for the virus. The result is similar to the experience of outpatients in Sri Lanka.17 During the endemic influenza season, detecting the virus made it possible to reduce the use of antibiotics by 20%, from 83.7 to 62.3%, despite a positive test for influenza. The high volume of patients and the fear of bacterial superinfection were important factors in the excessive use of antibiotics in that study.
The design in our study does not allow us to know the factors that influenced the treating physicians to continue prescribing antibiotics. In 39% of the files, some justification for its use was found due to bacterial superinfection, but no data of greater severity were found. Perhaps the most influencing factor was the hospitalisation of the patients. The antibiotics that were used the most as definitive treatment were cephalosporins (45%) and macrolides (17%).
In the analysed samples, we had a predominance of RSV in 25%, followed by influenza A with 18% and B with 9%; in 16% of the samples, more than one virus was detected. In a study conducted in our country on patients with influenza-like illness, the proportion of viruses identified was predominantly rhinovirus (36.5%), followed by influenza (22.6%), coronavirus (17.9%) and RSV (14.2%). Although the samples appear to correspond to this influenza, 47% were hospitalized.17 This reflects the wide circulation of respiratory viruses, which can condition moderate-to-severe symptoms that require hospital management. In particular, RSV and the influenza virus tend to cause more serious conditions. In the winter season, rhinoviruses predominate. However, in cases with community pneumonia or a respiratory infection requiring hospitalisation, the influenza virus, RSV, and metapneumovirus predominate.18,19 There is a discrepancy as to whether viral coinfections can cause a more serious course of infection. 18,19
Although antimicrobial resistance is currently one of the leading public health problems in the world,12 its prescription continues without reason. It is necessary to give the correct weight to the association of laboratory studies that detect viruses in patients with moderate to severe respiratory infections and to accept that in the highest percentage of these cases, the use of antimicrobials is not required. Some studies find respiratory viruses in patients with community pneumonia and patients with severe respiratory infections requiring intensive care management in which no bacterial association was demonstrated.20,21 The medical behaviour regarding their therapeutic decision in a private hospital shows that despite having carried out a study to search for viruses as the cause of acute respiratory infection, the results obtained were not used to generate a modification regarding the use of antibiotics.
It will be necessary to know the reasons that lead to the continuation of antimicrobial management, even in the presence of viral aetiology. Probably there is no confidence in laboratory studies because their sensitivity and specificity are unknown. There is also a fear of not offering a treatment ‘that can cure the infection’, primarily because few antiviral alternatives exist. There is intense family pressure because, in private hospitals, the doctor is the caregiver and assumes the decisions and responsibilities for their patient.
Greater dissemination of information is required on the participation of viruses in respiratory infections of moderate-to-severe evolution that does not require antimicrobial treatment. It probably takes time to accept the usefulness of the new diagnostic laboratory techniques that have recently been incorporated into hospitals.
ACKNOWLEDGMENTS
The authors would like to thank Dr. Al-kuraishy Haidar for his support during the study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
The study was approved by the Institutional Ethics Committee of Alkarama hospital and the University of Baghdad, School of Medicine (approval number; 11942, 21/2/2019).
AVAILABILITY OF DATA
All datasets generated or analysed during this study are included in the manuscript.
- Tamir SO, Shemesh S, Oron Y, Marom T. Acute otitis media guidelines in selected developed and developing countries: uniformity and diversity. Arch Dis Child. 2016;102(5):450-457.
Crossref - Burillo A, Bouza E. Use of rapid diagnostic techniques in ICU patients with infections. BMC Infect Dis. 2014;14(1):593.
Crossref - Bromwich M. Tonsillectomy or adenotonsillectomy versus nonsurgical management for obstructive sleep-disordered breathing in children. Paediatrics & Child Health. 2018;23(6):388-390.
Crossref - Abduljabbar HL, Hussein AA, Al-Mayah QS, Aufi IM. Phylogenetic analysis of respiratory syncytial virus isolated from children with respiratory tract infections in Baghdad city, Iraq. J Phys Conf Ser. 2019;1234:012082.
Crossref - Fauci A, Touchette N, Folkers G. Emerging Infectious Diseases: a 10-Year Perspective from the National Institute of Allergy and Infectious Diseases. Emerg Infect Dis. 2005;11(4):519-525.
Crossref - Romanin V, Agustinho V, Califano G, et al. Epidemiological situation of pertussis and strategies to control it. Argentina, 2002-2011. Arch Argent Pediatr. 2014;112(5):413-420.
Crossref - Korin Y, Kukol L. Viral pneumonia: features of diagnostics, clinical course and treatment. Med Alliance. 2020;8(4):32-39.
Crossref - Fowler D, Dise N, Sheppard L. Committee on air pollution effects research: 40 years of UK air pollution. Environ Pollut. 2016;208(Part B):876-878.
Crossref - Al-Jumaili A, Younus M, Kannan Y, Nooruldeen Z, Al-Nuseirat A. Pharmaceutical regulations in Iraq: from medicine approval to post-marketing. East Mediterr Health J. 2021;27(10):1007-1015.
Crossref - Rhedin S, Lindstrand A, Rotzen-Ostlund, et al. Clinical Utility of PCR for Common Viruses in Acute Respiratory Illness. J Pediatr. 2014;133(3):e538-e545.
Crossref - Ostrowsky B, Banerjee R, Bonomo R, et al. Infectious Diseases Physicians: Leading the Way in Antimicrobial Stewardship. Clin Infect Dis. 2018;66(7):995-1003.
Crossref - Hoffman SJ, Outterson K, Rottingen J-A, et al. An international legal framework to address antimicrobial resistance. B World Health Organ. 2015;93(2):66.
Crossref - Al-Samydai A, AL-Samydai MJ, Abu Hajleh MN. The De-marketing Strategies as a Vital Resolve of Antibiotics Misuse Dilemma. J Pure Appl Microbiol. 2021;15(4):1898-1906.
Crossref - Loeffelholz M, Pong D, Pyles R, et al. Comparison of the FilmArray Respiratory Panel and Prodesse Real-Time PCR Assays for Detection of Respiratory Pathogens. J Clin Microbiol. 2011;49(12):4083-4088.
Crossref - Yoshii Y, Shimizu K, Morozumi M, et al. Detection of pathogens by real-time PCR in adult patients with acute exacerbation of bronchial asthma. BMC Pulm. Med. 2017;17(1):150.
Crossref - Kaplan S. Coinfections in Hospitalized Children With Community-Acquired Pneumonia: What Does This Mean for the Clinician? J Infect Dis. 2018;218(2):173-175.
Crossref - Tillekeratne LG, Bodinayake CK, Nagahawatte A, et al. Use of Rapid Influenza Testing to Reduce Antibiotic Prescriptions Among Outpatients with Influenza-Like Illness in Southern Sri Lanka. Am J Trop Med Hyg. 2015;93(5):1031-1037.
Crossref - Noyola D, Hunsberger S, Valdes Salgado R, et al. Comparison of rates of hospitalization between single and dual virus detection in a Mexican cohort of children and adults with influenza-like illness. Open Forum Infect Dis. 2019;6(11):ofz424.
Crossref - Al-hassany HA, Albu-rghaif AH, A. Naji M. Tumor diagnosis by genetic markers protein P-53, p16, C-MYC, N-MYC, protein K-Ras, and gene her-2 Neu is this possible? Tumor diagnosis by genetic markers C-MYC, N-MYC, protein P-53, p16. Pak J Med Health Sci. 2021;15(8):2350-2354.
Crossref - Al Amad M. Severe Acute Respiratory Infections with Influenza and Non-Influenza Respiratory Viruses: Yemen, 2011-2016. IProc. 2018;4(1):e10599.
Crossref - Murthy S, Kissoon N. Management of severe viral infections in the pediatric intensive care unit. J Pediatr Intensive Care. 2015;03(04):205-216.
Crossref - Al-kuraishy H. M, Al-gareeb Ali I, Naji HA. Febuxostat Modulates Oxidative And Apoptotic Pathways In Acute Doxorubicin Induced Cardiotoxicity: An Experimental Animal Model Study. Asian J Pharm Clin. 2019;12(4):73-76.
Crossref - Alkuraishy HM, Al-Gareeb AI, Al-hussaniy HA. Doxorubicin-induced cardiotoxicity: molecular mechanism and protection by conventional drugs and natural products. Int J Clin Oncol Cancer Res. 2017;2(2):31-44.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.