ISSN: 0973-7510
E-ISSN: 2581-690X
A wide range of gastrointestinal (GI) illnesses is caused by foodborne bacteria that can arise from either a direct bacterial infection or bacterial toxin ingestion. The treatment of these infections has been hampered by the appearance of resistant strains. This current study aims to investigate the prevalence of Gastrointestinal tract (GIT) infections in Omani patients and their resistance pattern against commonly used antibiotics. Seven hundred and ninety fresh stool samples were obtained from Omani patients attending Sultan Qaboos University Hospital with GI manifestation from the 1st of June to the 30th of November 2019. Bacterial identification in stool samples was carried out by inoculation in culture media, microscopical examination and biochemical tests confirmed by MALDI. BD PhoenixTM. The antibiotics sensitivity testing was carried out by the Manual disk diffusion method and by MALDI. BD PhoenixTM. Out of 790 stool samples, 49 samples were positive for GIT bacterial infections. Salmonella spp. was the most prevalent isolate and more associated with children less than ten years old. Out of the 49 bacterial isolates, 3 (6.1%) were Clostridium difficili, 4 (8.2%) were Shigella flexneri, 5 (10.2%) were Campylobacter jejuni, and different Salmonella spp. serotypes were detected such as Salmonella Kentucky (8.2%), Salmonella enteritidis (6.1%), Salmonella infantis (4.1%), Salmonella welteverden (4.1%), Salmonella typhimurium (4.1%), Salmonella anatum (2.0%), Salmonella tesvia (2.0%), Salmonella Uganda (2.0%), Salmonella Arizona (2.0%) and (40.8%) of other Salmonella spp. serotypes. Eighty percent of isolated Campylobacter jejuni were resistant to Ciprofloxacin and Tetracycline. Salmonella spp. and Shigella flexneri were highly resistant to Amikacin, Gentamicin, and Cefuroxime. The low level of bacterial infection detected among screened patients in the present study indicates the excellent hand washing hygiene practice in reducing GIT infections among patients in Oman. This good hand washing hygiene practice is of great help in the efforts of controlling the spread of other severe diseases like COVID-19. However, detecting the emerging of antibiotic-resistant of GIT bacterial pathogens among patients in Oman, such as Salmonella and Shigella to a commonly used antibiotic such as Gentamicin, is alarming.
Gastrointestinal tract infections, tertiary care hospital, Oman
Diarrhoea resulting from GIT infections is a major health burden worldwide. Almost 4% of South Africa deaths during the year 2000 were deaths attributable to infectious diarrhoea, the fifth leading cause of death in this African country1. Among children under the age of five, diarrhea ranks as the second leading cause of preventable illness2.
Diarrhoea resulting from ingesting contaminated food or water. Person to person transmission can be the source of infection of various pathogenic bacteria, including Escherichia coli, Shigella spp., non-typhoidal Salmonella spp., Staphylococcus aureus, enterotoxigenic Bacteroides fragilis, and Campylobacter spp. and Enterococci faecalis1.
Enterococci faecalis have rapidly appeared as critical nosocomial pathogens, with increasing resistance to most, and some cases to all currently approved antibiotics3.
Both developed and less developed countries are affected by Gastrointestinal (GI) diseases, despite the strong relationship between GI illnesses and risk factors such as the absence of clean drinking water, poor sanitation, and other factors2.
The emergence of resistant microorganisms is always a potential risk when antibiotics are administered4. One study to detect the extensive antibiotic resistance of diarrheagenic Escherichia coli and Shigella spp., in local health centres of Isfahan province in Iran showed that ‘bacterial dysentery due to Shigella species is considered to be the most complicated form of diarrhoea and a large number of associated deaths are reported annually5. The burden of diarrheal diseases is far more in developing countries for example in a multi-centre Asian study, the annual incidence of shigellosis was 13.2/1000 children under the aged of five years6. Furthermore, the antimicrobial resistance of enteric bacteria is rapidly increasing in the developing world. In the past, antibiotic resistance was mostly seen in nosocomial infections due to the widespread use of antibiotics in hospitals, however recently, multidrug-resistant bacteria are commonly encountered in community-acquired infections, particularly GIT infections. The globalization of antibiotics resistance is a reality7.
Moreover, an Iranian study which was performed to identify virulence genes and antibiotic resistance pattern of the enterotoxigenic and enteroaggregative E. coli isolated from stool samples obtained from diarrheagenic patients reveals that Gentamicin had the lowest resistance zero % while trimethoprim had the highest resistance (79.8 %)8. In this study, the resistance of E. coli isolates to chloramphenicol, cefotaxime, sulphamethoxazole, ciprofloxacin, ampicillin, and tetracycline were 3.5 %, 7.01 %, 71.05 %, 10.5 %, 52.63 %, and 3.5 %, respectively8.
A recent study was carried out in Qatar on 76 of E. coli obtained from less than ten years old diarrheagenic hospitalized children in order to identify the phenotypic resistance profiles and ESBL genotype and virulence profiles of Enteroaggregative E. coli (EAEC) and Enteropathogenic E. coli (EPEC)9. This study, and by using the E-test method showed that (73.7 %) of isolates were resistant to at least one antibiotic9. The isolates demonstrated overall, high resistance rate (70 %) to the first-line antibiotics such as ampicillin, tetracycline (46.4 %), and sulfamethoxazole-trimethoprim (42.9 %). Further, 39.5 % of the isolates from the stool samples obtained from ten years old diarrheagenic hospitalized children in Qatar, were multidrug-resistant (MDR), with 22.4 % being ESBL producers. On the other hand, all isolates were susceptible to fosfomycin, carbapenem, colistin and amikacin9.
Several studies were performed to detect antibiotic resistance of Helicobacter pylori. One of these studies was carried out in China from the year 2013 to 2016 and showed that the overall resistance rates were as follows: furazolidone (1.49 %), amoxicillin (1.58 %), rifampicin (2.80 %), gentamicin (3.73 %), doxycycline (9.20 %), clarithromycin (22.73 %), levofloxacin (24.75 %), cefuroxime (25.20 %), moxifloxacin (38.53%), ciprofloxacin (43.20 %), ceftriaxone (49.60 %), azithromycin (85.60 %), tinidazole (87.20 %), metronidazole (92.53 %), ornidazole (94.27 %) and cefetamet (97.20 %)10.
A recent study was conducted to investigate the antimicrobial resistance rates and virulence gene profile among Shigella species isolated from stool samples obtained from Iranian children11. The children aged between 2 months and 14 years. In this study, 141 were isolated as Shigella spp., revealing S. sonnei as the most prevalent species (78.7 %), followed by S. flexneri (19.9 %) and S. boydii (1.4 % ).
Antimicrobial susceptibility testing in this Iranian study, showed that most of the isolates (99.29 %) were resistant to at least one antibiotic. A total of 101 (71.6 %) Shigella isolates were multi drug resistant (MDR) strains11.
The present study aims to assess the prevalence and characterization of GIT bacterial pathogens and detect its antibiotics resistance patterns among Omani patients attending Sultan Qaboos University Hospital (SQUH).
Specimens
This study was performed at SQUH, clinical microbiology laboratory. A total of seven hundred and ninety fresh stool samples was collected from patients having GI manifestation during the period between 1st of June and 30th of November 2019. Clinical data of each patient including sex, age, clinical history was collected. This project has been approved by the Medical Research Ethics Committee, College of Medicine & Health Sciences, Sultan Qaboos University, Muscat, Oman (SQU-EC/184/19).
Detection of pathogenic bacteria in stool samples using routine culture media
Briefly, 0.2 gram of fresh stool sample from each patient was mixed with one drop of saline and then smeared in a glass microscopic slide and heat-fixed on the hot plate. Once dried, the stool sample was examined under a light microscope to investigate for the presence of white blood cells, red blood cells. All findings were documented in the request form.
All stool samples were routinely cultured in Xylose Lysine Deoxycholate Agar (XLD), Hektoen Enteric agar (HE), Campylobacter agar (CAMPY), and selenite F broth. Moreover, stool sample inoculated in selenite F broth, were sub-cultured after 24 hours incubation at 37°C in XLD, HE and CAMPY agars. For detection of causative bacterial pathogens. XLD and HE agar plates were incubated for 24 hours at 37°C in aerobic conditions. CAMPY agar plates were incubated for 48 hours at 42°C in anaerobic conditions. Bloody stool samples were also cultured on a Sorbitol agar plate. After 24 hours of incubation of XLD and HE agar plate colonies with the yellow colour, which means that they ferment lactose, were considered as normal flora and reported as No Enteric Pathogen (NEP). Whereas, if any colorless colonies appear, subsequent identification tests were performed. Subsequent tests, including oxidase test, triple sugar iron (TSI) and urease. If oxidase negative results were obtained, urease and triple sugar iron (TSI) tests were carried out. Negative urease and TSI that show pink slant and yellow bottom with or without H2S formation (black color) suspect the presence of Salmonella or Shigella. The confirmation of bacterial identification was carried out by using MALDI system. Suspected colonies are sub-cultured in Cystine Lactose Electrolyte Deficient (CLED) agar plate (purity plate) for subsequent identification and antibiotic sensitivity test using automated BD PhoenixTM. Once Salmonella spp. is confirmed, a sample was sent to Central Public Health Laboratories (CPHL), Muscat, for serotyping. After 24 hours of incubation of stool samples, Campy agar plate is retrieved, Campylobacter spp. growing colonies appear as a grey in coloured colonies with a shape like a water droplet and have aluminium shin. Gram staining of Campylobacter spp. reveals curved gram-negative bacteria, which is oxidase positive. Identification is Campylobacter spp. was confirmed by MALDI system.
Assessment of antibiotics sensitivity of bacterial isolate
Antibiotics susceptibility test was carried out on an automated BD Phoenix system, which provides rapid, accurate, and reliable detection of known and emerging antimicrobial resistance. Antibiotics sensitivity of bacterial isolate results were reported according to the Clinical and Laboratory Standards Institute (CLSI) 2019 recommendations. Manual disk diffusion method was also used in the present study for antibiotics susceptibility testing for Campylobacter, including Erythromycin, Tetracycline, Ciprofloxacin in the blood agar plate.
Data analysis
Patient’s data were all documented in the Microsoft Excel program. Statistical analysis, resistance percentage, tables, and graphs were all done using the Microsoft Excel program 2010.
Clinical features
A total of 790 patients having GI manifestation was recruited in this study. Patients’ age ranged from less than 1 year to 92 years old (Table 1). Of 790 patients, 414 were females & 376 were males (Table 2).
Table (1):
Distribution of patients according to age.
Age groups (years) |
Number of cases |
Percentage |
|---|---|---|
(0-10) |
291 |
36.80% |
(11-20) |
60 |
7.60% |
(21-30) |
79 |
10% |
(31-40) |
111 |
14.10% |
(41-50) |
62 |
7.80% |
(51-60) |
60 |
7.60% |
(61-70) |
64 |
8.10% |
(71-80) |
46 |
5.80% |
(81-90) |
15 |
1.90% |
(91-100) |
2 |
0.30% |
Table (2):
Distribution of patients according to gender.
Gender |
Number of cases |
Percentage |
|---|---|---|
Females |
414 |
52.40% |
Males |
376 |
47.60% |
Pathogen occurrence
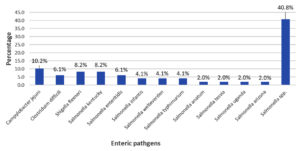
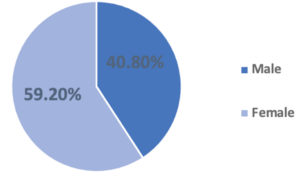
49 (6.2 %) stool samples out of 790 samples were positive for GIT infections. Out of 49 positive cases, 29 (59.2 %) were females, and 20 (40.8 %) were males. Out of the 49 bacterial isolates, 3 (6.1 %) were Clostridium difficili, 4 (8.2 %) were Shigella flexneri, 5 (10.2 %) were Campylobacter jejuni, and different Salmonella spp. serotypes were detected such as Salmonella Kentucky (8.2 %), Salmonella enteritidis (6.1 %), Salmonella infantis (4.1 %), Salmonella welteverden (4.1 %), Salmonella typhimurium (4.1 %), Salmonella anatum (2.0 %), Salmonella tesvia (2.0 %), Salmonella Uganda (2.0 %), Salmonella Arizona (2.0 %) and (40.8 %) of other Salmonella spp. serotypes. Fig. 1 shows the distribution of enteric bacterial pathogens detected in-patient attending SQUH with GI manifestations.
Association of GIT infection with gender
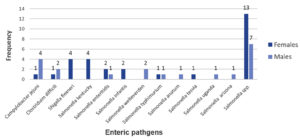
Out of the 29 bacterial isolates obtained from female stool samples, one sample (3.4%) was positive for Campylobacter jejuni, 1 (3.4 %) was positive for Clostridium difficili, 4 (13.8 %) were positive for Shigella flexneri, and 23 (79.3 %) were positive for different serotypes of Salmonella spp. Out of the 20 bacterial isolates obtained from male stool samples, 2 (10 %) of them were positive for Clostridium difficili, 4 (20 %) were positive for Campylobacter jejuni and 14 (70 %) were positive for different serotypes of Salmonella spp. Fig. 2 and 3 show the distribution of enteric bacterial pathogens in both females & males patients diagnosed with GI manifestations.
Association of GIT infection with age
The majority of GIT infections (24 patients) detected were from less than ten years old age group and more associated with Salmonella spp. infections, followed by patients of the age group of (41-50) years old. Table 3 shows the distribution of enteric bacterial pathogens in association with different age groups.
Table (3):
Distribution of enteric bacteria in association with age groups.
Age groups (years) |
Campylobacter jejuni |
Clostridium difficili |
Shigella flexneri |
Salmonella spp |
Total |
|---|---|---|---|---|---|
(0-10) |
2 |
0 |
1 |
22 |
24 |
(11-20) |
1 |
0 |
0 |
2 |
3 |
(21-30) |
2 |
0 |
1 |
0 |
3 |
(31-40) |
0 |
1 |
1 |
3 |
5 |
(41-50) |
0 |
1 |
0 |
8 |
9 |
(51-60) |
0 |
1 |
0 |
1 |
2 |
(61-70) |
0 |
0 |
1 |
1 |
2 |
(71-80) |
0 |
0 |
0 |
0 |
0 |
(81-90) |
0 |
0 |
0 |
0 |
0 |
(91-100) |
0 |
0 |
0 |
0 |
0 |
Table (4):
The prevalence of Antibiotic resistance of Campylobacter jejuni.
Enteric pathogen / Antibiotics |
Ciprofloxacin |
Erythromycin |
Tetracycline |
|---|---|---|---|
Campylobacter jejuni |
80% |
0% |
80% |
Antibiotics resistance patterns
Antibiotics sensitivity testing of Campylobacter jejuni, shows that 80% of Campylobacter jejuni were resistant to Ciprofloxacin and Tetracycline, and completely sensitive to Erythromycin. Salmonella spp. show a high percentage of resistance to Amikacin, Gentamicin, and Cefuroxime (91.9 %, 89.2 %, and 86.5 %, respectively). Shigella flexneri show increased resistance to Amikacin (75 %), Gentamicin (75 %), Cefuroxime (75 %), and Ampicillin (75 %). Table 4 shows the prevalence of antibiotic resistance of Campylobacter jejuni. Table 5 describes the prevalence of antibiotic resistance of Shigella flexneri and Salmonella spp. The prevalence of antibiotic resistance of Salmonella spp. serotypes are summarized in Table 6.
Table (5):
The prevalence of Antibiotic resistance of Shigella flexneri and Salmonella spp.
Enteric pathogen / Antibiotics |
AK |
CN |
ETM |
IPM |
MEM |
CXM |
CAZ |
CRO |
FEP |
AMP |
AMC |
TZP |
SXT |
CIP |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Shigella flexneri |
75% |
75% |
0% |
0% |
0% |
75% |
25% |
0% |
0% |
75% |
0% |
0% |
25% |
25% |
Salmonella spp |
91.9% |
89.2% |
0% |
0% |
0% |
86.5% |
13.5% |
16.2% |
13.5% |
35.1% |
2.7% |
0% |
10.8% |
13.5% |
AK: Amikacin, CN: Gentamicin, ETM: Ertapenem, IPM: Imipenem, MEM: Meropenem, CXM: Cefuroxime, CAZ: Ceftazidime CRO: Ceftriaxone FEP: Cefepime AMP: Ampicillin,
AMC: Amoxicillin-Clavulanate, TZP: Piperacillin-Tazobactam, SXT: Trimethoprim-Sulfamethoxazole, CIP: Ciprofloxacin.
Table (6):
The prevalence of antibiotic resistance of Salmonella spp. Serotyp.
Enteric pathogen / Antibiotics |
AK |
CN |
ETM |
IPM |
MEM |
CXM |
CAZ |
CRO |
FEP |
AMP |
AMC |
TZP |
SXT |
CIP |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Salmonella kentucky |
100% |
100% |
0% |
0% |
0% |
100% |
25% |
25% |
25% |
100% |
0% |
0% |
25% |
75% |
Salmonella enteritidis |
100% |
100% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
Salmonella infantis |
50% |
50% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
Salmonella welteverden |
100% |
100% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
Salmonella typhimurium |
100% |
100% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
Salmonella anatum |
100% |
100% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
Salmonella tesvia |
100% |
100% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
0% |
Salmonella uganda |
100% |
100% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
Salmonella arizona |
100% |
100% |
0% |
0% |
0% |
100% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
0% |
Salmonella spp. |
95% |
90% |
0% |
0% |
0% |
80% |
20% |
25% |
20% |
35% |
5% |
0% |
15% |
10% |
AK: Amikacin, CN: Gentamicin, ETM: Ertapenem, IPM: Imipenem, MEM: Meropenem, CXM: Cefuroxime, CAZ: Ceftazidime, CRO: Ceftriaxone, FEP: Cefepime, AMP: Ampicillin,
AMC: Amoxicillin-Clavulanate, TZP: Piperacillin-Tazobactam, SXT: Trimethoprim-Sulfamethoxazole, CIP: Ciprofloxacin.
In this study, among 790 stool samples of patients attending SQUH with GI manifestations, 49 (6.2 %) were positive for bacterial GIT infections. This finding was fewer in terms of prevalence when compared with another study conducted in Thailand and showed that 84.9 % of GIT infection cases are associated with bacterial enteric infection12.
In the present study, Salmonella spp. were the most prevalent bacterial isolate with a total of 37 (75.4 %), followed by Campylobacter jejuni 5 (10.2 %), Shigella flexneri 4 (8.2 %), Clostridium difficili 3 (6.1 %). These findings were consistent with a previous study performed at the University Hospital of Geneva in Switzerland13 to investigate the etiological agents of infectious diarrhoea in 7124 patient’s specimens. This Geneva hospital study detected 369 (2.6 %) Salmonella spp., 408 (2.9 %) Campylobacter spp., and 79 (0.6 %) Shigella spp.13
In the present study, the association between GIT infections and sex of patients was investigated. GIT infections were prevalent in females more than males. Out of 49 positive samples, 29 (59.2 %) were females, and 20 (40.8 %) were males. A previous study carried out in Qatar showed a higher percentage of positive GIT infections among males (55.9 %) than females (44.2 %). This finding is in agreement with other findings in the Gulf region14.
The correlation between GIT infections and age was also investigated in the present study, the majority of positive cases 24 (49 %) were of age 10 years and below and 22 (91.6 %) of them were associated with Salmonella spp. infection. A previous study showed that the risk of Salmonella increases with age and patients infected with Salmonella spp. tended to be relatively older with a range of 85.8 years14.
The present study showed a high resistance percentage (80 %) of Campylobacter jejuni to Ciprofloxacin and Tetracycline and (0%) resistance to Erythromycin. These results were consistent with a similar study from Belgium which showed high rates of resistance against Ciprofloxacin (55.8 %) and Tetracycline (49.7 %). Alternatively, lower resistance rates to Erythromycin (2 %)15.
The present study detected high resistance percentage (75 %) of Shigella flexneri against Amikacin, Gentamicin, Cefuroxime, and Ampicillin, which is incomparable with a previously published Iranian study which showed that most frequent antibiotic resistance Shigella flexneri was towards Co-trimoxazole (89 %), Ampicillin (77 %) and Ceftriaxone (51 %), except for Ampicillin16. In the present study, Salmonella spp. similarly to Shigella flexneri show a high resistance percentage to Amikacin, Gentamicin, and Cefuroxime. This high resistance rate could be due to increased prescription of such antibiotics.
The present study showed that salmonella spp. is the most prevalent enteric bacteria in patients attending SQUH with a percentage of (75.4%) and more associated with children of the age of 10 years and below. The second most prevalent was Campylobacter jejuni (10.2%) followed by Shigella flexneri (8.2%), then Clostridium difficili (6.1%). Both salmonella spp. and Shigella flexneri showed high resistance toward Amikacin, Gentamicin, and Cefuroxime. Campylobacter jejuni was highly resistant to Ciprofloxacin and Tetracycline.
This study demonstrated the antibiotic resistance patterns of bacterial pathogens isolated from fresh stool samples obtained from patients in Oman with GIT manifestations. It indicates the emerging of antibiotic-resistant of GIT bacterial pathogens such as Salmonella and Shigella to a commonly used antibiotic such as Gentamicin. This finding will be of great help to the health authorities in their monitoring of the antibiotic-resistant bacterial pathogens among cohort of patients in Oman.
ACKNOWLEDGMENTS
We are grateful to the head, the clinicians and the staff of the Microbiology Department, Sultan Qaboos University Hospital, Muscat, Oman for all of their help and support. The authors are grateful to the Central Public Health Laboratories, Muscat for the serotyping of the Salmonella spp.The authors are grateful to the Sultan Qaboos University ethical committee for allowing us to obtain patient data.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
AASA and RNA carried out the laboratory work. SAAB carried out the clinical supervision. MM conceived the presented idea and conducted the daily supervision of the project and wrote the manuscript. All authors reviewed and apporved the manuscript for publication.
FUNDING
None.
ETHICS STATEMENT
This project has been approved by the Medical Research Ethics Committee, College of Medicine & Health Sciences, Sultan Qaboos University, Muscat, Oman(SQU-EC/184/19).
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Kullin B, Meggersee R, D’Alton J, et al. Prevalence of gastrointestinal pathogenic bacteria in patients with diarrhoea attending Groote Schuur Hospital, Cape Town, South Africa. S A Med J. 2015;105(2):121-125.
Crossref - Fletcher SM, McLaws M-L, Ellis JT. Prevalence of gastrointestinal pathogens in developed and developing countries: systematic review and meta-analysis. J Public Health Res. 2013;2(1):42-53.
Crossref - Shepard BD, Gilmore MS. Antibiotic-resistant enterococci: The mechanisms and dynamics of drug introduction and resistance. Microbes Infect. 2002;4(2):215-224.
Crossref - Ebner W, Kropec-Hubner A, Daschner FD. Bacterial resistance and overgrowth due to selective decontamination of the digestive tract. Eur J Clin Microbiol Infect Dis. 2000;19(4):243-247.
Crossref - Sadeghabadi AF, Ajami A, Fadaei R, et al. Widespread antibiotic resistance of diarrheagenic Escherichia coli and Shigella species. J Res Med Sci. 2014;19(Suppl 1):S51-S55. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4078378/
- Seidlein LV, Kim DR, Ali M, et al. A Multicentre Study of Shigella Diarrhoea in Six Asian Countries: Disease Burden, Clinical Manifestations, and Microbiology. PLoS Med. 2006;3(9):e353.
Crossref - Pickering LK. Antimicrobial resistance among enteric pathogens. Sem Pediatr Infect Dis. 2004;15(2):71-77.
Crossref - Barati S, Boniadian M, Habibian R, Jostejou T. Antibiotic resistance of enterotoxigenic and entroaggrigative Escherichia coli isolated from gastroenteritis cases. Asian Journal of Biomedical and Pharmaceutical Sciences. 2012;2(14):54-58. https://www.alliedacademies.org/articles/antibiotic-resistance-of-enterotoxigenic-and-entroaggrigative-escherichia-coli-isolated-from-gastroenteritis-cases.pdf
- Eltai NO, Al Thani AA, Al Hadidi SH, Al Ansari K, Yassine HM. Antibiotic resistance and virulence patterns of pathogenic Escherichia coli strains associated with acute gastroenteritis among children in Qatar. BMC Microbiol. 2020;20(1):54.
Crossref - Shao Y, Wang B, Xu Q, Ye G, Lu R, Yang Y. Antibiotic resistance of Helicobacter pylori to 16 antibiotics in clinical patients. J Clin Lab Anal. 2017;32(4):e22339.
Crossref - Karimi-Yazdi M, Ghalavand Z, Shabani M, et al. High rates of antimicrobial resistance and virulence gene distribution among shigella spp. Isolated from pediatric patients in Tehran, Iran. Infect Drug Resist. 2020;13:485-492.
Crossref - Okada K, Wongboot W, Kamjumphol W, et al. Etiologic features of diarrheagenic microbes in stool specimens from patients with acute diarrhea in Thailand. Sci Rep. 2020;10:4009.
Crossref - Rohner P, Pittet D, Pepey B, Nije-Kinge T, Auckenthaler R. Etiological agents of infectious diarrhea: Implications for requests for microbial culture. J Clin Microbiol. 1997;35(6):1427-1432.
Crossref - Weam B, Abraham M, Doiphode A, et alfi. Foodborne Bacterial Pathogens Associated with the Risk of Gastroenteritis in the State of Qatar. Int J Health Sci. 2016;10(2):185-195.
Crossref - Elhadidy M, Ali MM, El-Shibiny A, et al. Antimicrobial resistance patterns and molecular resistance markers of Campylobacter jejuni isolates from human diarrheal cases. PLOS ONE. 2020;15(1):e0227833.
Crossref - Nikfar R, Shamsizadeh A, Darbor M, Khaghani S, Moghaddam M. A study of the prevalence of Shigella species and antimicrobial resistance patterns in paediatric medical center, Ahvaz. Iran. Iran J Microbiol. 2017;9(5):277-283. https://pubmed.ncbi.nlm.nih.gov/29296272/
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.