ISSN: 0973-7510
E-ISSN: 2581-690X
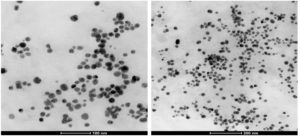
In the present study, a soil actinomycete was isolated from near the river Nile shoreline, Egypt. The identification of this isolate as Streptomyces griseorubens was performed using 16s rDNA. The sequence has been deposited in the Gene Bank with the accession number LC066679. Factors affecting the biogenesis of AgNPs were optimized by applying the Plackett-Burman factorial design. The maximum silver nanoparticles (AgNPs) biosynthesis (2.76 OD at 400 nm) was achieved in the trial No. 9 that contained the following ingredients (g/L): Starch (20); MgSO4 (0.05); K‚ HPO4 (1.0); NaNO3 (2.0); AgNO3 (1.0) mmol/l; PH (7); incubated at temperature 30°C for 72 hr. The biosynthesized nanoparticles were characterized using spectroscopic techniques. AgNPs showed the characteristic UV spectra at a wavelength range 300 – 600 nm and a characteristic absorption peak was recorded at the wavelength of 400 nm. For AgNPs with absorbance height ofH≈2.56 a.u. and peak width at half maximum absorbance (PWHM) H≈120 nm which remained constant over a long period of time indicating its stability. FTIR spectra showed the functional group of the biomaterials capping the AgNPs. EDX confirmed the formation of the metallic silver nanoparticles, indicating the presence of proteinaceous cabbing. TEM micrograph showed spherical AgNPs in shape with an average diameter of 22 nm. The biosynthesized AgNPs showed high stability up-to two weeks. The conjugate (AgNPs/cellulosic fiber (C)) of Luffa aeygptiaca and (AgNPs/activated carbon (AC)) was applied for drinking water treatment, which resulted in fecal Coliform bacteria removal with a success of 99.9% as a water treatment application.
Streptomyces griseorubens, AgNPs biosynthesis, AgNPs/cellulosic fiber, drinking water treatment application.
Disinfection of drinking water is a public and major health concern1. The traditional chemical disinfection such as chlorine gas is an effective tool against E. coli pathogens but may react with various water constituents resulting in harmful carcinogenic by-products such as Trihalomethane.2-4 In addition the emergence of new resistant strains to the chemical disinfection which show a high tolerance to these chemicals. The silver nanoparticles (AgNPs) have already been proved toxic to both aerobic and anaerobic bacteria5 and due to their fine and homogenized distributed particle size they show a high resistance to oxidation.6 They have proved a promising bactericidal effect against Escherchia coli and Staphylococcus aureas.7 The application of AgNPs for water treatment in conjunction with a suitable material as the activated carbon (AC) which is an effective adsorbent, with high surface area (500 – 1500 m².g–¹) and high porosity8, 9 is considered to be promising technologies and is being applied to various treatment processes to remove pollutants.10, 11 Generally, silver nanoparticles (AgNPs) can be synthesized by different methods, but the green synthesis approaches are more preferable than the conventional methods using chemical agents which may result in the environmental toxicity. Actinomycetes are the rich source of the potential bioactive compounds.12 AgNPs synthesis has been reported on the use of actinomycete members such as Thermomonospora sp., Streptomyces hygroscopicus, Streptomyces albogriseolus, Streptomyces sp. SS2 and Streptomyces sp. LK3.13-17
This study aimed at the isolation, identification of soil actinomycetes, investigating their ability for green AgNPs biogenesis and their conjugation to the granular activated carbon and a local cellulosic fiber (Luffa aeygptiaca) in an attempt for Coliform bacteria removal from drinking water as an indicator for water treatment.
Microorganisms and cultural conditions
Five actinomycetes isolates were maintained on starch-nitrate agar medium (Waksman, 1959) with the following ingredients (g/L): Starch 20; KNO3 2; K2 HPO4 1; MgSO4 .7H‚ O 0.5; NaCl 0.5; CaCO3 3; FeSO4 .7H2O 0.01; agar 20; 3.5 mmol/l AgNO3 2 ml and distilled water up to 1 L. Slopes were incubated for a period of 7 days at 30°C. The isolates were also stored as a spore suspension in 20% (v/v) glycerol at -20°C for further use18.
Inoculum preparation
Erlenmeyer flask (250 ml) containing 50 ml of starch-nitrate liquid medium containing (g/l): soluble starch 20; NaNO3 2; K‚ HPO2 1; MgSO4 ·7H2O 0.5 and distilled water up to 1 L were inoculated with three disks of 9 mm diameter taken from a 7-days old stock culture grown on starch nitrate agar medium. Culture was incubated for 72 h in a rotatory incubator at 200 rpm and 30°C to be used as a seed culture for subsequent experiments.19
The synthesis of AgNPs
In order to screen the isolates for their ability to synthesize AgNPs, the seed culture (5 ml) challenged with 50% (v/v) of AgNO3. 1 mmol/L and then incubated on an orbital shaker in a dark condition for 24 h at 30°C.20 After ending incubation period, the scanning for the UV spectra at a wavelength range 300 – 600 nm for the characteristic surface plasmon resonance (SPR) at wavelength of 400 nm. The isolate with a characteristic absorption peak at 400 nm, the highest peak absorbance height that relates to high concentration and the lowest Peek width at half maximum absorbance (PWHM) which measured as the full width at half-maximum absorbance that relates to nanoparticles homogeneity was then selected for further analysis. The cell free broth media without the addition of AgNO3 was maintained as a control.
The selected isolate identification
The pure colony of the promising isolate was identified using 16S rDNA gene sequence analysis21 at Sigma Scientific Services Co. (Cairo, Egypt). The obtained sequence was then compared to available sequence database in Gene Bank using the NCBI BLAST program and deposited in the Gene Bank (accession no LC066679).
Plackett-Burman experimental design
Plackett-Burman fractional factorial design22, 23 was used to investigate various environmental factors that affect the biogenesis of AgNPs. Seven independent variables were investigated for their effect on nanoparticles biogenesis. For each variable, a high (+), low (-) level was tested and (0) level as control. All trails were performed in duplicates and the average result was considered as the response. The main effect of each variable was determined with the following equation:
Exi = (SMi+ – SMi-) / N
Where Exi is the variable main effect, Mi+ and Mi- are the AgNPs biosynthesis in trails where the independent variable (xi) was present in high and low concentrations, respectively, and N is the number of trials divided by 2. A main effect figure with a positive sign indicates that the high concentration of this variable is near to the optimum and a negative sign indicates that the low concentration of this variable is near to the optimum. Using Microsoft Excel, statistical f-values for equal un-paired samples were calculated for determination of variable significance24.
Characterization of silver nanoparticles
For UV-vis spectral analyses was performed on CECIL, UV/Visible-2021 (Cambridge instrument from England) in the range 200 -1000 nm25. The biosynthesized AgNPs surface plasmon resonance (SPR) scanned at different time intervals to check its stability. For morphology, size and EDX analysis, the instrument HRTEM (Tecnai G 20, Super twin, double tilt) was used. In typical measurement diluted AgNPs solution was drop casted on a carbon coated cupper grid and measurements were taken to determine the AgNPs presence confirmation, size distribution and morphology. FTIR spectrum was performed on Perkin- ElmerBXII instrument in the range 500–4000 cm.–¹ FTIR was applied to identify the functional groups on the surface of the biosynthesized nanoparticles.
Application of biosynthesized AgNPs in water treatment
Granular activated carbon and cellulosic fiber were prepared for coating with silver nanoparticles.
The biosynthesized silver nanoparticles conjugated with cellulosic fiber of Luffa aeygptiaca (AgNPs/C), and granular activated carbon (AgNPs/GAC – particle size range 0.60 – 2.36 mm) supplied by Jacobi Carbons AB, Bredbandet 1, Varvsholmen SE- 392 30 Kalmar, Sweden. The GAC granules have been purified by the treatment with hot concentrated HNOƒ, hot distilled water, hot concentrated NHƒ solution and finally with hot distilled water. GAC was dried in an oven for 2 hrs at 110°C prior to its coating with AgNPs26. The cellulosic fiber has been boiled for 15 min, washed several times by distilled water and air dried prior to its coating with AgNPs. Both the GAC granules and cellulose fiber were impregnated in 250 ml of biosynthesized silver nanoparticles 40 ppm solution under vigorous stirring at room temperature and were kept overnight for contact. The conjugate was there analyzed by using SEM (the scanning electron microscope) Model Quanta 250 FEG (Field Emission Gun) attached with EDX unit (Energy Dispersive X-ray Analyses), with accelerating voltage 30 K.V., magnification14x up to 1000000 and resolution for Gun.1n., from FEI Company, Netherlands.
Removal of Thermotolerant Coliform
The prepared conjugate with, without AgNPs was examined as a filtration medium for the removal efficiency of Thermotolerant (fecal) Coliform from highly polluted water samples27 using a selective lactose medium (m-FC agar, Alpha Biosciences Inc. USA). The water flow adjusted at constant contact time (5 min) through the designed prototype filter column that contained the filtration medium (Fig 1). The percentage of removal efficiency was then calculated using the following equation:28
The removal%=100-(survivor count ÷ initial count)×100
Isolates potential of for green AgNPs synthesis
The challenged broth culture of five actinomycete isolates with 50% (v/v) of 1 mM AgNO3 solution resulted in a clear brown color for only three isolates number 1, 3 & 4 that indicated the formation of AgNPs29 with a characteristic plasmonic peak around 400 nm which is a clear indication for the formation of the AgNPs.30 The three isolates had the potential to reduce silver ions to silver nanoparticles. The maximum biosynthesis of silver nanoparticles by the isolate No. 3 reached after 2 hours. The surface plasmon peak absorbance (SPR) shape and values gave a clear indication about the distribution and the concentration of AgNPs.31 The isolate No. 3 recorded the highest absorbance peak which related to the highest yield of AgNPs and the lowest PWHM indicating the monodispersed nature of the biosynthesized AgNPs so it was selected for the mediation of biosynthesis of AgNPs and further analysis (Table No. 1). On the contrary with the other isolates 2 and 5 gave no color change or the characteristic plasmonic peak. The bioreduction happened which could be explained on the ground that the reduction of silver ions to metallic silver by proteins, reductive enzymes and secondary metabolites.32

Fig. 1. A designed prototype as a filter column containing the filtration medium for Coliform removal.
Table (1):
Silver nanoparticles peak absorbance height (at 400 nm) and PWHM
Isolate No. |
Peak height at 400 nm |
PWHM (nm) |
|---|---|---|
1 |
0.656 |
160 |
3 |
2.48 |
120 |
4 |
0.811 |
1000 |
Identification of the isolate No.3
The colonies color of the isolate No. 3 was white gray when grown on starch-nitrate agar medium. The strain produce jasmine odor and exhibited an antibacterial activity against the E. coli (ATCC 8739 and ATCC 25922).33 The Identification of isolate No. 3 that showed the highest potential for the mediation of AgNPs biosynthesis was performed by ribotyping of 16S rDNA. The developed tree based on the neighbor joining method for obtained 16S rDNA sequences depicted that the isolate occupies a distinct phylogenetic position within the elucidated radiation including representatives of the Streptomyces family (Fig 2). The results of 16S rDNA gene sequencing (1200bp) analysis revealed that this isolate corresponds to Streptomyces griseorubens. The isolate was deposited in the gene bank with the accession number LC066679.
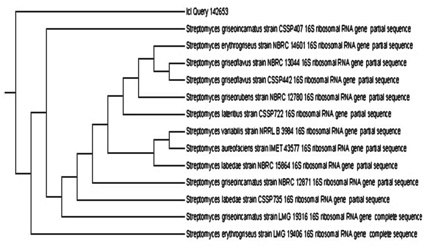
Fig. 2.The phylogenetic tree depicting the position of the isolate No.3 among well known isolates.
Optimization of the fermentation conditions affecting the mediated biosynthesizes of AgNPs by S. griseorubens
Seven culture variables were set to study the influence of culture medium conditions on the mediated AgNPs biogenesis as a response (Table 2), according to the Plackett-Burman design.
Table (2):
Plackett-Burman design matrix representing the coded values for 7 independent variables and the absorbance readings at 400 nm as a response reflecting the AgNPs concentration
Trial No. |
AgNO₃ X1 mol/L |
pH X2 |
Starch X3 g/L |
NaNO₃ X4 g/L |
K₂HPO₄ X5 g/L |
Incubation Temperature X6 (ͦC ) |
Incubation period X7 (hours) |
Absorbance (400 nm) |
|---|---|---|---|---|---|---|---|---|
1 |
1.5 |
8 |
25 |
1.5 |
1.5 |
25 |
48 |
0.634 |
2 |
0.5 |
8 |
25 |
2.5 |
0.5 |
35 |
48 |
0.272 |
3 |
0.5 |
6 |
25 |
2.5 |
1.5 |
25 |
96 |
1.404 |
4 |
1.5 |
6 |
15 |
2.5 |
1.5 |
35 |
48 |
0.703 |
5 |
0.5 |
8 |
15 |
1.5 |
1.5 |
35 |
96 |
0.975 |
6 |
1.5 |
6 |
25 |
1.5 |
0.5 |
35 |
96 |
1.000 |
7 |
1.5 |
8 |
15 |
2.5 |
0.5 |
25 |
96 |
1.172 |
8 |
0.5 |
6 |
15 |
1.5 |
0.5 |
25 |
48 |
0.887 |
9 |
1 |
7 |
20 |
2 |
1 |
30 |
72 |
2.760 |
The data (Fig 3) indicated that the presence of high levels of starch, high incubation temperature, high incubation period and AgNO3 concentration affected positively AgNPs synthesis by S. griseorubens. On the other hand, the presence of K2 HPO4, NaNO3 and pH at their lowest levels resulted in high AgNPs synthesis. The main effect pointed out these higher levels of starch, incubation temperature, incubation period and AgNO3 concentration with low levels of K2 HPO4, NaNO3 and pH supported the mediated biogenesis of AgNPs in the culture filtrate.
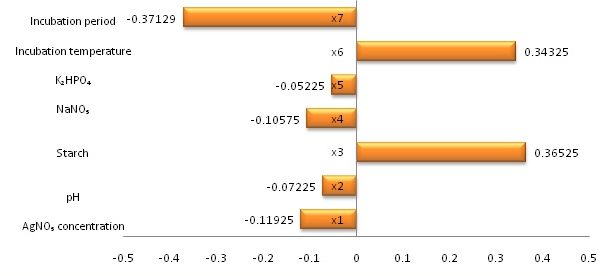
Fig. 3. Main effects for the variables affecting AgNPs biosynthesis
The statistically based experimental designs for optimizing nutrients, which involved simultaneously varying factors in a deliberate manner, are useful for identifying the important environmental conditions, media composition and the interactions between two or more factors.23,24,34 Furthermore, applying multi factorial experiment considers the interaction between the non-linear natures of the response in a very short experiment. Plackett-Burman experiments design was effective in the determination of the medium components, which had a significant effect on the mediated production of AgNPs.
Based on the obtained results, it can be concluded that the optimum medium for the mediated producing of AgNPs from the culture of S. griseorubens contained the following ingredients: (g/L): Starch (25); MgSO4 (0.05); K‚ HPO2 (0.5); NaNOƒ (1.5); AgNO3 (1.5) mmol/l; incubation period for 96 hours at 35°C and pH 6.0. The results revealed that the highest AgNPs yield (2.76 OD at 400 nm) was achieved for the trial No.9 which contained the following formula :(g/L): Starch (20); MgSO4 (0.05); K2 HPO4 (1.0); NaNO3 (2.0); AgNO3 (1.0) mmol/l; pH (7); incubation temperature (30°C) and incubation time (72 hr).
Characterization of the biosynthesized AgNPs
The identified isolate S. griseorubens mediated the biosynthesis of AgNPs giving a clear brown color with a characteristic absorption peak (at 400 nm). The stability of AgNPs was examined (Fig.4) using UV-Vis spectrometry measurements over a period of time (240 and 480 hours). The position and the intensity of the absorption peaks remained unchanged indicating high stability of the biosynthesized AgNPs.35 The FTIR results shown in Fig 5 indicated the functional groups; the band at 3419 cm–¹ is the O–H hydroxyl group, which may be responsible for reducing the metal ions into their respective nanoparticles. The alkynes group, C=C detected at 2088 cm–¹ that might be responsible for the stability of the synthesized metal nanoparticles. Also the N-H bending group around 1640 cm–¹ indicating the presence of amine group which could support the idea of a protein capping agents when the metal nanoparticles are formed in the solution.33 It is also contributes towards the stabilization of the AgNPs against the Van der Waals forces of attraction which may otherwise cause nanoparticles coagulation36.
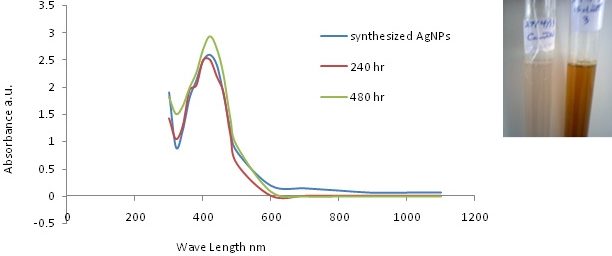
Fig. 4. UV-vis analysis for the biosynthesized AgNPs with the clear brown color and the control as white in the top – right corner
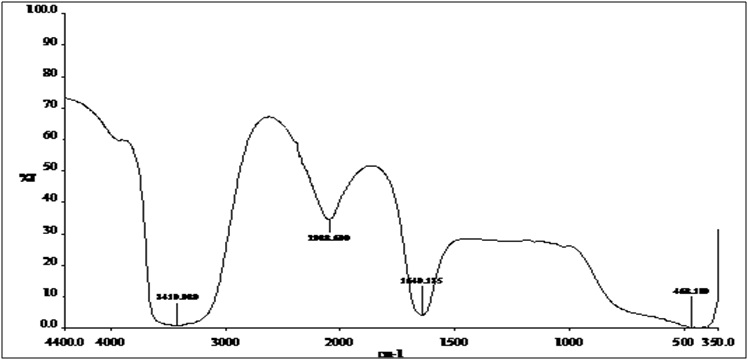
Fig. 5. FTIR spectrum recorded for the biosynthesized AgNPs
High Resolution Transmission Electron Microscopy visualization (HRTEM) was carried out to determine the morphology and the particle size of the AgNPs. A well dispersed spherical silver nanoparticles having the average size of 22 nm (Fig. 6). The capping nature of the synthesized AgNPs is revealed in the FTIR spectra and there is a minor aggregation in consistence with the broadening in the UV-vis spectra recorded at different time intervals.
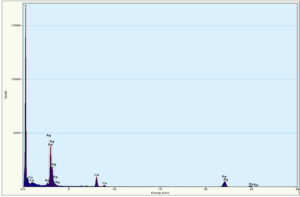
Energy dispersive X-ray spectroscopy (EDX) was performed to confirm the presence of elemental silver (Fig. 7) revealed the optical absorption band peaks up to 30 keV. The absorption band peak in the range of 3 – 4 keV was typical for the absorption of metallic silver crystallites that confirms the presence of the colloidal AgNPs in solution.37 There were other EDX peaks for C and Cu that derived from the C and Cu grid used.
Water treatment application of the biosynthesized AgNPs by S. griseorubens
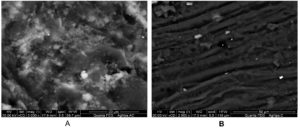
The conjugates of AgNPs/ GAC and AgNPs/C were visualized by the Scanning Electron Microscopy (SEM). The micrograph showed also AgNPs distribution and adsorption on the porous surface of the activated carbon and cellulosic fiber of Luffa aeygptiaca (Fig 8).
Fig. 8. SEM micrograph of conjugated AgNPs with A- activated carbon conjugated to AgNPs (AgNPs/GAC) at magnification of 5000 x. B- Cellulosic fiber of L. aeygptiaca conjugated to AgNPs (AgNPs/C) at magnification of 2500 x. The AgNPs appeared like white spots on the porous surface of the activated carbon and cellulosic fiber
The raw samples before the filtration process contained high counts of thermotolerant coliform (1000-1100 CFU/100mL). The results in Table 3 revealed that the granular activated carbon (GAC) actually removed 96% and cellulosic fiber (C) of L. aeygptiaca removed 93% from Thermotolerant coliform in raw water sample. After that the conjugation of the previously mentioned materials with AgNPs resulted in the granular activated carbon (AgNPs/GAC) disinfection efficiency which increased by 3.7% and natural cellulosic fiber of L. aeygptiaca (AgNPs/C) by 5.8 % due to the antibacterial nature of the biosynthesized silver nanoparticles5,38. Furthermore, upon using the filtration medium containing both conjugates (AgNPs/GAC and AgNPs/C) resulted in the reduction of thermotolerant coliform counts to almost near zero.
Table (3):
Efficiency of water disinfection by different filtration media at 5min contact time
Filtration medium |
Fecal Coliform count (CFU/100 mL) |
Fecal Coliform Removal % |
|---|---|---|
GAC |
40 |
96 |
AgNPs/GAC |
3 |
99.7 |
C |
70 |
93 |
AgNPs/C |
12 |
98.8 |
AgNPs/GAC and AgNPs/C |
1 |
99.9 |
The present work has provided a successfully mediated green biosynthesis of high stability, environment friendly AgNPs by S. griseorubens. The increased capabilities of activated carbon and cellulosic fiber to the conjugate AgNPs, for removing water Thermotolerant Coliform bacteria. The present route could be easily scaled up for the industrial applications to increase the yield of the eco-friendly AgNPs biogenesis extensively and designing cost-effective application for drinking water disinfection.
- Prashant, J., Pradeep T. Potential of silver nanoparticle-coated polyurethane foam as an antibacterial water filter. Biotechnol. Bioeng., 2005; 90(1):59-63.
- Li, Q., Mahendra, S., Lyon, D.Y., Bruent, L., Liga, M., Li, D., Alvarez, P.J. Antimicrobial nanomaterials for water disinfection and microbial control: potential applications and implications. Water Research. 2008; 42(18):4591-602.
- USEPA. Analysis of Trihalomethanes in drinking water by liquid/liquid extraction. 40 CFR part141, Appendix C. Part II 1980.
- USEPA. 1979. The analysis of Trihalomethanes in finished water by the purge and trap method 501.1. Environmental Monitoring & Support Lab., Environmental Research Center, Cincinnati, Ohio 45268.
- Chou, W., Yu, D., Yang, M. The preparation and characterization of silver-loading cellulose acetate hollow fiber membrane for water treatment. Polym. Adv. Technol., 2005; 16(8):600-607.
- Choi, O., Hu, Z. Size dependent and reactive oxygen species related nanosilver toxicity to nitrifying bacteria. Environ. Sci. Technol., 2008; 15(12):4583–4588.
- Sharma, V., Yngard, R., Lin, Y. Silver nanoparticles: green synthesis and their antimicrobial activities. Advances in Colliod and Interface Science., 2009; 145(1-2):83- 96.
- Crittenden, J.C., Vaitheeswaran, K., Hand, D.W., Howe, E.A., Aieta, E., Tate, C.H., McGuire, M.J., Davis, M.K. Removal of dissolved organic carbon using granular activated carbon. Water research. 1993; 27(4): 715-721.
- Cook, D., Newcombe, G., Sztajnbok, P. The application of powdered activated carbon for MIB and geosmin removal: predicting pac doses in four raw waters. Water Research. 2001; 35(5):1325-33.
- USEPA. US Environmental Protection Agency nanotechnology white paper. Science Policy Council, Washington DC (100/B-07/001) 2007.
- Korotta-Gamage, S.M., Sathasivan, A. A review: Potential and challenges of biologically activated carbon to remove natural organic matter in drinking water purification process. Chemosphere., 2017; 167:120-138.
- El-Naggar, M.Y., El-Aassar, S.A., Abdul-Gawad, S.M. Solid-State fermentation for the production of Meroparamycin by Streptomyces sp. strain MAR01. Journal of Microbiology. 2009; 19(5): 468-473.
- Sastry, M., Ahmad, A., Khan M.I., Kumar, R. Biosynthesis of metal nanoparticles using Fungi and Actinomycetes. Curr Sci., 2003; 85:162–170.
- Sadhasivam, S., Shanmugam, P., Yun, K. Biosynthesis of silver nanoparticles by Streptomyces hygroscopicus and antimicrobial activity against medically important pathogenic microorganisms. Colloids Surf. B. Biointerfaces., 2010; 81:358–362.
- Samundeeswari, A., Dhas, S.P., Nirmala, J., John, S.P., Mukherjee, A., Chandrasekaran, N. Biosynthesis of silver nanoparticles using actinobacterium Streptomyces albogriseolus and its antibacterial activity. Biotechnol. Appl. Biochem., 2012; 59(6):503–507.
- Mohanta, Y.K., Behera, S.K. Biosynthesis, characterization and antimicrobial activity of silver nanaoparticles by Streptomyces sp. SS2. Bioprocess Biosyst. Eng., 2014; 37(11):2263-2269.
- Karthik, L., Kumar, G., Kirthi, A.V., Rahuman, A.A., Rao, K.V. Streptomycessp. LK3 mediated synthesis of silver nanoparticles and its biomedical application. Bioprocess Biosyst. Eng. 2014; 37(2):261-267.
- El-Naggar, M.Y., El-Aassar, S.A., Abdul-Gawad, S.M. Meroparamycin production by newly isolated Streptomyces sp. Strain MAR01: Taxonomy, fermentation, purification and structural elucidation. Journal of microbiology., 2006; 44(4):432-438.
- El-Naggar, N.E., Abdelwahed, A.M. Application of statistical experimental design for optimization of silver nanoparticles biosynthesis by a nanofactory Streptomyces viridochromogenes. J. Microbiol., 2014; 52:53-63.
- Morgan, M.C., Boyette, M., Goforth, C., Katharine, V.S., Shermalyn, R. Biosynthesis, characterization and antimicrobial activity of silver nanoparticles by Streptomyces sp. SS2 Greece. J. Microb. Methods. 2009; 79:336–343.
- Biswas, M., Rahman, M.A., Khatun, M.H., Islam, M. Isolation and characterization of Streptomyces sp. ANBS-15 and antimicrobial activities of its secondary metabolites. Bangladesh Pharm. J., 2011; 14(1):15–20.
- Plackett, R.L., Burman, J.P. The design of optimum multifactorial experiments. Biometrika., 1946; 33: 305-325.
- Al-Sarrani, A.Q., El-Naggar, M.Y. Application of Plackett Burman factorial design to improve citrinin production in Monascus ruber batch cultures. Bot. Sci. (Formerly Bot. Bull. Acad. Sin.). 2006; 47(2):167-174.
- Youssef, A.S., El-Naggar, M.Y., El-Aassar, S.A., Beltagy, I.A. Optimization of culture conditions for â-Mannanase production by a local Aspergillus nigerisolate. Int. J. Agri. Biol., 2006; 8(4): 539-545.
- El-Naggar, N.E., Abdelwahed, A.M., Osama, M.M.. Fabrication of biogenic antimicrobial silver nanoparticles by Streptomyces aegyptia NEAE 102 as Eco-Friendly nanofactory. J. Microbiol. Biotechnol., 2014; 24(4):453–464.
- El-Aassar, A.M., Said, M.M., Abdel-Gawad, A.M., Shawky, H.A. Using silver nanoparticles coated on activated carbon granules in columns for microbiological pollutants water disinfection in Abu Rawash area, Great Cairo, Egypt. Australian Journal of Basic and Applied Sciences. 2013; 7(1):422-432.
- Margo, E., Hunt. Thermotolerant (Fecal) Coliform membrane filter procedure. Standard method 22nd edition, Microbiological examination, part 9000, 9222D (9 – 11) 2011.
- Brozel, V.S., Cloete, T.E. Effect of storage time and temperature on the aerobic plate count and on the community structure of two water samples. Water SA., 1991; 17(4):289-300.
- Pranay, J., Vibhor, A. Synthesis, Characterization and antimicrobial effects of silver nanoparticles from microorganisms-A review. International Journal of Nano and Material Sciences., 2012; 1(2):108-120.
- Traci, R., Jensen, George, C., Shatz, Richard, P., Duyne, V. Nanosphere lithography: Surface Plasmon Resonance spectrum of a periodic array of silver nanoparticles by Ultraviolet”Visible extinction spectroscopy and electrodynamic modeling. J. Phys. Chem. B. 1999; 103(13):2394-2401.
- Sosa, I.O., Noguez, C., Barrera, R.G. Optical properties of metal nanoparticles with arbitrary shape. J. Phys. Chem., 2003; 107:6269-6275.
- Morones, J.R., Elechiguerra, J.L., Camacho, A., Holt, K., Kouri, J.B., Ramirez, J.T., Yacaman, M. J. The bactericidal effect of silver nanoparticles. Nanotechnology., 2005; 16: 2346-2353.
- Jasmine, S., Krishnan, K. Antimicrobial activity of Streptomyces sp. VITBT7 and its synthesized silver nanoparticles against medically important fungal and bacterial pathogens. Scholars Research Library., 2013; 5 (3):192-200.
- Zhang, J., Marcin, C., Shifflet, M.A., Salmon, P., Brix, T., Greasham, R., Buckland, B., Chartrain M. Development of a defined medium fermentation process for physotigmine production by Streptomyces griseofuscu. Appl. Microbiol. Biotechnol., 1996; 44:568-75.
- Rita, E.J., Subbiayh, R. Biosynthesis of Silver Nanoparticles from Streptomyces olivaceous and its antimicrobial activity. International Journal of Pharma Research and Health Sciences, 2014; 2(2):166-172.
- Saifuddin, N., Wong, C.W., Yasumira, A.A., Yasumira, N. Rapid biosynthesis of silver nanoparticles using culture supernatant of bacteria with microwave irradiation. J. Chem., 2009; 6:61-70.
- Martinez-Castanon, G.A., Nino-Martinez, N., Martinez-Gutierrez, F., Martinez-Mendoza, J.R., Ruiz, F. Synthesis and antibacterial activity of silver nanoparticles with different sizes. Nanotechnology., 2008; 10(8):1343-1348.
- Priyaragini, S., Sathishkumar, S.R., Bhaskararao, K.V. Biosynthesis of silver nanoparticles using actinobacteria and evaluating its antimicrobial and cytotoxicity activity. International Journal of Pharmacy and Pharmaceutical Sciences, 2013; 5(2):709-712.
© The Author(s) 2017. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.