ISSN: 0973-7510
E-ISSN: 2581-690X
Genus Streptomyces under phylum actinobacteria has been recognized as a prolific source for production of bioactive secondary metabolites. Actinobacterial strain designated as SKCMM1 isolated from Pichavaram Mangrove forest sediment was identified as Streptomyces cavourensis using polyphasic taxonomic approach. This strain shares 99% sequence similarity with Streptomyces cavourensis NBRC 13026T. Ethyl acetate fraction of the strain SKCMM1 exhibited highest biological activity. FTIR and GCMS analysis of crude compounds isolated from the active ethyl acetate fraction states the presence of several phenolic, hydrocarbon and fatty acid compounds with various bioactivities. This study will be an attempt to understand that the strain holds significant antimicrobial activity against test pathogens and antiproliferative activity against HeLa cell lines (IC50 value of 8.9µg/ml).
Streptomyces, Polyphasic Taxonomy, Liquid – liquid extraction, Spectral analysis, Bioactive compounds
Actinobacteria are present in various ecological habitats such as soil, fresh water, backwater, lake, sewage and marine environment1. These bacteria are bounteous group of microorganisms that produce a wide array of secondary metabolites such as antibiotics, antitumor agents, immunosuppressive agents, vitamins, enzymes and even cosmetics2. On contemplating the microbes of the same region, the marine Actinobacteria are unique for their antibiotic production in fluctuating physical, chemical and biological factors3. In recent years the escalating diligence is towards the exploration of novel microbes that manufacture metabolites to combat drug resistant pathogenic microbes4-6.
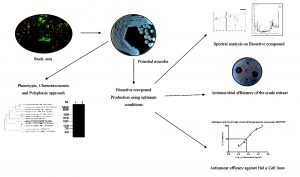
Marine ecosystem is a resplendent choice for the isolation of novel microbes that confer splendid outcome in identification, characterization and application of secondary metabolites7. Earlier, Streptomyces sp. VITMK1 was isolated from Pichavaram region of Tamil Nadu and it yielded a potent antibacterial compound diketopiperazine5. Similarly, ethyl acetate crude extract obtained from Streptomyces strain CH54-4 was reported for best antimicrobial and anti proliferative activity8. In the current study, Streptomyces SKCMM1 is isolated from Pichavaram Mangrove forest of Tamil Nadu and identified through polyphasic taxonomic characterization approach. The antagonistic ability of the isolate was assayed through shake flask fermentation and spectral studies (Fig. 1).
Isolation and Identification of Potential Actinomycete
Five marine sediment samples were collected from Pichavaram Mangrove forest, Tamil Nadu using a sterile borer at a depth of 10 to 15 cm (Table 1 and Figure 2). The serially diluted (10-1 to 10-5) samples were seeded on Starch Casein agar (SCA) (HiMedia, India) plates supplemented with antibiotics (Cycloheximide 50µg/ml and Nystatin 50µg/ml) and incubated at room temperature for 10 days. Characteristic tough leathery colonies of Actinomycetes with a clear zone of inhibition were selected and preserved in SCA slants for further use. Isolates were grown in 50 ml of starch casein broth by submerged culture method (32°C for 7 days), centrifugation was carried out at 8000 rpm for 15 min and the clear supernatant was challenged against selected bacterial pathogens (E.coli MTCC 1698, E.coli ESBL, Bacillus pumilis NCIM2327, Proteus vulgaris, Staphylococcus aureus MTCC 3160, Staphylococcus aureus (methicillin resistant), Shigella flexneri MTCC 1457) by well diffusion method. The potent Actinobacteria was selected based on the vast zone of inhibition and it was taken for further studies.
Phenotypic Characterization of Strain SKCMM1
Potential Actinobacterial strain SKCMM1 was characterized morphologically and physiologically as prescribed in International Streptomyces Project (ISP)9. The micro-morphology of strain was observed under light microscopy and Scanning Electron Microscopy (JEOL JSM-5610) after incubation at 28°C for 7 days. The pigmentation of aerial mycelium and structure of sporophores were observed on different ISP media 1 – 7, which are an ideal aspect in its classification. Colony morphology was recorded with respect to size, colour, aerial mycelium, substrate mycelium and pigmentation. Addition to that, biochemical and physiological examinations were performed according to standard methodology5, 10. Standard procedures were used to determine the cell wall amino acids11 and cell wall sugars12 of the potent Actinobacterial strain. Fatty acids extracted from isolate SKCMM1 were methylated and analyzed by GC-MS, using the standard sherlock microbial identification system13.
16s rRNA Gene Sequencing
Genomic DNA was extracted and subjected to polymerase chain reaction14 targeting 16S rRNA gene using the 5’-AGAGTTTGATCCTGGCTCAG-3’– forward primer and 5’TACGGCTACCTTGTTACGACT-3’ – reverse primer15. The reaction mixture contained 10X Taq buffer, 25 mM MgCl2, 4 mM dNTPs, Taq polymerase 0.5U, template DNA 5-10 ng and primers at 10 pmol concentration (Fermentas, Thermo Scientific, Vilnius, Lithuania). The PCR conditions comprised of initial denaturation at 94oC for 10 min followed by 23 cycles of denaturation at 94oC for 15 sec, annealing at 55oC for 15 sec, extension at 72oC for 1 min and a final extension of 72oC for 10 min carried out in Master thermal cycler (Eppendorf, Germany). Later, amplicon sequencing was done in ABI 3730xl cycle sequencer (Facility from Sciegenom, Cochin, India) and sequences were deposited in GenBank (NCBI). The dendrogram analysis was done in MEGA 4.0 to study the evolutionary similarity of SKCMM1 strain.
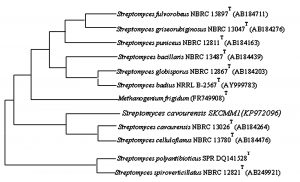
Fig. 4. Neighbor-joining phylogenetic tree based on 16S rRNA gene sequences showing the relationships of strain SKCMM1 with related species
Production and Extraction of Bioactive Compound using Streptomyces cavourensis SKCMM1
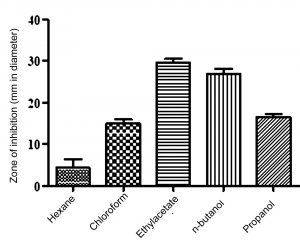
For production, 125 ml of Starch casein broth SKCMM1 (500 ml Erlenmeyer flask) was inoculated with 1 ml Streptomyces cavourensis and incubated at 28oC in a rotary shaker (180 rpm) for 7 days. Then, broth was filtered using Whatmann No 42 aseptically and stored at 4oC. In order to select the best choice of solvent for active compound extraction, an equal volume of various solvents (hexane, chloroform, ethyl acetate, n-butanol and propanol) along with filtrate were shaken well for 60-90 mins. The collected solvent fractions were concentrated using distillation column and the crude obtained was challenged against test bacterial strains. Based on the zone of inhibition results, ethyl acetate was selected as the best solvent for compound extraction (Figure 5). The fermented broth and ethyl acetate was treated at a ratio of 1:2 to extract bioactive molecules.
Fig. 5. Results of solvent extraction showing ethyl acetate as best choice of solvent for active compound extraction
Spectral analysis of ethyl acetate extract
The active extract dissolved in methanol was taken for UV Vis and FTIR analysis using Shimadzu – 1601 UV VIS and Shimadzu IR 8000 respectively. The functional groups present in the sample were detected in the spectral range of 400 to 4000 cm-1. The ethyl acetate sample extract was analyzed by GC-MS [Thermo GC – Trace Ultra Version 5.0]. In this method, a 30m×0.25 mm ZB 5 – MS capillary Standard Non-Polar column with a film thickness of 0.25µm was used. The carrier gas was helium which had a column flow rate of 1 mL/min (at a pressure of 105 kPa). Then, 1.0 µL of ethyl acetate sample was injected with the following column program 50°C@7°C/min to 200°C (3 min) @7°C/min to 280°C (15 min) [Scan range: 40 – 1000 m/z]. Every discrete constituent showed by GC MS tandem analysis was used to compare the compounds and functional groups identified with a standard compound of NIST library (Version 2005).
Anticancer activity of crude extract against HeLa cell lines
The HeLa cell line obtained from National Centre for Cell Science (NCCS), Pune was grown in RPMI-1640 containing 10% fetal bovine serum (FBS). Single cell suspensions were disintegrated from monolayer cells using trypsin-ethylene diaminetetraacetic acid (EDTA). By subsequent medium dilution, the viable cells with a final density of 1×105 cells/mL were acquired. The cell suspension (100µl) was incubated in 96 well plate (103 cells/well) for 24 hrs to achieve cellular adherence. Later, test samples were added in varying concentrations of 0 – 50 µg/ml dimethylsulfoxide (DMSO) to cells and kept for 48 hr incubation. Afterwards, 15 µl of MTT (5 mg/mL) in phosphate buffered saline (PBS) was added to each well and incubated at 37oC for 4 hrs. Finally, the medium with MTT was then flicked off and the formazan crystals were solubilized in 100 µl of DMSO to measure the absorbance at 570 nm using micro plate reader. The mean value IC50 was calculated by nonlinear regression analysis and graph was plotted between percentage of cell inhibition and Log10 concentration using GraphPad Prism software.
In the current investigation, five sediment samples collected from mangrove region were had slight variation in their physical parameters. Notably, the temperature of sediments ranged between 24oC to 29oC, while the pH varied from 7.5 to 8.2 which are considered to be optimum for Actinobacterial growth and endurance. Totally twelve actinobacterial isolates obtained from sediment samples were sorted out for superlative samples depending on temperature and pH conditions having potential microbial diversity (Table 1 and Figure 2). Generally, Actinobacteria living in these conditions are capable of growing in extreme and incessantly changing environment which is a quintessential representation for survival in microbial diversity. Also, studies conducted on Actinobacteria in mangrove soil by many eminent researchers suggest that this may actually be due to the better quality of the sediments, described in terms of structure, pH and humic substance8,16.
Table (1):
Characteristics of samples collected from Pichavaram Mangrove forest, India.
Sample no |
Latitude (N) |
Longitude(E) |
Temperature (o C) |
pH |
|---|---|---|---|---|
MS1 |
11° 30′ 11” |
79° 46′ 37” |
24 |
7.9 |
MS2 |
11° 28′ 37” |
79° 47′ 15” |
28 |
7.5 |
MS3 |
11° 27′ 51” |
79° 47′ 24” |
28 |
8.2 |
MS4 |
11° 27′ 22” |
79° 47′ 17” |
29 |
7.7 |
MS5 |
11° 27′ 24” |
79° 46′ 46” |
26 |
8.1 |
MS = Marine Sediment
In the well diffusion assay of current investigation, all the twelve isolates were active against selected clinically pathogenic bacterial strains. Especially the isolate named SKCMM1 showed an extraordinary activity against all the Gram positive and Gram negative reference bacterial strains used in this investigation. The isolate SKCMM1 showed conspicuous antimicrobial activity in terms of zone of inhibition (p value = 0.0005) in mm ranging from 8 to 27 against test pathogens (Table 2). For this reason, isolate Streptomyces SKCMM1 is focused for centralized assiduity. Generally, actinobacterial populations of mangrove forest are capable of tolerating a wide span of catastrophes, they have not been extensively explored for the registration of novel bioactive compounds. Thus, it is crucial to note that broad spectrum activity of marine microorganisms from hitherto unexplored habitats should be considered novel source for bioactive secondary metabolites.
Table (2):
Antimicrobial activity of the isolates using well diffusion assay.
SKCMM 1 |
SKCMM 2 |
SKCMM 3 |
SKCMM 4 |
SKCMM 5 |
SKCMM 6 |
SKCMM 7 |
SKCMM 8 |
SKCMM 9 |
SKCMM 10 |
SKCMM 11 |
SKCMM 12 |
|
|---|---|---|---|---|---|---|---|---|---|---|---|---|
1 |
12.3 ± 1.5 |
0 |
11.7 ± 1.2 |
0 |
13.3 ± 1.5 |
0 |
0 |
0 |
12.3 ± 0.6 |
8.7 ± 1.2 |
10.7 ± 1.2 |
12.7 ± 1.5 |
2 |
17.7 ± 1.2 |
7 ± 1.0 |
0 |
0 |
11.0 ± 1.7 |
0 |
0 |
11.3 ± 1.5 |
0 |
21.0 ± 1.0 |
0 |
8.3 ± 1.2 |
3 |
27.0 ± 1.0 |
9.3 ± 1.2 |
0 |
0 |
0.0 |
0 |
15.3 ± 1.2 |
0 |
6.0 ± 1.7 |
0.0 |
18.0 ± 0.0 |
12.0 ± 1.7 |
4 |
12.7 ± 1.5 |
0 |
0 |
7.7 ± 1.2 |
19.0 ± 1.0 |
24.0 ± 0.0 |
9.7 ± 1.2 |
0 |
0 |
23.0 ± 1.7 |
0 |
0 |
5 |
9.3 ± 1.2 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
7.3 ± 1.5 |
0 |
0 |
0 |
6 |
14.0 ± 0.0 |
0 |
0 |
0 |
0 |
16.0 ± 1.7 |
9.3 ± 2.5 |
0 |
0 |
0 |
0 |
0 |
7 |
11.7 ± 1.2 |
0 |
0 |
0 |
0 |
11.0 ± 1.7 |
0 |
0 |
0 |
0 |
0 |
0 |
1. Staphylococcus aureus MTCC 3160, 2. Bacillus pumilis NCIM 2327, 3. Staphylococcus aureus (methicillin resistant) 4. Escherichia coli MTCC 1698 5. Escherichia coli (ESBL), 6. Shigella flexneri MTCC 1457, 7. Proteus vulgaris
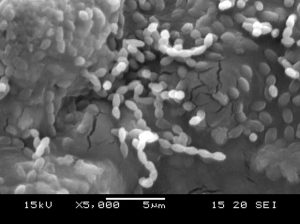
In the present investigation, morphology of potential isolate SKCMM1 was tested on various media such as Starch casein agar, Actinomycetes isolation agar, Nutrient agar and ISP (International Streptomyces Project) media (ISP 1 to 7). The isolate produced circular, pale green colour colonies on Starch casein agar media. The size of the colony measured from 0.5 to 0.75 mm in dm. Further morphological details are given in Table 3. The isolate SKCMM1 showed substantial growth on all the media used for the growth characterization (Table 4). SEM studies revealed well developed and unfragmented aerial and substrate mycelia of the strain SKCMM1 (Figure 3). The spore chain was in straight, rectiflexibiles and spore surface was very smooth, oval shaped and arranged in a long chain.
Table (3):
Phenotypic characterization of the isolate SKCMM1 on Starch Casein Agar.
Phenotypic characteristics |
SKCMM1 |
|---|---|
Colony morphology |
Circular, umbonate, entire |
Sporophore morphology |
Straight or Rectiflexibiles |
Spore surface |
Smooth |
Colour of aerial mycelium |
Pale green |
Colour of substrate mycelium |
Brown |
Spore mass |
Pale green |
Pigment |
Brown colour |
Table (4):
Cultural characteristics of isolate SKCMM1 on International Streptomyces Project media.
Media |
Growth |
Aerial Mycelium |
Substrate Mycelium |
|---|---|---|---|
ISP1 |
++++ |
Creamy White |
Brown |
ISP2 |
+++ |
Creamy White |
Mild Brown |
ISP3 |
++++ |
Pale Green |
Pale Yellow |
ISP4 |
++++ |
Pale Green |
White |
ISP5 |
++++ |
Pale Green |
Mild Brown |
ISP6 |
++++ |
Pale Green |
Dark Brown (Soluble Pigment Detected) |
ISP7 |
++++ |
Pale Green |
Blackish Brown |
NA |
++++ |
Creamy White |
Brown |
SCA |
++++ |
Pale Green |
Brown (Soluble Pigment Detected) |
AIA |
++++ |
Creamy White |
Brown |
Growth as per article; ++++ Excellent; ++ Moderate; +++ Good; + Poor; ISP- International Streptomyces Project; AIA- Actinomycete isolation agar; NA- Nutrient agar.
In chemotaxonomic characterization, no cell sugars were detected on chromatogram comparing with the standard lanes loaded with sugars such as galactose, glucose, arabinose, mannose, ribose and rhamnose. Also in cell wall amino acid analysis, two amino acids spots were detected in sample lane against the standards LL-2,6 diaminopimelic acid and glycine. Presence of LL-2,6 diaminopimelic acid and absence of standard cell wall sugars reveals the cell wall of strain SKCMM1 confirmed that cell wall belongs to type I category. The results were in high agreement with previous studies on Streptomyces sp identification8, 21. The cell wall fatty acids were extracted from isolate SKCMM1 and analyzed in GC MS (Table 5). Presence of Tetradecanoic acid, Pentadecanoic acid, Hexadecanoic acid and Heptadecanoic acid in the form of methyl esters confirming the cell wall belonged under Streptomyces13.
Table (5):
GC-MS analysis of Fatty acid methyl esters in cell wall of isolate SKCMM1.
Retention time |
Compound name |
Molecular Formula |
Molecular Weight |
Area % |
|---|---|---|---|---|
19.01 |
12-Methyltetradecanoic acid |
C16H32O2 |
256 |
56.60 |
19.62 |
Pentadecanoic acid, methyl ester |
C16H32O2 |
256 |
7.10 |
21.64 |
Hexadecanoic acid, methyl ester |
C17H34O2 |
270 |
21.30 |
23.03 |
Heptadecanoic acid, methyl ester |
C18H36O2 |
284 |
14.99 |
The isolate SKCMM1 was sequenced using primer specific for Streptomyces sp using 16s rRNA gene sequencing and deposited in GenBank (NCBI) as a strain named Streptomyces cavourensis SKCMM1 with the following accession number of KP972096. Streptomyces cavourensis SKCMM1 showed 99% homology with Streptomyces cavourensis strain NBRC 13026T on blast analysis. The GC content of the isolate was about 58.2% which is a typical characteristic of the Streptomyces. For dendrogram construction (using MEGA 4.0), 16s rRNA gene sequence of the isolate SKCMM1 and close neighbor of isolate sequence was included in the sequence – based comparative analysis that revealed high similarity with Streptomyces cavourensis NBRL 13026 (AB184264) and Streptomyces cavourensis NBRL 13780 (AB184476) (Figure 4).
Active compound separation from the fermented broth showed that ethyl acetate is a best choice of solvent (p value = 0.0009) (Figure 5). Similar findings have been reported by various investigators upon using ethyl acetate as a best choice of solvent for extraction of bioactive compounds5,22. The active fraction of ethyl acetate fraction was significantly activity against reference bacterial strains used in this study (Table 6). The condensed crude is considered as partially purified bioactive compound and taken for spectral studies.
Table (6):
Antimicrobial activity of ethyl acetate crude of SKCMM1 against reference bacterial strains.
| Name of the Organism | Zone of Inhibition(mm) | |
|---|---|---|
| Gram Positive | ||
| Staphylococcus aureus MTCC 3160 | 34.3 ± 1.52a | |
| Bacillus pumilis NCIM 2327 | 25.0 ± 1.73a | |
| Staphylococcus aureus (methicillin resistant) | 32.0 ± 1.73a | |
| Gram Negative | ||
| Escherichia coli MTCC 1698 | 24.3 ± 3.1a | |
| Escherichia coli (ESBL) | 18.3 ± 1.2a | |
| Shigella flexneri MTCC 1457 | 18.6 ± 1.5a | |
| Proteus vulgaris | 19.0 ± 2.0a | |
a = p< 0.05
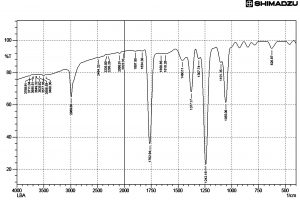
Fig. 6. FT IR Spectrum of active ethyl acetate extract of Streptomycetes cavourensis SKCMM1 fermented broth
FT IR spectral studies play a major role in the identification of active compounds. The data obtained through this study will disclose the nature of compounds and functional groups present in the sample. The FT IR results showed 26 elevated peaks in the range between 3800 cm-1 and 600 cm-1 (Figure 6). The peak values indicated the presence of functional groups such as C=C alkenes, C=O group of acids, C=C alkynes, -C-H stretch (alkane H) and O-H (hydrogen bond, intermolecular, polymeric association). The spectral data of FT-IR reveals the presence of large broader where peaks were observed at 1241cm-1 and 1762cm-1 indicating the presence of alcohols, ester, ethers and carboxylic acid groups. The two peaks between 1055cm-1 and 1377cm-1 indicated the presence of primary alcohols and nitro compounds respectively in the sample. The presence of a peak at 2989cm-1 indicated alkanes. The presence of 4 peaks between 3200cm-1 to 3600cm-1 range indicated the presence of monomeric alcohols and phenols.
The ethyl acetate extract of Streptomycetes cavourensis SKCMM1 was subjected to gas chromatography-mass spectrometry analysis. The identification of compound is based on the peak area and mass by charge ratio. This peak area is directly proportional to the quantity of the compound present in the extract. The volatile metabolic profiling is expressed as chromatogram and the results were tabulated (Table 7). The GC MS results indicated the presence of more than 27 different compounds notably 1, 4-Dioxane-dimethanol (13.06%), 1, 3-Bisbenzo thiophene (8.66%), 1,3,5-trimethyl-2-octadecyl cyclohexane (7.58%), 2,4,6-Decatrienoic acid (2.74%), a-Amyrin (2.24%), Cyclopropane carboxamide (0.8%).
Table (7):
GC-MS analysis of the ethyl acetate extract of Streptomycetes cavourensis SKCMM1 fermented broth.
Classification |
Compound |
% |
|---|---|---|
Aniline Compound |
2,4-Dimethoxy- tri phenylamine |
0.52 |
Fatty acid |
Pentanoic acid |
1.76 |
2,4,6-Octatrienoic acid |
2.74 |
|
Hydroxy Compound |
1,4-Dioxane-2,6-dimethanol |
13.06 |
Caryophyllenyl alcohol |
0.67 |
|
3-Cyclohexen-1-ol |
0.67 |
|
Geosmin |
2.48 |
|
Hydrocarbons |
1,5-Bis pentane |
0.47 |
Docosane |
0.50 |
|
Cycloheptanone |
1.12 |
|
Benzene |
2.12 |
|
Cycloalkane compound |
Cyclohexane |
7.58 |
Cyclopropane |
1.11 |
|
Aromatic compound |
1,3-Bisbenzo thio phene |
8.66 |
Carboxamide |
Cyclopropane carboxamide |
0.8 |
Unknown compounds |
55.74 |
1, 4-Dioxane-dimethanol compound contain two -OH stretch at 2nd and 6th position and FTIR investigation (two larger boarder peaks observed at 1241cm-1 and 1762cm-1) also supports the presence of alcohol groups in the partially crude extract. Recently, 1, 4-Dioxane-dimethanol has been filed for its anti-diabetic activity23. The second major compound resulted during mass spectroscopic study is 1, 3-Bisbenzo thiophene. Thiophene is a heterocyclic compound containing of planar five membered ring structure. Several, thiophene derivative compounds have been reported for biological activities such as antibacterial activity24 and antiviral efficiency25. Thus 1, 3-Bisbenzo thiophene could also be an bioactive thiophene compound against the test pathogens.
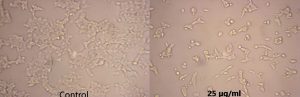
Fig. 7. Anticancer activity of the crude extract of Streptomyces cavourensis SKCMM1 against cancer cell lines
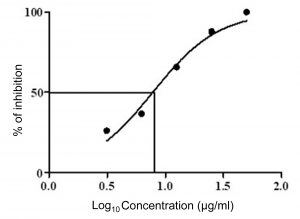
Fig. 8. Dose response curves (GraphPad Prism) of crude extract of Streptomyces cavourensis SKCMM1 against cancer cell lines
The antiproliferative efficiency of SKCMM1 strain against HeLa cell lines has an enormous IC50 value of 8.9 µg/ml (R2 = 0.9732) (Figure 7 and 8). ANCI (American National Cancer Institute) endorses that IC50 value of any crude compound falling down to 30 µg/ml as a potent drug28. Some of the most dynamic bioactive compound observed during spectral studies were a-Amyrin and Cyclopropane carboxamide despite its small proportion in the crude extract. a-Amyrin belongs to the triterpene class of compounds and has been reported for antimicrobial, antifungal and anti-inflammatory26. The presence of a-Amyrin in the crude could be the reason for the anti-inflammatory activity expressed against HeLa cell lines. Similarly, some of the structural derivatives of cyclopropane carboxamide has been reported for antifungal activity27. In this, most of them were previously reported for various biological activities such as antimicrobial, anti-inflammatory and cytotoxic activities. Therefore, it is suggested that further investigation may explore the significance of these unnoticed compounds.
In the present investigation, it is discernible that marine Actinobacterial populations from Pichavaram Mangrove region harbor plentiful biological activities particularly antimicrobial compounds production. These regions have magnificently concealed therapeutic and economic biological potential which is highly praised and agreed with the perception of other investigators4,16-20.
As myriad of studies have been conducted on bacteria to look out for novel bioactive compounds, geographical locations such as mangrove forest may offer a potent microbial diversity for scrutinization. As an attempt to identify an effective microbe, here a study was conducted at Pichavaram mangrove forest, Tamil Nadu which culminated intensely propitious Streptomyces cavourensis SKCMM1 strain which was isolated and characterized through polyphasic taxonomy. Secondary metabolites proved to have broad spectrum antibacterial and antiproliferative efficiency against HeLa cell lines. Spectroscopic studies of partially purified extract enunciated several activities antimicrobial, antioxidant and cytotoxic compounds. A kindling approach for assessing purification of such compounds may exhibit their connotation in pharmaceutical industries.
Acknowledgements
I would like to thank the management of Sri Krishna Arts and Science College, Coimbatore for providing necessary facilities to carry out this work.
Conflict of Interest
The author(s) declare that there is no conflict of interest.
- Williams ST, Goodfellow M, Alderson G, Wellington EM, Sneath PH, Sackin MJ. Numerical classification of Streptomyces and related genera. J Gen Microbiol. 1983;129(1 983):1743–813.
- Da Rocha AB, Lopes RM, Schwartsmann G. Natural products in anticancer therapy. Curr Opin Pharmacol [Internet]. 2001;1(4):364–9.
- Okazaki T, Okami Y. Actinomycetes in Sagami Bay and their antibiotic substances. J Antibiot (Tokyo). 1972; 25(8): 461–6.
- Mangamuri U, Muvva V, Poda S, Naragani K, Munaganti RK, Chitturi B, et al. Bioactive metabolites produced by Streptomyces Cheonanensis VUK-A from Coringa mangrove sediments: isolation, structure elucidation and bioactivity. 3 Biotech [Internet]. Springer Berlin Heidelberg; 2016 Jun 13; 6(1):63.
- Manimaran M, Gopal JV, Kannabiran K. Antibacterial Activity of Streptomyces sp. VITMK1 Isolated from Mangrove Soil of Pichavaram, Tamil Nadu, India. Proc Natl Acad Sci India Sect B Biol Sci [Internet]. 2017 Jun 2; 87(2):499–506.
- Vela Gurovic MS, Olivera NL. Antibacterial producing actinomycetes from Extra Andean Patagonia. J Arid Environ [Internet]. Elsevier Ltd; 2017 Sep; 144: 216–9.
- Fiedler HP, Bruntner C, Bull AT, Ward AC, Goodfellow M, Potterat O, et al. Marine actinomycetes as a source of novel secondary metabolites. Antonie van Leeuwenhoek, Int J Gen Mol Microbiol. 2005; 87(1):37–42.
- Srivibool R, Jaidee K, Sukchotiratana M, Tokuyama S, Pathom-Aree W. Taxonomic characterization of Streptomyces strain CH54-4 isolated from mangrove sediment. Ann Microbiol. 2010; 60(2):299–305.
- Shirling EB, Gottlieb D. Methods for characterization of Streptomyces species. Int J Syst Bacteriol. 1966; 16(3): 313–40.
- Flowers TH, Williams ST. The influence of pH on the growth rate and viability of neutrophilic and acidophilic streptomycetes. Microbios [Internet]. 1978; 18(73–74): 223–8. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27702
- Becker B, Lechevalier MP, Gordon RE, Lechevalier H a. Rapid Differentiation Between Nocardia and Streptomyces By Paper Chromatography of Whole-Cell Hydrolysates. Appl Microbiol [Internet]. 1964; 12: 421–3.
- Suput J, Lechevalier MP, Lechevalier H a. Chemical composition of variants of aerobic actinomycetes. Appl Microbiol [Internet]. 1967; 15(6): 1356–61.
- Sasser M. Identification of Bacteria by Gas Chromatography of Cellular Fatty Acids. Tech Note. 2001;101(February):1–6.
- Nikodinovic J, Barrow KD, Chuck JA. High yield preparation of genomic DNA from Streptomyces. Biotechniques. 2003;35(5):932–6.
- McVeigh HP, Munro J, Embley TM. Molecular evidence for the presence of novel actinomycete lineages in a temperate forest soil. J Ind Microbiol Biotechnol. 1996; 17: 197–204.
- Ramesh S, Mathivanan N. Screening of marine actinomycetes isolated from the Bay of Bengal, India for antimicrobial activity and industrial enzymes. World J Microbiol Biotechnol. 2009; 25(12): 2103–11.
- Sujatha P, Bapi Raju KVVSN, Ramana T. Studies on a new marine streptomycete BT-408 producing polyketide antibiotic SBR-22 effective against methicillin resistant Staphylococcus aureus. Microbiol Res. 2005; 160(2): 119–26.
- Karthik L, Gaurav K, Rao KVB, Rajakumar G, Rahuman AA. Larvicidal, repellent, and ovicidal activity of marine actinobacteria extracts against Culex tritaeniorhynchus and Culex gelidus. Parasitol Res. 2011; 108(6): 1447–55.
- Meena B, Rajan LA, Vinithkumar NV, Kirubagaran R. Novel marine actinobacteria from emerald Andaman & Nicobar Islands: a prospective source for industrial and pharmaceutical byproducts. BMC Microbiol [Internet]. 2013; 13(1):145.
- Valliappan K, Sun W, Li Z. Marine actinobacteria associated with marine organisms and their potentials in producing pharmaceutical natural products. Vol. 98, Applied Microbiology and Biotechnology. 2014. p. 7365–77.
- Phongsopitanun W, Thawai C, Suwanborirux K, Kudo T, Ohkuma M, Tanasupawat S. Streptomyces chumphonensis sp. nov., isolated from marine sediments. Int J Syst Evol Microbiol [Internet]. 2014; 64(Pt 8):2605–10.
- Parthasarathi S, Sathya S, Bupesh G, Manikandan M, Kim CJ, Manikandan T, et al. Isolation, characterization and extraction of antimicrobial compound from marine actinomycete Streptomyces hygroscopicus BDUS 49. Res J Biotechnol. 2013; 8(3): 40–8.
- Del SP. Drugs for diabetes [Internet]. Google Patents; 2002. Available from: https://encrypted.google. com/patents/WO200203 0867A2?cl=ja
- Khalil AM, Berghot MA, Gouda MA. Synthesis and antibacterial activity of some new thiazole and thiophene derivatives. Eur J Med Chem. 2009; 44(11):4434–40.
- Churkin YD, Panfilova L V, Boreko EI, Timofeeva MM, Votyakov VI. Biologically active thiophene derivatives. IV. Synthesis and antiviral activity of unsaturated ketones of the thiophene series. Pharm Chem J [Internet]. 1982; 16(2): 103–5. Available from: https://doi.org/10.1007/BF00762027
- Vazquez LH, Palazon J, Navarro-Ocaña A. The Pentacyclic Triterpenes , α, β-amyrins: A Review of Sources and Biological Activities. In: Phytochemicals-A Global Perspective of their Role in Nutrition and Health. InTech; 2012.
- Liu XH, Shi YX, Ma Y, Zhang CY, Dong WL, Pan L, et al. Synthesis, antifungal activities and 3D-QSAR study of N-(5-substituted-1,3,4-thiadiazol-2-yl)cyclopropanecarboxamides. Eur J Med Chem. 2009; 44(7): 2782–6.
- Zheng Z, Zeng W, Huang Y, Yang Z, Li J, Cai H, et al. Detection of antitumor and antimicrobial activities in marine organism associated actinomycetes isolated from the Taiwan Strait, China. FEMS Microbiol Lett. 2000; 188(1): 87–91.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.