Recent instances of novel biological circuits that enable cells to gain biosynthetic skills demonstrate synthetic biology’s therapeutic potential. Synthetic biology is a branch of biology whose primary role is to build completely functional biological systems from the smallest basic elements such as DNA, proteins, and other organic molecules to complex bacteria. This review briefly mentions some novel way of synthetic strategies like bacterial modelling, two-component systems, synthetic peptide, and synthetic flavonoids used for targeting biofilm and drug-stable microbial communities. Bacterial modelling was mainly done in Escherichia coli and Mycoplasma using different strategies like introducing quorum sensing devices and CRISPR-mediated editing. Synthetic peptides are also one of the extensively studied ongoing areas which are produced from natural peptides taking as a template and altering amino acid position. Flavonoids are produced by two-step reaction and molecular hybridization methods. This kind of synthetic approach reported significant biofilm dispersion and lethal effects on clinically relevant bacteria like Pseudomonas aeruginosa, Staphylococcus aureus, E. coli, Acinetobacter baumannii, and Streptococcus species and Klebsiella pneumonia.
Synthetic Peptides, Bacterial Modelling, Engineered Phage, Modified Flavonoids, Bacterial Two-component System
Biofilm is one of the unsolved mysteries of medical science, which leads to the death of so many patients in medical hospitals.1 Out of total infections, 80% of infections reported in hospitals have involvement of biofilm. Biofilm can develop on both internal and external surfaces of the human body even though it makes significant complications in surgical implants like prosthetic heart valves, urinary catheters, and joint implants. Biofilm involved in commonly reported healthcare infections is endocarditis, cystic fibrosis, periodontitis, rhinitis, kidney infections etc.2 The most significant biofilm bacteria involved in implants are Staphylococcus aureus & Staphylococcus epidermidis.2,3 50-70% of catheter-mediated infections and 40-50% of prosthetic heart valve infections are caused by S.aureus & Staphylococcus epidermidis. Candida biofilm formation reported a 50% mortality rate in the United States. The most prevalent biofilm-producing bacteria reported in hospital fields are Enterococcus faecalis, Staphylococcus aureus, Staphylococcus epidermidis, Streptococcus viridans, E.coli, Klebsiella pneumoniae, Proteus mirabilis and Pseudomonas aeruginosa.3 Biofilm Extracellular Polymeric substances (EPS) consist of polysaccharides, teichoic acid, proteins and extracellular DNA. Due to the presence of thick EPS materials in the biofilm bacterial cells inside the biofilm are 100-1000 times more resistant to antibiotics.4 To overcome this situation physicians, use a higher concentration of drugs leading to the emergence of antimicrobial resistance (AMR).5-8 The metabolic rate inside and the surface of biofilm is different, coming to deeper metabolic rate is low compared to the surface. This gives a prolonged exposure to antibiotics less than a lethal dose to the inner side of biofilm-producing bacteria. This leads to the gradual development of antibiotic resistance. Disassociation of this bacteria from biofilm leads to the development of new AMR bacteria.9 Autoinducers play significant roles in the AMR process.10 The concentration of autoinducers depends on the population density and stimuli from the environment, when the concentration of autoinducers crosses the threshold leads to activation of some transcriptional factors and produces proteins helpful for adaptation favourable to environmental changes and virulence pathways.11,12 However, the actual autoinducer-mediated mechanism of AMR is still a lacunae and lacks proper strategies to target bacterial cells inside the biofilm. At present, little research focuses on developing new technology and strategy to target bacteria in the biofilm. But till now no proper methodology has been identified to deal with the mysteries of biofilm. In this review, we are focusing on the engineered bacterial systems, synthetic peptides, two-component systems (TCS), flavonoids, and modified phage which are showing some promising results in the application of degrading the biofilm (Table 1).
Table (1):
Synthetic units and their mechanisms.
| Name | Target | Mechanism | Application | Reference |
|---|---|---|---|---|
| Synthetic peptides | ||||
| DRGN1 Natural resource: Komodo dragon Production method: Solid-phase China peptide method |
P.aeruginosa S.aureus |
DRGN1 acts on the cytoplasmic membrane causing leakage of the intracellular components |
|
33 |
| FLEUCIN K59 Natural resource: Bombina orientalis Production method: Fmoc-solid phase method |
E.coli S.aureus A.baumannii |
Cell membrane damage and leakage of contents |
|
41 |
| GEMNIPEPTIDE Production method: Solid-phase method |
E.coli | Cell membrane damage and leakage of contents |
|
45 |
| Art-175+KZ-144 (Recombinant E. coli) |
Pseudomon as aureginosa, Acinetobac ter baumanii, E.coli | Enzymatic degradation of the PG layer and osmotic lysis |
|
47 |
| KBI-3221 (Streptococcus mutans) |
Streptococcus species | Quorum sensing inhibition |
|
45 |
| FLAVANOIDS | ||||
| CICI-Flav Production method: Two-step reaction |
E. coli K.pneumoniae |
Disruption of cell membrane integrity |
|
62 |
| FLUORINATED CHALCONE-1,2,3- TRIAZOLE Production method: Molecular hybridization |
E.coli |
|
57 | |
| ENGINEERED BACTERIAL SYSTEM | ||||
| E.coli | P.aureginosa | Antimicrobial peptide, dnase, micr ocin mediated killing |
|
13 |
| Mycoplasma | S.aureus | Enzyme mediated biofilm degradation(break down N-acetyl D glucosamine) |
|
20 |
| ENGINEERED BACTERIOPHAGES | ||||
| Modified T7 phage | E.coli | Dispersin B mediated distruction of biofilm |
|
69 |
| K1F phage | E.coli | Endosialoidase produced by phage make E.coli susceptible to Autophagy by capsular k antigen alteration |
|
70 |
Synthetic Biology Strategies
Bacterial modeling
Targeting biofilms using engineered bacterial strains is becoming popular among researchers due to the failure of antibiotics and the emergence of AMR in bacterial communities.13 Recent research has shown that commensal bacteria might be used as delivery methods for anti-virulence factors to treat bacterial and viral illnesses.14 The challenges faced in bacterial modelling are the movement of microbes toward the target cells and attacking the target by overcoming their antimicrobial strategies. To overcome this limitation, scientists reprogram the chemotaxis response of biofilm-destroying bacteria and selectively swim to the target pathogen. The scientists selected E.coli as a model organism to target biofilm-producing P.aeruginosa.13 For recognition and targeting of biofilm, introducing a quorum sensing device to E.coli has the ability to sense acyl-homoserine lactone is a quorum-sensing molecule produced by P.aeruginosa and achieved the colonization of E.coli near the target and release antimicrobial peptides, microcin nuclease DNase 1.13,15 Pathogen specific movement of E.coli attained by controlling cheZ gene responsible for smooth swimming and act as agonistic phosphatase of cheY helping in cell tumbling by regulating the ratio of this two genes chemotaxis is modifiable known as pseudo taxis.16-19 A similar kind of study is done on Mycoplasma pneumonia.20 The interest behind opting Mycoplasma as a model are the extensive availability of datasets, easily understandable metabolic gene networks and limitation of horizontal gene transfer, weak recombination ability, and lack of cell wall.20,21 It is very difficult for the host immune system to recognize without a cell wall and also it can directly release the target attacking substance to their environment both of these advantages make M. pneumoniae hide in the host and attack pathogen causing problems to the host.20,22 M. pneumoniae is an infectious pathogen in humans to make them unlethal, destroyed the bacterial proteins (P90, P30, P40) involved in binding the human sialo glycoproteins of respiratory epithelial cells also done CRISPR mediated editing in their genome to prevent Community-Acquired Respiratory Distress Syndrome toxin.23,24 Identified peptide signal that controls and boosts transcription and translation of M. pneumoniae, for activating the gene platform introduced responsible for the production of dispersin Lysostaphin.25 Using this synthetically engineered bacteria against S.aureus reported significant biofilm destabilization property due to the production of dispersin B20. Dispersin B has the potential to break N acetyl D glucosamine substance present on S .aureus biofilm.26
Bacterial two-component system modeling and possibilities
Two-component systems (TCS) are found in bacteria which are commonly involved in environmental sensing and response mechanism.27 Recent studies in some clinical pathogens identified TCS have significant roles in biofilm formation and antimicrobial resistance of organisms like Cronobacter sakazakii, Vibrio cholerae, P.aeruginosa etc.28,29 TCS consists of a sensory histidine kinase and response regulator that will control the downstream gene clusters accordingly to external stress. The final output of TCS activation will help the bacteria to survive the current situation or uptake a particular compound or something which will be favorable for bacteria.29 Novel synthetic approaches target the TCS with the help of rDNA technology for different purposes like bioremediation, targeted bacterial killing etc.
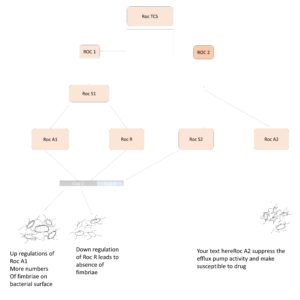
A better understanding of mechanisms involved in TCS and their signaling pathways will require finding and developing alternative strategies in the synthetic biology approach. Using the possibilities of TCS engineering there are lots of studies is conducted in biosensor development but there are not many studies reported against biofilm and antimicrobial resistance. But rolls of some TCS in biofilms are well studied for example PhoP/PhoQ in Cronobacter.28 PhoP/PhoQ is commonly found in many gram-negative bacteria and has a crucial role in environmental stress resistance and biofilm formation.30 Deletion studies on PhoPQ reported a significant reduction in biofilm biomass and viability of C. sakazakii. The results are produced by growing both wild-type bacteria and mutant lack PhoPQ two-component system on Glass/silicon wafers. Observations from Microscopy and FESEM state mutant type show difficulties in progression from microcolony to entire biofilm. The data observed from the TEM flagellar assembly of the mutant is also altered. There are lots of transposons of C.sakazakii is involved in flagellar assembly and especially mot A and motB which are involved in the motor activity of flagella but mutant strains show a significant reduction in mot A and motB gene.28 So, if we can develop a drug that collapses this TCS cascade will be a future scope to overcome biofilm. In P.aeruginosa there is different TCS involved in the assembly of extracellular appendages and production of extracellular polysaccharides and antibiotic susceptibility. Roc1 system is the first TCS well studied for cup genes in P.aeruginosa. These cup clusters are regulated by Roc 1 locus consisting of three genes encoding sensor kinase and response regulators which have a significant role in biofilm maturation. TCS of P.aeruginosa also shows some similarities to Bordetella species BvgSar system involved in resistance. RocA1 and RocR are conventional response regulator and shows the opposite effect on cup genes RocA1 activate fimbria expression and Roc R suppress it but the mechanism behind RocR is still not clearly understood. But it is strongly believed that it should affect the relative affinity of RocS1. Similar to Roc 1 locus there is another locus Roc 2 which is involved in the efflux pump and makes the bacteria more susceptible to drugs.29 However, the bacteria inside the biofilm show more resistance to drugs but here the signaling cascade support biofilm formation along with making bacteria more susceptible to the mechanism behind this still not well studied (Figure 1). Studies on PhoP/PhoQ revealed they have potential roles in human antimicrobial peptide sensing.31 The CAMP passes the outer membrane of Salmonella and comes in contact with PhoP/PhoQ two-component system cations that have a crucial role in this process that will lead to the activation downstream of this TCS. The activation and repression of the genes are regulated by some ions. There are not that many studies done in TCS biofilm targeting so the future aspects of this kind of study are very useful to target biofilm producers in vivo conditions. Proteolytic enzymes from Mycobacterium reported significant biofilm dissolving nature.32 So, the future of TCS in biofilm relies on using the rDNA technology to introduce the genes responsible for metalloprotease enzymes in TCS downstream portion so that the model bacteria sense the antimicrobial peptide or some components in the biofilm of pathogenic bacteria which leads to activation of proteolytic enzyme gene and biofilm dispersion of target bacteria. Similar to bacterial modeling changes can be made in TCS but there are not that many studies are done yet. Figure 1 illustrates the role of Roc TCS in fimbriae development. Figure 2 shows the two-component system modeling in bacteria.
Figure 1. Roc TCS system in Pseudomonas aeruginosa, Roc 1 and Roc 2 are two locus of the Roc TCS system which involved in cup gene regulation. Roc A1 and Roc R are response regulators which regulate the expression of fimbriae on the surface of bacteria involved in biofilm formation. Roc A1 upregulates and increase the expression of fimbriae on the bacterial surface and Roc R suppresses the Cup C gene and reduces the fimbriae number on the bacterial surface. RocR mechanism of suppression is still unknown. Roc 2 locus is involved in the efflux pump regulation
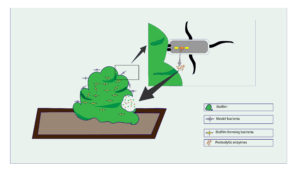
Figure 2. Modern concept of biofilm targeting using model bacteria. Modified bacteria are able to sense particular components in biofilm which may be polysaccharides or nucleic acid or protein or AMP that activate the model bacteria Two-component system (TCS). this Indirectly activates the genes which is introduced by rDNA technology and leads to the production of biofilm destruction of the target pathogen
Synthetic peptide
Antimicrobial peptides (AMP) are multifunctional compounds with a lot of medical applications.33 There are different anti-microbial peptides that are identified with a potential role in bactericidal, fungicidal, and virucidal activities.34 AMPs have also been reoffered to as host defense peptides in the past.35AMPS is suspected to be the first-line innate immune response of the host.36 AMPs are able to interfere with the pathways inside the bacterial cell without changing its membrane integrity.37 Certain AMPs like nisin have been demonstrated to destroy MRSA resistant to vancomycin.38 Compared to conventional antibiotics, antimicrobial peptide resistance development is very difficult for bacteria.39 Synthetic peptides are produced from naturally occurring peptide modifications like rearranging the amino acids.33
Some peptides are able to boost the wound healing process, combining this potential with an antimicrobial effect is very useful for the treatment of wounds.40,33 Synthetic peptides from reptile-like Komodo dragon reported showing significant wound healing, and antibiofilm activities. DRGN-1 is produced from a natural peptide named VK25 present in Komodo dragon by changing the amino acid position in VK25. DRGN1 is produced by an artificial production called the solid-phase china peptide method. DRGN1 acts on the cytoplasmic membrane and causes the leakages of intracellular components. DRGN1 reported significant antibiofilm properties against P.aeruginosa and S.aureus.33 Feleucin K59 is a synthetic peptide derived from feleucin K3 isolated from the skin of Bombina orientalis. Basically, feleucin k3 is an AMP containing three amino acid residues which makes this peptide more convenient to study.41 Studies using feleucin k3 showed it has potent antibiofilm and antimicrobial activity against MDR bacteria by modifying its fourth residue of leucine replaced by alanine will result in enhanced activity against P.aeruginosa with enhanced antibiofilm activity leads to the realization of the fourth residue in feleucin control the antibacterial activity.42 Because of the potential metabolic harm in repairing membrane components, and physical disruption of the cell membrane, it makes it difficult for bacteria to evolve drug resistance.43,44 Feleucin K59 is produced by artificial synthesis known as the Fmoc solid-phase method. Four lysine groups containing Gemini peptide showed excellent antimicrobial activities against E.coli.45 But FLEUCIN K59 shows some limitations in in vivo studies like toxicity, and hemolytic properties so there are optimization and standardizations required for future in vivo trials in humans. Gemini peptide is also produced by the solid-phase method. The mechanism of action is initially high in polar peptide can self-assemble into rods and bind to bacterial membrane then it dissociates into monomer and penetrate into the membrane and cause lysis of membrane.41 Distinct polarity amino acids are predicted to provide peptide amphiphiles with a different self-assembling structure and membrane penetration capability, both of which are important to dispersing biofilms.45 Endolysin’s terminals were fused with anti-microbial peptides to improve the entry to the peptidoglycan layer.46 Fusing sheep antimicrobial peptide called Art-175 with endolysin KZ-144 could kill P. aeruginosa, Acinetobacter baumannii, and E. coli. The mechanism of this peptide is reported as enzymatic degradation of peptidoglycan layer and osmotic lysis of membrane.47 CWR11 is a designed arginine tryptophan-rich peptide with a high antibacterial action against a broad spectrum of microorganisms via membrane disruption and superior salt resistance. Tethering CWR11 to a model polydimethylsiloxane surface shows antibiofilm and bactericidal properties.48 Engineered peptides like tryptophan and arginine-rich peptides have improved antimicrobial activity and overcome salt sensitivity problems.49,50 Studies on quorum sensing peptides have presented another strategy for disrupting peptides.51 A competence-stimulating peptide is a quorum-sensing peptide present in S.mutans but a higher concentration of CSP leads to the death of S.mutans. Engineered Analogue of CSP known as KBI-3221 is effective in reducing biofilm in several streptococcus species.51 The synergistic application of synthetic peptides and antibiotics is reported to be effective against biofilms, peptide will enhance the uptake of the antibiotic which leads to lethal effects in bacteria.52 The arrangement of various AMPs in a functional complex may help them to fight together. Due to the simultaneous existence of four separate families of AMPs like defensins, cecropins, diptericin, and proline-rich peptides, natural complex fly larvae immune peptide 7 isolated from Calliphora vicina maggots has been proven to ensure broad spectrum antibiofilm action.53 Table 2 shows the efficacy of synthetic peptide.
Table (2):
Synthetic units and their antimicrobial and antibiofilm properties.
| No | Synthetic unit | organism | Antimicrobial activity | Antibiofilm activity | Reference | ||
|---|---|---|---|---|---|---|---|
| Synthetic peptides | |||||||
| 1 | • DRGN 1 • VK25 (Template for DRGN1) |
P.aureginosa S.aureus P.aureginosa S.aureus |
EC50 4.46µM EC50 2.63µM EC50 17.7µM EC50 >65µM |
MBIC(Minimum biofilm inhibitory concentration) 25µg/ml MBIC 25µg/ml |
33 | ||
| 2. | • FLEUCIN K59 | E.coli S.aureus A.baumannii |
MIC 8µg/ml MIC 4µg/ml MIC 4µg/ml |
MBIC50 1µg/ml MBIC50 2µg/ml MBIC50 2µg/m |
MBIC90 8µg/ml MBIC90 4µg/ml MBIC90 4µg/ml |
41 | |
| 3. | •GEMINIPEPTIDE 12-(Arg)4-12 |
E.coli S.aureus |
MIC90 5.5 µM MIC90 5.6 µM |
MBIC90 50 µM MBIC90 50 µM |
45 | ||
| 4. | • Art-175+KZ-144 | Pseudomon as aureginosa | MIC50 4µg/ml |
MIC90 10µg/ml |
Not available | 47 | |
| Synthetic flavonoids | |||||||
| 5 | • CICI Flavonoid | Gram-positive Gram-negative |
Not available | MBIC 0.97µg/ml MBIC 0.48µg/ml |
62 | ||
| 6. | • FLUORINATED CHALCONE-1,2,3- TRIAZOLE | E.coli | MIC 0.0034µM/ML |
Not available | 57 | ||
Synthetic flavonoids
Flavonoids are substances naturally occurring in the plant kingdom that have the potential to act as anti-bacterial, anti-fungal, and anti-inflammatory.54 Several structural factors are involved in the antimicrobial properties of flavonoids like coplanarity, the presence of carboxyl group, and hydroxyl group.55 Synthetic flavonoids target the cell membrane structures, affect permeability, and inhibit bacterial metabolism.56 Synthetic flavanol like fluorinated chalcone-1,2,3-trazoles shows an antimicrobial effect by making covalent interactions with DNA topoisomerase.57 2-alkyl-3-imidazolylchromanones inhibit a key enzyme involved in ergosterol biosynthesis.58 Flavonoids present in grape wine such as quercetin, fisetin, kaempferol, apigenin, and chrysin efficient in inhibiting the production of S. aureus biofilms.59 some flavonoids are able to induce cell-to-cell communications in biofilm.60 Flavanol morin is reported to show antibiofilm properties against Listeria monocytogenes, flavonoids like phloretin are inactive to planktonic bacteria but show potent activity against biofilm formation this shows the significance of flavonoids in the antibiofilm treatment.61 Synthetic flavonoids like CICI-flav reported showing significant antibiofilm activity against E.coli, K.pneumonia. CICI-flav is a synthetic sulfur-containing tricyclic flavonoid with chlorine as a halogen substituent at the benzopyran core. CICI-flavonoid is produced by a two-step reaction and its mode of action is to disruption of cell membrane integrity.62 The efficacy of flavonoids is shown in Table 2.
Modeling possibilities in Bacteriophage
Bacteriophages and bacteriophage therapy are extensive studies undergoing area in modern research. The antibiotic crisis and multidrug-resistant strain emergence renewed the interest in phage studies are noticeable in recent years. Bacteriophages, with their ability to rapidly infect and overcome bacterial resistance, have proved a long-term strategy for combating bacterial infections, particularly in biofilms.63 The binding between bacteria and phages is based on receptors, showing significant specificity.64 The significands of phages and modified synthetic phages can produce biofilm dispersing enzymes like polysaccharidases to their surroundings that will help the phage attach to bacteria inside biofilm and allow the drug to enter biofilm.65 Phages can be synthetically modified to produce EPS-degrading enzymes via depolymerase synthesis.66 Indeed, extracellular proteins and polysaccharides are two major components of most EPS matrices; hence, the proteolytic enzyme (protease) and polysaccharides enzyme are two major EPS degrading enzymes that may be used for biofilm detachment.67 These enzymes, however, are not environmentally stable, and high pH, temperature, or salt concentrations may denature them, resulting in a reduction in enzymatic activity.63
There are various methods available to edit and modify phage genomes like Traditional recombination-based technique, Bacteriophage recombination of electroporated DNA(BRED), CRISPR-Cas-Based editing, and Rebooting phages using assembled phage genomic DNA.68
Traditional recombination-based technique
Simultaneously the host cells get co-infected with two parental phages will lead to the exchange of nucleotide sequences (Homologous recombination). Then the produced progenies were screened for desired phenotypes followed by purification. There is also some modified method available such as homologous recombination with plasmid. The limitation of this method is inability to do specific modifications to the targeted site and also this is a time-consuming process.68
Bacteriophage recombination of electroporated DNA(BRED)
BRED is also a homologous recombination-based method but it is done with the help of phage-mediated recombination systems like the RecE/RecT system. Rec recombination system consists of Gam, Exo, and bet genes. Gam gene shows inhibitory effect on E.coli RecBCD exonuclease to prevent degradation of double-stranded DNA substrate. Exo gene has a role in making ds DNA into a single strand. Bet gene is involved in the incorporation of removed ssDNA into recombination site on the phage genome. Compared to the previous method the frequency of homologous recombination is higher in the BRED method.68
CRISPR-Cas-Based Phage Engineering
In CRISPR-mediated editing, the components of the CRISPR-Cas 9 complex are cloned to the plasmid of the host first. Followed by the formation of the CRISPR-Cas9 complex which specifically attaches to the target site on the phage genome and makes double-stranded DNA break during phage infection. There is also a donor plasmid present on the host cell the mutation was introduced. The DNA break introduced by CRISPR-Cas 9 is then repaired by recombination with the donor plasmid and generates mutants of interest.68
Rebooting phage Using Assembled Phage Genomic DNA
In this method, the phage genome is isolated and assembled in vitro and introduce mutations with the help of polymerase cycling assembly, Polymerase chain reaction, and introduce host cell and generate mutant phage.68
Synthetic biology of phages using modular designs to generate more efficient phages for bacterial biofilm eradication is another unique strategy that has recently been researched. By expressing dispersin B as an EPS depolymerase and degrading enzyme, genetically engineered phage T7 was able to destroy bacterial biofilm. As a consequence, as compared to the parent T7 strain or dispersin B enzyme alone, the modified T7 phage has shown promising results in greatly enhancing in vivo destruction of E.coli biofilm. T7 phages have some limitations to replicating on E. coli cells containing F plasmid to overcome this, researchers incorporated gene 1.2 from T3 Phage to the BciI site of the T7 phage. The T7 phage is synthetically modified by adding dspB gene responsible for dispersin B is incorporated under the control of a strong T710 promoter. The mechanism of action behind modified T7 phage is expressing the dspB gene during infection intracellularly leads to the formation of dipercin B and which will be released out while cell lysis leads to the dispersion of biofilm. After validation studies, they reported 99% success in removing biofilm with synthetically produced phage T7.69
Engineered phage K1F also shows promising results on intracellular E.coli in human epithelial cells during urinary tract infections. The K1 antigens of some E.coli help them to survive intracellularly during phagocytosis because of their structural similarities to human tissue components. To overcome this limitation, scientist engineered a phage named K1F. Phage K1F is similar to the T7 phage at the genome scale but instead of T7 tail fiber protein, K1F phages have Endo sialidase enzyme with in tail structure which enables the phage to degrade the K1 antigen of E.coli.70
Future perspective
The development of antibiotic-resistant bacterial emergence and biofilm formation make difficulties in treatments that force science to find novel ways to treat them. Scientists may use synthetic biology based novel approaches that have the capabilities to overcome the ability of bacteria to develop resistance by changing their genotype or phenotype in the future. The new bacterial strains, peptides, and flavonoids produced by the possibilities of synthetic biology reported significant biofilm degradation and bacterial killing properties. The future optimization and attenuation of this kind of model will help humans to survive antibiotic stewardship and help to survive antibiotic resistance due to biofilm. Similarly, synthetic flavonoids and peptides show significant antimicrobial and antibiofilm activities. By the optimization of this peptide’s toxic and haemolytic properties, we can substitute the antibiotics with them. Synthetic biology strategies and their future optimizations on humans will definitely pave the way for decreased morbidity and mortality in hospital settings and their prevention in future.
The emergence of antibiotic-resistance due to the biofilm communities make synthetic biology strategies more popular in microbiology. There are four major strategies are mainly described in this review such as bacterial modelling, TCS possibilities, synthetic peptides, synthetic flavonoids and recombinant phage. The synthetic approach of bacterial modelling is commonly done in E.coli and Mycoplasma by introducing a quorum sensing device for transforming E.coli and CRISPR-mediated editing used for Mycoplasma both are targeted the biofilm and killing effect. Synthetic peptides like VK25, FLEUCIN K59, GEMINIPEPTIDE, Art175+KZ-144, and KBI-3221 are shown antimicrobial properties against Pseudomonas aeruginosa, E.coli, and Acinetobacter, Streptococcus etc. CICI-flav, Fluorinated calcone-123 triazole shows significant antibiofilm properties against E.coli and Klebsiella pneumonia respectively. Synthetic phage modified T7 phage and K1F phage shows excellent biofilm dispersion properties towards E.coli.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
PAA conceptualized, initiated, performed the literature search, and wrote the original draft. LR were involved in the topic conceptualization and critical reviewing. TS, AM, ACS, VC, KK, MSS critically revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Pinero-Lambea C, Ruano-Gallego D, Fernandez LA. Engineered bacteria as therapeutic agents. Curr Opin Biotechnol. 2015;35:94-102.
Crossref - Khatoon Z, McTiernan CD, Suuronen EJ, Mah TF, Alarcon EI, Alarcon Bacterial EI. Bacterial biofilm formation on implantable devices and approaches to its treatment and prevention. Heliyon. 2018;4(12):e01067.
Crossref - Chen M, Yu Q, Sun H. Novel strategies for the prevention and treatment of biofilm related infections. Int J Mol Sci. 2013;14(9):18488-18501.
Crossref - Jamal M, Tasneem U, Hussain T, Andleeb S. Bacterial Biofilm: Its Composition, Formation and Role in Human Infections. Research & Reviews: Journal of Microbiology and Biotechnology. 2015;4(3):1-14. https://www.rroij.com/open-access/bacterial-biofilm-its-composition-formation-and-role-in-human-infections.php?aid=61426
- Bjarnsholt T. The role of bacterial biofilms in chronic infections. APMIS. 2013;121:1-58.
Crossref - Jamal M, Ahmad W, Andleeb S, et al. Bacterial biofilm and associated infections. J Chin Med Assoc. 2018;81(1):7-11.
Crossref - Sunarintyas S. Bioadhesion of biomaterials. Advanced Structured Materials. 2016;58:103-125.
Crossref - Vickery K, Allan J, Jacombs A, Valente P, Deva A. Prevention of Implantable Medical Device Failure (IMD) Associated with Biofilm Infection. Am J Infect Control. 2011;39(5):E45.
Crossref - Schillaci D, Spano V, Parrino B, et al. Pharmaceutical Approaches to Target Antibiotic Resistance Mechanisms. J Med Chem. 2017;60(20):8268-8297.
Crossref - Castillo-Juarez I, Maeda T, Mandujano-Tinoco EA, et al. Role of quorum sensing in bacterial infections. World J Clin Cases. 2015;3(7):575.
Crossref - Duplantier M, Lohou E, Sonnet P. Quorum sensing inhibitors to quench P. Aeruginosa pathogenicity. Pharmaceuticals. 2021;14(12):1262.
Crossref - Khalifa ABH, Moissenet D, Thien HV, Khedher M. Les facteurs de virulence de Pseudomonas aeruginosa: Mecanismes et modes de regulations. Ann Biol Clin. 2011;69(4):393-403.
Crossref - Hwang IY, Tan MH, Koh E, Ho CL, Poh CL, Chang MW. Reprogramming microbes to be pathogen-Seeking killers. ACS Synth Biol. 2014;3(4):228-237.
Crossref - Goh YL, He HF, March JC. Engineering commensal bacteria for prophylaxis against infection. Curr Opin Biotechnol. 2012;23(6):924-930.
Crossref - Saeidi N, Wong CK, Lo TM, et al. Engineering microbes to sense and eradicate Pseudomonas aeruginosa, a human pathogen. Mol Syst Biol. 2011;7.
Crossref - Huang C, Stewart RC. CheZ Mutants with Enhanced Ability to Dephosphorylate CheY, the Response Regulator in Bacterial Chemotaxis. Biochim Biophys Acta. 1993;1202(2):297-304
Crossref - Kuo SC, Koshland DE. Roles of Che Y, CheZ Gene Products in Controlling Flagellar Rotation in Bacterial Chemotaxis of Escherichia Coli. J Bacteriol. 1987;169(3):1307-1414.
Crossref - Topp S, Gallivan JP. Guiding bacteria with small molecules and RNA. J Am Chem Soc. 2007;129(21):6807-6811.
Crossref - Sinha J, Reyes SJ, Gallivan JP. Reprogramming bacteria to seek and destroy an herbicide. Nat Chem Biol. 2010;6(6):464-470.
Crossref - Garrido V, Pinero-Lambea C, Rodriguez-Arce I, et al. Engineering a genome-reduced bacterium to eliminate Staphylococcus aureus biofilms in vivo. Mol Syst Biol. 2021;17(10):e10145.
Crossref - Guell M, van Noort V, Yus E, et al. Transcriptome complexity in a genome-reduced bacterium. Science. 2009;326(5957):1268-1271.
Crossref - Sukhithasri V, Nisha N, Biswas L, Anil Kumar V, Biswas R. Innate immune recognition of microbial cell wall components and microbial strategies to evade such recognitions. Microbiol Res. 2013;168(7):396-406.
Crossref - Chaudhry R, Kumar Varshney A, Malhotra P. Adhesion Proteins of Mycoplasma Pneumoniae. Front Biosci. 2007;12:690-699.
Crossref - Pinero-Lambea C, Garcia-Ramallo E, Martinez S, Delgado J, Serrano L, Lluch-Senar M. Mycoplasma pneumoniae Genome Editing Based on Oligo Recombineering and Cas9-Mediated Counterselection. ACS Synth Biol. 2020;9(7):1693-1704.
Crossref - Yus E, Yang JS, Sogues A, Serrano L. A reporter system coupled with high-throughput sequencing unveils key bacterial transcription and translation determinants. Nat Commun. 2017;8(1):368.
Crossref - Kaplan JB. Therapeutic Potential of Biofilm-Dispersing Enzymes. Int J Artif Organs. 2009;32(9):545-554.
Crossref - Ravikumar S, Baylon MG, Park SJ, Choi J il. Engineered microbial biosensors based on bacterial two-component systems as synthetic biotechnology platforms in bioremediation and biorefinery. Microb Cell Fact. 2017;16(1):62.
Crossref - Ma Y, Zhang Y, Shan Z, Wang X, Xia X. Involvement of PhoP/PhoQ two-component system in biofilm formation in Cronobacter sakazakii. Food Control. 2022;133(Part A):18621.
Crossref - Mikkelsen H, Sivaneson M, Filloux A. Key two-component regulatory systems that control biofilm formation in Pseudomonas aeruginosa. Environ Microbiol. 2011;13(7):1666-1681.
Crossref - Park SY, Groisman EA. Signal-specific temporal response by the Salmonella PhoP/PhoQ regulatory system. Mol Microbiol. 2014;91(1):135-144.
Crossref - Otto M. Bacterial Sensing of Antimicrobial Peptides. Contrib Microbiol. 2009;16:136–149.
Crossref - Saggu SK, Jha G, Mishra PC. Enzymatic degradation of biofilm by metalloprotease from microbacterium sp. Sks10. Front Bioeng Biotechnol. 2019;7:192.
Crossref - Chung EMC, Dean SN, Propst CN, Bishop BM, van Hoek ML. Komodo dragon-inspired synthetic peptide DRGN-1 promotes wound-healing of a mixed-biofilm infected wound. NPJ Biofilms Microbiomes. 2017;3:9.
Crossref - Bahar AA, Ren D. Antimicrobial peptides. Pharmaceuticals. 2013;6(12):1543-1575.
Crossref - Brown KL, Hancock REW. Cationic host defense (antimicrobial) peptides. Curr Opin Immunol. 2006;18(1):24-30.
Crossref - Zasloff M. Antimicrobial Peptides of Multicellular Organisms. 2002:415.
Crossref - Brogden KA. Antimicrobial peptides: Pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol. 2005;3(3):238-250.
Crossref - Brumfitt W, Salton MRJ, Hamilton-Miller JMT. Nisin, alone and combined with peptidoglycan-modulating antibiotics: Activity against methicillin-resistant Staphylococcus aureus and vancomycin-resistant enterococci. J Antimicrob Chemother. 2002;50(5):731-734.
Crossref - Fjell CD, Hiss JA, Hancock REW, Schneider G. Designing antimicrobial peptides: Form follows function. Nat Rev Drug Discov. 2012;11(1):37-51.
Crossref - Steinstraesser L, Hirsch T, Schulte M, et al. Innate defense regulator peptide 1018 in wound healing and wound infection. PLoS One. 2012;7(8):e39373.
Crossref - Guo X, Rao J, Yan T, et al. Feleucin-K3 Analogue with an α-(4-Pentenyl)-Ala Substitution at the Key Site Has More Potent Antimicrobial and Antibiofilm Activities in Vitro and in Vivo. ACS Infect Dis. 2021;7(1):64-78.
Crossref - Xie J, Li Y, Li J, et al. Potent effects of amino acid scanned antimicrobial peptide Feleucin-K3 analogs against both multidrug-resistant strains and biofilms of Pseudomonas aeruginosa. Amino Acids. 2018;50(10):1471-1483.
Crossref - Hoffmann JA, Kafatos FC, Janeway Jr CA, B Ezekowitz RA. Translating Advances in Human Genetics into Public Health Action: A Strategic Plan. World Health Organization; 1999:283.
- Senyurek I, Paulmann M, Sinnberg T, et al. Dermcidin-derived peptides show a different mode of action than the cathelicidin LL-37 against Staphylococcus aureus. Antimicrob Agents Chemother. 2009;53(6):2499-2509.
Crossref - Qi R, Zhang N, Zhang P, et al. Gemini Peptide Amphiphiles with Broad-Spectrum Antimicrobial Activity and Potent Antibiofilm Capacity. ACS Appl Mater Interfaces. 2020;12(15):17220-17229.
Crossref - Wang T, Zheng Y, Dai J, et al. Design SMAP29-LysPA26 as a Highly Efficient Artilysin against Pseudomonas Aeruginosa with Bactericidal and Antibiofilm Activity. Microbiol Spectr. 2021;9(3):e0054621.
Crossref - Briers Y, Walmagh M, Grymonprez B, et al. Art-175 is a highly efficient antibacterial against multidrug-resistant strains and persisters of Pseudomonas aeruginosa. Antimicrob Agents Chemother. 2014;58(7):3774-3784.
Crossref - Lim K, Chua RRY, Saravanan R, et al. Immobilization studies of an engineered arginine-tryptophan-rich peptide on a silicone surface with antimicrobial and antibiofilm activity. ACS Appl Mater Interfaces. 2013;5(13):6412-6422.
Crossref - Pasupuleti M, Schmidtchen A, Chalupka A, Ringstad L, Malmsten M. End-tagging of ultra-short antimicrobial peptides by W/F stretches to facilitate bacterial killing. PLoS One. 2009;4(4).
Crossref - Li X, Saravanan R, Kwak SK, Leong SSJ. Biomolecular engineering of a human beta defensin model for increased salt resistance. Chem Eng Sci. 2013;95:128-137.
Crossref - LoVetri K, Madhyastha S. Antimicrobial and Antibiofilm Activity of Quorum Sensing Peptides and Peptide Analogues Against Oral Biofilm Bacteria. 2010:383-392.
Crossref - Dosler S, Mataraci E. In vitro pharmacokinetics of antimicrobial cationic peptides alone and in combination with antibiotics against methicillin resistant Staphylococcus aureus biofilms. Peptides (NY). 2013;49:53-58.
Crossref - Grassi L, Maisetta G, Esin S, Batoni G. Combination strategies to enhance the efficacy of antimicrobial peptides against bacterial biofilms. Front Microbiol. 2017;8(DEC).
Crossref - Sarbu LG, Bahrin LG, Babii C, Stefan M, Birsa ML. Synthetic flavonoids with antimicrobial activity: a review. J Appl Microbiol. 2019;127(5):1282-1290.
Crossref - Alcaraz LE, Blanco SE, Puig ON, Tomas F, Ferretti FH. Antibacterial activity of flavonoids against methicillin-resistant Staphylococcus aureus strains. J Theor Biol. 2000;205(2):231-240.
Crossref - Ikigai H, Nakae T, Hara Y, Shimamura T. Bactericidal Catechins Damage the Lipid Bilayer.; 1993;1147(1):132-6.
Crossref - Lal K, Yadav P, Kumar A, Kumar A, Paul AK. Design, synthesis, characterization, antimicrobial evaluation and molecular modeling studies of some dehydroacetic acid-chalcone-1,2,3-triazole hybrids. Bioorg Chem. 2018;77:236-244.
Crossref - Emami S, Banipoulad T, Irannejad H, et al. Imidazolylchromanones containing alkyl side chain as lanosterol 14a-demethylase inhibitors: Synthesis, antifungal activity and docking study. J Enzyme Inhib Med Chem. 2014;29(2):263-271.
Crossref - Cho HS, Lee JH, Cho MH, Lee J. Red wines and flavonoids diminish Staphylococcus aureus virulence with anti-biofilm and anti-hemolytic activities. Biofouling. 2015;31(1):1-11.
Crossref - Vikram A, Jayaprakasha GK, Jesudhasan PR, Pillai SD, Patil BS. Suppression of bacterial cell-cell signalling, biofilm formation and type III secretion system by citrus flavonoids. J Appl Microbiol. 2010;109(2):515-527.
Crossref - Lee JH, Regmi SC, Kim JA, et al. Apple flavonoid phloretin inhibits Escherichia coli O157:H7 biofilm formation and ameliorates colon inflammation in rats. Infect Immun. 2011;79(12):4819-4827.
Crossref - Babii C, Mihalache G, Bahrin LG, et al. A novel synthetic flavonoid with potent antibacterial properties: In vitro activity and proposed mode of action. PLoS One. 2018;13(4).
Crossref - Motlagh AM, Bhattacharjee AS, Goel R. Biofilm control with natural and genetically-modified phages. World J Microbiol Biotechnol. 2016;32(4):1-10.
Crossref - Orlova E v. Bacteriophages and Their Structural Organisation. www.intechopen.com
- Wittebole X, de Roock S, Opal SM. A historical overview of bacteriophage therapy as an alternative to antibiotics for the treatment of bacterial pathogens. Virulence. 2014;5(1):226-235.
Crossref - Yan J, Mao J, Xie J. Bacteriophage polysaccharide depolymerases and biomedical applications. BioDrugs. 2014;28(3):265-274.
Crossref - Loiselle M, Anderson KW. The use of cellulase in inhibiting biofilm formation from organisms commonly found on medical implants. Biofouling. 2003;19(2):77-85.
Crossref - Chen Y, Batra H, Dong J, Chen C, Rao VB, Tao P. Genetic engineering of bacteriophages against infectious diseases. Front Microbiol. 2019;10:954.
Crossref - Lu TK, Collins JJ. Dispersing Biofilms with Engineered Enzymatic Bacteriophage.; 2007. www.pnas.orgcgidoi10.1073pnas.0704624104
- Moller-Olsen C, Ho SFS, Shukla RD, Feher T, Sagona AP. Engineered K1F bacteriophages kill intracellular Escherichia coli K1 in human epithelial cells. Sci Rep. 2018;8(1):17559.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.