ISSN: 0973-7510
E-ISSN: 2581-690X
Different solvents were used for sumac extractions to find different active compounds in each fraction, and these fractions were tested on Staphylococcus aureus (which isolated from soft white cheese), the (MIC) minimum inhibitory concentration of each fraction were measured then sub-inhibitory concentration (SIC) was used. We found that all fractions can inhibit quorum sensing in Staphylococcus aureus in a different ratio. By using Real-time PCR found that the different sumac fractions can inhibit the expression of tested genes (Sea, Seb, AgrA, RNAIII, and Hla). Furthermore, most sumac extracted fractions have the ability to decreasing biofilm and growth curve in Staphylococcus aureus significantly, while other fractions decreased them non-significantly.
Staphylococcus aureus, sumac extract, quorum sensing inhibitor, Biofilm, Growth curve.
A distinguished human pathogen Staphylococcus aureus which causes numerous infections (acute and chronic) such as pneumonia, pericarditis, and sepsis. This bacteria also produce food poisoning by secretion enterotoxins1. Quorum sensing (QS), an essential regulatory system of bacteria that depend on the production and detection of extracellular materials called autoinducers, have been shown to control virulence factors in many pathogens2. In S. aureus, the Agr (accessory gene regulator) QS is the prevalent and well-studied virulence regulator and is responsible for increased expression of virulence genes, inclusive many enzymes, and toxins that are critical for the infection establishment3. S. aureus express and secret virulence factors that responsible of bacterial pathogenicity is under control of (Agr) quorum -sensing system4. AgrA (response regulator) and AgrC (histidine kinase signal transduction) are components of (TCST) two-component signal transduction system of Agr system5. In the external environment AIP (autoinducing peptide) activate and phosphorylate AgrC. The phosphate group transferred to AgrA, then the active AgrA lead to virulence factor expression and formation of biofilm, this due to interaction between AgrA with RNAII and RNAIII6. The four genes (AgrA, AgrB, AgrC, and AgrD) and two units of RNAII and RNAIII, are comprises of S. aureus quorum-sensing system7. The trans-autophosphorylation induced by AgrC, which cause AgrA activation, this cause activation of P2 and P3 for RNAII and RNAIII respectively8. The quorum sensing system Agr in staphylococci comprises of (2) components (RNAII and RNAIII)9. Phosphorylated AgrA triggers The P2 and P3 promoter. is bound to the autoinducer the Agr operon by the phosphorylated AgrA. Besides triggering the P2 promoter. The expression of RNAIII is under control of the P3 promoter10. Production of a-toxin is under control of RNAIII11.
The use and Recognition of natural products have a long history. In the report of the World Health Organization (WHO), in developing countries around 80% of people confirm that the natural products have benefits for health nursing. Due to rising microorganisms resistance to ordinary drugs, the use and recognize of natural products were increased12,13. Generally referred to as Elm-Leaved Sumac or Tanner’s Sumac with the scientific name Rhus coriaria L. is a small trees to deciduous shrub which grow one to five meters. Its leave accommodates nine to fifteen bushy leaflets. The term Sumac is used in preferred as a common call for numerous of Rhus plant species, however the most, not unusual species is Rhus coriaria L14. An solvent (ethanol) extracts of the sumac fruit decreased production of biofilm. Restricted decreasing of bacterial growth was at doses 128–512_g-ml-1 15. Great phytochemical research has been done on black horehound, and numerous phenylpropanoid glycosides were diagnosed and related to slight decreasing growth by S. aureus16. The invention of QS represents a novel goal for antimicrobial drugs and potential plant-produced QS-regulating agents directed towards plant-related bacterial communication systems would possibly provide a unique technique to controlling bacterial infection in human beings and animals17. The prevailing study is to understand the impact of sumac extracts on QS circuits in S. aureus and to examine the effect of subinhibitory concentration of sumac fractions on the expression some QS & virulence factor genes, biofilm formation and growth of the S. aureus.
Isolation and identification Staphylococcus aureus from soft white cheese
Staphylococcus aureus isolated and identified using FDA’s Bacteriological Analytical Manual USA 200118.
Sumac extractions
Different solvents were used for the extraction of the active substances from sumac are selected primarily based on the polarity of the solute, from low polar to high polar, as follows: Hexane – Chloroform – Ethylacetate – Butanol – Ethanol – Water19.
Weight 50g of ground sumac seeds mixed with 100ml of Hexane put it in an ultrasonic bath (1.5 hours at 37° C), then filtered the mixture we get extract part one (fraction 1) using rotary evaporator to evaporate hexane, the residue will be dried at room temperature and used for next solvent extraction.
The same technique used for the other solvents to get the extracts {part (fraction 2) from chloroform, part (fraction 3) from ethylacetate, part (fraction 4) from butanol, part (fraction 5) from ethanol, and part (fraction 6) from water}20.
Preparation of Concentration of the sumac extracts
One gram of each sumac fraction was dissolved in 5 ml of 10% DMSO, mixed well by vortex and different concentrations prepared from 2-150 mg /ml.
Preparation of Bacterial Inoculum
A single colony of S. aureus was inoculated into 5ml of the BHI broth medium and incubated overnight at 37°C, the inoculum adjusted by McFarland for each independent test.
Determining minimum inhibitory concentration (MIC) of Sumac extracts
96-well microplates was used to assay different concentrations of sumac extracts (0, 2, 4, 5, 10, 15, 20, 25, 50, 75, 100, 125 and 150 mg/ml) by mixing with tryptone soy broth. All wells inoculated by 10µl of previous activated culture of S. aureus, overnight incubated at 37°C Then the MIC was determined, the absorbency of them were measured before and after incubation by ELISA reader at wave length 490 nm21.
Enterotoxin detection assay by RPLA kit
Staphylococcal enterotoxin test kit (RPLA kit) was used for determining Staphylococci enterotoxins A (Sea), B (Seb), C (Sec), and D (Sed) in culture supernatants (after growing bacteria in trypton soy broth which contain sub-mic sumac extracts concentration) by passive agglutination22.
Determining gene expression by using real_time PCR
Effect of sumac extracts on expression each of (Sea, Seb, AgrA, RNAIII, and Hla gene) were tested by using Real-time PCR, the total RNA extracted from the bacterial culture which growing in trypton soy broth contain sub-mic sumac extracts comparing them with control culture of Staphylococcus aureus.
The decontaminated RNA for DNA microarray evaluation were used for real-time PCR (RT-PCR). cDNA was made from 1g of RNA using high-capacity RNA-to-cDNA master mix (Applied Biosystems). For quantifiable real-time PCR, magnification was performed with Power Sybr green master mix in first step plus thermal cycler (Applied Biosystems). The primers had been obtained from other researchers as shown in table 1. Thirty cycles had been run with denaturation at 95°C for 15 s, annealing at 55°C for the 30s, and extension at 60°C for 45s23.
Table (1):
Primers used for determining gene expression.
| Primer | Sequence | Target Gene | Reference |
|---|---|---|---|
| Sea Fw | GGTTATCAATGTGCGGGTGG | Sea | -24 |
| Sea Rv | CGGCACTTTTTTCTCTTCGG | ||
| Seb Fw | GTATGGTGGTGTAACTGAGC | Seb | -24 |
| Seb Rv | CCAAATAGTGACGAGTTAGG | ||
| hla Fw | ATGGTGAATCAAAATTGGGG | hla | -25 |
| hla Rv | GTTGTTTGGATGCTTTTC | ||
| RNA III Fw | AATACATAGCACTGAGTCCAAGG | RNA III | -26 |
| RNA III Rv | TGGATTATCGACACAGTGAACA | ||
| Agr A Fw | TGATAATCCTTATGAGGTGCTT | Agr A | -26 |
| Agr A Rv | CACTGTGACTCGTAACGAAAA | ||
Effect of sumac extracts on biofilm formation by Staphylococcus aureus
Microtiter dish plate method was used to detect biofilm formation before and after growing bacteria in trypton soy broth contain sub-mic sumac extracts and comparing it with control culture, by using the method of OToole27.
Effect of sumac extract on growth curve of S. aureus
Effects of sumac extract on growth curve was determined by growing bacteria in a microtiter plate which contains trypton soy broth plus sub-mic sumac extracts, then measuring optical density in each hour in comparison with control culture28.
Statistical Analysis
Spss program 20 was used to compare between means (for biofilm and growth curve) by One way ANOVA and Duncan test.
From 31 isolates of Staphylococcus aureus, one isolate which is an enterotoxin A & B producing strain (tested by RPLA-KIT) was chosen and different sumac fractions were tested on it.
As shown in table 2 the mic of sumac extract fraction 1, 2, 3, 4, 5, and 6 were 75, 25, 2, 5, 5, and 25 mg/ml respectively; the concentrations were different because the fractions contain different active ingredients.
Table (2):
Effect of different sumac fractions on production of enterotoxin A and B in Staphylococcus aureus.
Fraction 6 |
Fraction 5 |
Fraction 4 |
Fraction 3 |
Fraction 2 |
Fraction 1 |
Control |
Enterotoxin |
|---|---|---|---|---|---|---|---|
– |
– |
– |
– |
– |
+ |
+++ |
Sea |
– |
– |
– |
– |
– |
+ |
+++ |
Seb |
By RPLA test, we get that the all sumac extract fractions caused inhibition of enterotoxin expression as shown in table (3).
Table (3):
Minimum inhibitory concentration of different sumac fractions against Staphylococcus aureus.
| Absorbency of bacterial culture ( extract concentration mg/ml) | Extracts | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 150 | 125 | 100 | 75 | 50 | 25 | 20 | 15 | 10 | 5 | 2 | |
| 0 | 0 | 0 | 0 | 0.128 | 0.311 | 0.756 | 1.0878 | 1.442 | 1.963 | 2.145 | Fraction 1 |
| 0 | 0 | 0 | 0 | 0 | 0 | 0.209 | 0.438 | 0.974 | 1.568 | 2.223 | Fraction 2 |
| 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | Fraction 3 |
| 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.541 | Fraction 4 |
| 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.389 | Fraction 5 |
| 0 | 0 | 0 | 0 | 0 | 0 | 0.254 | 0.611 | 1.042 | 1.456 | 2.187 | Fraction 6 |
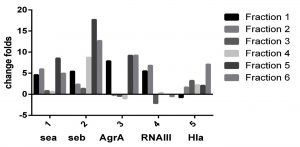
By RT-PCR it was evident that the different sumac extract fractions have different effect on expression of tested gene (Sea, Seb, AgrA, RNAIII, and Hla). As evident in Fig.1 Sea and Seb gene inhibited by all sumac extract fractions, and AgrA inhibited by fraction 1, 5, and 6, while RNAIII not expressed by fraction 5 and don’t affected by fraction 3, and 6, also Hla don’t inhibit by fraction 1.
Fig. 1. Effect of different sumac fractions on different gene expression in Staphylococcus aureus By RT_PCR.
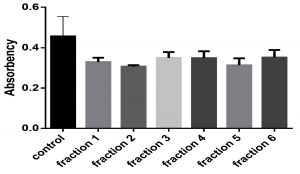
Sumac extract caused decreasing biofilm formation by Staphylococcus aureus, according to one way ANOVA test (P ֲ 0.05), fraction 1, 2, and 5 decreased biofilm significantly while fraction 3, 4, and 6 caused decreasing biofilm non significantly comparing with control, as shown in Fig. 2.
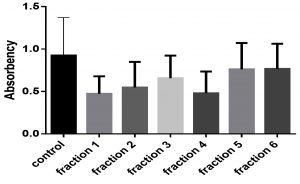
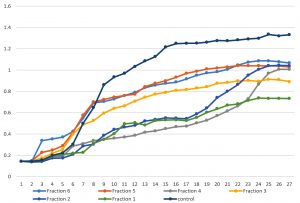
Fig. 3 and 4 showed effects of sumac extract on growth curve of S. aureus, as shown all extract fractions have an effect on the growth curve, fraction 1, 2, 3, and 4 significantly shortened the growth curve but fraction 5, and 6 have non-significant effect on growth curve, this results obtained by one way ANOVA test (P < 0.05).
Fig. 4. Growth curve shape of Staphylococcus aureus before and after growing it with different sumac fractions.
It¼s clear in our results that the sumac extract blocked some virulence factors and decreased the others, furthermore we note that the sumac extract has a role in biofilm disassembly and changing Staphylococcus aureus behaviors.
The DNA microarray and quantitative real-time PCR (qRT-PCR) data studies had shown the decreasing expression of the QS regarding genes in several pathogenic bacteria, which lead to a reduction in their virulence properties20. Natural products from plants include favorable tools for the bacterial pathogenesis management and bacterial modulations. Previous studies on anti-QS activities or QSI of natural products in bacteria have concentrated mainly on elucidating the decreasing of expressions of well-established determine induced genes of QS17. The attenuation of bacterial virulence due to reduction of expression QS genes cause the prevention of bacterial pathogenicity18. The reduction of expression QS genes and the level of signaling molecules that affect the production of virulence factors provide more insight into why these natural products were used in the past and how they can be used in the future to control the microbial infections21.
The lively compound of sumac which is gallic acid. Borges and co-workers worked on the mechanism of action of gallic acid towards distinctive kind of microorganism, gallic acid induced irreversible modifications inside the membrane properties via hydrophobicity modifications, a lower of anions, and prevalence of locally rupture or formation pores within the cell membranes which cause leakage of vital components of the cell29.
1, 2-dioxo-6- hydroxycyclohexadiene-4-carboxilic acid is the other active quinones compound of sumac. The quinine compounds present some free radicals and can react with nucleophilic amino acids in the protein and produce stable complexes which cause loss it’s function and rapturing the cell. The quinone oxidization targets are the surface-exposed adhesion, polypeptides in cell wall, and membrane-bound enzyme31.
Between 56 Palestinian plants tested, found that the sumac have the most antibacterial activity toward Probionibacterium acnes (MIC 6 mg.ml-1, MBC 6 mg.ml-1), S. aureus (MIC 4 mg.ml-1, MBC 6 mg/ml), E. coli (MIC 6 mg/ml, MBC 8 mg/ml) and Pseudomonas aeruginosa (MIC 4 mg.ml-1and MBC 6 mg.ml-1)32. Although R. coriaria is a particularly rich source of phenolic compounds33.
ACKNOWLEDGMENTS
This study was supported by Department of Biology, College of Science, Salahaddin University-Erbil, Iraq as a part of PhD thesis results.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
FA designed research; AS conducted research, analysed data; and wrote the paper; FA edited the paper; AS and FA had primary responsibility for final content. Both authors read and approved the final manuscript.
FUNDING
This work was supported by the Ministry of Higher Education and Scientific Research, Kurdistan Region, Iraq.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
- Chambers HF, Deleo FR. Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat. Rev. Microbiol., 2009; 7(9): 629-41.
Crossref - Reuter K, Steinbach A, Helms V. Interfering with Bacterial Quorum Sensing. Perspect Medicin Chem., 2016; 8: 1-15.
Crossref - Le KY, Otto M. Quorum-sensing regulation in staphylococci-an overview. Frontiers in Microbiology. 2015; 6: 1174.
Crossref - Cheung AL, Nishina KA, Trotonda MP, Tamber S. The SarA protein family of Staphylococcus aureus. Int. J. Biochem. Cell Biol., 2008; 40(3): 355-61.
Crossref - Zehnpfennig B, Urbatsch IL, Galla HJ. Functional reconstitution of human ABCC3 into proteoliposomes reveals a transport mechanism with positive cooperativity. Biochemistry, 2009; 48(20): 4423-30.
Crossref - Matthew Thoendel JSK, Caralyn E. Flack, and Alexander R. Horswill. Peptide Signaling in the Staphylococci. Chemical Reviews, 2011; 111: 117-51.
Crossref - George Cisar EA, Geisinger E, Muir TW, Novick RP. Symmetric signalling within asymmetric dimers of the Staphylococcus aureus receptor histidine kinase AgrC. Mol. Microbiol., 2009; 74(1): 44-57.
Crossref - Kavanaugh JS, Thoendel M, Horswill AR. A role for type I signal peptidase in Staphylococcus aureus quorum sensing. Mol. Microbiol., 2007; 65(3): 780-98.
Crossref - Richard P. Novick HFR, Steven J.Projan JK, Moghazeh BKaS. Synthesis of staphylococcal virulence factors is controlled by a regulatory RNA molecule. The EMBO Journal, 1993; 12(10): 3967-75.
Crossref - Dunman PM, Murphy E, Haney S, Palacios D, Tucker-Kellogg G, Wu S, et al. Transcription profiling-based identification of Staphylococcus aureus genes regulated by the agr and/or sarA loci. Journal of Bacteriology, 2001; 183(24): 7341-53.
Crossref - Palombo EA. Traditional Medicinal Plant Extracts and Natural Products with Activity against Oral Bacteria: Potential Application in the Prevention and Treatment of Oral Diseases. Evid Based Complement Alternat Med., 2011; 2011: 680354.
Crossref - Mohapatra DP, Thakur V, Brar SK. Antibacterial efficacy of raw and processed honey. Biotechnol. Res. Int., 2011; 2011: 917505.
Crossref - Nasar-Abbas SM, Halkman AK. Antimicrobial effect of water extract of sumac (Rhus coriaria L.) on the growth of some food borne bacteria including pathogens. International Journal of Food Microbiology, 2004; 97(1): 63-9.
Crossref - Pereira JA, Oliveira I, Sousa A, Valentao P, Andrade PB, Ferreira IC, et al. Walnut (Juglans regia L.) leaves: phenolic compounds, antibacterial activity and antioxidant potential of different cultivars. Food Chem. Toxicol., 2007; 45(11): 2287-95.
Crossref - Didry N, Seidel V, Dubreuil L, Tillequin F, Bailleul F. Isolation and antibacterial activity of phenylpropanoid derivatives from Ballota nigra. Journal of Ethnopharmacology, 1999; 67: 197-202.
Crossref - Max Teplitski JBR, and Wolfgang D. Bauer. Plants Secrete Substances That Mimic Bacterial N-Acyl Homoserine Lactone Signal Activities and Affect Population Density-Dependent Behaviors in Associated Bacteria. MPMI, 2000; 13(6): 637-48.
Crossref - FDA. Bacteriological Analytical Manual. 2001.
- Altemimi A, Lakhssassi N, Baharlouei A, Watson DG, Lightfoot DA. Phytochemicals: Extraction, Isolation, and Identification of Bioactive Compounds from Plant Extracts. Plants (Basel), 2017; 6(4).
Crossref - Harborne JB. Phytochemical Methods. Chapman and Hall,London EC4P 4EE. 1980.
Crossref - Eloff JN. A sensitive and quick microplate method to determine the minimal inhibitory concentration of plant extracts for bacteria. Planta Medica, 1998; 64(8): 711-3.
Crossref - Sally A. Rose PBaMFS. Detection of staphylococcal enterotoxins in dairy products by the reversed passive latex agglutination (SET-RPLA) kit International Journal of Food Microbiology, 1989; 8: 65-72.
Crossref - Jakobsen TH, van Gennip M, Phipps RK, Shanmugham MS, Christensen LD, Alhede M, et al. Ajoene, a sulfur-rich molecule from garlic, inhibits genes controlled by quorum sensing. Antimicrobial Agents and Chemotherapy, 2012; 56(5): 2314-25.
Crossref - Muyiwa Ajoke Akindolire OOBaCNA. Detection of Antibiotic Resistant Staphylococcus aureus from Milk: A Public Health Implication. International Journal of Environmental Research and Public Health, 2015; 12.
Crossref - Zhang H, Zheng Y, Gao H, Xu P, Wang M, Li A, et al. Identification and Characterization of Staphylococcus aureus Strains with an Incomplete Hemolytic Phenotype. Frontiers in Cellular and Infection Microbiology, 2016; 6: 146.
Crossref - Da F, Yao L, Su Z, Hou Z, Li Z, Xue X, et al. Antisense locked nucleic acids targeting agrA inhibit quorum sensing and pathogenesis of community-associated methicillin-resistant Staphylococcus aureus. Journal of Applied Microbiology, 2017; 122(1): 257-67.
Crossref - O’Toole GA. Microtiter dish biofilm formation assay. Journal of visualized experiments : JoVE, 2011; (47).
Crossref - Zwietering MH, Jongenburger I, Rombouts FM and Riet KVT. Modeling of the Bacterial Growth Curve. Applied and Environmental Microbiology, 1990; 56(6): 1875-81.
- Mahdi Ahmadian-Attari M, Amini M, Farsam H, Amin G, Fazeli MR, Monsef Esfahani HR, et al. Isolation of Major Active Antibacterial Compounds of Sumac Fruit (Rhus coriaria L.). International Journal of Enteric Pathogens, 2016; 4(4): 1-5.
Crossref - Borges A, Ferreira C, Saavedra MJ, Simoes M. Antibacterial activity and mode of action of ferulic and gallic acids against pathogenic bacteria. Microb. Drug Resist., 2013; 19(4): 256-65.
Crossref - Ali-Shtayeh M S, Al-Assali A A, Jamous R M. Antimicrobial activity of Palestinian medicinal plants against acne-inducing bacteria. African Journal of Microbiology Research, 2013; 7(21): 2560-2573.
Crossref - Kossah R, Nsabimana C, Zhang H, Chen W. Optimization of Extraction of Polyphenols from Syrian Sumac (Rhus Coriaria L.) and Chinese Sumac (Rhus typhina L.) Fruits. Research Journal of Phytochemistry, 2010; 4(3): 146-153.
Crossref
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.