ISSN: 0973-7510
E-ISSN: 2581-690X
Bacteriocins are of great interest as potential antimicrobial agents against various types of bacteria, fungi, and viruses. Isolates of microorganisms derived from natural sources were used in the current study, including lactic acid bacteria and other antagonistic microorganisms. The species of the microorganisms were determined using 16S rDNA and ITS nrDNA analyses. E. coli, S. enterica, S. aureus, P. aeruginosa, B. mycoides, A. faecalis, P. vulgaris, S. flexneri , L. monocytogenes, C. albicans, A. flavus, and P. citrinum were used as pathogenic and opportunistic strains. It was found that 11 strains of antagonistic microorganisms have significant antimicrobial activity against all pathogenic and opportunistic microorganisms. The antimicrobial properties of these microorganisms are currently under study.
Bacteriocins, Lactic acid bacteria, Antagonistic microorganisms, Isolates, Antimicrobial activity, Pathogenic microorganisms
The discovery of antibiotics was a major breakthrough in treating infectious diseases, significantly improving the quality of life and longevity worldwide. However, after several years of antibiotic use, resistance to antimicrobials emerged. Since then, its spread has become a severe health problem.1Gram-positive and gram-negative bacteria are increasingly showing multidrug resistance to the main antibiotics and drugs used in clinical practice, such as drug-resistant methicillin-resistant Staphylococcus aureus, resulting in higher morbidity and mortality than usual.2-4 Multidrug resistance of enterococci to vancomycin has also been reported. Drug resistance has placed a burden on patient health and the economies of hospitals and communities, indicating the need for developing new antibiotic drugs. Additionally, rational use of existing antibiotics could alleviate this problem.5
Many natural resources, including plants, animals, and microorganisms, have been studied.6 Combining chemical and biotechnology tools, these resources can provide several compounds that may prove to be promising antimicrobial agents.7-9 Bacteria have also been studied in natural resources for their ability to resist other bacteria. The discovery of the first bacteriocin in 1925 made it possible to develop an entire field of research, including several studies aimed at identifying novel antimicrobial compounds of microbial origin in the subsequent decades.10 Bacteriocins are of increasing interest as potential antibacterial agents against various bacteria, fungi, and viruses.11-13 Naturally resistant structures such as bacterial biofilm can also be similarly exploited.14-15 These natural peptides are synthesized by ribosomes and produced by bacteria living in competitive microbial environments and can be used to kill other bacteria, especially those that are closely related to the biofilm producers.16-19 Consequently, the considerable variety of bacteriocins provides a wide range of activity.20-22 Ribosomes produce bacteriocins and convert them into molecules unlike synthetic antibiotics,23 such as lipopeptides and glycopeptides.23
Although bacteriocins are produced in a gram-positive form,24 most registered bacteriocins are gram-negative,25 especially Lactobacillus.12,26 These microbial compounds are predominantly bacterial in origin, and some studies have shown that virtually any bacteria can produce bacteriocins.27,28 Due to the diversity of bacteria producing bacteriocins, numerous bacteriocins have been identified, and some bacteria can produce more than one bacteriocin.29 These bacteriocins can be used in various fields of biotechnology, industry, and pharmaceuticals.22,30,31
The food industry is one of the main areas for potential bacteriocin use.32 Due to their probiotic properties, some LABs and their metabolites are generally considered safe for use in the food industry.33 Therefore, LABs can inhibit the biological capacity of competing flora,30 especially that of foodborne pathogens such as Listeria monocytogenes, Clostridium, Staphylococcus, and E. coli. Lactic acid bacteria (LAB) can be used in multiple ways, including fermentation and food preservation.34-36As bacteriocins are readily degraded by proteolytic enzymes, they can be considered safe for human use, e.g., proteases in the gastrointestinal tract of mammals.33
Unlike antibiotics, bacteriocins can act on particular pathogens without affecting other symbiotic flora .37 The European Union has now authorized the use of the bacteriocin nisin as a food preservative; by the World Health Organization nisin has been registered as a food additive under the number E23438
However, the development of bacteriocins as biological preservatives is still of great interest. Bacteriocins have been widely studied, particularly in relation to the food industry. For example, it has been shown that bacteriocins can improve the biological safety of vegetable products39, and they can be utilized in effective packaging.40 Various methods can be used to add bacteriocins to foods. Bacteriocins can be added via direct inoculation or indirectly as lactic acid bacteria that produce bacteriocin products, as purified bacteriocins for food preservation, and the addition of LAB-containing fermentation products as food components.41-42
Another important potential use of bacteriocins is in combating antibiotic-resistant bacteria.12,43 Drug-resistant bacteria reduce antibiotic efficacy, a major problem in societies due to the challenges in developing alternative antibiotic treatment methods.5,44-47 Due to the presence of a survival factor in bacteria, bacteriocins can antagonize antibiotic-resistant bacteria, and the mechanisms they use to overcome resistance are unique.19 Pneumococci, Pseudomonas aeruginosa, and Escherichia coli are resistant to penicillin and have attracted considerable attention due to their pathogenic potential.12,19 Molecular diversity, specificity of antimicrobial mechanisms, and potential synergism with other drugs are among the advantages that make bacteriocins attractive pharmacological targets. However, some disadvantages, such as sensitivity to proteolytic enzymes and possible adverse effects on mammalian cells, exist.48,49 Bacteriocins have enormous potential to replace antibacterial compounds or could be used in combination with antibiotics.11 In vivo studies have shown that bacteriocins can be effective against recurrent infections (mycobacteria, streptococci, etc.).
This study aims to evaluate the antimicrobial potential of antagonistic microorganisms.
Isolation of antagonistic microorganisms
Isolates of microorganisms from water, soil, plants, and fresh vegetables (Kemerovo region, Russia) were used as they contain a large number of LAB strains. Samples were collected and stored under sterile conditions at 4°C before delivery to the laboratory.
The samples were weighed on a technical scale (10 g) by pouring with a spatula on sterile paper to isolate the microorganisms. A flask containing MRS medium was placed in a water bath and heated until the medium was completely melted. The molten nutrient medium was poured under heat into sterile Petri dishes (10 mL each), mixed by careful swaying, and left to solidify. A sample of soil water, crushed plant parts, or vegetables was placed in a flask containing 90 mL of sterile water and gently shaken periodically in a circular motion for 5 min.
All 20 isolates were cultured as described below. The resulting suspension (1 mL) was transferred using a sterile pipette into a smaller volume flask. Nine milliliters of sterile water was added and slightly shaken, after which 1 mL of the resulting mixture was transferred to another flask. From the test tubes of the third and fourth dilutions, 1 mL of the suspension was transferred to sterile petri dishes containing liquid agar. To optimize the conditions for bacterial growth, the nutrient medium was acidified with 0.4% lactic acid. After inoculation, the dishes were wrapped in sterile envelopes and incubated at 23–25°C for 5 to 10 d. Pure cultures were obtained from individual colonies of lactobacilli and stored on MRS medium.
16S rDNA and ITS nrDNA sequencing and phylogeny
DNA extraction was performed using the DNeasy Ultra Clean Microbial Kit (Qiagen, Hilden, Germany). Samples were amplified using qPCR mix-HS Mix (Eurogen) in duplicate. The annealed regions of the primers coincided with the standard primer 1492R 5′-CGGTTACCTTGTTACGACTT-3, primer 27F 5′-AGAGTTTGATCMTGGCTCAG-3′, primer ITS3 F 5′-GCATCGATGAAGAACGCATTAGC-3′, and primer ITS4 RCCGATGATGATTGATT GATGATTGATTGATTGATTGATTGATT GATTGATT GATTGATTGATITS4 RCCGATGATGATCITS4 RCCGATGATGATTGATTGATTGATTGATTGAT TGATTGATTGATITS4. Amplification was carried out using a BioRad C1000 thermal cycler (BioRad Laboratories, Hercules, CA, USA) as follows: denaturation at 98 °C for 30 s, annealing at 57 °C for 30 s, and extension at 72 °C for 90 s. The libraries were purified using the Cleanup Mini kit (Eurogen). The quality of the purified libraries was checked by agarose gel electrophoresis, and the library concentrations were measured using a Qubit 2.0 Fluorometer (Thermo Fisher Scientific, Waltham, MA, USA).
Capillary sequence analysis was performed using a Big Dye Terminator Cycle Sequencing kit (Applied Biosystems, Foster City, CA, USA) with a Genetic Analyzer (Applied Biosystems). The sequence reads obtained were assembled into contigs using CLC Genomics Workbench software. Bioinformatic analysis was conducted using local algorithms and information from GenBank and the Ribosomal Database Project. Analysis of the phylogenetic sequences of the 16SrRNA and pDNA genes was performed using MEGA X.
Physiological and biochemical properties of microorganisms
The physiological and biochemical properties of bacteria were evaluated using the API 50 test system (Pro-mix, Moscow, Russia). Additionally, M21 medium with analyzed carbohydrate (10 g/L) and bromocresol purple dye (BCP; 32 mg/L) were used. Sugar utilization was confirmed by the color change of the colonies and nutrient medium to yellow.
The ability of bacteria to degrade esculin was determined using a medium consisting of 0.5% casein peptone, 0.3% meat extract, 0.05% iron citrate, 0.1% esculin, and 1.5% agar 1.5 at pH 6.6. Microorganisms were cultivated by incubation for 12 h at 37 °C. For esculin fermentation, medium darkening around the colonies and loss of fluorescence were observed.
Pathogenic microorganisms
Test strains of pathogenic microorganisms were purchased from the State Collection of Pathogenic Microorganisms and Cell Cultures (GKPM-Obolensk, Russia). Pathogenic effects were determined using E. coli ATCC 25922, S. enterica ATCC 14028, S. aureus ATCC 25923, P. aeruginosa B6643, B. mycoides EMTC 9, A. faecalis EMTC 1882, P. vulgarisella ATCC 63, P. vulgarisella ATCC 7644, C. albicans EMTC 34, A. flavus ATCC 9643, and P. citrinum ATCC 9849.
Cultivation of test microorganism strains
Media for cultivating pathogenic microorganisms were prepared immediately before testing. Bacterial strains were cultivated for 24 h at 30–37 °C in special media51. The optical density and number of bacteria were determined at 595 nm.
Screening of isolated microorganisms for antimicrobial activity
Screening for the detection of microorganisms with antagonistic properties was carried out using the spot method according to Lucke and Schillinger with some modifications50. Briefly, the cultures were placed on plates (1.2% agar) and incubated for 24 h at 30 °C to release metabolites and multiply bacteria. Approximately 5 ´ 107 CFU/mL of indicator strains important in food industry (E. coli, S. enterica, S. aureus, P. aeruginosa, B. mycoides, A. faecalis, S. flexneri, L. monocytogenes, and C. albicans and P. citrinum) were inoculated into 100 mL of soft trypticase soy agar containing 0.7% agar and poured over a plate in which the isolated LAB were grown. After a 24-hour incubation at 37 °C, the inhibition of microorganisms from the edge of the zone was measured using a caliper and expressed in millimeters.
Statistical analysis
Experiments were performed in three sequences, experimental values were expressed as mean ± standard error, and statistical processing was performed using Microsoft Office Excel 2007 (Microsoft Corporation, Redmont, WA, USA) and Statistica 10.0 (StatSoft, Tulsa, OK, USA). Equality of means was tested using the median test and the nonparametric Kruskal-Wallis test; if p d” 0.05, then the differences were considered significant.
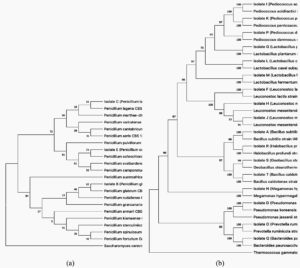
Identification of microorganisms based on 16S rDNA sequencing is presented in Table 1. The LAB position according to the 16S rDNA and ITS nrDNA sequences is shown in the phylogenetic tree. Bordetella pertussis and Saccharomyces cerevisiae were investigated separately using MEGA-5; the starting position for 1,000 replicas is shown at the nodes of the phylogenetic tree. The scale bar corresponds to 0.05 units of the number of base substitutions per site. The phylogenetic trees are shown in Figure 1.
Table (1):
Results of the identification of microorganisms depending on the isolation sources.
Isolate |
Isolation source |
Species |
Isolate |
Isolation source |
Species |
|---|---|---|---|---|---|
A |
I |
B. subtilis |
K |
III |
P. pentosaceus |
B |
I |
P. glabrum |
L |
II |
L. casei |
C |
II |
P. lagena |
M |
II |
L. fermentum |
D |
I |
P. koreensis |
N |
I |
B. hypermegas (M. hypermegale) |
E |
III |
P. ochrochloron |
O |
I |
B. ruminicola
(P. ruminicola) |
F |
IV |
L. lactis |
P |
III |
P.s damnosus |
G |
IV |
L. plantarum |
Q |
I |
Bacteroides paurosaccharolyticus |
H |
III |
L. mesenteroides |
R |
V |
H. profundi |
I |
I |
P. acidilactici |
S |
III |
G. stearothermophilus |
J |
II |
L. mesenteroides |
T |
I |
B. caldotenax |
I – soil; II – fruits and vegetables; III -natural water bodies; IV – food wastes; V — wastewater
Fig. 1. Phylogenetic tree showing the relative position of fungi isolates based on ITS nrDNA (a) and on 16S rDNA (b) sequences, using the neighbor-joining method
The ability to ferment various carbohydrates, including sugars, alcohols, and organic acids, underlies the distinguishing characteristics of bacteria. In the present study, the ability of microorganisms to use various organic substances as carbon and nitrogen sources was investigated.
Table (2):
Physiological and biochemical properties of microorganisms isolated from natural sources of the Kemerovo region.
| Substance | Samples | |||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | |||||
| Control | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| Gentiobiose | − | − | − | − | − | − | + | + | − | + | − | − | + | + | + | − | − | − | − | − | ||||
| Erythritol | − | − | − | − | − | − | − | − | − | − | − | − | + | + | + | − | − | − | − | − | ||||
| L-rham−se | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| D-fructose | − | − | − | − | + | + | + | + | − | + | − | + | + | + | + | + | + | − | − | + | ||||
| D-sucrose | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | − | − | − | ||||
| D-arabi−se | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| Potassium gluconate | − | − | − | − | − | − | + | + | − | + | − | + | + | + | + | − | − | − | − | − | ||||
| Methyl-βD-xylopyra−side | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| Esculin | − | − | − | − | + | + | + | + | − | + | − | − | + | + | + | + | + | − | − | + | ||||
| Amygdalin | − | − | − | − | − | − | + | + | − | + | − | − | + | + | + | − | − | − | − | − | ||||
| Salicin | − | + | + | + | − | − | + | + | + | + | + | − | + | + | + | − | − | + | + | − | ||||
| L-sorbose | − | + | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| Glycogen | − | − | − | − | − | − | − | − | − | − | − | − | + | + | + | − | − | − | − | − | ||||
| Dulcitol | − | + | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| I−sitol | − | + | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| D-glucose | + | + | + | − | + | + | + | + | − | + | − | + | + | + | + | + | + | + | + | + | ||||
| D-sorbitol | − | + | + | − | − | − | − | − | − | − | − | + | + | + | + | − | − | − | − | − | ||||
| D-raffi−se | − | + | + | + | − | − | + | + | + | + | + | − | + | + | + | − | − | − | − | − | ||||
| Methyl-αD-glucopyra−side | − | + | + | − | − | − | − | − | − | − | − | + | − | − | − | − | − | − | − | − | ||||
| D-adonitol | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| D-cellobiose | − | + | + | − | − | − | + | + | + | + | − | + | + | + | + | − | − | + | + | − | ||||
| D-melezitose | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| D-lactose | + | + | + | − | + | + | − | + | − | + | − | + | − | − | − | − | − | − | − | − | ||||
| Starch | + | + | + | − | − | − | − | − | − | − | − | − | + | + | + | − | − | + | + | − | ||||
| D-arabitol | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| Inulin | − | − | − | − | − | − | − | − | − | − | − | + | − | − | − | − | − | − | − | − | ||||
| D-fucose | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| D-trehalose | − | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | − | − | + | ||||
| Xylitol | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| Potassium 2-ketogluconate | − | − | − | − | − | − | + | − | − | − | − | − | − | − | − | − | − | − | ||||||
| D-tura−se | − | − | − | − | − | − | − | + | − | + | − | + | − | − | − | − | − | − | − | − | ||||
| D-melibiose | − | − | − | + | + | + | − | + | + | + | + | − | − | − | − | + | + | − | − | − | ||||
| D-lyxose | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
| D-mannitol | + | + | + | − | − | − | + | + | − | + | − | + | + | + | + | − | − | − | − | − | ||||
| L-arabi−se | − | + | + | + | − | − | + | + | + | + | + | − | + | + | + | − | − | − | − | − | ||||
| Glycerol | − | + | + | − | − | − | − | − | − | − | − | − | + | + | + | − | − | − | − | − | ||||
| D-xylose | − | − | − | + | − | − | − | + | + | + | + | − | − | − | − | − | − | − | − | − | ||||
| Methyl-αD-man−pyra−side | − | + | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | ||||
“+”– positive result; “−” – negative result. Isolates A – B. subtilis; B – P. glabrum; C – P. lagena; D – P. koreenis; E – P. ochrochloron; F – L. lactis; G – L. plantarum; H – L. mesenteroides; I – P. acidilactici; J – L. mesenteroides; K – P. pentosaceus; L – L. casei; M – L. fermentum; N – B. hypermegas; O – B. ruminicola; P – P. damnosus; Q – B. paurosaccharolyticus; R – H. profundi; S – G. stearothermophilus; T – B. caldotenax.
The physiological and biochemical properties of the microorganisms isolated from natural sources are presented in Table 2. Isolate A (B. subtilis) ferments D-glucose, D-mannitol, D-maltose, D-lactose, D-sucrose, starch. Isolate B (P. glabrum) and isolate C (P. lagena) actively ferment glycerol, L-arabinose, D-glucose, D-mannose, L-sorbose, dulcitol, inositol, D-mannitol, D-sorbitol, methyl-aD-mannopyranoside, methyl-aD-glucopyranoside, salicin, D-cellobiose, D-maltose, D-lactose, D-sucrose, D-trehalose, D-raffinose, and starch. Isolate E (P. ochrochloron) and isolate F (L. lactis) ferment, D-glucose, D-fructose, D-mannose, esculin, D-maltose, D-lactose, D-melibiose, D-sucrose, D-trehalose. Isolate G (L. plantarum) ferments L-arabinose, D-glucose, D-fructose, D-mannose, D-mannitol, amygdalin, esculin, salicin, D-cellobiose, D-sucrose, D-trehalose, D-raffinose, gentiobiose, potassium gluconate, potassium 2-ketogluconate. Isolates H and J ferment L-arabinose, D-xylose, D-glucose, D-fructose, D-mannose, D-mannitol, amygdalin, esculin, salicin, D-cellobiose, D-lactose, D-melibiose, D-sucrose, D-trehalose, D-raffinose, gentiobiose, D-turanose, potassium gluconate.
Table (3):
Results of determining the antimicrobial activity of microorganisms isolated from natural sources.
| Sample | Lysis zone diameter, mm | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| I | II | III | IV | V | VI | VII | VIII | IX | X | XI | XII | |
| Control 1 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 |
| Control 2 | 21.0±1.1 | 24.0±1.2 | 19.0±1.0 | 22.0±1.1 | 25.0±1.3 | 23.0±1.2 | 20.0±1.0 | 22.0±1.1 | 24.0±1.2 | 18.0±0.9 | 25.0±1.33 | 21.0±1.1 |
| A | 18.0±0.9 | 20.0±1.0 | 17.0±0.9 | 20.0±1.0 | 22.0±1.1 | 20.0±1.0 | 17.0±0.9 | 18.0±0.9 | 19.0±1.0 | 15.0±0.8 | 21.0±1.1 | 15.0±0.8 |
| B | 0.0±0.0 | 0.0±0.0 | 5.0±0.3 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 7.0±0.4 | 4.0±0.2 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 |
| C | 6.0±0.3 | 10.0±0.5 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 8.0±0.4 | 0.0±0.0 | 0.0±0.0 | 4.0±0.2 | 5.0±0.3 |
| D | 12.0±0.6 | 5.0±0.3 | 18.0±0.9 | 0.0±0.0 | 0.0±0.0 | 17.0±0.9 | 15.0±0.8 | 10.0±0.5 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 |
| E | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 6.0±0.3 | 0.0±0.0 | 0.0±0.0 | 4.0±0.2 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 |
| F | 20.0±1.0 | 22.0±1.1 | 17.0±0.9 | 21.0±1.1 | 24.0±1.2 | 21.0±1.1 | 18.0±0.9 | 20.0±1.0 | 21.0±1.1 | 16.0±0.8 | 23.0±1.2 | 19.0±1.0 |
| G | 19.0±1.0 | 18.0±0.9 | 15.0±0.8 | 19.0±1.0 | 22.0±1.1 | 21.0±1.1 | 17.0±0.9 | 20.0±1.0 | 23.0±1.2 | 16.0±0.8 | 22.0±1.1 | 18.0±0.9 |
| H | 17.0±0.9 | 20.0±1.0 | 16.0±0.8 | 19.0±1.0 | 22.0±1.1 | 20.0±1.0 | 18.0±0.9 | 19.0±1.0 | 21.0±1.1 | 15.0±0.8 | 21.0±1.1 | 18.0±0.9 |
| I | 20.0±1.0 | 21.0±1.1 | 17.0±0.9 | 19.0±1.0 | 23.0±1.2 | 21.0±1.1 | 18.0±0.9 | 19.0±1.0 | 21.0±1.1 | 16.0±0.8 | 24.0±1.2 | 18.0±0.9 |
| J | 15.0±0.8 | 18.0±0.9 | 14.0±0.7 | 17.0±0.9 | 20.0±1.0 | 17.0±0.9 | 15.0±0.8 | 16.0±0.8 | 20.0±1.0 | 13.0±0.7 | 20.0±1.0 | 18.0±0.9 |
| K | 21.0±1.1 | 20.0±1.0 | 17.0±0.9 | 19.0±1.0 | 24.0±1.2 | 22.0±1.1 | 18.0±0.9 | 18.0±0.9 | 21.0±1.1 | 16.0±0.8 | 24.0±1.2 | 19.0±1.0 |
| L | 18.0±0.9 | 19.0±1.0 | 15.0±0.8 | 10.0±0.5 | 20.0±1.0 | 17.0±0.9 | 15.0±0.8 | 17.0±0.9 | 22.0±1.1 | 16.0±0.8 | 22.0±1.1 | 17.0±0.9 |
| M | 15.0±0.8 | 18.0±0.9 | 14.0±0.7 | 19.0±1.0 | 21.0±1.1 | 17.0±0.9 | 15.0±0.8 | 16.0±0.8 | 21.0±1.1 | 15.0±0.8 | 20.0±1.0 | 18.0±0.9 |
| N | 12.0±0.6 | 10.0±0.5 | 0.0±0.0 | 14.0±0.7 | 0.0±0.0 | 9.0±0.5 | 11.0±0.6 | 13.0±0.7 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 |
| O | 7.0±0.4 | 11.0±0.6 | 0.0±0.0 | 12.0±0.6 | 0.0±0.0 | 7.0±0.4 | 10.0±0.5 | 15.0±0.8 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 |
| P | 17.0±0.9 | 22.0±1.1 | 16.0±0.8 | 20.0±1.0 | 23.0±1.2 | 19.0±1.0 | 18.0±0.9 | 21.0±1.1 | 20.0±1.0 | 16.0±0.8 | 21.0±1.1 | 18.0±0.9 |
| Q | 15.0±0.8 | 13.0±0.7 | 0.0±0.0 | 16.0±0.8 | 0.0±0.0 | 12.0±0.6 | 14.0±0.7 | 17.0±0.9 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 |
| R | 0.0±0.0 | 0.0±0.0 | 11.0±0.6 | 0.0±0.0 | 14.0±0.7 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 | 10.0±0.5 | 0.0±0.0 | 0.0±0.0 | 0.0±0.0 |
| S | 20.0±1.0 | 22.0±1.1 | 18.0±0.9 | 19.0±1.0 | 22.0±1.1 | 20.0±1.0 | 18.0±0.9 | 21.0±1.1 | 19.0±1.0 | 16.0±0.8 | 24.0±1.2 | 18.0±0.9 |
| T | 18.0±0.9 | 23.0±1.2 | 17.0±0.9 | 20.0±1.0 | 21.0±1.1 | 22.0±1.1 | 19.0±1.0 | 21.0±1.1 | 23.0±1.2 | 15.0±0.8 | 20.0±1.0 | 17.0±0.9 |
Control 1 – culture medium without any additions; control 2 – Ciprofloxacin; isolates A – B. subtilis; B – P. glabrum; C – P. lagena; D – P. koreenis; E – P. ochrochloron; F – L. lactis; G – L. plantarum; H – L. mesenteroides; I – P. acidilactici; J – L. mesenteroides; K – P. pentosaceus; L – L. casei; M – L. fermentum; N – B. hypermegas; O – B. ruminicola; P – P. damnosus; Q – B. paurosaccharolyticus; R – H. profundi; S – G. stearothermophilus; T – B. caldotenax. Test-cultures I – E. coli; II – S. enterica; III – S. aureus; IV – P. aeruginosa; V – B. mycoides; VI – A. faecalis; VII – P. vulgaris; VIII – S. flexneri; IX – L. monocytogenes; X – C. albicans; XI – A. flavus; XII – P. citrinum.
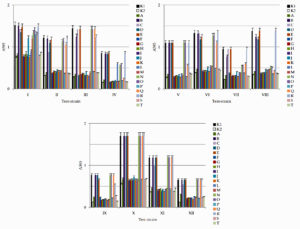
All microorganisms were isolated using one type of medium under similar conditions, which were optimal for their cultivation. The antimicrobial properties of lactic acid bacteria and other antagonistic microorganisms are presented in Table 3 and Fig. 2.
Fig. 2. Results of determination of antimicrobial activity against test-cultures I – E. coli; II – S. enterica; III – S. aureus; IV – P. aeruginosa; V – B. mycoides; VI – A. faecalis; VII – P. vulgaris; VIII – S. flexneri; IX – L. monocytogenes; X – C. albicans; XI – A. flavus; XII – P. citrinum in a liquid nutrient medium isolates of microorganisms: K1 – control (culture medium without any additions); K2 – ciprofloxacin; isolates A – B. subtilis; B – P. glabrum; C – P. lagena; D – P. koreenis; E – P. ochrochloron; F – L. lactis; G – L. plantarum; H – L. mesenteroides; I – P. acidilactici; J – L. mesenteroides; K – P. pentosaceus; L – L. casei; M – L. fermentum; N – B. hypermegas; O – B. ruminicola; P – P. damnosus; Q – B. paurosaccharolyticus; R – H. profundi; S – G. stearothermophilus; T – B. caldotenax. Test-cultures I – E. coli; II – S. enterica; III – S. aureus; IV – P. aeruginosa; V – B. mycoides; VI – A. faecalis; VII – P. vulgaris; VIII – S. flexneri; IX – L. monocytogenes; X – C. albicans; XI – A. flavus; XII – P. citrinum
Data analysis (Fig. 2, Table 3) showed that isolate A (B. subtilis) exhibited antimicrobial activity against all pathogenic and opportunistic microorganisms, with the greatest antimicrobial activity against B. mycoides (22.0 ± 1.1), S. enterica (20.0 ± 1.0), P. aeruginosa (20.0 ± 1.0), A. faecalis (20.0 ± 1.0), and A. flavus (21.0 ± 1.1). Isolate B (P. glabrum) was only slightly active against S. aureus (5.0 ± 0.3), P. vulgaris (7.0 ± 0.4), and S. flexneri (4.0 ± 0.2). Isolate C (P. lagena) exhibited moderate to low antimicrobial activity against E. coli (6.0 ± 0.3), S. enterica (10.0 ± 0.5), S.flexneri (8.0 ± 0.4), A. flavus (4.0 ± 0.2), and P. citrinum (5.0 ± 0.3). Isolate D (P. koreenis) exhibited moderate antimicrobial activity against E. coli (12.0 ± 0.6), S. enterica (5.0 ± 0.3), S. aureus (18.0 ± 0.9), A. faecalis (17.0 ± 0.9), P. vulgaris (15.0 ± 0.8), and S. flexneri (10.0 ± 0.5). Isolate E (P. ochrochloron) had low antimicrobial activity against A. faecalis (6.0 ± 0.3) and L. monocytogenes (4.0 ± 0.2). Isolate F (L. lactis) exhibited antibacterial activity against all pathogenic and opportunistic bacteria, with the greatest antimicrobial activity against B. mycoides (24.0 ± 1.2), A. flavus (23.0 ± 1.2), and S. enterica (22.0 ± 1.1). Isolate G (L. plantarum) showed antibacterial potential against all pathogenic and opportunistic bacteria; its greatest antimicrobial activity was against L. monocytogenes (23.0 ± 1.2), B. mycoides (22.0 ± 1.1), A. flavus (22.0 ± 1.1), and A. faecalis (21.0 ± 1.1). Isolate H (L. mesenteroides) exhibited antibacterial activity against all pathogenic and opportunistic bacteria; its greatest antimicrobial activity was against B. mycoides (22.0 ± 1.1), L. monocytogenes (21.0 ± 1.1), and A. flavus (21.0 ± 1.1). Isolate I (P. acidilactici) showed antibacterial potential against all pathogenic and opportunistic bacteria with the highest antimicrobial activity against A. flavus (24.0 ± 1.2), B. mycoides (23.0 ± 1.2), A. faecalis (21.0 ± 1.1), S. enterica (21.0 ± 1.1), and L. monocytogenes (21.0 ± 1.1). Isolate J (L. mesenteroides) showed antibacterial activity against all pathogenic and opportunistic bacteria with the highest activity against B. mycoides (20.0 ± 1.0), L. monocytogenes (20.0 ± 1.0), and A. flavus (20.0 ± 1.0). Isolate K (P. pentosaceus) showed antibacterial activity against all pathogenic and opportunistic bacteria; its greatest antimicrobial activity was against B. mycoides (24.0 ± 1.2), A. flavus (24.0 ± 1.2), and A. faecalis (22.0 ± 1.1). Isolate L (L. casei) showed antibacterial potential against all pathogenic and opportunistic bacteria; its greatest antimicrobial activity was against L. monocytogenes (22.0 ± 1.1) and A. flavus (22.0 ± 1.1). Isolate M (L. fermentum) exhibited antibacterial potential against all pathogenic and opportunistic bacteria, with the greatest antimicrobial activity against B. mycoides (21.0 ± 1.1) and L. monocytogenes (21.0 ± 1.1). Isolate N (B. hypermegas) exhibited moderate antimicrobial activity against E. coli (12.0 ± 0.6), S. enterica (10.0 ± 0.5), P. aeruginosa (14.0 ± 0.7), A. faecalis (9.0 ± 0.5), P. vulgaris (11.0 ± 0.6), and S. flexneri (13.0 ± 0.7). Isolate O (B. ruminicola) exhibited moderate antimicrobial activity against E. coli (7.0 ± 0.4), S. enterica (11.0 ± 0.6), P. aeruginosa (12.0 ± 0.6), A. faecalis (7.0 ± 0.4), P. vulgaris (10.0 ± 0.5) and S. flexneri (15.0 ± 0.8). Isolate P (P. damnosus) exhibited antibacterial potential against all pathogenic and opportunistic bacteria, with the greatest antimicrobial activity against B. mycoides (23.0 ± 1.2), S. enterica (22.0 ± 1.1), S. flexneri (21.0 ± 1.1), and A. flavus (21.0 ± 1.1). Isolate Q (B. paurosaccharolyticus) exhibited moderate antimicrobial activity against E. coli (15.0 ± 0.8), S. enterica (13.0 ± 0.7), P. aeruginosa (16.0 ± 0.8), A. faecalis (12.0 ± 0.6), P. vulgaris (14.0 ± 0.7,) and S.flexneri (17.0 ± 0.9). Isolate R (H. profundi) exhibited moderate antimicrobial activity against Staphylococcus aureus (11.0 ± 0.6), Bacillus mycoides (14.0 ± 0.7), and Listeria monocytogenes (10.0 ± 0.5). Isolate S (G. stearothermophilus) exhibited antibacterial potential against all pathogenic and opportunistic bacteria, with the greatest antimicrobial activity against Aspergillus flavus (24.0 ± 1.2), B. mycoides (22.0 ± 1.1), S. enterica (22.0 ± 1.1), and S. flexneri (21.0 ± 1.1). Isolate T (B. caldotenax) exhibited antibacterial potential against all pathogenic and opportunistic bacteria; the greatest antimicrobial activity was against S. enterica (23.0 ± 1.2), L. monocytogenes (23.0 ± 1.2), and A. faecalis (22.0 ± 1.1).
Table (4):
Statistical analysis results.
| Isolates | Test cultures | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| I | II | III | IV | V | VI | VII | VIII | IX | X | XI | XII | |
| A | − | − | + | + | + | + | + | + | + | + | + | + |
| B | − | − | − | − | − | − | − | − | − | − | − | − |
| C | − | − | − | − | − | − | − | − | − | − | − | − |
| D | − | − | + | − | − | − | + | − | − | − | − | − |
| E | − | − | − | − | − | − | − | − | − | − | − | − |
| F | + | + | + | + | + | + | + | + | + | + | + | + |
| G | + | + | + | + | + | + | + | + | + | + | + | + |
| H | + | + | + | + | + | + | + | + | + | + | + | + |
| I | + | + | + | + | + | + | + | + | + | + | + | + |
| J | − | + | + | + | + | + | + | + | + | + | + | + |
| K | + | + | + | + | + | + | + | + | + | + | + | + |
| L | + | + | + | + | + | + | + | + | + | + | + | + |
| M | − | + | + | + | + | + | + | + | + | + | + | + |
| N | − | − | − | − | − | − | ± | + | − | − | − | − |
| O | − | − | − | − | − | − | ± | + | − | − | − | − |
| P | − | + | + | + | + | + | + | + | + | + | + | + |
| Q | − | − | − | + | ± | − | ± | + | − | − | − | − |
| R | − | − | − | − | − | − | − | − | − | − | − | − |
| S | + | + | + | + | + | + | + | + | + | + | + | + |
| T | + | + | + | + | + | + | + | + | + | + | + | + |
“+” no statistically significant differences in values in comparison with the antibiotic; “-” no statistically significant differences in values in comparison with the control; «±» – statistically significant differences in values in comparison with both antibiotic and control. Test-cultures I – E. coli; II – S. enterica; III – S. aureus; IV – P. aeruginosa; V – B. mycoides; VI – A. faecalis; VII – P. vulgaris; VIII – S. flexneri; IX – L. monocytogenes; X – C. albicans; XI – A. flavus; XII – P. citrinum.
The largest number of strains of lactic acid bacteria and other antagonistic microorganisms were isolated from the soil and natural water bodies (Table 1), including B. subtilis, P. glabrum, P. koreenis, P. ochrochloron, L. mesenteroides, P. acidilactici, P. pentosaceus, B. hypermegale, B. ruminicola, P. damnosus, B. paurosaccharolyticus, G. stearothermophilus, and B. caldotenax.
A culture liquid without metabolites of isolated microorganisms, without antimicrobial characteristics, was considered as a control. The antibiotic ciprofloxacin has antimicrobial properties. Statistical analysis was performed using the nonparametric Kruskal-Wallis test. The median test for two independent samples for each test culture established the presence or absence of antimicrobial characteristics by confirming or refuting the equality of the means in the samples when compared with the antibiotic or control. The absence of statistically significant differences (p < 0.05) from the control was considered to be due to the absence of antibacterial characteristics, and the absence of statistically significant differences with the antibiotics was considered as the presence of antibacterial properties. Statistically significant differences between the control and antibiotic groups were interpreted as mild antibacterial characteristics of the isolate (Table 4).
The isolates F–I, K, L, S, and T, which affected all test cultures under consideration, were the most promising for further research. These isolates included both lactic acid bacteria and other microorganisms. All lactic acid bacteria isolated from natural sources exhibited antimicrobial efficacy.
The results of our studies are consistent with those presented by other authors.52-54 In the present study,52 LAB strains were characterized, and their antimicrobial activity was investigated. Of the 800 isolates, 20 inhibited the growth of enterotoxigenic E. coli and S. enterica. Based on the 16S rRNA sequence analysis, 20 isolates were identified as L. casei (7), L. paracasei (2), L. plantarum (4), L. rhamnosus (2), E. avium (3), E. faecium (1), and E. lactis (1). The natural ability of L. plantarum and L. rhamnosus to inhibit P. aeruginosa growth has been established51. In a previous study,54 lactic acid was isolated from 32 samples of lactic acid bacteria, and 13 (13/32) of the best Lactobacillus isolates were selected by preliminary selection as potential probiotics with antimicrobial activity against pathogenic bacteria. All Lactobacillus isolates were then characterized in vitro for their antimicrobial activity against pathogens. The isolates were resistant to all investigated pathogens, including E. coli (opportunistic bacterium causing gastroenteritis in humans), S. enterica (opportunistic bacterium causing gastroenteritis in humans), P. vulgaris (opportunistic bacterium causing acute intestinal infections in humans), S. flexneri (causative agent of bacterial dysentery in humans), and L. monocytogenes (causative agent of listeriosis in humans).
Out of the 20 tested microorganisms, 11 exhibited high antimicrobial activity against all tested strains of pathogenic and opportunistic microorganisms. Among the isolated strains, some demonstrated limited antimicrobial activity against only gram-positive or gram-negative test strains, and this requires further study.
As the number of antibiotic-resistant strains among pathogenic and opportunistic microorganisms continues to grow, the study of bacteriocins that are safe for humans as alternative antimicrobial substances is very timely. Non-toxicity, biological safety, and the possibility of combining bacteriocins with other antimicrobial agents (antibiotics, bacteriophages, etc.) will determine their use alone or combined with drugs for antimicrobial therapy. Bacteriocin production is an important characteristic of lactic acid bacteria strains, which should be considered when developing drugs for correcting dysbiotic conditions. Considering the significant potential of bacteriocins and their growing demand, methods for their selection and subsequent chemical synthesis are currently under development.55 Recently, information on the possibility of using bacteriocins to fight oncological diseases has been made available. Research in this direction continues, and several research groups, including ours, are studying the possibility of using bacteriocins in treating oncological and infectious bacterial diseases as an alternative to antibiotics.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This research was funded by The Ministry of Science and Higher Education of the Russian Federation within the framework of the Federal Targeted Program “Research and Development in Priority Areas of Development of the Scientific and Technological Complex of Russia for 2014-2020”, Agreement No. 075-02-2018-1934 of December 20, 2018; Agreement No. 075-15-2019-1383 of June 18, 2019 (unique project identifier RFMEFI57418X0207).
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Gould IM. Antibiotic resistance: the perfect storm. Int J Antimicrob Agents. 2009;34:S2-S5.
Crossref - Xia J, Gao J, Kokudo N, Hasegawa K, Tang W. Methicillin-resistant Staphylococcus aureus antibiotic resistance and virulence. Biosci Trends. 2013;7:113-121.
- Giamarellou H, Poulakou G. Multidrug-resistant gram-negative infections. Drugs. 2009;69:1879-1901.
Crossref - Calfee DP. Methicillin-resistant Staphylococcus aureus and vancomycin-resistant Enterococci, and other Gram-positives in healthcare. Curr Opin Infect Dis. 2012;25:385-394.
Crossref - Duval RE, Grare M, Demoré B. Fight against antimicrobial resistance: we always need new antibacterials but for right bacteria. Molecules. 2019;24:3152.
Crossref - Mazurek-Popczyk J, Pisarska J, Bok E, Baldy-Chudzik K. Antibacterial activity of bacteriocinogenic commensal escherichia coli against zoonotic strains resistant and sensitive to antibiotics. Antibiotics. 2020;9(7):411.
Crossref - Planson AG, Carbonell P, Grigoras I, Faulon JL. Engineering antibiotic production and overcoming bacterial resistance. Biotechnol J. 2011;6:812-825.
Crossref - Butler MS, Blaskovich MA, Cooper MA. Antibiotics in the clinical pipeline in 2013. J Antibiot. 2013;66:571-591.
Crossref - Mbaveng AT, Sandjo LP, Tankeo SB, et al. Antibacterial activity of nineteen selected natural products against multidrug resistant Gram-negative phenotypes. SpringerPlus. 2015;4:823.
Crossref - Gratia A. Sur un remarquable exemple d’antagonisme entre deux souches de coilbacille. CR Seances Soc Biol Fil. 1925;93:1040-1041.
- Akerey B, Le-Lay C, Fliss I, Subirade M, Rouabhia M. In vitro efficacy of nisin Z against Candida albicans adhesion and transition following contact with normal human gingival cells. J Appl Microbiol. 2009;107:1298-1307.
Crossref - Hassan M, Kjos M, Nes IF, Diep DB, Lotfipour F. Natural antimicrobial peptides from bacteria: Characteristics and potential applications to fight against antibiotic resistance. J Appl Microbiol. 2012;113:723-736.
Crossref - Torres NI, Noll KS, Xu S, et al. Safety, formulation and in vitro antiviral activity of the antimicrobial peptide subtilosin against herpes simplex virus type 1. Probiotics Antimicrob Proteins. 2013;5:26-35.
Crossref - Park SC, Park Y, Hahm KS. The role of antimicrobial peptides in preventing multidrug-resistant bacterial infections and biofilm formation. Int J Mol Sci. 2011;12:5971-5992.
Crossref - Yasir M, Willcox MDP, Dutta D. Action of antimicrobial peptides against bacterial biofilms. Materials. 2018;11:2468.
Crossref - Cascales E, Buchanan SK, Duché D, et al. Colicin biology. Microbiol Mol Biol Rev. 2007;71:158-229.
Crossref - Martínez B, Rodríguez A, Suárez E. Antimicrobial peptides produced by bacteria: the bacteriocins. In Villa TG, Vinas M (eds.), New weapons to control bacterial growth. Springer International Publishing, Cham, Switzerland, 2016:15-38.
Crossref - Ikryannikova LN, Kurbatov LK, Gorokhovets NV, Zamyatnin AA Jr. Contact-dependent growth inhibition in bacteria: do not get too close! Int J Mol Sci. 2020;21(21):7990.
Crossref - Cotter PD, Ross RP, Hill C. Bacteriocins – A viable alternative to antibiotics? Nat Rev Microbiol. 2013;11:95-105.
Crossref - Heng NCK, Wescombe PA, Burton JP, Jack RW, Tagg JR. The diversity of bacteriocins in gram-positive bacteria. In Riley MA, Chavan MA (eds.), Bacteriocins: ecology and evolution. Springer Berlin Heidelberg, Berlin/Heidelberg, Germany, 2007:45-92.
Crossref - Perez RH, Zendo T, Sonomoto K. Novel bacteriocins from lactic acid bacteria (LAB): Various structures and applications. Microb Cell Factories. 2014;13:S3.
Crossref - Yang SC, Lin CH, Sung CT, Fang JY. Antibacterial activities of bacteriocins: Application in foods and pharmaceuticals. Front Microbiol. 2014;5:241.
Crossref - Kleinkauf H, von Döhren H. Review nonribosomal biosynthesis of peptide antibiotics. In Christen P, Hofmann E (eds.), EJB Reviews 1990. Springer, Berlin/Heidelberg, Germany, 1990:151-165.
Crossref - Jack RW, Tagg JR, Ray B. Bacteriocins of gram-positive bacteria. Microbiol Mol Biol Rev. 1995;59:171-200.
Crossref - Chavan MA, Riley MA. Molecular evolution of bacteriocins in gram-negative bacteria. In Riley MA, Chavan MA, editors. Bacteriocins: ecology and evolution. Springer, Berlin/Heidelberg, Germany, 2007:19-43.
Crossref - Klaenhammer TR. Bacteriocins of lactic acid bacteria. Biochimie. 1988;70:337-349.
Crossref - O’Connor PM, O’Shea EF, Guinane CM, O’Sullivan O, Cotter PD, Ross RP, Hill C. Nisin H is a new nisin variant produced by the gut-derived strain Streptococcus hyointestinalis DPC6484. Appl Environ Microbiol. 2015;81:3953-3960.
Crossref - Majeed H, Lampert A, Ghazaryan L, Gillor O. The weak shall inherit: bacteriocin-mediated interactions in bacterial populations. PLoS ONE. 2013;8:e63837.
Crossref - Cintas LM, Casaus MP, Herranz C, Nes IF, Hernández PE. Review: bacteriocins of lactic acid bacteria. Food Sci Technol Int. 2001;7:281-305.
Crossref - Simons A, Alhanout K, Duval RE. Bacteriocins, antimicrobial peptides from bacterial origin: overview of their biology and their impact against multidrug-resistant bacteria. Microorganisms. 2020;8(5):639.
Crossref - Balciunas EM, Castillo Martinez FA, Todorov SD, de Melo Franco BDG, Converti A, de Souza Oliveira RP. Novel biotechnological applications of bacteriocins: A review. Food Control. 2013;32:134-142.
Crossref - Deegan LH, Cotter PD, Hill C, Ross P. Bacteriocins: Biological tools for bio-preservation and shelf-life extension. Int Dairy J. 2006;16:1058-1071.
Crossref - Zacharof MP, Lovitt RW. Bacteriocins produced by lactic acid bacteria a review article. APCBEE Procedia. 2012;2:50-56.
Crossref - Abee T, Krockel L, Hill C. Bacteriocins: Modes of action and potentials in food preservation and control of food poisoning. Int J Food Microbiol. 1995;28:169-185.
Crossref - Ahmad V, Khan MS, Jamal QMS, Alzohairy MA, Al Karaawi MA, Siddiqui MU. Antimicrobial potential of bacteriocins: In therapy, agriculture and food preservation. Int J Antimicrob Agents. 2017;49:1-11.
Crossref - Barbosa AAT, Mantovani HC, Jain S. Bacteriocins from lactic acid bacteria and their potential in the preservation of fruit products. Crit Rev Biotechnol. 2017;37:852-864.
Crossref - Rea MC, Sit CS, Clayton E, et al. A posttranslationally modified bacteriocin with a narrow spectrum of activity against Clostridium difficile. Proc Natl Acad Sci USA. 2010;107:9352-9357.
Crossref - Molloy EM, Ross RP, Hill C. ‘Bac’ to the future: Bioengineering lantibiotics for designer purposes. Biochem Soc Trans. 2012;40:1492-1497.
Crossref - Settanni L, Corsetti A. Application of bacteriocins in vegetable food biopreservation. Int J Food Microbiol. 2008;121:123-338.
Crossref - Santos JCP, Sousa RCS, Otoni CG, et al. Nisin and other antimicrobial peptides: Production, mechanisms of action, and application in active food packaging. Innov Food Sci Emerg Technol. 2018;48:179-194.
Crossref - Chen H, Hoover DG. Bacteriocins and their food applications. Compr Rev Food Sci Food Saf. 2003;2:82-100.
Crossref - Beshkova D, Frengova G. Bacteriocins from lactic acid bacteria: Microorganisms of potential biotechnological importance for the dairy industry. Eng Life Sci. 2012;12:419-432.
Crossref - Singh N, Abraham J. Ribosomally synthesized peptides from natural sources. J Antibiot. 2014;67:277-289.
Crossref - Michael CA, Dominey-Howes D, Labbate M. The antimicrobial resistance crisis: causes, consequences, and management. Front Public Health. 2014;2:145.
Crossref - Bush K, Courvalin P, Dantas G, et al. Tackling antibiotic resistance. Nat Rev Microbiol. 2011;9:894-896.
Crossref - Freire-Moran L, Aronsson B, Manz C, et al. Critical shortage of new antibiotics in development against multidrug-resistant bacteria-Time to react is now. Drug Resist Updat. 2011;14:118-124.
Crossref - Toussaint KA, Gallagher JC. b-Lactam/b-Lactamase inhibitor combinations: from then to now. Ann Pharmacother. 2015;49:86-98.
Crossref - Allen HK, Trachsel J, Looft T, Casey TA. Finding alternatives to antibiotics. Ann NY Acad Sci. 2014;1323:91-100.
Crossref - Tenea GN, Hurtado P, Ortega C. A novel weissella cibaria strain utngt21o isolated from wild solanum quitoense fruit: genome sequence and characterization of a peptide with highly inhibitory potential toward gram-negative bacteria. Foods. 2020;9(9):1242.
Crossref - Zimina M, Babich O, Prosekov A, et al. Overview of global trends in classification, methods of preparation and application of bacteriocins. Antibiotics. 2020;9(9):553.
Crossref - Prosekov AY, Dyshlyuk LS, Milentyeva IS, et al. Antioxidant and antimicrobial activity of bacteriocin-producing strains of lactic acid bacteria isolated from the human gastrointestinal tract. Progress in nutrition. 2017;19(1):67-80.
- Wang X, Wang W, Lv H, et al. Probiotic potential and wide-spectrum antimicrobial activity of lactic acid bacteria isolated from Infant Feces. Probiotics & Antimicro Prot. 2021;13:90-101.
Crossref - Chappell TC, Nair NU. Engineered lactobacilli display anti-biofilm and growth suppressing activities against Pseudomonas aeruginosa. NPJ Biofilms Microbiomes. 2020;6:48.
Crossref - Prabhurajeshwar C, Chandrakanth K. Evaluation of antimicrobial properties and their substances against pathogenic bacteria in-vitro by probiotic Lactobacilli strains isolated from commercial yoghurt. Clin Nutr Exp. 2019;23:97-115.
Crossref - Zaslavskaya MI, Makhrova TV, Aleksandrova NA, et al. Prospects for using bacteriocins of normal microbiota in antibacterial therapy (review). Bacteriocins of Microbiota and Antibiotic Therapy. 2019;11(3):136-144.
Crossref
© The Author(s) 2021. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.