ISSN: 0973-7510
E-ISSN: 2581-690X
Cookeina sulcipes and C. tricholoma are a cup fungi (Ascomycota) collected in Saraburi, Thailand. The fungi have been isolated, cultured and confirmed as respective species. For morphology, both Cookeina sp. are white mycelium and the growth rate on potato dextrose agar (PDA) result present C. sulcipes is faster than C. tricholoma. The molecular characterization from a rapid and simple DNA extraction method that is modified based on thermolysis method, The DNA extraction is finish in thirty minutes and efficiency to continuous with polymerase chain reaction (PCR) amplification to fungi species level identification. The DNA sequence from internal transcribed spacer (ITS) gene regions by universal primer pairs ITS5/ITS4 is effective to confirm Cookeina species level that C. sulcipes has 616 bp and C. tricholoma has 570 bp. Including, DNA sequence of large subunit (LSU) gene regions by universal primer pairs LROR/LR5 is generate that C. sulcipes has 912 bp and C. tricholoma has 906 bp. The cultures are screened for antibacterial activity by agar plug diffusion method and found that both isolates have been no activity against test strains (Bacillus subtilis, Escherichia coli, Kocuria rhizophila (Micrococcus luteus), Pseudomonas aeruginosa, Staphylococcus aureus and S. epidermidis). In a preliminary screening test of enzymes involved in plant biomass breakdown by agar plate method, both Cookeina sp. show cellulolytic and hemicellulolytic enzymatic activity, and manganese peroxidase (MnP) productivity. In contrast, only C. sulcipes had additional laccase activity. Neither isolate generate pectinolytic and lignin peroxidase (LiP) activities. Thus, Cookeina spp. proved the potentiality to break down lignocelluloses.
Antibacterial Activity, Cup Fungi, DNA Extraction, Lignocellulolytic Enzymes, rDNA
Genus Cookeina is widely distributed in tropical and subtropical regions, including Thailand.1-4 The genus is generally known as a cup fungus, and its common name is “champagne mushroom” because the fruiting body has a cup shape, similar to Basidiomycota. However, the taxonomy of Cookeina spp. in MycoBank (https://www.mycobank.org/) and The National Center for Biotechnology Information (NCBI, https://www.ncbi.nlm.nih.gov/) online database recognizes them in Asocomycota, Pezizomycetes, Sarcoscyphaceae. At present, Index Fungorum has twenty-eight species of Cookeina (http://www.indexfungorum.org/Names/Names.asp) and on NCBI only eleven species have record. Recently, a new species of Cookeina was recorded ‒ 2016, C. garethjonesii on dead stems in Yunnan1; 2017, C. cremeirosea on woody debris in American Samoa5; 2021, C. colombiana on dead wood of Angiospermae in Colombia.3 Three species are so far found in Thailand ‒ C. speciosa, C. sulcipes and C. tricholoma.1,4 Previous systematic studies of the genus Cookeina, based on morphology and molecular characterization of internal transcribed spacer (ITS) and large subunit (LSU) of the nuclear ribosomal DNA (rDNA) regions, confirmed the taxonomy of the species level.1,2,4 Nowadays, nucleotide information on NCBI has one hundred ninety-eight sequences from the ITS, LSU, small subunit (SSU) ribosomal RNA, RNA polymerase II (RPB2) and elongation factor 1-alpha (EF1-α) gene. The top three species of DNA sequences data are C. tricholoma (58 sequences), C. speciosa (43 sequences) and C. indica (17 sequences). However, C. sulcipes only ten sequences on NCBI database.
For fungi, several methods have been developed and used for DNA extraction, including the standard or modified CTAB extraction method, lysis buffer, SDS method, etc. The first step is to break the fungal cell wall with liquid nitrogen, micropestle, glass bead, sonicator or homogenizer.6-12 Recently, Wingfield and Atcharawiriyakul compared five available simple and rapid methods (NaOH-Tris, Ultra-simple, TE buffer, water, and IRRI method) for extraction of DNA in molecular identification.13 From a user point of view, the mycelium was frozen in liquid nitrogen and ground into a fine powder as an initial step. Pilo, et al.10 presented a method for rapid genomic DNA extraction from filamentous fungi without buffer, using a sonication bath, which required only an hour to accomplish. Therefore, a rapid and simple method for fungal DNA extraction would have significant benefits.
C. sulcipes and C. tricholoma are reported to be edible.14 However, to date, there has been no report on their biological activity. In nature, Cookeina species grow on dead bark or wood indicating that one major function is as decomposer and aid in lignocellulose breakdown. Primary plant cell wall components are 25-50% hemicellulose, 10-35% pectin, 9-25% cellulose and 10% proteins and the secondary cell wall is 40-80% cellulose, 10-40% hemicellulose and 5-25% lignin.15 Involving, lignocellulosic biomass can be obtained from various sources or industries as a wastes product such as agricultural industry waste, plantations waste, organic waste, etc.16,17 Therefore, the enzymes involved in plant biomass breakdown or cell wall degradation are cellulolytic, hemicellulolytic and ligninolytic from brown-, white- and soft-rot fungi,16,18,19 and even pectinolytic enzymes from soft-rot fungi.18,20 This study focused on a more simple, rapid, and efficient method for identifying fungal species and determining antibacterial and enzymatic activities for future study.
Collection and fungal isolation
Specimens of C. sulcipes and C. tricholoma were collected in Saraburi, Thailand, and brought to the laboratory in a plastic box. Fresh fruiting bodies (ascomata) were divided into two parts. The first part was used to study the morphology and the second to isolate the fungus in culture. The fungi were isolated by spore shooting and single spore isolation methods.21 Spores were placed in a Petri dish with water agar (WA) medium, and the dishes were incubated at room temperature (30 ± 2°C) overnight. Germinated spores were placed on a potato dextrose agar (PDA) plate, and colonies were purified by the hyphal tip transfer method. The pure cultures were subcultured into PDA plates and used to study morphological characterization, molecular identification and as stock culture to maintain at the Thailand Mushroom Culture Collection (TMCC), Biotechnology Research and Development Office, Department of Agriculture, Thailand, for future study.
Morphological identification
Fresh fruiting bodies of C. sulcipes and C. tricholoma were cut with a razor blade and thin sections were observed under a light microscope. Morphological identification is done by observing the macroscopic and microscopic characters, together with colony characteristics, colour and growth pattern. The colony dimensions of the fungi were evaluated by transferring 5-mm diameter agar plugs from the periphery of 5-day-old cultures grown at room temperature. The colony characteristics and colour were determined after 7 days of incubation on PDA at room temperature in triplicate. The colony growth rate was determined at 4 and 7 days by measuring the colony diameter (ø) on PDA with a digital vernier caliper and weighing the dry mycelium of each isolate grown on potato dextrose broth (PDB).
Rapid DNA extraction
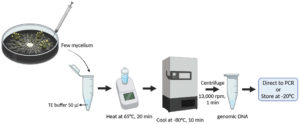
The genomic DNA extraction was done based on the thermolysis method.22 Both isolates were cultured on PDA plates for 5-7 days and incubated at room temperature. The mycelial mass was harvested using a flat-tipped needle and transferred into 50 µl of Tris-EDTA (TE) buffer, pH 8 in a 1.5 ml microcentrifuge tube and incubated at 65°C in a water bath or heat block for 20 min. Subsequently, they were incubated at -80°C for 10 min and finally, centrifuged at 13,000 rpm for 1 min. The supernatant containing genomic DNA was directed to polymerase chain reaction (PCR) amplification or stored at -20°C until use (Figure 1).
Figure 1. Schematic representation of rapid genomic DNA extraction based on thermolysis. Created in BioRender.com
PCR amplification and nucleotide sequencing
PCR was amplified based on ITS and LSU of the nuclear rDNA region using a primer pair of ITS5/ITS423 and LROR/LR5,24,25 respectively. PCR used a 25 µl reaction containing 14.3 μl nuclease-free water, 2.5 μl 10× standard Taq reaction buffer, 4 μl dNTP (1.25 mM each), 1μl of each primer
(5 μM), 0.2 μl Taq DNA polymerase (1 U/μl, New England Biolabs, USA), and 2 μl DNA template. The standard PCR program was run as in our previous study.26 PCR products were checked on 1% agarose electrophoresis gels in 0.5× tris borate EDTA (TBE) buffer for 30 min at 100 volts. Further, agarose gel was stained with ethidium bromide. PCR products were sequenced following the Barcode Taq sequencing (BTSeq) technique based on Next-Generation sequencing (NGS), Illumina Hiseq by Celemics, Inc. Korea. The consensus sequences obtained were edited and aligned using BioEdit version 7.2. All sequences of ITS and LSU were submitted to GenBank, The National Center for Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov/) (Table 1). The sequences were compared to find similarities in the GenBank database using the Nucleotide Basic Local Alignment Search Tool (Nucleotide BLAST or blastn) program. Finally, sequence datasets were done by Phylogeny.fr (http://www.phylogeny.fr/index.cgi), following the workflow of advance mode as MUSCLE multiple alignments, Gblocks alignment curation and maximum likelihood (ML) method with 100 bootstrapping procedures to construct the phylogenetic tree.
Table (1):
GenBank accession numbers of Cookeina in this study.
| Species name | Isolate | GenBank accession number | |
|---|---|---|---|
| ITS | LSU | ||
| C. sulcipes | NTKMITL20-049 | OM574929 | OM540427 |
| C. tricholoma | NTKMITL20-047 | OM574928 | OM540426 |
Antibacterial activity by agar plug diffusion method
The agar plug diffusion method for preliminary screening of antibacterial activity.27 That both Cookeina sp. isolates were screened for six human pathogenic bacteria (Gram-positive: Bacillus subtilis TISTR 1248, Kocuria rhizophila (Micrococcus luteus) TISTR 2374, Staphylococcus aureus TISTR 746 and S. epidermidis TISTR 2141, Gram-negative: Escherichia coli TISTR 074 and Pseudomonas aeruginosa TISTR 2370) obtained from the Thailand Institute of Scientific and Technological Research (TISTR) culture collection, Bangkok, Thailand. The bacteria were activated in nutrient broth (NB) for 24 hours at 37°C. Then, the bacterial solutions were diluted with 0.85% normal saline to an optimum concentration (0.08-0.13) monitoring absorption at 625 nm as a starter.28 A sterile cotton swab was dipped into the bacteria solution and swabbed on the PDA Petri dish. The fungal isolate was cultured for 5 days on a PDA Petri dish at room temperature and a 5 mm diameter fungal agar plug was cut and transferred to the surface of PDA Petri dish previously smeared with bacteria. WA media and sterile fungal plug on PDA Petri dish were used as negative and positive controls. All antibacterial activity analyses were run in duplicates of four plugs in two Petri dishes. After incubating the co-culture plate at room temperature for 24-72 h, the antimicrobial activity was estimated by measuring inhibition zone or clear zone diameters.
Enzyme activities by agar plate method
The fungal isolates were subcultured on a PDA plate and incubated at room temperature for 5 days. Then 5 mm diameter plugs were transferred to agar plates for preliminary screening of enzyme activity in plant biomass degradation.
Cellulolytic activity
For cellulolytic activity, a modified method was used as follows: Each fungal plug was placed on the plates of carboxymethylcellulose agar (CMCA) (0.2% NaNO3, 0.1% K2HPO4, 0.05% MgSO4•7H2O, 0.05% KCl, 0.2% peptone, 0.2% CMC, and 1.7% agar) in triplicate and the Petri dishes were incubated at room temperature for 4 days. Following incubation, CMCA plates were flooded with 5 ml of Gram’s iodine (1% KI and 0.5% I2) stain for 5 min.29 Then, the agar surface was washed with distilled water. The cellulolytic activity of the fungi was analyzed by measuring a clear zone of hydrolysis around colonies. The hydrolytic capacity (HC) was determined by the diameter of the clear zone/diameter of the fungal colony (mm).30
Hemicellulolytic activity
Hemicellulolytic activity used a modified method similar to cellulolytic activity: a fungal plug was transferred onto xylan agar (0.2% NaNO3, 0.1% K2HPO4, 0.05% MgSO4•7H2O, 0.05% KCl, 0.2% peptone, 0.2% xylan and 1.7% agar) plates in Petri dishes, in triplicate, and incubated at room temperature for 4 days. The Petri dish was flooded with 5 ml Gram’s iodine, stained for 5 min and surface-washed with distilled water.29 The hemicellulolytic activity of the fungi was observed as a clear zone of hydrolysis and analyzed as in the cellulolytic activity method.
Pectinolytic activity
For pectinolytic activity, a modified method was as follows: The fungal plug was transferred onto a pectin agar medium (0.5% pectin, 0.1% yeast extract, and 1.5% agar, pH 5) in triplicate and the Petri dishes were incubated at room temperature for 4 days. Following, the Petri dish was flooded with 5 ml of 1% Cetyltrimethylammonium bromide (CTAB), stained for 10 min and surface-washed with distilled water.31 The pectinolytic activity of the fungi was observed as a clear zone of hydrolysis and analyzed as in the cellulolytic activity method.
Ligninolytic activity
Laccase (Lac)
The laccase activity was measured by a modified method following Shrestha et al.32: In triplicate, fungal plug of each isolate was placed on a PDA plate supplemented with 1-naphthol (0.005%, Merck), in a Petri dish and incubated at room temperature, in the dark, for 4 days. After incubation, the appearance of a purple colour zone around the colony indicated laccase enzyme production.
Lignin peroxidases (LiP)
For lignin peroxidase activity, the method was modified from Delila et al.33: In triplicate, the fungal plug was transferred to a PDA plate, supplemented with methylene blue (0.0065%, Sigma-Aldrich), and incubated at room temperature, in the dark, for 14 days. A clear zone around the colony, caused by the disappearance of the blue colour of the dye, confirmed the presence of peroxidase or decolorization of methylene blue dyes.
Manganese peroxidase (MnP)
For manganese peroxidase activity, the modified method was as follows: In triplicate, an inoculated fungal plug on a PDA plate was supplemented with phenol red indicator (0.008%, Merck),34 and incubated at room temperature, in the dark, for 18 days. A colour change from yellow to red confirmed the presence of the peroxidase.
Statistical analysis
The triplicates for growth rate and HC in diameter values were analyzed using SPSS, version 28. Means ± standard error (SE) were reported. Statistical analysis used one-way ANOVA, followed by Duncan’s test, p < 0.01 was considered significant.
Molecular identification and phylogenetic analyses
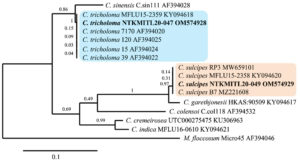
The result from PCR product and DNA sequences on both gene regions (ITS and LSU) shows that the genomic DNA from rapid extraction, which has not been tested for DNA purity or concentration and agarose gel electrophoresis was efficiently used directly for PCR amplification. DNA sequence length (without primers) of C. sulcipes NTKMITL20-049 and C. tricholoma NTKMITL20-047 on ITS was 616 bp and 570 bp and LSU were 912 bp and 906 bp, respectively. Further, the ITS phylogenetic tree was analyzed and constructed to support identification compared to morphology characteristics. The data analysis on ITS of Cookeina sp. with 615 characters that included alignment gap were parsimony informative with ingroup and one outgroup (Microstoma floccosum). Phylogeny presents C. sulcipes NTKMITL20-049 in the same clade as C. sulcipes, whereas C. tricholoma NTKMITL20-047 in C. tricholoma clade (Figure 2).
Figure 2. Phylogeny of Cookeina sp. base on internal transcribed spacer (ITS). C sulcipes and C. trocholoma with reference taxa and outgroup (Microstoma floccosum) from the maximum likelihood (ML)
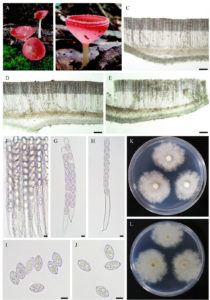
Figure 3. Cookeina sulcipes. A-B apothecia on dead wood, C-E cross section of apothecium, F-H asci with ascospore, I-J ascospores. Scale bars: C-E = 100 µm, F-J = 10 µm
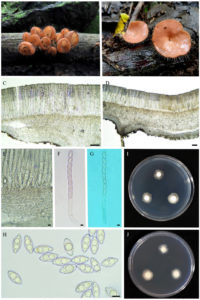
Figure 4. Cookeina tricholoma. A-B apothecia on dead wood, C-D cross section of apothecium, E base of asci, F-G asci with ascospores, H ascospore. Scale bars: C-D = 100 µm, E-H = 10 µm
Taxonomy
Cookeina sulcipes (Berk.) Kuntze, Revisio generum plantarum 2: 849 (1891) (Figure 3)
Index Fungorum number: IF 123427; MycoBank number: MB 123427
Basionym: Peziza sulcipes Berk., London Journal of Botany 1 (3): 141. 1842. MycoBank MB 230885
Type: Peziza sulcipes Berk., London Journal of Botany 1(3): 141. 1842. (Holotype K(M) 35303)
≡ Pilocratera sulcipes (Berk.) Sacc. & Traverso, Sylloge Fungorum 20: 413. 1911. MycoBank MB 216740]
≡ Trichoscypha sulcipes (Berk.) Cooke. MycoBank MB 188721
≡ Cookeina notarisiana (Bagnis) Kuntze: 849. 1891. MycoBank MB 580254
:– Habitat saprobic on dead wood. Sexual stage: Ascomata an apothecium with short hairs emerging from the exterior and rim region, pink to orange when fresh, Asci cylindrical, long pedicellate, 8-spored, Ascospores oval, 21–30 × 13–16 μm (x ̅ = 25 × 14 μm, n = 40), smooth and transparent. Colonies on PDA are white or cream above and reverse, flat elevation, and irregular form with lobate margin. At room temperature, they reached a diameter of 43 mm after 4 days and 90 mm after 7 days and, on PBD medium the average dry mycelial mass was 0.04 mg after 4 days and 0.06 mg after 7 days.
Material examined – THAILAND. Saraburi, 4 October 2020, N. Tangthirasunun; living culture = NTKMITL20-049.
Cookeina tricholoma (Mont.) Kuntze, Revisio generum plantarum 2: 849 (1891) (Figure 4)
Index Fungorum number: IF 121551; MycoBank number:MB 121551
Basionym: Peziza tricholoma Mont., Annales des Sciences Naturelles Botanique 2: 77. 1834. MycoBank MB 169162
Type: not indicated
Lectotype Cookeina tricholoma (Mont.) Kuntze. 1891. (Designated by Korf, Mycologia 64(5): 967. 1972). Registration Identifier 597346
≡ Pilocratera tricholoma (Mont.) Henn. MycoBank MB 224863
≡ Trichoscypha tricholoma (Mont.) Cooke, Syll. fung.: 160. 1889. MycoBank MB 188903
≡ Lachnea tricholoma (Mont.) Pat. & Gaillard, Bulletin de la Societe Mycologique de France 4 (3): 98. 1889. MycoBank MB 812614
:– Habitat saprobic on dead wood. Sexual stage: Ascomataan apothecium with long hairs, orange when fresh, Asci cylindrical, long,8-spored, pedicellate, Ascospores, oval, smooth, hyaline, 26–31 × 12–15 μm (x ̅ = 29 × 13 μm, n = 40). Colonies on PDA medium are white or cream above and reverse of Petri dish, flat elevation, and filamentous form and margin. After 4 days at room temperature, on PDA plates, the average diameter was 28 mm and, after 7 days, 45 mm and, on PBD medium, the average of dry mycelial mass was 0.04 mg after 4 days and 0.04 mg after 7 days.
Material examined – THAILAND. Saraburi, 4 October 2020, N. Tangthirasunun; living culture = NTKMITL20-047.
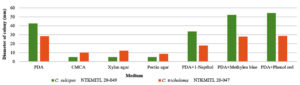
Finally, the growth rate (colony diameter) comparison of C. sulcipes NTKMITL20-049 and C. tricholoma NTKMITL20-047 at room temperature for 4 days on seven agar mediums (PDA, CMCA, xylan agar, pectin agar, and PDA with 1-naphthol or methylene blue or phenol red). The result was presented C. sulcipes NTKMITL20-049 was growth faster than C. tricholoma NTKMITL20-047 on four mediums (PDA, PDA with 1-naphthol, PDA with methylene blue, and PDA with phenol red). Whereas C. tricholoma faster NTKMITL20-047 was growth faster than C. sulcipes NTKMITL20-049 on three mediums (CMCA, xylan agar, and pectin agar) (Table 2 and Figures 5 and 7).
Table (2):
Growth of Cookeina sp. on agar medium after 4 days.
| Species | Medium | Colony diameter (ø) (mm) | |||
|---|---|---|---|---|---|
| R1 | R2 | R3 | Mean | ||
| C. sulcipes NTKMITL20-049 | PDA | 43 | 45 | 40 | 43 ± 1.42 |
| CMCA | 5 | 5 | 5 | 5 ± 0.00 | |
| Xylan agar | 5 | 5 | 5 | 5 ± 0.00 | |
| Pectin agar | 5 | 5 | 5 | 5 ± 0.00 | |
| PDA+1-Napthol | 33 | 35 | 32 | 34 ± 0.94 | |
| PDA+Methylene blue | 53 | 52 | 51 | 52 ± 0.55 | |
| PDA+Phenol red | 54 | 53 | 54 | 54 ± 0.32 | |
| C. tricholoma NTKMITL20-047 | PDA | 28 | 28 | 30 | 28 ± 0.66 |
| CMCA | 10 | 10 | 10 | 10 ± 0.17 | |
| Xylan agar | 12 | 12 | 12 | 12 ± 0.10 | |
| Pectin agar | 9 | 9 | 9 | 9 ± 0.10 | |
| PDA+1-naphthol | 17 | 18 | 18 | 18 ±0.27 | |
| PDA+Methylene blue | 28 | 27 | 28 | 28 ± 0.29 | |
| PDA+Phenol red | 29 | 28 | 29 | 29 ± 0.28 | |
*R = Replicate, Mean ± SE (p < 0.01)
Figure 5. Growth rate chart at 4 days on PDA, CMCA, xylan agar, pectin agar, and PDA with 1-naphthol or methylene blue or phenol red
Antibacterial activity
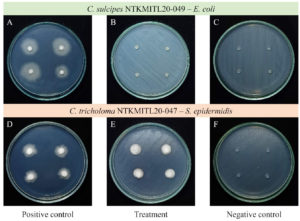
The preliminary result of antibacterial activity shows that both isolates of Cookeina present no activity. There are no clear zones seen with six bacterial strains (B. subtilis, E. coli, K. rhizophila, P. aeruginosa, S. aureus, and S. epidermidis) that represented as Gram-negative and Gram-positive bacteria in this study (Figure 6).
Figure 6. Antibacterial activity of Cookeina sp. on PDA Petri dish. A C. sulcipes, B C. sulcipes incubate with Escherichia coli, C agar plug incubates with E. coli, D C. tricholoma, E C. tricholoma, incubate with Staphylococcus epidermis, and F agar plug incubates with S. epidermis
Enzyme activities
The result of screening for enzyme activities on the agar plate method is presented as in Table 3 and Figure 7. Both isolates of Cookeina sp. showed similar activity on cellulolytic and hemicellulolytic enzymes by the sight of the clear zone. On manganese peroxidase, C. sulcipes was stronger than C. tricholoma by display of red and orange colour, respectively. However, the fungi exhibited no activity on pectinolytic (no clear zone) and lignin peroxidases (no decolorization). There was no decoloration of media from blue to purple colour. For laccase activity, C. sulcipes showed purple colouration on the reverse of Petri dish.
Table (3):
Enzyme activity of Cookeina species.
| Enzyme activity | C. sulcipes NTKMITL20-049 | C. tricholoma NTKMITL20-047 | ||||||
|---|---|---|---|---|---|---|---|---|
| R1 | R2 | R2 | Mean | R1 | R2 | R2 | Mean | |
| Hydrolytic capacity: HC (mm) | ||||||||
| Cellulolytic | 4.77 | 4.84 | 4.97 | 4.86 ± 0.06 | 1.88 | 1.98 | 1.90 | 1.92 ± 0.03 |
| Hemicellulolytic | 5.30 | 5.29 | 5.31 | 5.30 ± 0.01 | 1.69 | 1.58 | 1.58 | 1.62 ± 0.04 |
| Pectinolytic | ND | ND | ||||||
| Laccase | Yes | ND | ||||||
| Lignin peroxidases | ND | ND | ||||||
| Manganese peroxidase | ++ (Red) | + (Orange) | ||||||
*Mean ± SE (p < 0.01), ND = not detected
C. sulcipes and C. tricholoma are commonly found in the Central, Northeast, Northern and Southern parts of Thailand, besides other tropical and subtropical countries. C. sulcipes has earlier been described from a material collected from Prachuap Khiri Khan province of Thailand.1 C. tricholoma has been recorded in Brazil, China (Yunan province), Puerto Rico and Venezuela, besides Thailand.1,2,4 The rapid genomic DNA extraction method used in this study is unique in that it used only a very little quantity of mycelia and without liquid nitrogen13 or homogenizer (e.g., sonication,9,10 FastPrep-24™,11 pestle motor mixer or micro pestle,12 etc.) to breaking fungal cell wall. This method is faster, simple and does not demand many materials or equipment. Compared with standard fungal DNA extractions such as CTAB method, SDS method, lysis buffer method, etc.7,8,13,22 or DNA extraction/isolation kit, the method followed in this study is simple, cost saving, and quicker. Both species of Cookeina have been reported to be edible.14 On screening against six bacterial strains, the Cookeina species studied showed no activity. Genus Cookeina is generally found on dead wood. This study confirmed the presence of hydrolytic hemi- and cellulolytic enzymes and laccase and manganese peroxidase that involve in the degradation of plant cell wall.
The most widely studied cellulolytic fungi are Aspergillus sp., Humicola sp., Penicillium sp., and Trichoderma sp. of these, Trichoderma sp. is the most suitable candidate for cellulase production and utilization in the industry.35 Similar is with pectinase production in agro-industrial waste inhabitant fungi such as Aspergillus spp. and Lentinus edodes. Whereas P. chrysogenum, T. harzianum and Schizophyllum commune are important pectinase producing fungi.36 Laccase and manganese peroxidase useful in biodegradation are present in Irpex lacteus, Marasmius sp., Mucor circinelloides, and Pleurotus ostreatus. Similarly, decolorization of wood is carried out by Coprinopsis cinerea, Geotrichum candidum, Phanerochaete chrysosporium, and Trametes species.37 Reports of fungi from the same family Sarcoscyphaceae include Phillipsia sp. SXS629 (on the dead tree trunk, Brazil) in polyphenol oxidase and carboxymethyl cellulase (CMCase) activity. Some of these fungi can grow with increase of biosolids or wastewater treatment plant (WWTP) concentration.38
The rapid DNA extraction is appropriate for filamentous fungi for species identification by PCR, based on ITS, LSU and other regions. This method is simple, convenient, inexpensive, and saves time. In this study, C. sulcipes and C. tricholoma are no bacterial activity. Nevertheless, the Cookeina sp. also a source of lignocellulolytic enzyme, deserves further attention in the future.
ACKNOWLEDGMENTS
The authors would like to thank Department of Biology, King Mongkut’s Institute of Technology Ladkrabang for their support. Authors would also like to thank Asst. Prof. Thanarak Chantaraprasit for their fieldwork, Matchima Siddhidesananda, Thi Thu Huong Luong and Chanaphon Jadtanim for helping us with lab work and Prof. Jayarama Darbhe Bhat for suggestions.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
NT conceptualized the study, performed visualization, software work, formal analysis, investigation. NT and SP applied methodology, performed validation, collected resources and performed data curation. NT administrated the project and funding acquisition. NT wrote the manuscript. SP performed supervision. NT and SP reviewed and edited the manuscript. Both the authors read and approved the final manuscript for publication.
FUNDING
This work was supported by King Mongkut’s Institute of Technology Ladkrabang under Grant KREF186308.
DATA AVAILABILITY
Not applicable.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
- Ekanayaka AH, Hyde KD, Zhao Q. The genus Cookeina. Mycosphere. 2016;7(9):1399-1413.
Crossref - Iturriaga T, Pfister DH. A monograph of the genus Cookeina (Ascomycota, Pezizales, Sarcoscyphaceae). Mycotaxon. 2006;95:137-180.
- Raymundo T, Montes-Fuentes G, Valenzuela R. Cookeina colombiana (Sarcoscyphaceae, Ascomycota) a new species from the department Cordoba, Colombia. Acta Bot Mex. 2021;128:e1763.
Crossref - Weinstein RN, Pfister DH, Iturriaga T. A phylogenetic study of the genus Cookeina. Mycologia. 2002;94(4):673-682.
Crossref - Kropp BR. Cookeina cremeirosea, a new species of cup fungus from the South Pacific. Mycoscience. 2017;58(1):40-44.
Crossref - Doyle JJ, Doyle JL. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin. 1987;19(1):11-15.
- Tripathy SK, Maharana M, Ithape DM, et al. Exploring rapid and efficient protocol for isolation of fungal DNA. Int J Curr Microbiol Appl Sci. 2017;6(3):951-960.
Crossref - Tan SC, Yiap BC. DNA, RNA, and protein extraction: The past and the present. J Biomed Biotechnol. 2009;2009:574398.
Crossref - Rana S, Kumar K, Singh M, Lal R. Optimization of protocol for DNA isolation from filamentous fungi and its identification. Int. J Basic Appl Sci. 2018;8(5):225-231. https://www.researchgate.net/publication/331318706_Optimization_of_protocol_for_
DNA_isolation_from_filamentous_fungi_and_its_identification. Accessed June 23, 2022. - Pilo P, Tiley AMM, Lawless C, Karki SJ, Burke J, Feechan A. A rapid fungal DNA extraction method suitable for PCR screening fungal mutants, infected plant tissue and spore trap samples. Physiol Mol Plant Pathol. 2022;117:101758.
Crossref - Aamir S, Sutar S, Singh SK, Baghela A. A rapid and efficient method of fungal genomic DNA extraction, suitable for PCR based molecular methods. Plant Pathol & Quar. 2015;5(2):74-81.
Crossref - Chi MH, Park SY, Lee YH. A quick and safe method for fungal DNA extraction. Plant Pathol J. 2009;25(1):108-111.
Crossref - Wingfield LK, Atcharawiriyakul J. Evaluation of simple and rapid DNA extraction methods for molecular identification of fungi using the internal transcribed spacer regions. Asia Pac J Sci Technol. 2021;26(2):APST260201.
Crossref - Boa E. Wild Edible Fungi: A Global Overview of Their Use and Importance to People. Food and Agriculture Organization (FAO); 2004. https://www.fao.org/publications/card/en/c/709ef43f-313d-5098-9178-d974989bfe72/. Accessed April 6, 2022.
- Hernandez-Beltran JU, Lira IOHD, Cruz-Santos MM, Saucedo-Luevanos A, Hernandez-Teran F, Balagurusamy N. Insight into pretreatment methods of lignocellulosic biomass to increase biogas yield: Current state, challenges, and opportunities. Appl Sci. 2019;9:3721.
Crossref - Suryadi H, Judono JJ, Putri MR, et al. Biodelignification of lignocellulose using ligninolytic enzymes from white-rot fungi. Heliyon. 2022;8(2):e08865.
Crossref - Asmoro NW, Afriyanti A, Ismawati I. Ekstraksi selulosa batang tanaman jagung (Zea mays) metode basa. Jurnal Ilmiah Teknosains. 2018;4(1):24-28.
Crossref - Saini S, Sharma KK. Fungal lignocellulolytic enzymes and lignocellulose: A critical review on their contribution to multiproduct biorefinery and global biofuel research. Int J Biol Macromol. 2021;193(Part B):2304-2319.
Crossref - Sheng Y, Lam SS, Wu Y, et al. Enzymatic conversion of pretreated lignocellulosic biomass: A review on influence of structural changes of lignin. Bioresour Technol. 2021;324:124631.
Crossref - Patidar MK, Nighojkar S, Kumar A, Nighojkar A. Pectinolytic enzymes-solid state fermentation, assay methods and applications in fruit juice industries: A review. 3 Biotech. 2018;8(4):199.
Crossref - Senanayake IC, Rathnayaka AR, Marasinghe DS, et al. Morphological approaches in studying fungi: Collection, examination, isolation, sporulation and preservation. Mycosphere. 2020;11(1):2678-2754.
Crossref - Zhang YJ, Zhang S, Liu XZ, Wen HA, Wang M. A simple method of genomic DNA extraction suitable for analysis of bulk fungal strains. Lett Appl Microbiol. 2010;51(1):114-118.
Crossref - White TJ, Bruns T, Lee S, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, eds. PCR Protocols: A Guide to Methods and Applications. Academic Press Limited. 1990:315-322.
Crossref - Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol. 1990;172(8):4238-4246.
Crossref - Rehner SA, Samuels GJ. Taxonomy and phylogeny of Gliocladium analysed from nuclear large subunit ribosomal DNA sequences. Mycol Res. 1994;98(6):625-634.
Crossref - Tangthirasunun N, Silar P, Bhat DJ, et al. Morphology and phylogeny of Pseudorobillarda eucalypti sp. nov., from Thailand. Phytotaxa. 2014;176:251-259.
Crossref - Balouiri M, Sadiki M, Ibnsouda SK. Methods for in vitro evaluating antimicrobial activity: A review. J Pharm Anal. 2016;6(2):71-79.
Crossref - Clinical and Laboratory Standards Institute. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Approved Standard-Ninth Edition. CLSI document M07-A9. Clinical and Laboratory Standards Institute, Wayne, Pennsylvania. 2012. https://www.researchgate.net/file.PostFileLoader.html?id=564ceedf
- Neethu K, Rubeena M, Sajith S, et al. A novel strain of Trichoderma viride shows complete lignocellulolytic activities. Adv Biosci Biotechnol. 2012;03(08):1160-1166.
Crossref - Reanprayoon P, Pathomsiriwong W. Tropical soil fungi producing cellulase and related enzymes in biodegradation. J Appl Sci. 2012;12(18):1909-1916.
Crossref - Sunitha VH, Devi DN, Srinivas C. Extracellular enzymatic activity of endophytic fungal strains isolated from medicinal plants. World J Agric Sci. 2013;9(1):01-09.
Crossref - Shrestha P, Joshi B, Joshi J, Malla R, Sreerama L. Isolation and physicochemical characterization of laccase from Ganoderma lucidum-CDBT1 isolated from its native habitat in Nepal. Biomed Res Int. 2016;2016:3238909.
Crossref - Delila L, Susanti E, Sanjaya EH. Isolation and screening of indigenous fungus producing lignin peroxidase from the cocoa plantation in Sepawon Kediri regency Indonesia. KnE Life Sci. 2017;3(5):131.
Crossref - Sánchez-Corzo LD, Álvarez-Gutiérrez PE, Meza-Gordillo R, Villalobos-Maldonado JJ, Enciso-Pinto S, Enciso-Sáenz S. Lignocellulolytic enzyme production from wood rot fungi collected in Chiapas, Mexico, and their growth on lignocellulosic material. J Fungi. 2021;7:450.
Crossref - Imran M, Anwar Z, Irshad M, Asad MJ, Ashfaq H. Cellulase production from species of fungi and bacteria from agricultural wastes and its utilization in industry: A review. Adv Enzyme Res. 2016;4:44-55.
Crossref - Haile S, Ayele A. Pectinase from microorganisms and its industrial applications. Sci World J. 2022;2022:1881305.
Crossref - El-Gendi H, Saleh AK, Badierah R, Redwan EM, El-Maradny YA, El-Fakharany EM. A comprehensive insight into fungal enzymes: Structure, classification, and their role in mankind’s challenges. J Fungi. 2022;8:23.
Crossref - Naves LR, da Silva LL, da Cunha EL, et al. Filamentous fungi as promising agents for the biodegradation of biosolids compounds. Front J Soc Technol Environ Sci. 2019;8(2):35-51.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.