Majority of the global population have been affected by food-borne diseases, and Staphylococcus aureus is one of the causes of this disease. S. aureus can be transmitted through contaminated food and is a risk to universal human health because of its ability to produce toxins-staphylococcal enterotoxins. Additionally, methicillin resistant Staphylococcus aureus (MRSA) furthered public health concerns. Although MRSA has been identified in food worldwide, little information is available on this topic locally and internationally. This review presents information on MRSA that was collected as evidence of such infections globally and in Saudi Arabia.
Saudi Arabia, Food-born disease, MRSA, Antibiotics, PVL, SCCmec, MecA
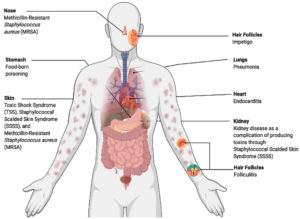
Staphylococcal species are opportunistic pathogens that can cause various types of infections with varying levels of severity. S. saprophyticus is the second main cause of urinary tract infections, and S. simulans, S. warneri, and S. haemolyticus can cause device-related infections. Additionally, S. aureus is a serious human pathogen that can cause lethal diseases such as pneumonia, endocarditis, toxic shock syndrome, and sepsis (Figure 1). Moreover, S. aureus and S. epidermidis can cause nosocomial infections in indwelling devices.1 S. aureus can cause alternative diseases, particularly methicillin-resistant Staphylococcus aureus (MRSA), which forms approximately 13–74% of S. aureus infections.
MRSA infection and colonization can be negatively affected by factors such as obesity; however, the mechanism has not yet been revealed.2 According to origin of the outbreaks, MRSA is divided into two groups: healthcare-associated MRSA (HA-MRSA) and community-associated MRSA (CA-MRSA).3 CA-MRSA clones differ from country-to-country; e.g., ST8 (USA300) and ST1 (USA400) clones were found essentially in the USA and Canada, whereas ST80 clones were found in Europe.4 Although the MRSA USA300 clone was mainly found in the USA and Canada, it was also infrequently detected in wild pig meat in Germany, Switzerland, and Spain.5 The probability of MRSA infection in patients colonized with CA-MRSA (13%) are lower than those with HA-MRSA (37%). Hence, patients colonized with HA-MRSA exhibit longer recovery time than those colonized with CA-MRSA.6 Additionally, the major cause of this infectious disease is that S. aureus is resistant to several types of antibiotics, and the susceptibility of HA-MRSA to antibiotics is lower than that of CA-MERSA.7,8
Livestock is considered a major source of staphylococci species, particularly S. aureus, causing livestock-associated MRSA (LA-MRSA). Thus, LA-MRSA can occur by direct contact with animals or by use of animal products obtained from animals such as lambs, calves, and goats. However, the spread rate of S. aureus was lower in calves (12.5%) and goats (1 %) than in lambs (30%). LA-MRSA was first discovered in France and the Netherlands and was found in pig populations in several countries of Europe. Furthermore, CA-MRSA, HA-MRSA and LA-MRSA clones vary between countries; e.g., the CC398 clone was detected in Europe and the USA, whereas the CC9 clone was found in Asia.5, 9
Pathogenesis of Staphylococcal Disease and Virulence Factors
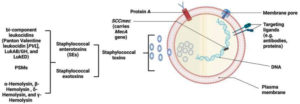
Genetically, one of the many types of staphylococcal cassette chromosome mec complex gene (SCCmec) would be sufficient to induce an MRSA infection. Elements of SCCmec carry mecA gene, which encodes penicillin-binding protein 2a (PBP2a) that is responsible for bacterial resistance to methicillin, as shown in Figure 2. PBP2a has low similarity to beta-lactamases, which are responsible for resistance against extended-spectrum cephalosporins, monobactams, penicillins, and carbapenems.10,11,12
Furthermore, the Panton-Valentine Leukocidin (PVL) gene is a genetic marker of MRSA (first identified in 2003) (Figure 2). Strains that produce PVL may cause skin infection or serious diseases such as pneumonia. However, PVL gene was more prevalent in CA-MRSA than in HA-MRSA and was found in approximately 60%–100% of CA-MRSA strains. Consequently, PVL develops the pathogenicity of CA-MRSA strains, and the prevalence of PVL between strains relies on mobile genetic elements, particularly bacteriophages.8,13,14 S. aureus also causes Staphylococcus scalded skin syndrome by producing the epidermolytic toxins A and B, which affect the skin all over the body. It commonly affects children aged <5 years, in addition to adults with kidney disease.15
Some species of Staphylococci can cause a very prevalent disease called food-borne poisoning that occurs after consumption of contaminated food. It is mainly caused by S. aureus or occasionally by other Staphylococcus species, such as S. intermedius. Symptoms (vomiting, fever, nausea, diarrhea, dizziness, low blood pressure, and headache) present 30 min to 8 h post consumption of contaminated food.16 S. aureus live in various habitats and can contaminate foods such as sandwich fillings, salads, milk, sausages, canned meat, cheeses, and cooked meals.17 However, staphylococci can be killed by different methods for food preservation, such as cooking food with high heat and pasteurization.18 Bacteria can also contaminate food during food preparation and milk processing or storage. Consequently, Staphylococci produce enterotoxins, which include more than 20 types of toxins.16
Statement of the Problem
MRSA infections spread rapidly throughout Saudi Arabia. A group of clinicians reported that the percentage of such infections in King Fahad Medical City in Riyadh reached 50% during 2011, and there are no sufficient studies addressing this problem.19 Although MRSA in Saudi Arabia was discovered in the 1990s, the number of studies on MRSA infection in Saudi Arabia is limited compared to that in other countries around the world.20 According to PubMed, there were only 141 studies conducted on MRSA in Saudi Arabia over past 5 years (2015–2019), of which majority have been conducted on HA-MRSA. Nevertheless, until May 2014, there were only 12 articles on MRSA infections, of which ten were published in Riyadh and Dammam.10 Despite conducting investigations and discussions on HA-MRSA, there is still a shortage in the number of studies on HA-MRSA in children, and surprisingly, HA-MRSA is primarily found in men than in women.21 Based on the prevalence of CA-MRSA, the number of patients significantly increased from 9.9 per 10,000 in 2001 to 67 per 10,000 in 2008.21,22 Furthermore, between 2000 and 2008, the spread rate of CA-MRSA in the eastern region of Riyadh increased six-fold.19 Furthermore, CA-MRSA samples collected from Qatif city, Saudi Arabia, showed that CA-MRSA percentages increased gradually from 23% in 2006 to 60% in 2015.23 There is still a lack of studies that aimed to investigate LA-MRSA.24
Mrsa in the Capital City of Saudi Arabia
According to the Center of Disease Control, two in 100 individuals develop MRSA infections, and 33% of persons colonize with Staphylococcus.25 S. aureus strains (n=135) have been collected from main public health centers and hospital laboratories in Riyadh city between 2008 and 2009 to examine the presence of CA-MRSA infections. As a result, scientists have announced that the CA-MRSA infection rate was amplified in Riyadh city.22 Al Yousef et al. reported that 94% of MRSA strains have been detected in Riyadh.26 In addition, all 37 isolates of S. aureus, which were collected from King Khalid University Hospital in Riyadh (2007), were MRSA, and only two of them carried mecA gene. 27
Moreover, SCCmec type IV was tested in Riyadh to determine the type of strains; subsequently, CC22-MRSA-IV was found to be a common strain associated with strains collected from India and the Middle East rather than that from the Western European countries.28 Senok et al. obtained 117 samples of MRSA infections over a 6-year period (2009–2015) in Riyadh city and examined them to identify the most derived strains related to CA-MRSA; CC5, CC22, CC6, and CC80 (13, 12, 15, and 35 strains, respectively) were the most commonly identified strains. This study identified three new MRSA clones in Saudi Arabia ( CC15-MRSA).29
Alaklobi et al. estimated the growth of CA-MRSA levels among children in Riyadh city, which ranged from 0%–9%, and 164/824 samples in this study were colonized with S. aureus; however, only 38 samples were MRSA, and 23% of the MRSA infections were reported in children.30 In addition, another study examined 100 raw retail meat samples: 24 lamb (mutton), 24 camel, 23 beef, and 29 poultry (chicken parts) in Riyadh. Of the 100 meat samples, 25 were contaminated with S. aureus. While meat samples from poultry assets were recorded to have the lowest contamination among the tested samples, the meat sample from camel had the highest level of contamination.24 Moreover, 80 camel milk samples were collected from different places throughout Riyadh city, and strains of bacteria that carried the 16s RNA gene similar to many pathogens such as MRSA were found.31
MRSA in Different Cities of Saudi Arabia
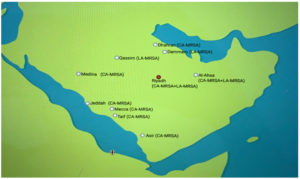
As shown in Figure 3, MRSA’s types have been spotted in variant cities of Saudi Arabia such as Dhahran, Dammam, Medina, Jeddah, Taif, Asir, etc. In 2019; moreover, Farah et al. reported that there was a high level of multidrug resistance among seven different locations and 12 hospitals in Saudi Arabia. During the same period, HA-MRSA cases declined significantly from 0.17/1000 to 0.03/1000.25 In Saudi Arabia, HA-MRSA was first discovered, followed by CA-MRSA and LA-MRSA. However, CA-MRSA has been reported to be less serious than HA-MRSA; hence, researchers in Saudi Arabia are more focused on HA-MRSA.32,33 In addition, investigators have tended to use PVL as an indicator of CA-MRSA presence, which has been recorded to be present at high levels in Riyadh since 2012.19,32
Additionally, a group of scientists in Riyadh and Taif focused on CA-MRSA and found that since SCCmec IV and V can easily be transmitted among organisms, they are predominant than SCCmec I, II, and III.22,34
Owing to the Hajj season, 31 studies were conducted in Mecca, Mina, and Medina, and the authors examined food workers, hospitalized patients, and pilgrims between 2000 and 2015. Subsequently, a sharp increase in MRSA infections in pilgrims was observed (from 1% in 2000 to 63% in 2015). The rate of MRSA isolated from food handlers has also increased remarkably (i.e., from 0% in 2001 to 20% in 2014).35Additionally, certain restaurants in Makkah city have tested and reported that the food samples were contaminated with S. aureus. 33 In addition, many food samples (e.g., raw milk and cheese) in Makkah City were found to have a significant incidence of multidrug-resistant S. aureus and coagulase-negative staphylococci.36
The western region of Saudi Arabia has been similarly investigated by El amin and Faidah, and the most isolated MRSA infections (87%) were from soft tissue, skin, and wounds. In addition, the prevalence of HA-MRSA was 53%, whereas that of CA-MRSA was 31.5%.37 Approximately 5% of MRSA infections were also reported in Madinah city by Ghanem et al. during the time of fourteen-month. Most of them were isolated from men (237 MRSA strains), including 37 recovered strains from nasal swabs, 120 strains from sputum, and 124 strains from wound swabs. Moreover, MRSA isolated from catheter tips and pus samples was higher in women than in men, and the rate of MRSA infections reached a peak in the summer season throughout the Hajj term (51%), which decreased during other seasons: 21% in autumn, 18.5% in winter, and 10% in spring.38
Apart from Mecca city, different cities in Saudi Arabia have been examined for MRSA infections; e.g., a study conducted in Qatif city from January 1, 2006, to December 31, 2015, highlighted that 25/100,000 patients were diagnosed with MRSA infection, and 20%–35% of all patients in the examined city were children.23 Moreover, MRSA was detected in 101 samples from public health centers and large hospital laboratories in Jeddah city between August 2009 and May 2011. MRSA strains carry mecA gene and are positive for PVL; therefore, the common type of SCCmec in that study was type V, followed by type III (42.5% and 39%, respectively).39 Four different strains of S. aureus (CC22, CC8, CC5, and CC80) were isolated from 206 nasal swabs (103 from Saudi Arabia and 103 from Egypt). The prevalence of MRSA in Saudi Arabia was lower than that in Egypt (12% and 15%, respectively). In this study, mecA gene was positive, and the most common types of SCCmec were IVa and V.40
A recent study in Jeddah city, conducted by Al-Zahrani et al. recorded the highest ratio of MRSA infections (CA-MRSA comprised majority of the infections). During Hajj, pilgrims travel from Jeddah to Mecca, which lead to a rise in MRSA cases (>61%), and these cases majorly include foreigners (from Pakistan, Sudan, Syria, Yamen, Somalia, Egypt, and Chad).41
In addition, a 7-year study from 2001 to 2008 was conducted in the eastern region of Saudi Arabia, which revealed that the rate of CA-MRSA infections increased from 9.9/10,000 in 2001 to 67/10,000 in 2008. This investigation was divided into two stages. The presence of CA-MRSA was 67 (20%) and 176 (59%) in the first (2001–2004) and second stages (2005–2008), respectively.42 Furthermore, Dhahran city in Saudi Arabia was among the places that attracted the attention of researchers for further investigation of MRSA. Khanfar et al. reported 878 cases of MRSA infections from the Saudi Aramco Dhahran Health Center between 2004 and 2009; 777 (88%) and 101(11.5%) cases were CA-MRSA and HA-MRSA infections, respectively.43 Approximately 30% of MRSA strains have been discovered in approximately 90 minced meat and vegetable sandwiches collected from fast-food canteens in Al-Ahsa city.44
Throughout the southern region of Saudi Arabia, from Asir Central Hospital and Abha General Hospital, 210/9831 infectious samples contained S. aureus. The samples were collected from three dissimilar sources (ten from the hospital environment, 100 from community-acquired infections, and 100 from hospital-acquired infections). Hence, the hospital environment is a serious threat to all hospital staff, visitors, and hospitalized patients in terms of MRSA dissemination.45
Several CA-MRSA + HA-MRSA and LA-MRSA infections have been reported in several cities of Saudi Arabia. For example, 20 isolates from 775 nasal swabs and 235 milk samples taken from a goat farm in eastern Saudi Arabia were MRSA. The maximum prevalence of MRSA in this study was 2%, and most of them were from mastitic milk (9%) and a few were from normal milk (0.6%).46 Furthermore, in Al-Hasa city, 20 out of 187 fresh raw camel meat samples consisted of S. aureus strains, and three of 20 isolates were reported as MRSA.47
In Qassim city, 90/400 animal-derived samples (from goat, sheep, cow, and camels) were identified as LA-MRSA (57%). The smallest level of LA-MRSA isolates was from cow and goat samples, whereas the highest level was from sheep and camel samples.48
Reasons of Mrsa Infection
Bhedi et al. stated that the presence of MRSA infections in poultry meat is attributed to many factors, such as dropping meat handling levels in selling shops, hygiene, and sales practices.49 Globally, food handlers are also considered a major cause of food contamination; e.g., in Dhaka city, street food is served with bare hands. Thus, approximately 50–70% of ready-to-eat food (RTE) handlers are transmitters of S. aureus. Moreover, 32% and 1% of food handlers of Zimbabwe and Hong Kong, respectively, are infected with S. aureus. Another study in China confirmed that 3–6% of Hong Kong butchers were affected by LA-MRSA ST9.50,51
This literature review revealed that the rate of MRSA prevalence has significantly increased worldwide, with the emergence of MRSA in food handlers, food samples, and animal-derived food. In Saudi Arabia, studies have emphasized that HA-MRSA is more serious, even though the susceptibility of HA-MRSA to antibiotics is lower than that of CA-MERSA. Lastly, it is worth mentioning that more studies on MRSA are required because of their impact on public health.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Otto M. Staphylococcus colonization of the skin and antimicrobial peptides. Expert review of dermatology 2010; 5(2):183-195.
Crossref - Narayanan N, Adams CD, Kubiak DW, et al. Evaluation of treatment options for methicillin-resistant Staphylococcus aureus infections in the obese patient. Infection and drug resistance 2019; 12, 877-891.
Crossref - Dweba CC, Zishiri OT, El-Zowalaty ME. Methicillin-resistant Staphylococcus aureus: livestock-associated, antimicrobial, and heavy metal resistance. Infection and Drug Resistance 2018; 11:2497-2509.
Crossref - Huang YC, Ho CF, Chen CJ, Su LH, Lin TY. Comparative molecular analysis of community-associated and healthcare-associated methicillin-resistant Staphylococcus aureus isolates from children in northern Taiwan. Clinical Microbiology and Infection 2008; 14(12), 1167-1172.
Crossref - Mama OM, Gómez-Sanz E, Ruiz-Ripa L, Gómez P, Torres C. Diversity of staphylococcal species in food producing animals in Spain, with detection of PVL-positive MRSA ST8 (USA300). Veterinary microbiology 2019; 233, 5-10.
Crossref - Reich PJ, Boyle MG, Hogan PG, et al. Emergence of community-associated methicillin-resistant Staphylococcus aureus strains in the neonatal intensive care unit: an infection prevention and patient safety challenge. Clinical microbiology and infection 2016;22(7), 645-e1.
Crossref - Osman K, Alvarez-Ordóñez A, Ruiz L, et al. Antimicrobial resistance and virulence characterization of Staphylococcus aureus and coagulase-negative staphylococci from imported beef meat. Annals of clinical microbiology and antimicrobials 2017; 16(1),35.
Crossref - Loewen K, Schreiber Y, Kirlew M, Bocking N, Kelly L. Community-associated methicillin-resistant Staphylococcus aureus infection: Literature review and clinical update. Canadian Family Physician 2017; 63(7):512-520.
- Butaye P, Argudín MA, Smith TC. Livestock-associated MRSA and its current evolution. Current Clinical Microbiology Reports 2016; 3(1), 19-31
- Al Yousef SA, Taha EM. Methicillin-Resistant Staphylococcus aureusin Saudi Arabia: Genotypes distribution review. Saudi Journal of Medicine & Medical Science 2016; 4(1), 2-8.
Crossref - Vindel A, Trincado P, Cuevas O, Ballesteros C, Bouza E, Cercenado E. Molecular epidemiology of community-associated methicillin-resistant Staphylococcus aureus in Spain: 2004–12. Journal of Antimicrobial Chemotherapy 2014; 69(11), 2913-2919.
Crossref - Drawz SM, Bonomo RA. Three decades of beta-lactamase inhibitors. Clin Microbiol Revo 2010; 123(1), 160-201.
Crossref - Motamedi H, Abadi SSR, Moosavian SM, Torabi M. The association of Panton-Valentine leukocidin and mecA genes in Methicillin-Resistant Staphylococcus aureusisolates from patients referred to Educational Hospitals in Ahvaz, Iran. Jundishapur Journal of Microbiology 2015;8(8).
Crossref - Kong EF, Johnson JK, Jabra-Rizk MA. Community-associated methicillin-resistant Staphylococcus aureus: an enemy amidst us. PLoS pathogens 2016; 12(10), e1005837.
Crossref - Mishra AK, Yadav P, Mishra A. A systemic review on staphylococcal scalded skin syndrome (SSSS): a rare and critical disease of neonates. The Open Microbiology Journal 2016; 10, 150-159.
Crossref - Hennekinne JA, De Buyser ML, Dragacci S. Staphylococcus aureus and its food poisoning toxins: characterization and outbreak investigation. FEMS Microbiology Reviews 2012; 36(4), 815-836.
Crossref - Paprella A, Serio A, Rossi C, Mazzarrino G, Francesco CCL. Food-borne transmission of staphylococci. In V. Savini (Ed.), Pet-To-Man Travelling Staphylococci: A world in progress 2018; pp. 77-87. London, United Kingdom: Elsevier Academic Press.
Crossref - Bergdoll MS, Wong AL. Staphylococcus detection. In B. Caballero, L. Trugo, & P. M. Finglas (Eds.), Encyclopedia of food sciences and nutrition 2003; 2nd ed., pp. 5551-5556. California, USA: Academic Press.
Crossref - Monecke S, Skakni L, Hasan R, et al. Characterisation of MRSA strains isolated from patients in a hospital in Riyadh, Kingdom of Saudi Arabia. BMC Microbiology. 2012; 12(1).
Crossref - Iyer AP, Baghallab I, Albaik M, Kumosani T. Nosocomial infections in Saudi Arabia caused by methicillin resistance Staphylococcus aureus (MRSA). Clinical Microbiology: Open Access 2014; 3(3), 146.
Crossref - Alrabiah K, Al Alola S, Al Banyan E, Al Shaalan M, Al Johani S. Characteristics and risk factors of hospital acquired–methicillin-resistant Staphylococcus aureus (HA-MRSA) infection of pediatric patients in a tertiary care hospital in Riyadh, Saudi Arabia. International Journal of Pediatrics and Adolescent Medicine 2016; 3(2), 71-77.
Crossref - Moussa IM, Hessan AM. Rapid detection of community acquired-methicillin resistance Staphylococcus aureus recovered from King Saudi Arabia. African Journal of Microbiology Research 2010; 4(24), 2804-2810.
- Al-Hamad AM, Alfaraj AA, Altowaileb J, et al. Incidence and antibiotic susceptibility of MRSA infections in a Saudi Arabian Hospital: a 10-year surveillance study. The Journal of Infection in Developing Countries. 2018; 12(6), 454-461.
Crossref - Raji MA, Garaween G, Ehricht R, Monecke S, Shibl AM, Senok A. Genetic characterization of Staphylococcus aureusisolated from retail meat in Riyadh, Saudi Arabia. Frontiersin Microbiology 2016; 7(911).
Crossref - Farah SM, Alshehri MA, Alfawaz TS, Alasmeri FA, Alageel AA, Alshahrani DA. Trends in antimicrobial susceptibility patterns in King Fahad Medical City, Riyadh, Saudi Arabia. Saudi Medical Journal. 2019; 40(3), 252-259.
Crossref - Yousef SA, Mahmoud SY, Eihab MT. Prevalence of methicillin-resistant Staphylococcus aureus in Saudi Arabia: systemic review and meta-analysis.African Journal of Clinical 2013; 14(3): 146-154.
Crossref - Moussa I, Shibl AM. Molecular characterization of methicillin-resistant Staphylococcus aureus recovered from outpatient clinics in Riyadh, Saudi Arabia. Saudi Med J 2009;30(5), 611-617. https://pubmed.ncbi.nlm.nih.gov/19417957/
- Senok A, Somily A, Raji A, et al. Diversity of methicillin-resistant Staphylococcus aureus CC22-MRSA-IV from Saudi Arabia and the Gulf region. International Journal of Infectious Diseases 2016;51, 31-35.
Crossref - Senok A, Ehricht R, Monecke S, Al-Saedan R, Somily A. Molecular characterization of methicillin-resistant Staphylococcus aureus in nosocomial infections in a tertiary-care facility: emergence of new clonal complexes in Saudi Arabia. New microbes and new infections 2016; 14, 13-18.
Crossref - Alaklobi F, Aljobair F, Alrashod A, et al. The prevalence of community-associated methicillin-resistant Staphylococcus aureus among outpatient children in a tertiary hospital: A prospective observational study in Riyadh, Saudi Arabia. International Journal of Pediatrics and Adolescent Medicin 2015.; 2 (3-4), 136-140.
Crossref - Hirad AH, Ahmad J, Alkhedhairy AA, Bahkali AH, Khan ST. Bacterial isolates exhibiting multidrug resistance, hemolytic activity, and high 16S rRNA gene similarity with well-known pathogens found in camel milk samples of Riyadh region. Apmis 2018;126(3), 215-226.
Crossref - Deurenberg RH, Stobberingh EE. The evolution of Staphylococcus aureus. Infection, genetics and evolution 2008;8(6), 747-763.
Crossref - Todd ECD. Foodborne disease and food control in the Gulf States. Food control. 2017; 73(part B), 341-366.
Crossref - Eed EM, Ghonaim MM, Hussein YM, Al-Shehri SS, Khalifa AS. Molecular characterisation of Panton–Valentine leucocidin-producing methicillin-resistant Staphylococcus aureus clones isolated from the main hospitals in Taif, KSA. Indian Journal of Medical Microbiology 2016; 34(4), 476-482.
Crossref - Center for Infectious Disease Research and Policy. Antibiotic resistance at the Hajj; MERS in Riyadh; Plague cases; Cat-linked tularemia case; HPV vaccine impact; H5N8 in Belgium 2017. retrieved from
http://www.cidrap.umn.edu/news-perspective/2017/06/news-scan-jun-27-2017 - Abulreesh HH, Organji SR. The Prevalence of Multidrug-resistant Staphylococci in Food and the Environment of Makkah, Saudi Arabia. Research Journal of Microbiology 2011; 6(6), 510-523.
- El Amin NM, Faidah HS. Methicillin-resistant Staphylococcus aureus in the western region of Saudi Arabia: prevalence and antibiotic susceptibility pattern. Annals of Saudi medicine 2012;32(5), 513-516.
Crossref - Ghanem A, Bahashwan SA, El Shafey HM, et al. Antimicrobial resistance pattern of MRSA strains isolated from patients of a hospital in Madinah, Kingdom of Saudi Arabia. African Journal of Microbiology Research 2018;12(47), 1044-1049.
Crossref - Moussa IMI, Kabli SA, Hemeg HA, Al-Garni SM, Shibl AM. A novel multiplex PCR for molecular characterization of methicillin resistant Staphylococcus aureus recovered from Jeddah, Kingdom of Saudi Arabia. Indian Journal of Medical Microbiology 2012;30(3), 296-301.
Crossref - Shady HMA, Bakr AEA, Hashad ME, Alzohairy MA. Staphylococcus aureus nasal carriage among outpatients attending primary health care centers: a comparative study of two cities in Saudi Arabia and Egypt. Brazilian Journal of Infectious Diseases 2015;19(1), 68-76.
Crossref - Al-Zahrani IA, Azhar EI, Jiman-Fatani AA, et al. Impact of mass migrations on the clonal variation of clinical Staphylococcus aureus strains isolated from the Western region of Saudi Arabia. Journal of infection and public health 2019;12(3), 317-322.
Crossref - Bukharie HA. Increasing threat of community-acquired methicillin-resistant Staphylococcus aureus. The American journal of the medical sciences 2010;340(5), 378-381.
Crossref - Khanfar H, Senok A, Anani A, Zinkevich V. Methicillin-resistant Staphylococcus aureus transmission in a low-prevalence healthcare setting. Journal of Infection and Public Health 2012;5(4), 311-316.
Crossref - Al-Humam NA. Detection of Escherichia coli, Salmonella spp. and Staphylococcus aureus in Ready-to-Eat Food in Al-Ahsa Province, Saudi Arabia. J Nutr 2019; 9(2), 1-6.
- Hamid ME. Resistance pattern of coagulase positive Staphylococcus aureus clinical isolates from Asir region, Kingdom of Saudi Arabia. Journal of Microbiology and Antimicrobials 2011;3(4), 102-108.
- El-Deeb W, Fayez M, Elmoslemany A, Kandeel M, Zidan K. Methicillin resistant Staphylococcus aureus among goat farms in Eastern province, Saudi Arabia: Prevalence and risk factors. Preventive veterinary medicine 2018;156, 84-90.
Crossref - El-Ghareeb W, Almathen F, Fayez M, Alsultan R. Methicillin resistance Staphylococcus aureus (MRSA) in camel meat: Prevalence and antibiotic susceptibility. Slovenian Veterinary Research 2019;56(22), 249-256.
- Alzohairy MA. Colonization and antibiotic susceptibility pattern of methicillin resistance Staphylococcus aureus (MRSA) among farm animals in Saudi Arabia. African Journal of Bacteriology Research 2011; 3(4), 63-68.
- Bhedi KR, Nayak JB, Brahmbhatt MN, Roy A, Mathakiya RA, Rajpura RM. Detection and Molecular Characterization of Methicillin-Resistant Staphylococcus aureus Obtained From Poultry and Poultry House Environment of Anand District, Gujarat, India. International Journal of Current Microbiology and Applied Sciences. 2018; 7(2), 867–872.
Crossref - Islam MA, Parveen S, Rahman M, et al. Occurrence and characterization of methicillin resistant Staphylococcus aureus in processed raw foods and ready-to-eat foods in an urban setting of a developing country. Frontiers in microbiology 2019;10(503).
Crossref - Cuny C, Layer F, Hansen S, Werner G, Witte W. Nasal Colonization of Humans with Occupational Exposure to Raw Meat and to Raw Meat Products with Methicillin-Susceptible and Methicillin-Resistant Staphylococcus aureus. Toxins 2019;11(4).
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.