ISSN: 0973-7510
E-ISSN: 2581-690X
Patients with coronavirus disease-19 (COVID-19) present as mildly, moderately, or severely and critically ill. Cytokine storm is responsible for fatal pneumonia and acute respiratory distress syndrome. Interferon-γ-induced protein-10 (IP-10) and chemokine ligand-7 (CCL-7) are chemokines that play a role in the chemotaxis of inflammatory cells and the release of pro-inflammatory cytokines. In this study, we assessed the serum levels of IP-10 and CCL-7 chemokines in COVID-19 patients and their correlation with disease severity and prognosis. The serum levels of CCL-7 and IP-10 were assessed in 67 COVID-19 patients and 10 healthy controls. Serum samples were collected and examined for these two markers using direct enzyme-linked immunosorbent assay. Patients were divided into two groups according to their disease severity. Serum levels of the test markers were compared between patients and controls, and between patients with different disease severities and correlated with other clinical and laboratory parameters. CCL-7 and IP-10 levels were significantly higher in patients than in controls and in severe than in mild/moderate cases. The receiver operating characteristic curve analysis of the two markers showed better performance of the combined markers as predictors of disease severity (area under the curve = 0.792). The results of our study suggest a potential role of IP-10 and CCL-7 as predictors of COVID-19 severity.
Pandemic, COVID-19, IP-10, CCL-7
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) is a novel coronavirus that causes a severe acute respiratory syndrome that is currently named as coronavirus disease-19 (COVID-19). It frequently induces fatal inflammatory responses and acute lung injuries. As of January 7, 2022, 298,915,721 confirmed COVID-19 cases have been reported worldwide, with an estimated mortality rate of 1.8%.1
Coronaviruses are a group of diverse, enveloped, positive-sense, and single-stranded RNA viruses.2 Currently, seven known coronaviruses infect humans, including SARS-CoV-2. Domestic or wild animals, such as bats, civil cats, camels, and rodents, are recognized as reservoirs that transmit the virus to humans.2
COVID-19 has a clinical presentation that ranges from mild to moderate in majority of the cases. However, some patients suffer a hyper-inflammatory syndrome triggered by massive cytokine/chemokine production, referred to as cytokine storm, which results in severe pneumonia and acute respiratory distress syndrome. Studying the pathogenesis of cytokine storms is helpful for developing efficient treatment modalities.3
In patients with COVID-19, the counts of CD4+, CD8+ T lymphocytes, and natural killer cells are lower than expected in severe cases.4 On the contrary, monocyte and macrophage counts are high, which explains the increased levels of pro-inflammatory cytokines, including tumor necrosis factor-α (TNF-a) and interleukins (IL)-6, 1, and 8 that are responsible for the cytokine storm.5
Previous studies have reported higher serum levels of multiple cytokines and chemokines, including TNF-a, IL-1, 6, 10, and 17, interferon-g (IFN-g), IFN-g inducible protein-10 (IP-10), granulocyte-monocyte colony-stimulating factor, granulocyte colony-stimulating factor (G-CSF), and chemokine ligand-7 (CCL-7)] in patients with COVID-19. Their serum levels were significantly correlated with disease severity and poor prognosis, and were higher in severe and critically ill patients.6-11 These results suggested that measuring the serum levels of different cytokines and chemokines may serve as predictors of disease severity and respiratory failure in COVID-19.7
Chemokines play a vital role in immunity against viruses by recruiting both innate and adaptive immune cells. However, the excessive expression of chemokines can lead to inflammation and acute respiratory distress syndrome (ARDS), which is a common complication following COVID-19.12 Therefore, studying chemokine levels in patients helps to understand the immunopathogenesis of COVID-19 and their utility as severity and prognostic markers.
IP-10/CXC ligand-10 belongs to the chemokine family. IFN-g induces its secretion from different cells, such as leukocytes and endothelial and epithelial cells. IP-10 participates in chemotaxis, apoptosis, and cell growth inhibition.13 IP-10 is a chemokine that plays an important role in viral infections. Some studies reported that IP-10 is released within a few days of SARS infections, where it suppresses myeloid progenitor cell proliferation, resulting in lymphopenia observed in SARS-CoV infections.14 A Study of chemokine profiles in COVID-19 patients showed an upregulation of IP-10 levels, when compared with non-COVID patients.15
Chemokine (C-C motif) ligand-7 (CCL-7) is a chemokine formerly known as monocyte-chemotactic protein-3 (MCP-3). Along with other chemokines, it is released upon macrophage activation, enhancing further chemotaxis of mononuclear macrophages, leading to overproduction of pro-inflammatory cytokines.16
Multiple studies have suggested that a higher level of CCL-7 in COVID-19 patients was related to disease severity, exacerbated lung injury, and fatal prognosis.8,9
Immune dysregulation is a target of different therapeutic agents. Several TNF-blocking antibodies and monoclonal antibodies targeting IL-6 and IL-1 receptor antagonists (IL-1 ra and IL-2ra) have been successfully used to treat hospitalized COVID-19 patients.3,17,18
Purpose of the study
To evaluate the utility of measuring the serum levels of IP-10 and CCL-7 chemokines as severity and prognostic markers in COVID-19 patients.
Study population
This study was conducted on 67 COVID-19 patients selected from the outpatient clinics and isolation wards of the Ain-Shams University Hospitals from April to July, 2021. There were 37 men and 30 women with a mean age 53.84 ± 15.96. Patients were categorized as follows: Group I included 33 patients with mild to moderate disease (mean age± standard deviation (SD) =49.00 ± 14.65), and group II included 34 severe and critically ill patients (mean age± SD = 58.53 ± 15.97).
COVID-19 diagnosis and patient classification were performed according to the Ain-Shams clinical guide for COVID-19. Diagnosis was confirmed by detecting SARS-CoV-2 RNA in nasopharyngeal swabs using polymerase chain reaction test. Ten healthy persons of matching age and gender were included in the control group. Consent was obtained from all the study participants. Patients on immunosuppressive drugs or with chronic infections (e.g., hepatitis B and C viral infections), cancers, or immunological disorders were excluded from this study.
We have recorded relevant clinical and demographic data and laboratory tests results, such as C-reactive protein (CRP), white blood cell (WBC) count, alanine aminotransferase, aspartate aminotransferase, creatinine, and D-dimer levels were recorded. Peripheral blood samples were collected from all the participants under aseptic conditions. Samples were transferred into serum separation tubes, and the separated serum was stored at -80 °C for the quantitative measurement of CCL-7 and IP-10.
Measurement of serum CCL-7 and IP-10 serum levels
The serum CCL-7 and IP-10 levels were measured using sandwich enzyme linked immunosorbent assay (ELISA) using ELISA Kits (Bioassay Technology Laboratory, England/China, Cat. No: E0126Hu for CCL-7 and Bioassay Technology Laboratory, England/Cina, Cat. No: E0412Hu for IP-10). The procedures were performed according to the manufacturer’s instructions. The results were read and interpreted using a digital and analog system plate reader (Roma, Italy). A standard curve was constructed for each assay. The concentrations of CCL-7 and IP-10 in the tested samples were determined and expressed in pg/mL.
Statistical analysis
Statistical analyses were performed using IBM SPSS (Statistical Package for the Social Sciences) version 26.0. Descriptive statistical tests were used to calculate median, mean ± SD, frequency, and interquartile range. The analytical statistics used were unpaired Student’s t-test, chi-square test, correlation coefficient (r) test, and Mann–Whitney U test. Receiver operating characteristic (ROC) curve analysis was used to determine the cut-off values, sensitivity, and specificity for CCL-7 and IP-10 as indicators of severity.
Demographic and clinical data of the enrolled 67 patients are presented in Table 1.
Table (1):
Demographic and clinical data of the patient group.
Demographic and clinical data |
|
|---|---|
Sex Males, n (%) Females, n (%) |
37 (55.2%) 30 (44.8%) |
Age, years Mean ± SD |
53.84 ± 15.96 |
Co-morbidities within cases: n (%) Diabetes mellitus Hypertension Ischemic heart diseases Chest diseases Renal disorders |
50 (74.6%) 26 (38.8%) 3 (4.5%) 3 (4.5%) 5 (7.5%) |
Clinical outcomes: Discharged n (%) Died n (%) |
22 (64.7%) 12 (35.3%) |
A significant difference was found between the ages of patients in both the groups (p = 0.013). CRP and D-dimer levels were significantly higher in group II than in group I (p =0.001 and 0.005, respectively) (Table 2).
Table (2):
Comparison of age and laboratory parameters between patient groups I and II.
| COVID-19 groups (n. 67) | Test value | P-Value | ||
|---|---|---|---|---|
| Group I (n.33) Median (IQR) |
Group II (n. 34) Median (IQR) |
|||
| Age | 49.00 ± 14.65 | 58.53 ± 15.97 | -2.543* | 0.013 |
| Laboratory parameters (reference interval) | ||||
| White cell count (4-11 103/uL) |
10.6 (6 – 13.9) | 10.1 (6.5 – 15.6) | 0.570 | 0.569 |
| CRP (0-6 mg/L) | 20.2(10 – 48.7) | 90.2 (23 – 165.5) | 4.536 | 0.001* |
| D-dimer (0-0.5 μg FEU/mL) | 0.8(0.35 – 0.93) | 1.84 (1 – 5.6) | 4.536 | 0.005* |
| Creatinine (0.6-1.2 mg/dL) | 1.4 (0.9 – 2.1) | 1 (0.5 – 1.5) | 1.262 | 0.207 |
| ALT (7-52 IU/L) | 28 (8 – 37) | 29 (17 – 40) | 0.262 | 0.207 |
| AST (13-39 IU/L) | 28 (8 – 37) | 29 (17 – 40) | 0.672 | 0.5 |
IQR: Interquartile range; *: Significant; Boldfaced data are significant
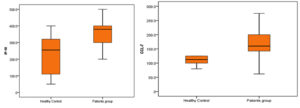
The Mann–Whitney test revealed a highly significant elevation in serum levels of CCL-7 and IP-10 in the studied patients compared to that in the control group, as shown in Table 3 and Fig. 1 (P = 0.001 and 0.003, respectively).
Table (3):
Serum levels of CCL-7 and IP-10 in the patient and control groups.
| Healthy Control No. 10 |
Patients Group No. 67 |
Mann-Whitney Test Value |
p-Value | Sig. | |
|---|---|---|---|---|---|
| CCL-7 | |||||
| Median (IQR) | 112.5 (100 – 125) | 160 (140 – 200) | -3.422 ≠ | 0.001 | HS |
| Range | 12.5 – 170 | 62 – 350 | |||
| IP-10 | |||||
| Median (IQR) | 255 (110 – 320) | 380 (300 – 400) | -2.941 ≠ | 0-03 | HS |
| Range | 50 – 400 | 200 – 1300 | |||
IQR: Interquartile Range; HS: Highly Significant; Bolded Data are Significant.
The difference between the serum levels of CCL-7 and IP-10 in group I and II (P=0.000 and 0.026, respectively) was statistically significant (Table 4).
Table (4):
Serum levels of CCL-7 and IP-10 in the patient group.
| Group I | Group II | Mann–Whitney test value | P-value | Sig. | |
|---|---|---|---|---|---|
| No. = 33 | No. = 34 | ||||
| CCL-7 | |||||
| Median (IQR) | 147 (125 – 165) | 190 (150 – 250) | -3.976 | 0.000 | HS |
| Range | 62 – 250 | 110 – 350 | |||
| IP-10 | |||||
| Median (IQR) | 330 (250 – 400) | 380 (300 – 500) | -2.227 | 0.026 | S |
| Range | 200 – 480 | 200 – 1300 |
IQR: Interquartile range; HS: Highly significant; S: Significant
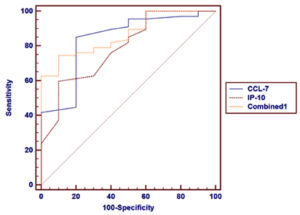
Table 5 and Fig. 2 show the ROC curve analysis of CCL-7 and IP-10 serum levels in patients with COVID-19 and healthy controls. The AUC value was higher in the combination of two markers (0.865) than in individual marker use (0.837 and 0.789 for CCL-7 and IP-10, respectively).
Table (5):
ROC curve analysis of CCL-7 and IP-10 markers in COVID-19 patients and healthy controls.
Variables |
Cut off point |
AUC |
Sensitivity |
Specificity |
+PV |
-PV |
|---|---|---|---|---|---|---|
CCL-7 |
>125 |
0.837 |
85.07 |
80.00 |
96.6 |
44.4 |
IP-10 |
>320 |
0.789 |
59.70 |
90.00 |
97.6 |
25.0 |
Combined |
0.865 |
74.63 |
90.00 |
98.0 |
34.6 |
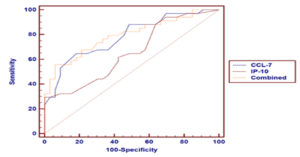
Table 6 and Fig. 3 show the ROC curve analysis of both CCL-7 and IP-10 serum levels as predictors of COVID-19 severity. The AUC value was higher in the combination of two markers (0.792) than in individual marker use (0.782 and 0.658 for CCL-7 and IP-10, respectively).
Table (6):
ROC curve analysis of CCL-7 and IP-10 severity markers in COVID-19 patients.
Variables |
Cut off point |
AUC |
Sensitivity |
Specificity |
+PV |
-PV |
P-value |
|---|---|---|---|---|---|---|---|
CCL-7 |
>170 |
0.782 |
64.71 |
81.82 |
78.6 |
69.2 |
|
IP-10 |
>480 |
0.658 |
29.41 |
100.00 |
100.0 |
57.9 |
0.018 |
Combined |
0.792 |
55.88 |
93.94 |
90.5 |
67.4 |
AUC: Area Under the Curve; +PV: Positive Predictive Value; -PV: Negative Predictive Value
Tables 7 and 8 show a statistically significant correlation between CCL-7 and IP-10 levels in all patient groups and in group II. In addition, there was a statistically significant correlation between IP-10 and D-dimer levels in all patient groups and in group II.
Table (7):
Correlation between CCL-7 and other laboratory parameters.
| CCL-7 | ||||||
|---|---|---|---|---|---|---|
| All cases | Group I | Group II | ||||
| r | p-value | r | p-value | R | p-value | |
| IP10 | 0.374* | 0.002 | 0.212 | 0.237 | 0.353* | 0.041 |
| CRP | 0.232 | 0.059 | 0.171 | 0.341 | -0.097 | 0.585 |
| D. dimer | 0.222 | 0.071 | 0.080 | 0.657 | -0.051 | 0.776 |
| Total WBC count | 0.146 | 0.410 | -0.195 | 0.452 | 0.389 | 0.122 |
| Creatinine | 0.080 | 0.653 | 0.456 | 0.066 | 0.027 | 0.917 |
| ALT | 0.117 | 0.510 | 0.116 | 0.656 | 0.042 | 0.873 |
| AST | 0.208 | 0.238 | 0.381 | 0.131 | -0.133 | 0.612 |
*: Significant
Table (8):
Correlation between IP-10 and other laboratory parameters.
| IP-10 | ||||||
|---|---|---|---|---|---|---|
| All cases | Group I | Group II | ||||
| r | p-value | r | p-value | R | p-value | |
| CCL-7 | 0.374* | 0.002 | 0.212 | 0.237 | 0.353* | 0.041 |
| CRP | -0.018 | 0.884 | -0.212 | 0.236 | -0.184 | 0.298 |
| D. dimer | 0.358* | 0.003 | 0.227 | 0.204 | 0.340* | 0.049 |
| Total WBC count | 0.048 | 0.787 | -0.114 | 0.663 | 0.104 | 0.690 |
| Creatinine | -0.054 | 0.763 | -0.062 | 0.814 | 0.027 | 0.919 |
| ALT | -0.092 | 0.603 | 0.040 | 0.878 | -0.321 | 0.209 |
| AST | -0.098 | 0.581 | 0.032 | 0.902 | -0.277 | 0.282 |
*: Significant
In the present study, the mean age of the patients was 53.84 ± 15.96 and 55.2% of them were men. The mean age of the patients was significantly higher in group II (58.53 ± 15.97) than in group I (49.00 ± 14.65). This is similar to the results of Yang et al.10 who found in their study that 81.8% of total and 68% of severe patients were above 60 years of age, while 64.3% of patients with moderate disease were between 16 and 59 years.
Our results revealed a difference between the patient and control groups regarding serum levels of CCL-7 and IP-10 (p = 0.001 and 0.003, respectively). The levels of CCL-7 and IP-10 were significantly higher in group II patients (p= 0.000 and 0.026, respectively) than in group I.
These results are consistent with those of Yang et al.10 who correlated the disease severity of COVID-19 with cytokine levels. Among the 48 tested cytokines, they reported a positive correlation between IP-10, CCL-7, IL-1ra, hepatocyte growth factor (HGF), monokine induced by interferon-gamma (MIG), and macrophage inflammatory protein-1 alpha (MIP-1a) with disease severity (p < 0.0001). Similarly, they found a significant elevation in CCL-7 level in severe patients than in healthy controls. The expression levels of IL-6, IL-1ra, IL-2ra, IL-18, IL-10, IFN-g, and macrophage colony-stimulating factor (M-CSF) were not significantly correlated with the Murray scores.
Some studies revealed the upregulation of IP-10 in COVID-19 patients, when compared with non-COVID-19 people.15,19,20 Moreover, IP-10 was found to be lower in patients with bacterial ARDS than in those with non-COVID-19 viral ARDS. Therefore, IP-10 could be used as a biomarker for viral infections.15 Previous studies have also reported that IP-10 levels increase with disease severity; therefore, it can be a potential predictor of disease severity and outcome.13,19,21 In contrast, Moustafa et al.20 showed that there was no significant difference between the IP-10 levels in moderate and severe cases.
In this study, we investigated the correlation between the tested chemokines, IP-10 and CCL-7, and different clinical and laboratory parameters in both patient groups (using Spearman’s correlation coefficient and the Mann–Whitney test). A significant correlation was found between IP-10 and D-dimer levels in all patient groups and in the group of severe cases (p = 0.003 and 0.049, respectively). However, no statistically significant association was observed between CCL-7 levels and any of the clinical or laboratory parameters of the patients.
Similar results have been reported in some previous studies, which found an increased IP-10 level in patients with elevated D-dimer levels, when compared to those with low D- dimer levels.8,9 Therefore, they concluded that IP-10 inhibited the recovery of endothelial cells. Therefore, an anti-IP-10 antibody is under trial to prevent cardiovascular events.10 Thus, targeting IP-10 may be an efficient therapeutic modality for COVID-19 patients with thrombosis.
Our results revealed no statistically significant association between the tested markers IP-10 or CCL-7 and the disease outcome.
Yang et al.,10 in their study, examined the expression levels of different cytokines and chemokines. They reported that the expression levels of IP-10, CCL-7, and HGF were the highest in critically ill patients, followed by severe and moderate patients throughout the study. Conversely, expression levels of IL-1ra M-CSF were mainly detected in the early stage of disease progression, while the expression levels of MIG, and MIP-1a were higher in the later stages of disease progression. Only few differences were found in IL-6, IL-18, IL-13, IL-1b, cutaneous T-cell–attracting chemokine, and G-CSF levels, between early and late stages of the disease.
They also correlated the dynamic expression of six tested cytokines with the different prognoses in eight critically ill patients. They reported that the levels of IP-10, CCL-7, and other cytokines were significantly high at the time of initial assessment and throughout the course of the disease in two fatal cases and in the other four patients who remained critically ill. However, the cytokine levels decreased significantly with time in two patients who were discharged from the hospital. They concluded that the cytokine storm was significantly more evident in severe cases than in mild cases.
In the present study, ROC curve analysis of CCL-7, IP-10, and a combination of both revealed AUC values of 0.837, 0.789, and 0.865, respectively in patients and controls. On comparison of the two severity groups of patients in our study, ROC curve analysis revealed AUC values of 0.782, 0.658, and 0.792 for CCL-7, IP-10, and their combination, respectively.
Similarly, Yang et al.,10 in their study compared two groups of COVID-19 patients (ARDS and non-ARDS). ROC curve analysis revealed AUC values of 0.803, 0.97 and 0.99 for CCL-7, IP-10, and their combination, respectively in ARDS and non-ARDS groups. These results suggested that the combination of these two markers may be an excellent biomarker for disease severity and progression. ROC curve analysis of other cytokines revealed variable AUC values ranging from 0.565 to 0.866 for IL-1ra.
In a study conducted by Han et al.,7 the authors investigated the role of a group of cytokines as diagnostic markers of COVID-19. ROC curve analysis was performed for a group of cytokines in 102 COVID-19 patients and healthy controls. The cytokines tested were IFN-g, IL-2, IL-10, IL-6, IL-4, TNF-α, and CRP. The area under the ROC curve of CRP was the largest among all cytokines (0.955), followed by IL-10 (0.841) and IFN-g (0.704).
To assess the values of the tested cytokines as markers of disease severity, Han et al.,7 performed an ROC curve analysis in patients with severe and critical COVID-19. The area under the ROC curve of IL-6 was the largest among all cytokines (0.826) and the lowest was that of IL-4 (0.627).
Neoptrin, an immune activation marker, is a possible diagnostic and prognostic marker in COVID-19. It is detected in many inflammatory conditions, cancers, and autoimmune disorders. Its production is mainly enhanced by IFN-g. Robertson et al.22 reported higher levels of both neoptrin and IFN-g in severely ill COVID-19 patients than in patients with mild disease.
Various chemokines have been evaluated as diagnostic or prognostic markers for COVID-19. One of these cytokines is CCL-17. An Egyptian study conducted at Ain-Shams University by Moustafa et al.20 showed a statistically significant difference between the CCL-17 levels of healthy controls and a group of COVID-19 patients. However, they found no significant difference between CCL-17 levels of the moderate and severe groups of patients. The authors proposed the use of CCL-17 as a diagnostic marker. In contrast to their results, our results suggest that IP-10 and CCL-7 could be useful predictive markers of disease severity.
This study suggests the potential utility of IP-10 and CCL-7 as indicators of COVID-19 severity. The findings of this study can be used in future large scale studies. Further studies can be conducted to validate the use of these chemokines in COVID-19 diagnosis.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
RAH, MAK, HGE, HMT, SHG, DHA, YMA participated in study design and selection of the patients. HGE, HMT and SHG collected samples from patients. RAH, MAK, YMA and DHA performed ELISA on the samples. RAH, MAK, HGE, HMT, SHG, DHA, YMA analyzed the data, wrote the manuscript, read and approved the final manuscript for publication.
FUNDING
None.
ETHICS STATEMENT
This study was approved by the Ethical Committee regulation of Faculty of Medicine,
Ain Shams University, Cairo, Egypt with reference number FMASU R 74/2021.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
INFORMED CONSENT
Written informed consent was obtained from the participants before enrolling in the study.
- World Health organization. 2022. https://covid19.who.int/
- Zhong J, Tang J, Ye C, Dong L. The immunology of COVID-19: is immune modulation an option for treatment? Lancet Rheumatol. 2020;2(7):e428-e436.
Crossref - Rahmati M, Moosav MA. Cytokine-Targeted Therapy in Severely ill COVID-19 Patients: Options and Caution. EJMO. 2020;4(2):179-181.
Crossref - Diao B, Wang C, Tan Y, et al. Reduction and Functional Exhaustion of T Cells in Patients With Coronavirus Disease 2019 (COVID-19). Front Immunol. 2020;11:827;1-7.
Crossref - Soy M, Keser G, Atagunduz P, et al. Cytokine storm in COVID-19: pathogenesis and overview of anti-inflammatory agents used in treatment. Clin Rheumatol. 2020;39(7):2085-2094.
Crossref - Costela-Ruiz VJ, Illescas-Montesa R, Puerta-Puertac JM, Ruiz C, Melguizo-Rodriguez L. SARS-CoV-2 infection: The role of cytokines in COVID-19 disease. Cytokine and Growth Factor Reviews. 2020;54:62-75.
Crossref - Han H, Ma Q, Li C , Liu R, Zhao L, et al. Profiling serum cytokines in COVID-19 patients reveals IL-6 and IL-10 are disease severity predictors. Emerging Microbes & Infections. 2020;9(1):1123-1130.
Crossref - Girija AS, Shankar EM, Larsson M. Could SARS-CoV-2-Induced hyperinflammation Magnify the Severity of Coronavirus Disease (CoViD-19) Leading to Acute Respiratory Distress Syndrome? Front Immunol. 2020;11:1206.
Crossref - Xu ZS, Shu T, Kang L, et al. Temporal profiling of plasma cytokines, chemokines and growth factors from mild, severe and fatal COVID-19 patients. Signal Transduct Target Ther. 2020;5(1):100.
Crossref - Yang Y, Shen C, Li J, et al. Plasma IP-10 and MCP-3 levels are highly associated with disease severity and predict the progression of COVID-19. J Allergy Clin Immunol. 2020:146(1):119-127.
- Wu D, Yang X. TH17 responses in cytokine storm ofCOVID-19: An emerging target of JAK2inhibitor Fedratinib. J Microbiol Immunol Infect. 2020;53(3):368- 370.
Crossref - Melchjorsen J, Sorensen LN, Paludan SR. Expression and function of chemokines during viral infections: from molecular mechanisms to In vivo function. J Leukoc Biol. 2003;74(3):331-343.
Crossref - Chen Y, Wang J, Liu C, et al. IP-10 and MCP-1 as biomarkers associated with disease severity of COVID-19. Mol Med. 2020;26(1):97.
Crossref - Zhou J, Chu H, Li C, et al. Active replication of middle east respiratory syndrome coronavirus and aberrant induction of inflammatory cytokines and chemokines in human macrophages: implications for pathogenesis. J Infect Dis. 2014:209(9):1331-1342.
Crossref - Hue S, Beldi-Ferchiou A, Bendib I, et al. Uncontrolled innate and impaired adaptive immune responses in patients with COVID-19 ARDS. Am J Respir Crit Care Med. 2020;202(11):1509-1519.
Crossref - Ye Q, Wang B, Mao J. The pathogenesis and treatment of the ‘Cytokine Storm’ in COVID-19. J Infect. 2020;80(6):607-613.
Crossref - Duret PM, Sebbag E, Mallick A, et al. Recovery from COVID-19 in a patient with spondyloarthritis treated with TNF alpha inhibitor etanercept. Ann Rheum Dis. 2020;79(9):1251-1252.
Crossref - Pacha O, Sallman MA, Evans SE. COVID-19: a case for inhibiting IL-17? Nat Rev Immunol. 2020;20(6):345-346.
Crossref - Yasin MM, El-Asady RS, Elsheikh NG, Mokhtar MM. Interferon- gamma-induced protein-10 (IP-10) and natural killer cell count as predictive factors for the severity of COVID-19. Microbes and Infectious Diseases. 2021;2(4):4:623-631
Crossref - Moustafa NM, Mohamed RA, Elsaid RG, Mahmoud FM. Diagnostic Value of Serum Level of Interleukin 33 (IL-33), C-C Motif Chemokine Ligand 17 (CCL17) and Interferon Gamma Inducible Protein-10 (IP-10) in Coronavirus Disease 2019 (COVID-19) Patients. Egypt J Med Microbiol. 2022;31:23-30.
Crossref - Khalil BA, Elemam NM, Maghazachi AA. Chemokines and chemokine receptors during COVID-19 infection. Comput Struct Biotechnol J. 2021;19:976-988.
Crossref - Robertson J, Gostne JM, Nilsson S, Andersson L, Fuchs D, Gisslen M. Serum neopterin levels in relation to mild and severe COVID-19. BMC Infect Dis. 2020;20:1-6.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.