ISSN: 0973-7510
E-ISSN: 2581-690X
Rising demand in the aquaculture sector tends towards finding innovative ways to promote better yield and profitability. Benthic soil microbiota can provide an insight into the potent opportunistic fish pathogens as well as probiotics present in the aquaculture system. This study reports the seasonal diversity and abundance of fifteen culturable pathogenic bacterial strains belonging to the genera of Comamonas, Aeromonas, Providencia, Klebsiella, Escherichia, Acinetobacter, Serratia, Stenotrophomonas, Staphylococcus, and Enterobacter along with nine probiotic strains native to genera of Bacillus and Pseudomonas isolated from an aquaculture farm benthic soil, located in East Kolkata Wetlands, West Bengal, India. Strains are isolated using traditional microbial culture methods and tested for their antimicrobial susceptibility against commonly available antibiotics. 16S rDNA analysis was done for the identification of the strains and the establishment of their phylogenetic relationships. Among the isolates, B. pumilus strain S8 in the pre-monsoon sample, E. coli strain M2aR1 in the monsoon sample, and A. hydrophila strain P6dF1 in the post-monsoon sample were the most abundant having MPN counts of 275±21 x 106 CFU/gram dry soil, 278±18 x 106 CFU/gram dry soil, and 321±28 x 106 CFU/gram dry soil respectively. Data on the temporal diversity, abundance, and drug-susceptibility of prokaryotic fish-pathogens and probiotics can be used to formulate measures for sustainable aquaculture practices with reduced maintenance costs.
Aquaculture, Fish pathogens, probiotics, 16S rDNA analysis, Antibiogram
Fish production in India has come a long way from being dependent on natural sources (rivers, lakes, and ponds) to well-contemplated aquaculture farms and earned huge success after the implementation of composite fish culture techniques, leading to species diversification and enhanced yield1. The blue revolution was brought in India by the incorporation of techniques like polyculture and induced fish breeding in static ponds or tanks2. India stands second in terms of farmed fish production globally, with more than 5.6 million tonnes of finfish (2015-2016) from inland aquaculture farms only3. Among the Indian states, Andhra Pradesh, and West Bengal are major producers in aquaculture farming4.
The immense problem to these farms is contributed by the inert fish feeds, which affect the maintenance of farm5, harm the environment6, escalate the production cost3, and raises fish prices in the market7. One of the solutions to this problem is microbial-based aquaculture systems, which may incur better sustenance of the farm with minimal input and reduced costs6. Prokaryotic microbiota of the soil and water plays various functions in fish farming, including valuable roles as probiotics, prebiotics, biomarkers, sentinels, and sometimes direct food for cultured fishes8–10 along with detrimental ones like opportunistic pathogens11–14. Prokaryotic pathogens cause several infectious diseases resulting in loss of revenue for the fish farm by causing fish morbidity15. Unfortunately, these pathogens are detected only when an outbreak occurs, and sometimes it is too late to respond. Commercially available antibiotics are used regularly, creating drug-resistant bacterial strains16. Benthic soil microcosm can be utilized to act as a natural feed for the aquaculture farms and hence play a part in its sustenance17. To implement bioaugmentation techniques, a detailed exploration of the microbial food chain, pathogenic transmission, and presence of probiotics are needed for formulating steps leading to the betterment of yield18.
East Kolkata Wetlands, in the eastern part of the Kolkata, is designated as a Ramsar site that supplies a major fraction of the fish demand of the city19. These inland freshwater aquaculture farms are one of a kind and serve two-fold purposes. Firstly these farms are mainly sewage fed, and act as natural sinks for waste recycling and attenuation of floods. Secondly, these water bodies are utilised for a thriving culture of several fish species20. The sewage fed aquaculture ponds, locally known as bheris, are unique in their operating procedure as they utilize municipal waste products as a fish feed with the occasional addition of inert feeds21. However, the seasonal dynamics of the physicochemical properties of water and benthic soil play crucial roles in the microbiome of the benthic soil of aquaculture farms22,23. Though several references are available on the description of the microbial communities of East Kolkata Wetlands, seasonal dynamics of the benthic soil prokaryotic microbiota emphasizing on the variation of probable fish pathogens and probiotics are scarce.
Media and Chemicals
All chemicals and media were procured from Himedia (India) and Sigma Aldrich Chemical (USA). Tryptone Soya Agar (TSA) was used as enrichment media for the isolation of strains. Routine subculture and the permanent stocks were made in Nutrient agar (NA). Antibiotic Susceptibility test was done on Mueller Hinton Agar (MHA). Culture media were subjected to sterilization at 120°C temperature and 20 psi pressure for 15 minutes before inoculation.
Sample collection
Benthic soil samples were acquired aseptically from an aquaculture farm in East Kolkata Wetlands (Lat – 22.5699° Long- 88.4394°) from the top surface of the soil below 60cm of the water column. Samples of three seasons in the year 2019 viz. Pre-monsoon sample (April), Monsoon sample (August), and Post monsoon sample (December) were chosen for prokaryotic analyses in this study.
Isolation of bacterial strains
Isolation of culturable bacterial strains was done according to the method given by Vieira and Nahas 200524, with some minor alterations. 5gms (wet weight) of each soil sample was first diluted and homogenized in sterile water (50 ml) with intermittent cooling in an ice bath for 30 minutes. The homogenate was then passed through a sterile 2mm mesh. Filtrates were serially diluted up to 108 fold, and 100 µl from the last four dilutions (105 to 108) of each filtrate were spread on TSA plates (in triplicates), which were then incubated at 37°C for 48 hours. A dry weight of soil was calculated by incubating 50 grams of benthic soil in 105°C for a period of 8 hours, and the final weight was used to calculate the wet weight to dry weight conversion factor25.
Most Probable Number Count
Colonies that appeared on Each TSA plate after 48 hours of incubation, were marked based on colony morphology (shape, size, colour, transparency, margin contour, and surface topology) and were analysed for the Most Probable Number (MPN) count was done according to the method given by Janssen et al. 200226 with some minor alterations. Conversion factors of wet weight to dry weight of each soil sample and dilution factor of the inoculum were considered in the MPN count. The count was taken as a mean along with standard deviation for each dilution (from the triplicate count) and results were expressed in CFU x 106/gram dry soil. For the process of purification, colonies were transferred to NA plates and repeated subcultures were done until the strains were free from conglomeration. Each purified strain was stored permanently in NA stab cultures at 4°C and 80% glycerol stock at -20°C.
Biochemical characterization
A total of fourteen biochemical tests, including determination of Gram Character, Methyl Red (MR), tests for gelatinase, Triple Sugar Iron (TSI), degradation of starch, Voges- Proskauer (VP), indole production, citrate, oxidase, motility tests, etc. were performed with each purified strains and they were characterized following the methods of Bergey’s Manual of Systematic Bacteriology27.
Antibiogram
Antibiogram was prepared by testing all the isolated strains with a multitude of antibiotics by disk diffusion method28 using commercially available disks with a total of thirteen antibiotics viz. A, Ampicillin (10 µg); Cf, Ciprofloxacin (5 µg); Co, Cotrimoxazole (25 µg); E, Erythromycin (15 µg), Fr, Furazolidone (100 µg); C, Chloramphenicol (30 µg); G, Gentamycin (30 µg); K, Kanamycin (30 µg), Of, Ofloxacin (5 µg), Na, Nalidixic acid (30 µg). S, Streptomycin (10 µg); T, Tetracycline (30 µg); VA, Vancomycin (30 µg) on MHA. Cell concentrations from overnight grown culture (in MHB) of each isolate were measured spectrophotometrically at 660 nm (Systronics 2202 Double Beam Spectrophotometer, Systronics, India) and inoculums were spread on sterile MHA (1×109 CFU/ml). Antibiotic discs were placed 15 cm apart on the spread plates followed by incubation at 37°C for 24 hours29. Strains were marked as susceptible, intermediate, or resistant by measurement of the diameter of the inhibitory zones to a particular antibiotic that was matched with the manufacturer’s interpretive table as per the recommendations given by the National Committee for Clinical Laboratory Standards30. An ATCC strain (Escherichia coli ATCC 25922) was used as a quality control.
Molecular Characterization
Isolation of Genomic DNA and 16S rRNA gene amplification
Isolation of Genomic DNA from the cultured bacterial strains was done using a bacterial genomic DNA isolation kit (GCC Biotech, India). Genomic DNA was electrophoresed through 0.8% agarose gel and was measured spectrophotometrically at 260 nm wavelength to estimate purity. The samples were subjected to 16S rRNA gene amplification using a Gradient Thermal Cycler (Biorad Laboratories, USA). Universal primer 27f (5′- AGAGTTTGATCMTGGCTCAG-3′) and 1492r (5′-GGTTACCTTGTTACGACTT-3′) encompassing V1-V9 hypervariable region of 16S rDNA were used for PCR amplification in conjunction with 2X PCR MasterMix (Thermo Fisher Scientific, India). The PCR amplification was carried out for 30 cycles and the amplicons (1.4 kbp approx.) were tested in 1.5% agarose gel using Ethidium Bromide stain. Purification of the amplicons was done using the Agarose Gel purification kit (NEB, USA) and sequencing was done from Xcelris Labs Ltd. (Ahmedabad, India).
In – silico analysis
Sequences were analysed in-silico using BLASTn (NCBI database), the hits were recorded for finding the nearest neighbour with the highest Max score. The aligned sequences were obtained in the FASTA format for downstream analyses in MEGA 731. CLUSTALW algorithm was used for alignment analyses and the data thus obtained were considered for phylogenetic analyses using Neighbor-Joining method32. The evolutionary distance was calculated by Maximum Likelihood Composite method33 and the branch lengths were calculated based on base substitution per site. Bootstrap tests of 500 replicates were done to construct the consensus tree to represent the evolutionary history of the taxa analysed34. Nearest four neighbours of each isolate from the BLASTn hits were used to create the phylogenetic tree. The partial 16S rDNA sequences of all strains were deposited into the GENBANK database.
Statistical analysis
Experiments on the enumeration of bacterial MPN were performed in triplicate and mean values were indicated along with standard deviation (SD). SPSS 17.0 (SPSS Inc., Chicago, USA) was used for the Statistical analyses.
Isolation of bacterial strains
In total, two hundred and forty-eight culturable bacterial strains were isolated and purified from the aquaculture benthic soil samples collected from three seasons. Molecular phylogenetic analyses revealed that several bacterial strains were common in samples from more than one season. Amongst the isolates, only twenty-four strains were revealed to have either presumptive pathogenic or probiotic values to fish and are reported in this communication. In the pre-monsoon season sample, a total of six strains (Bacillus flexus strain S1a, Aeromonas punctata strain S2, Bacillus pumilus strain S8, Bacillus subtilis strain S8a1, Comamonas aquatica strain SAC1 and Bacillus cereus strain SWA6a) were isolated. Cumulatively, eight strains were found in the monsoon season sample (Bacillus thuringiensis strain M10, Aeromonas enteropelogenes strain M11, Escherichia coli strain M2aR1, Pseudomonas aeruginosa strain M2F1, Bacillus flexus strain M3, Acinetobacter junii strain M5fR1, Serratia marcescens strain M5hR1 and Stenotrophomonas maltophilia strain M6aR1). Finally, from the post-monsoon season sample, another ten strains were isolated (Escherichia coli strain P1bR1, Providencia vermicola strain P2fR1, Enterobacter cloacae strain P4cR1, Bacillus flexus strain P5a, Klebsiella pneumoniae strain P5aR1, Serratia marcescens strain P5bR1, Bacillus cereus strain P6b1, Aeromonas hydrophila strain P6dF1, Comamonas aquatica strain P7, and Staphylococcus aureus strain P8d3a1). Isolate
B. flexus strain S1a, A. punctata strain S2,
B. pumilus strain S8, B. subtilis strain S8a1 were found in the samples from both the pre-monsoon and monsoon season whereas, E. coli strain M2aR1, B. flexus strain M3 and S. marcescens strain M5hR1 strains were found to occur in the monsoon and post-monsoon samples.
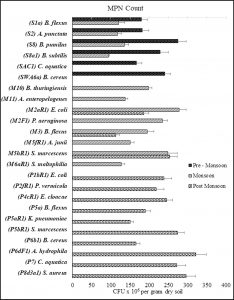
MPN count
The MPN counts of bacterial strains isolated from all three seasons are depicted in Figure 1. Isolate B. pumilus strain S8 in the pre-monsoon sample, isolate E. coli strain M2aR1 in the monsoon sample, and A. hydrophila strain P6dF1 in the post-monsoon sample were the most abundant having MPN counts of 275±21 x 106 CFU/gram dry soil, 278±18 x 106 CFU/gram dry soil, and 321±28 x 106 CFU/gram dry soil respectively. Strains were found to vary drastically in abundance in different seasons revealed by the difference in seasonal abundances of B. flexus strain S1a, A. punctata strain S2, B. pumilus strain S8, B. subtilis S8a1, E. coli strain M2aR1, B. flexus strain M3, and S. marcescens strain M5hR1. Coliform bacteria belonging to genus Escherichia, Klebsiella, Serratia, and Enterobacter, were found as the most abundant groups during the monsoon (25.87%), and the post-monsoon (45.92%) season samples. The values of the MPN count of all bacterial isolates collected from all seasons ranged from 113 to 321 x 106 CFU/gram dry soil.
Biochemical Characterization
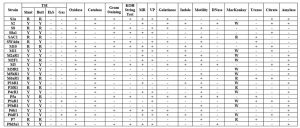
The biochemical characteristics of all the isolates are compiled in Table 1. Among the isolates, nine were Gram positives and rest were Gram-negative bacteria. Fermentation of sugar as analysed by TSI tests revealed a varying degree of fermentation potentiality amongst the isolates, in both aerobic and anaerobic conditions. All the strains tested were DNase negative and were unable to produce H2S in TSI media.
Table 1. Biochemical Test results of the Isolates
 Antibiogram
Antibiogram
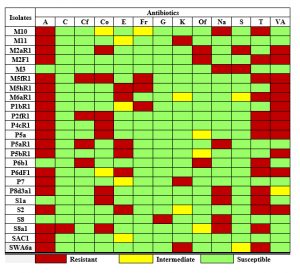
The results of the antibiogram of the bacterial strains are given in Table 2
(in a color-coded format). Isolate S. marcescens strain M5hR1 was most resistant against six antibiotics followed by P. vermicola strain P2fR1, B. subtilis S8a1, E. coli strain M2aR1, which were resistant against five different antibiotics each. Thirteen strains showed suppressed growths in Erythromycin, Furazolidone, Kanamycin, Ofloxacin, and Vancomycin of which four strains showed intermediate results in Erythromycin. Nearly all of the isolates were susceptible to Cotrimoxazole and Gentamycin except B. pumilus strain S8 and B. subtilis S8a1 respectively.
Table 2. Antibiogram results of twenty-four isolates from the soil sample against thirteen different Antibiotics. The table is colour coded according to the legend given below
[A – Ampicillin, C – Chloramphenicol, Cf – Ciprofloxacin, Co – Cotrimoxazole, E – Erythromycin, Fr – Furazolidone, G – Gentamycin, K – Kanamycin, Of – Ofloxacin, Na – Nalidixic acid, S – Streptomycin, T – Tetracycline, VA – Vancomycin.]
Fig. 2. Tree showing the phylogenetic relationship amongst the fish pathogenic strains. A total of 75 nucleotide sequences were used involving 180 positions in the final dataset
Molecular Characterization and phylogenetic analyses
BLASTn results revealed the identity of the isolates and taxonomic names were assigned based on the nearest neighbour of the NCBI database. Table 3 depicts the BLASTn analysis of the bacterial strains. Phylogenetic analysis of the fifteen pathogenic strains is given in Figure 2 and the optimal sum of the branch length was found to be 3.522. The phylogenetic relationship of the nine probiotic isolates is shown in Figure 3 with an optimal sum of branch length 1.478. Both the trees are drawn to scale and are shown as legend in the figure.
Table (3):
The assigned taxonomic name and Max Score of the twenty-four bacterial strains along with GenBank accession numbers.
Isolate |
Assigned Taxonomic Name |
Max Score |
GenBank Accession Number |
|---|---|---|---|
M10 |
Bacillus thuringiensis |
1565 |
MT254902 |
M11 |
Aeromonas enteropelogenes |
1790 |
MT254904 |
M2aR1 |
Escherichia coli |
1724 |
MT279647 |
M2F1 |
Pseudomonas aeruginosa |
1502 |
MT254901 |
M3 |
Bacillus flexus |
1714 |
MT279648 |
M5fR1 |
Acinetobacter junii |
1729 |
MT279649 |
M5hR1 |
Serratia marcescens |
1744 |
MT279650 |
M6aR1 |
Stenotrophomonas maltophilia |
1716 |
MT279675 |
P1bR1 |
Escherichia coli |
1766 |
MT254905 |
P2fR1 |
Providencia vermicola |
1775 |
MT256362 |
P4cR1 |
Enterobacter cloacae |
1729 |
MT279676 |
P5a |
Bacillus flexus |
1838 |
MT279679 |
P5aR1 |
Klebsiella pneumoniae |
1772 |
MT254946 |
P5bR1 |
Serratia marcescens |
1759 |
MT279682 |
P6b1 |
Bacillus cereus |
1692 |
MT279685 |
P6dF1 |
Aeromonas hydrophila |
1186 |
MT254947 |
P7 |
Comamonas aquatica |
1434 |
MT254973 |
P8d3a1 |
Staphylococcus aureus |
933 |
MT279691 |
S1a |
Bacillus flexus |
1725 |
MT279688 |
S2 |
Aeromonas punctata |
1555 |
MT254982 |
S8 |
Bacillus pumilus |
1620 |
MT254983 |
S8a1 |
Bacillus subtilis |
1482 |
MT254984 |
SAC1 |
Comamonas aquatica |
1236 |
MK849930 |
SWA6a |
Bacillus cereus |
1555 |
MF554643 |
Fig. 3. Tree showing the phylogenetic relationship amongst the probiotic strains. A total of 45 nucleotide sequences were used involving 709 positions in the final dataset
Probiotic and pathogen analyses
From several strains isolated from the enrichment cultures, post identification it was found that fifteen of the isolates (S2, SAC1, M5fR1, M11, M2aR1, M5hR1, M6aR1, P6dF1, P7, P1bR1, P5aR1, P8d3a1, P5bR1, P4cR1, and P2fR1) are opportunistic pathogens of Indian major carps, exotic carps, pangasiid catfishes and cichlid fishes cultivated in the farm., whereas nine isolates (S8, S8a1, S1a, SWA6a, M2F1, M10, M3, P6b1, and P5a) have possible probiotic values related to aquaculture.
With the help of online literature databases, a review was done highlighting the pathogenic effect of the isolates on fish species cultivated in the farm and also to ascertain the efficacy of the species isolated as probiotics. Data obtained from the literature survey is summarised for the pathogens and probiotics in Table 4 and Table 5 respectively. Notably, among the isolates, three species are belonging to genus Aeromonas were found that cause Aeromoniasis, which is by far the worst affecting disease, causing high fish morbidity in aquaculture fish cultivation scenario and is advised by several citations35–39. However, it is also appealing that among the probiotics isolated, several of them are reported to impart resistance against the pathogens isolated from the same source as well as boosting the immunity of the cultured fish specimens40–43.
Table (4):
List of bacterial strains isolated in this study that are earlier reported to cause pathogenesis among common fish species cultivated in freshwater aquaculture farms.
Sl. No. |
Isolate |
Identified as |
Pathogen of |
Pathogenesis |
Reference |
|---|---|---|---|---|---|
1 |
M11 |
Aeromonas enteropelogenes |
L. rohita |
• Motile Aeromonad septicemia • Ulceration in gastrointestinal mucosa • Produce putative toxins that enhances the chance of other opportunistic infections |
37,39 |
2 |
M2aR1, P1bR1 |
Escherichia coli |
All major carps |
• Found in intestinal tracts, gills muscle, and skin. • Indicator of water pollution • Transfers antibiotic resistance to several other fish pathogens in the intestinal milieu. |
44,59 |
3 |
P2fR1 |
Providencia_ vermicola |
L. rohita |
• Ulceration and reddish colouration in abdominal surfaces, the base of the pectoral fins and also on the cephalic region |
60 |
4 |
P5aR1 |
Klebsiella_ pneumoniae |
C. carpio, L. rohita |
• Haemorrha-gic subcutaneous ulcer and red spots in the body |
46,47 |
5 |
P6dF1 |
Aeromonas hydrophila |
Wide variety of freshwater fishes including all Indian major carps and exotic carps. |
•Motile Aeromonad septicemia • Tail/ fin rot • Epizootic Ulcerative Syndrome |
35,38,39 |
6 |
P7, SAC1 |
Comamonas aquatica |
L. rohita |
• Tail fin rot. |
61 |
7 |
S2 |
Aeromonas punctata |
C. carpio Labeo rohita |
•Haemorrhagic septicemia • Epizootic Ulcerative Syndrome |
36 |
8 |
M5fR1 |
Acinetobacter junii |
L. rohita, Hypophthal- michthys molitrix |
•Emerging fish pathogen • Causes septicemia leading to death |
62 |
9 |
M5hR1,P5br1 |
Serratia marcescens |
Variety of fishes |
• A probable pathogen of fishes causes high morbidity. • Opportunistic pathogen derived from highly polluted environment |
63 |
10 |
M6aR1 |
Stenotrophomonas maltophilia |
Catfishes |
• Surface discoloration with hemorrhages. • Tail fin rot, inflammation of internal organs |
64 |
11 |
P8d3a1 |
Staphylococcus aureus |
Hypophthal- michthys molitrix |
• Causes high morbidity due to severe eye infection and ultimately damaging the brain |
65 |
12 |
P4cR1 |
Enterobacter cloacae |
Mugil cephalus |
• Causes high fish morbidity. Pathogenicity caused by unspecified cationic factors. |
45,66 |
Table (5):
List of bacterial strains isolated in this study that are earlier reported to act as probiotics among common fish species cultivated in freshwater aquaculture farms.
Sl. No. |
Isolate |
Identified as |
Pathogenesis |
Reference |
|---|---|---|---|---|
1 |
M10 |
Bacillus thuringiensis |
• Maintenance of water quality. • Better conversion of organic matters by progressing the biogeochemical cycles. • Shows antimicrobial activity against several fish pathogens. • Possess immunostimulant activity against other pathogens • Immunity boosting activity when part of fish diet |
9,51,54,67–69 |
2 |
M2F1 |
Pseudomonas aeruginosa |
• Provide disease resistance and immunity against several pathogens • Can be used in conjunction with other bacterial pathogens with enhanced effect |
43,70,71 |
3 |
S8 |
Bacillus pumilus |
• Imparts resistance to A. hydrophila infections • Have a modulatory effect on the gut microbiota on dietary supplementation • It improves the growth rate and health of several fish species. |
42,58,72,73 |
4 |
S8a1 |
Bacillus subtilis |
• Enhances growth rate, boost immunity and impart disease resistant when used as a dietary supplement • Disease resistance against Vibriosis. • Produces a wide range of antibacterial substances and imparts resistance against Aeromoniasis. |
10,74–76 |
5 |
S1a, M3, P5a |
Bacillus flexus |
• Imparts disease resistance against pathogens • Produce cellulolytic and lipolytic enzymes and contributes to biogeochemical cycles |
41,77,78 |
6 |
SWA6a, P6b1 |
Bacillus cereus |
• Imparts resistance to A. hydrophila infections in L. rohita. • Can be used as a dietary supplement is several types of aquaculture |
10,79 |
This study reveals several pathogens and probiotic species present in the benthic soil of the aquaculture farm under study. The farm cultivates Indian and exotic major carps along with some catfishes and cichlid fishes and there is ample chance that any of these fish species may get affected by the pathogens revealed in this study. The presence of pathogens in a fish farm is never evaluated until or unless a disease breakout occurs with high fish morbidity44. Mass fish morbidity is reported caused by pathogens like A. hydrophila10,35,45, A punctata36, K. pneumonia46,47 causing a huge loss in revenue. Bacterial isolates identified in this study revealed that the benthic soil sample collected from post-monsoon season harbors a huge number of opportunistic pathogens with A. hydrophila being the most abundant. It is also noteworthy that, in comparison with the other two seasons, diversity and abundance of the opportunistic pathogens are much higher in the post-monsoon season sample as the occurrence of new genera of pathogens viz. Klebsiella, Enterobacter and Providencia were noticed along with strains of Escherichia and Serratia. This high abundance of diverse opportunistic pathogens could be attributed to the mixture of urban sewage and rainwater runoff from adjacent areas during monsoon and post-monsoon seasons.
Several antibiotic-resistant strains have been reported by scientists due to the indiscriminate antibiotic use in aquaculture farms48,49 and also pathogens like E. coli can transmit the resistance to other pathogens possibly through horizontal gene transfer44. Since the study reveals the antibiotic susceptibility of the strains, indiscriminate use of antibiotics could be avoided and specific treatment measures can be implemented.
The prokaryotic probiotics present in soil and water also boost the immunity of culturable fishes8,50–53 against several pathogens, and thus, steps could be formulated for the aid of the probiotics. These probiotic bacteria can also act as a dietary supplement for the fish specimens, mitigating the problem raised by inert fish feeds43,54–56. Probiotic species also engage in the continuance of biogeochemical cycles by several enzymes that help in the biodegradation of detritus materials in the benthos18. Probable probiotic species isolated in this study produces enzymes like cellulase57, lipase (unpublished data), and hence could be forerunners in the maintenance of the aquatic ecosystem. These probiotics are often consumed from the soil by bottom feeders thus circulating them along the food chain and could alter the gut microcosm of fish beneficially58. Further, it was revealed that the presence of probiotics is maximum is pre-monsoon season and declines rapidly from the monsoon season and thereafter. This is because the farm practices liming and the addition of organic fertilizers once annually in the pre-monsoon season leading to the amplification of probiotics. The monsoon rains bring a lot of agricultural and domestic runoffs as well as sewage overloads reducing the quality of water and soil, thus diminishing the number of probiotics and leading to the growth of pathogens. Coliforms are also found abundant in the soil during monsoon and post-monsoon season. This is probably the first report that provides a more in-depth insight into the culturable microcosm from an aquaculture farm of East Kolkata Wetlands, emphasizing their roles as fish pathogens and probiotics. This project could lead to early detection of pathogens and formulation of remedial measures even before the onset of the fish pathogenesis. This could be done by formulating a suitable bioaugmentation program to reinforce the growth of probiotics and eradication of the pathogens ensuring sustainable and profitable aquaculture.
ACKNOWLEDGMENTS
The authors acknowledge infrastructural assistance of the Department of Zoology, University of Burdwan and Department of Life Sciences, Presidency University, Kolkata. This article contains data from a thesis to be submitted by the first author for the partial fulfilment of the Ph.D. degree from the University of Burdwan, India.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
The contribution of the authors are as follows: AD: Conceptualization, Methodology, Investigation, and Writing – Original Draft, SM: Conceptualization, Writing – Review & Editing, GCS: Supervision, NCS: Supervision and Project administration.
FUNDING
None.
ETHICS STATEMENT
Not applicable
AVAILABILITY OF DATA
The datasets generated are included in this manuscript and 16S rRNA gene sequences of the bacterial strains are available at GenBank (https://www.ncbi.nlm.nih.gov/genbank/) with the following accession numbers MT254902, MT254904, MT279647, MT254901, MT279648, MT279649, MT279650, MT279675, MT254905, MT256362, MT279676, MT279685, MT254946, MT279682, MT279679, MT254947, MT254973, MT279691, MT279688, MT254982, MT254983, MT254984, MK849930, MF554643.
- Chandra Das P. Multi-species farming of major and minor carps for enhancing fish production in freshwater aquaculture Protocol development for production of stunted fingerlings of carp View project. Indian J Fish. 2016;63(2):55-61.
Crossref - Hassan MA, Puthiyottil M, Karnatak G, Sharma AP. Toward the Blue Revolution in India : Prospects for Inland Open Waters. World Aquaculture. 2017:2015-2018.
- Jayasankar P. Present status of freshwater aquaculture in India – A review. Indian J Fish. 2018;65:157-165.
Crossref - Patra BC, Kar A, Bhattacharya M, Parua S, Shit PK. Freshwater fish resource mapping and conservation strategies of West Bengal, India. Spatial Information Research. 2017;25:635-645.
- Piedrahita RH. Reducing the potential environmental impact of tank aquaculture effluents through intensification and recirculation. Aquaculture. 2003;206(1-4):35-44.
Crossref - Martinez-Cordova LR, Emerenciano M, Miranda-Baeza A, Martinez-Porchas M. Microbial-based systems for aquaculture of fish and shrimp: An updated review. Reviews in Aquaculture. 2015;7(2):131-148.
Crossref - Krishnan M, Birthal P, Singh ND. Aquaculture Development in India : Problems and Prospects Aquaculture Development in India : Problems and Prospects. National Centre For Agricultural Economics And Policy Research. 2014.
- Balcazar JL, de Blas I, Ruiz-Zarzuela I, Cunningham D, Vendrell D, Muzquiz JL. The role of probiotics in aquaculture. Vet Microbiol. 2006;114(3-4):173-186.
Crossref - Gupta A, Gupta P, Dhawan A. Dietary supplementation of probiotics affects growth, immune response and disease resistance of Cyprinus carpio fry. Fish and Shellfish Immunology. 2014;41:113-119.
- Kumar R, Mukherjee SC, Prasad KP, Pal AK. Evaluation of Bacillus subtilis as a probiotic to Indian major carp Labeo rohita (Ham.). Aquaculture Research. 2006;37(2):1215-1221.
Crossref - Irianto A, Austin B. Probiotics in aquaculture. J. Fish Dis. 2002;25:633-642.
Crossref - Martinez-Porchas M, Vargas-Albores F. Microbial metagenomics in aquaculture: a potential tool for a deeper insight into the activity. Rev Aquac. 2017;9:42-56.
Crossref - Ezemonye LIN, Ogeleka DF, Okieimen FE. Lethal toxicity of industrial detergent on bottom dwelling sentinels. International Journal of Sediment Research. 2009;24:479-483.
Crossref - Haenen O, Nl OH. MAJOR BACTERIAL DISEASES AFFECTING AQUACULTURE FMM/RAS/298: Strengthening capacities, policies and national action plans on prudent and responsible use of antimicrobials in fisheries. Aquatic AMR Workshop. 2017;1:10-11.
- Mishra SS, Das R, Choudhary P, et al. Present status of Fisheries and Impact of Emerging Diseases of Fish and Shellfish in Indian Aquaculture. Journal of Aquatic Research and Marine Sciences. 2017:5-26.
- Cabello FC. Heavy use of prophylactic antibiotics in aquaculture: A growing problem for human and animal health and for the environment. Environ Microbiol. 2006;8(7):1137-1144.
Crossref - Porchas-Cornejo MA, Martinez-Cordova LR, Ramos-Trujillo L, Hernandez-Lopez J, Martinez-Porchas M, Mendoza-Cano F. Effect of promoted natural feed on the production, nutritional, and immunological parameters of Litopenaeus vannamei (Boone, 1931) semi-intensively farmed. Aquaculture Nutrition. 2011;17:1-16.
Crossref - Moriarty DJW. The role of microorganisms in aquaculture ponds. Aquaculture. 1997:333-349.
Crossref - Ghosh S. Wastewater-Fed aquaculture in East Kolkata Wetlands: State of the art and measures to protect biodiversity. Wastewater Management Through Aquaculture. Springer Singapore; 2018:119-137.
- Bhattacharya S, Ganguli A, Bose S, Mukhopadhyay A. Biodiversity, traditional practices and sustainability issues of East Kolkata Wetlands: A significance Ramsar site of West Bengal, (India). Research & Reviews in BioSciences. 2012;6(2):340-347.
- Dey D, Banerjee S. Ecosystem and Livelihood Support: The Story of East Kolkata Wetlands. Environment and Urbanization Asia. 2013;4:325-337.
Crossref - Sarkar S, Ghosh PB, Mukherjee K, Sil AK, Saha T. Sewage treatment in a single pond system at East Kolkata Wetland, India. Water Sci Technol. 2009;60(2):2309-2317.
- Pal S, Manna S, Aich A, Chattopadhyay B, Mukhopadhyay SK. Assessment of the spatio-temporal distribution of soil properties in East Kolkata wetland ecosystem (A Ramsar site: 1208). Journal of Earth System Science. 2014;123:729-740.
Crossref - Vieira FCS, Nahas E. Comparison of microbial numbers in soils by using various culture media and temperatures. Microbiological Research. 2005;160(2):197-202.
Crossref - Anon. Soil Survey Standard Test Method Soil Moisture Content. Standards Association of Australia. 1990:1-5.
- Janssen PH, Yates PS, Grinton BE, Taylor PM, Sait M. Improved culturability of soil bacteria and isolation in pure culture of novel members of the divisions Acidobacteria, Actinobacteria, Proteobacteria, and Verrucomicrobia. Appl Environ Microbiol. 2002;68:2391-2396.
Crossref - Garrity GM, Holt JG. The Road Map to the Manual. Bergey’s Manual® of Systematic Bacteriology. New York, NY: Springer New York; 2001:119-166.
- Bauer AW, Kirby WMM, Sherris JC, Turck M. Am. In vitro evaluation of antibacterial potential of Annona squamosa against bovine mastitis. J Clin Pathol. 1966;45:493-496.
- Sutton S, Ph D. Measurement of Cell Concentration in Suspension by Optical Density. The Microbiology Network. 2006:4-6.
- Kiehlbauch JA, Hannett GE, Salfinger M, Archinal W, Monserrat C, Carlyn C. Use of the National Committee for Clinical Laboratory Standards guidelines for disk diffusion susceptibility testing in New York State Laboratories. J Clin Microbiol. 2000;38:3341-3348.
Crossref - Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol. 2016;33:1870-1874.
Crossref - Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406-425.
Crossref - Tamura K, Nei M, Kumar S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA. 2004;101:11030-11035.
Crossref - Felsenstein J. Confidence Limits On Phylogenies: An Approach Using The Bootstrap. Evolution. 1985;39:783-791.
Crossref - Aoki T. Motile Aeromonads (Aeromonas hydrophila). Fish Diseases and Disorders. Viral. 1999;3(B):427–453.
- Toor HS, Sehgal HS, Sehdev RS. A case study of acute fish diseases in tanks loaded with high levels of organic manures. Aquaculture. 1983;35:277-282.
Crossref - Ramesh D, Souissi S. Antibiotic resistance and virulence traits of bacterial pathogens from infected freshwater fish, Labeo rohita. Microb Pathog. 2018;116:113-119.
Crossref - Sharma M, Shrivastav A, Sahni Y, Pandey G. Overviews of The Treatment and Control of Common Fish Diseases. International Research Journal of Pharmacy. 2012;3:123-127.
- Austin B, Austin DA, Austin B, Austin DA. Aeromonadaceae Representatives (Motile Aeromonads). Bacterial Fish Pathogens. 2012:119-146.
- Kumar R, Mukherjee SC, Prasad KP, Pal AK. Evaluation of Bacillus subtilis as a probiotic to Indian major carp Labeo rohita (Ham.). Aquaculture Research. 2006;37(12):1215-1221.
Crossref - Alamri SA. Biodegradation of microcystin-RR by Bacillus flexus isolated from a Saudi freshwater lake. Saudi J Biol Sci. 2012;19(4):435–440.
Crossref - Sun Y-Z, Yang H-L, Ma R-L, Lin W-Y. Probiotic applications of two dominant gut Bacillus strains with antagonistic activity improved the growth performance and immune responses of grouper Epinephelus coioides. Fish Shellfish Immunol. 2010;29:803-809.
Crossref - Giri SS, Sen SS, Sukumaran V. Effects of dietary supplementation of potential probiotic Pseudomonas aeruginosa VSG-2 on the innate immunity and disease resistance of tropical freshwater fish, Labeo rohita. FFish Shellfish Immunol. 2012;32:1135-1140.
Crossref - Del Rio-Rodriguez RE, Inglis V, Millar SD. Survival of Escherichia coli in the intestine of fish. Aquaculture Research. 1997;28:257-264.
Crossref - Austin, Brian, Dawn A. Austin, Brian Austin A, Dawn A. Austin. Bacterial fish pathogens. Springer. 2012.
Crossref - Das A, Acharya S, Behera BK, et al. Isolation, identification and characterization of Klebsiella pneumoniae from infected farmed Indian Major Carp Labeo rohita (Hamilton 1822) in West Bengal, India. Aquaculture. 2018;482:111-116.
Crossref - Oliveira RV, Peixoto PG, Ribeiro D de C, et al. Klebsiella pneumoniae as a main cause of infection in nishikigoi Cyprinus carpio (carp) by inadequate handling. Brazilian Journal of Veterinary Pathology. 2014;7:86-88.
- Singh AK, Rathore G, Singh V, et al. Bacterial resistance to oxytetracycline in different life stages of Indian freshwater carp aquaculture system. International Journal of Microbiology Research. 2009;1(1):25-34.
- Abraham TJ. Food safety hazards related to emerging antibiotic resistant bacteria in cultured freshwater fishes of Kolkata, India. Adv J Food Sci Technol. 2011;3:69-72.
- Nayak SK, Mukherjee SC. Screening of gastrointestinal bacteria of Indian major carps for a candidate probiotic species for aquaculture practices. Aquaculture Research. 2011;42(7):1034-1041.
Crossref - Mukherjee A, Dutta D, Banerjee S, et al. Potential probiotics from Indian major carp, Cirrhinus mrigala. Characterization, pathogen inhibitory activity, partial characterization of bacteriocin and production of exoenzymes. Res Vet Sci. 2016;108:76-84.
Crossref - Ghosh S, Sinha A, Sahu C. Isolation of putative probionts from the intestines of Indian major carps. Israeli Journal of Aquaculture – Bamidgeh. 2007;59(1):127-132.
- Boyd, Claude E., Wood CW, Thunjai. T. Aquaculture pond bottom soil quality management. Pond Dynamics/Aquaculture Collaborative Research Support Program, Oregon State University. 2002.
- Suguna P, Binuramesh C, Abirami P, et al. Immunostimulation by poly-a hydroxybutyrate-hydroxyvalerate (PHB-HV) from Bacillus thuringiensis in Oreochromis mossambicus. Fish Shellfish Immunol. 2014;36(1):90-97.
Crossref - Dawood MAO, Koshio S. Recent advances in the role of probiotics and prebiotics in carp aquaculture: A review. Aquaculture. 2016;454:243-251.
Crossref - Reneshwary C, Rajalakshmi M, Marimuthu K, Xavier R. Dietary administration of Bacillus thuringiensis on the cellular innate immune response of African catfish (Clarias gariepinus) against Aeromonas hydrophila. Eur Rev Med Pharmacol Sci. 2011;15(1):53-60.
- De A, Mukherjee S, Sadhukhan GC, Saha NC. An insight into the celluloytic potential of three strains of Bacillus spp. Isolated from benthic soil of aquaculture farms in East Kolkata Wetlands, India. J Pure Appl Microbiol. 2018;12(3):1597-1605.
Crossref - Sun YZ, Yang HL, Ma RL, Song K, Lin WY. Molecular analysis of autochthonous microbiota along the digestive tract of juvenile grouper Epinephelus coioides following probiotic Bacillus pumilus administration. J Appl Microbiol. 2011;110(4):1093-1103.
Crossref - Dutta C, Panigrahi AK, Sengupta C. Prevalence of Pathogenic Bacteria in Finfish and Shellfish Obtained from Domestic Markets of West Bengal, India. Frontiers in Environmental Microbiology. 2015;1(2):14-18.
- Ramkumar R, Ravi M, Jayaseelan C, et al. Description of Providencia vermicola isolated from diseased Indian major carp, Labeo rohita (Hamilton, 1822). Aquaculture. 2014;420-421,193-197.
Crossref - Sanyal KB, Mukherjee D, Guchhait A, Dash G. Phenotypic and Molecular Identification of Bacterial Species in Indian Major Carps and Exotic Carps from South 24 Parganas, West Bengal, India. Int J Curr Microbiol Appl Sci. 2018;7(1):534-547.
- Malick RC, Bera AK, Chowdhury H, et al. Identification and pathogenicity study of emerging fish pathogens Acinetobacter junii and Acinetobacter pittii recovered from a disease outbreak in Labeo catla (Hamilton, 1822) and Hypophthalmichthys molitrix (Valenciennes, 1844) of freshwater wetland in West Bengal, India. Aquaculture Research. 2020;51(6).
Crossref - Baya AM, Toranzo AE, Lupiani B, Santos Y, Hetrick FM. Serratia marcescens: a potential pathogen for fish. J Fish Dis. 1992;15(1):15-26.
Crossref - Abraham TJ, Paul P, Adikesavalu H, Patra A, Banerjee S. Stenotrophomonas maltophilia as an opportunistic pathogen in cultured African catfish Clarias gariepinus (Burchell, 1822). Aquaculture. 2016;450:168-172.
- Shah KL, Tyagi BC. An eye disease in silver carp, Hypophthalmichthys molitrix, held in tropical ponds, associated with the bacterium Staphylococcus aureus. Aquaculture. 1986;55(1):1-4.
Crossref - Sekar VT, Santiago TC, Vijayan KK, et al. Involvement of Enterobacter cloacae in the mortality of the fish, Mugil cephalus. Lett Appl Microbiol. 2008;46(6):667-672.
Crossref - Balcazar JL, Blas I de, Ruiz-Zarzuela I, Cunningham D, Vendrell D, Muzquiz JL. The role of probiotics in aquaculture. Vet. Microbiol. 2006;114(3-4):173-186.
Crossref - Patel AK, Ahire JJ, Pawar SP, Chaudhari BL, Chincholkar SB. Comparative accounts of probiotic characteristics of Bacillus spp. isolated from food wastes. Food Res Int. 2009;42(4):505-510.
Crossref - Mancuso M. Probiotics in Aquaculture. Journal of Fisheries & Livestock Production. 2014;02.
Crossref - Giri SS, Sukumaran V, Dangi NK. Characteristics of Bacterial Isolates from the Gut of Freshwater Fish, Labeo rohita that May be Useful as Potential Probiotic Bacteria. Probiotics and Antimicrobial Proteins. 2012;4(4):238-242.
Crossref - Giri SS, Sukumaran V, Sen SS, Jena PK. Effects of dietary supplementation of potential probiotic Bacillus subtilis VSG1 singularly or in combination with Lactobacillus plantarum VSG3 or/and Pseudomonas aeruginosa VSG2 on the growth, immunity and disease resistance of Labeo rohita. Aquaculture Nutrition. 2014;20(2):163-171.
Crossref - Yang HL, Xia HQ, Ye YD, Zou WC, Sun YZ. Probiotic Bacillus pumilus SE5 shapes the intestinal microbiota and mucosal immunity in grouper Epinephelus coioides. Dis Aquat Organ. 2014;111(2):119-127.
Crossref - Thy HTT, Tri NN, Quy OM, et al. Effects of the dietary supplementation of mixed probiotic spores of Bacillus amyloliquefaciens 54A, and Bacillus pumilus 47B on growth, innate immunity and stress responses of striped catfish (Pangasianodon hypophthalmus). Fish Shellfish Immunol. 2017;60:391-399.
Crossref - Touraki M, Karamanlidou G, Karavida P, Chrysi K. Evaluation of the probiotics Bacillus subtilis and Lactobacillus plantarum bioencapsulated in Artemia nauplii against vibriosis in European sea bass larvae (Dicentrarchus labrax, L.). World J Microbiol Biotechnol. 2012;28(6):2425-2433.
Crossref - Vaseeharan B, Ramasamy P. Control of pathogenic Vibrio spp. by Bacillus subtilis BT23, a possible probiotic treatment for black tiger shrimp Penaeus monodon. Lett Appl Microbiol. 2003;36(2):83–87.
Crossref - Liu C-H, Chiu C-H, Wang S-W, Cheng W. Dietary administration of the probiotic, Bacillus subtilis E20, enhances the growth, innate immune responses, and disease resistance of the grouper, Epinephelus coioides. Fish Shellfish Immunol. 2012;33(4):699-706.
Crossref - Sarjito, Haditomo AHC, Ariyati RW, Sabdaningsih A, Desrina, Prayitno SB. Screening of potential isolate candidates probiotic against Aeromonas hydrophila from Boyolali, Indonesia. Journal of Physics: Conference Series. IOP Publishing; 2019:12147.
- Cai Y, Yuan W, Wang S, et al. In vitro screening of putative probiotics and their dual beneficial effects: To white shrimp (Litopenaeus vannamei) postlarvae and to the rearing water. Aquaculture. 2019;498:61-71.
- Nayak SK. Probiotics and immunity: A fish perspective. Fish Shellfish Immunol. 2010;29(1):2-14.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.