ISSN: 0973-7510
E-ISSN: 2581-690X
Actinomycetes are a suitable microbial group for the synthesis of lignocellulose-degrading enzymes. Enzymes that may degrade organic material, including cellulose, hemicellulose, and lignin, are released by actinomycetes. The aim of this research was to isolate actinomycetes from Rajkot, Gujarat, India’s soil and evaluate the activity of their cellulase and Xylanase enzymes. Starch Casein Agar (SCA) was used to identify a total of 30 isolates of actinomycetes. A qualitative plate assay (CMC-Na, Congo red) revealed that the highest zone of catalysis for MMD1 was 36 mm. Five strains were discovered to be effective for quantitative quantification of endoglucanase utilising filter paper and CMC as substrates: MMD1, MMD2, MMD3, MMD4, and MMD8. Following MMD 1 (endoglucanase 5.4 IU; FPase 4.4 IU), MMD 2 (endoglucanase 4.5 IU; FPase 3.4 IU) has demonstrated considerable endoglucanase and FPase activity. Beechwood xylan was used to treat sugarcane bagasse in order to test Xylanase, and 45% of the xylan (hemicellulose) fraction was obtained. MMD1 and MMD2 measured the xylanase enzyme activity (4.8IU and 4.2IU) in quantitative and qualitative assays (34 mm and 22 mm for BWX and 32 mm and 14 mm for agro-waste xylan). The strain MMD1 was identified as Streptomyces chartreusis through morphological, biochemical, and finally molecular characterization by 16S rRNA sequencing. It was then submitted to NCBI GenBank with the accession number MT254830.
Actinomycetes, Lignocelluloses, Endoglucanase, FPase, Sequencing
One of the most abundant renewable non-fossil carbon sources on earth is lignocellulose, an essential component of plant cell walls. It is widely distributed in waste from agriculture, fruits and vegetables, forests, and cities. Using less expensive lignocellulosic material as the major carbon source and other energy sources can lead to the economic and environmentally friendly production of bioethanol and other bioproducts. Lignocellulose as one of the most abundant renewable biomasses from green and dry vegetation usually comprised of cellulose, hemicellulose, and lignin.1 The hydrolysis of cellulose is carried out by endoglucanases (EC 3.2.1.4), b-glucosidase (EC 3.2.1.21) cellobiohydrolase (EC. 3.2.1.91) and exoglucanases (EC 3.2.1.74) respectively.2 Besides myriad microbial sources as lignocelluloses degraders3,4 actinomycetes a saprobe3 and a unique Gram-positive, filamentous prokaryote with high GC content and extracellular enzyme, secondary metabolites secretion properties was also studied as a resource organism for lignocellulose degradation by many researchers5 and the search still in progress either in culture dependent6 or independent7 manner. Efficient hemicellulose degraders generally reduce the need of commercial xylanase other hemicellulose degrading enzymes with specific substrate specificity. The prime aspects which had influenced to carry out the present study was an exploration for an efficient lignocellulose degrading actinobacteria especially Streptomycetes capable enough with multi lignocellulose synthesis potential and enhanced degradation ability of the renewable wastes abating pollution and be used as significant strain of choice in biofuel production in a cost-effective manner.
Sample collection
The two different soil samples were collected from agricultural field longitude 22.2344° N, Latitude 70.8004° E near Rajkot, Gujarat, India. Overnight, soil samples were air-dried and then incubated at 50°C.
Isolation of actinomycetes
Actinomycetes were isolated using serial dilution method on Starch Casein Agar Medium. 10 ml of distilled water taken in test tube, 1 gram of soil was weighed and thoroughly mixed. Then, it was serially diluted up to a 10-5 dilution. After that, 0.1 ml taken from the dilution of 10-3 and 10-5 and spread on the Starch Casein Agar (SCA) plate. The plates were incubated for 7–14 days at 30°C. Actinomycete isolates that have grown on SCA media Containing (g/L) Soluble Starch-10.0, Casein-0.30, MgSO4.7H2O-0.05, KNO3 -2.0, K2HPO4 -2.0, NaCl-2.0, FeSO4.7H2O-0.01, CaCO3 -0.02, Agar 18.0 Final pH at 30°C: 7.3±0.2 have been chosen based on their morphology. The chosen isolates were separated by further streaking on additional SCA plates. Then, for further testing, a pure colony of actinomycetes was kept in 10% glycerol at -80°C.8
Screening of Cellulase producing actinomycetes strain
Qualitative screening was performed with the Carboxy Methyl Cellulose (CMC)(g/L) Containing carboxymethyl cellulose (CMC) 10.0, K2HPO4 -2.0, peptone 10.0, MgSO4.7H2O-0.3, (NH4)2SO4-2.5, gelatine 2.0 and agar 18 and pH was adjusted at 6.8-7.2, By the point inoculation 30 Plates were incubated at 30°C for 7 days after catalytic zone was observed and intensified with consecutive flooding by 1% (w/v) Congo-red solution9 for 15 minutes and destaining with 1 M NaCl for 15 minutes.10 The area surrounding the selected colonies was measured.11
Synthesis of cellulase enzyme
Cellulase Producing Isolates are (MMD1, MMD2, MMD3, MMD4 and MMD8). This production culture assay for checking endo glucanase, FPase activity. These organisms were inoculated liquid culture containing 1% CMC, 0.1% K2HPO4, 1% NaNO3, 0.05% MgSO4.7H2O, 0.1% KCL,0.05% Yeast extract (pH 7). The developing culture was then moved into the production medium of the same composition and incubated for a week at 30°C and 120 rpm in an orbital shaker.10 The Production media was centrifuge for 10 minutes at 10,000 rpm to get cell free supernatant. This cell-free supernatant is used for cellulase activity. At every interval of 24 hr crude enzyme extracts of all the selected isolates were undergone assay for CMCase and FPase.11
Cellulase assay
The Cellulase activity was carried out using the method described by Maravi and Kumar12 performed by a reaction mixture was made up of 0.1 ml of crude enzyme extract and 1 ml of 0.5% CMC that had been produced in 50 mM phosphate buffer (pH 7). To create a volume of 2.0 ml The tube was filled with 0.9 ml of 50 mM phosphate buffer pH 7.0. The mixture was then further incubated in a water bath for 30 minutes at 50°C and then add 3.0 ml of dinitrosalicylic acid (DNSA)13 reagents to stop the reaction. For FPase Whatman filter paper (No.1) strip was used as a substrate the absorbance was recorded at 540 nm. Unit of endoglucanase or Fpase activity defined as amount of enzyme that liberated 1 µmol of glucose per minute. Unit of endoglucanase or Fpase activity defined as amount of enzyme that liberated 1µmol of glucose per minute. The absolute amount of glucose (0.5 mg/ml) plotted against 540 nm was used to create a linear graph of glucose for the enzyme unit calculation. A glucose standard curve was created using a standard formula to measure the concentration of released glucose and the activity of the enzymes.11
CMCase Activity= 0.185/(Enzyme concentrtion to release 0.5 mg glucose) × units ml–
Enzyme activity (FPU/ml)= (Sugar Concentration)/(MW of glucoseXT) × H/E
Where MW of Glucose is (180g/mol), incubation time t (minutes), and E is volume of enzyme (ml), H is total volume of enzyme- substrate (ml)
Determination of lignocellulose Composition in Sugarcane bagasse and ground nutshell
Sugar cane bagasse (SCB) and ground nutshell were collected from APMC Yard (Rajkot, Gujarat). Approximately 1 gm (Weight A) of SCB and ground nutshell was dried. The sample was allowed to air dry before being combined with 150 ml of distilled water and benzene alcohol. The combination was then refluxed at 100°C for one hour in a water bath. After filtered out, 300 ml of hot water was applied to rinse the reflux products. After the filtering procedure, Once the residue achieved a constant weight (weight B), it was dried in an oven. After adding 150 ml of 1 N H2SO4 to the dry residue, the mixture was refluxed once more for one hour at 100°C in a water bath. After the reflux products were filtered, the residue was cleaned with distilled water until the pH was neutral. After that, the residue was dried until it reached a constant weight (Weight C). After the addition of the dried residue, 100 ml of 72% H2SO4 were added, and the combination steeped at room temperature (27°C) for four hours. After then, the residue was filtered again. After adding 150 ml of 1 N H2SO4, the filtered residue was refluxed for one hour at 100°C in a water bath. The reflux products were filtered and then washed with distilled water after their pH reached a neutral level. After the residue reached a consistent weight, it was dried in an oven at 105°C (Weight D). The cellulose and hemicellulose content of ground nutshell and SCB can be calculated using the following formulas.
The obtained pellet was then used for Xylanase screening, both qualitatively and quantitatively.12
Hemicellulose(%)= b – c/a × 100%
Cellulose(%)= c-d /a × 100%
Screening of xylanase producing actinomycetes strain
Qualitative screening was performed with the Beech wood xylan (SRL) Containing (g/L) KH2PO4 3.0; CoCl2·6H2O 0.02, MgSO4·7H2O 0.5; CaCl2·2H2O 0.1; K2HPO4 2.0; FeSO4·7H2O 0.05; MnSO4·4H2O 0.02; and NH4Cl 10.0 agar 18 grams supplemented with Beech wood Xylan 1% (w/v) and pH was adjust at 6.8 to 7.0. xylan-degrading Potent colonies were detected by point inoculation and the Plates were incubated at 30°C for 7 days. The zone of hydrolysis was observed by Congo-red solution 0.1% (w/v) for 30 minutes and destaining with 1 M NaCl for 15 minutes. According to the prescribed procedure, the catalytic zone was measured.12
Synthesis of xylanase enzyme
Xylanase Producing Isolates (MMD1, MMD2, MMD3, MMD4 and MMD8). This production culture assay for checking Xylanase activity. These organisms were inoculated liquid culture containing (g/L) KH2PO4 3.0; K2HPO4 2.0; KH2PO4 3.0; CaCl2·2H2O 0.1; FeSO4·7H2O 0.05; MnSO4·4H2O 0.02; CoCl2·6H2O 0.02; MgSO4·7H2O 0.5 and NH4Cl 10.0 and 1% w/v Beech wood xylan as sole carbon source. The developing culture was then moved into the production medium of the same composition and incubated for a week at 30°C and 120 rpm in an orbital shaker. The Production media was centrifuge for 10 minutes at 10,000 rpm to get cell free supernatant. This cell-free supernatant is used as Xylanase activity. At every interval of 24 hr crude enzyme extracts of all the selected isolates were undergone assay for Xylanase.
Xylanase assay
The reaction mixture used in the experiment consisted of 1 ml of 0.5% Beech Wood Xylan and SCB that had been prepared in 50 mM phosphate buffer (pH 7) and 0.1 ml of crude enzyme extract. 0.9 ml of 50 mM phosphate buffer pH 7.0 was added to the tube to make a volume of 2.0 ml. The mixture was then incubated in a water bath for 30 minutes at 50°C. To stop the reaction, 3.0 ml of dinitrosalicylic acid (DNSA)13 reagent was added to the reaction tube.
Biochemical characterization of MMD1
A few significant biochemical tests, including IMViC (Indole, Methyl red, Voges Proskaur, Citrate utilization), hydrolysis of starch, gelatine liquefication, and nitrate reduction,14,15 was carried out to identify the genus of MMD1 comparing with criteria stated in Bergey’s Manual of Systematic Bacteriology.16
Identification and 16S rRNA sequence analysis and phylogenetic tree construction of the Potent Actinomycetes strain MMD1.
The Gram’s Staining was used to determine the morphology of the isolated actinomycetes. The Molecular identification of the Cellulase and Xylanase producing actinomycetes isolates was carried out by 16S rRNA Gene sequencing. The 16S rRNA sequence analysis was performed by Gene explore Diagnostics and Research Centre Pvt Ltd. in Ahmadabad. Following established protocols, a number of procedures were carried out, including DNA extraction from MMD1, PCR amplification, sequencing, BLAST analysis against the GenBank database (the first fifteen sequences were chosen and aligned using multiple sequence alignment software based on maximal identity score), and inferring evolutionary history using p-distance and the neighbour joining method. 2,17-19 The phylogenetic tree was generated and drawn to scale using the maximum likelihood approach, which measures branch lengths in the same units as evolutionary distances. Positions with holes and incomplete data were all removed. MEGA7 was used to conduct evolutionary analysis and determine evolutionary divergence between sequences. The proportion of duplicate trees in which the related taxa clustered together is shown next to the branches in the bootstrap test (1000 repetitions).19 Streptomyces sp. MMD1 16S rRNA sequence was uploaded to NCBI GenBank with a modification in coding as F (19)2.
Isolation of actinomycetes
In the current study, from two different agriculture soil sites, a total of 30 Actinomycetes strains were isolated.
Screening of Cellulase producing actinomycetes strain by rapid plate assay
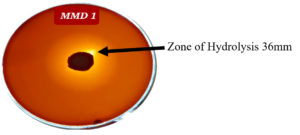
The most effective five actinomycetes strains (MMD1, MMD2, MMD3, MMD4, MMD8) developed hydrolytic zones on CMC-Congo red agar plates and were chosen for quantitative analysis out of 30 actinomycetes strains (Table 1). Compared to other strains, the MMD1 strain shown the largest zone of cellulose breakdown (36 mm) (Figure 1). The MMD7 strain shown the lowest zone of cellulose breakdown (0.04 mm).
Table (1):
Zone of hydrolysis for cellulase and Xylanase synthesis potent strains of actinomycetes
| Actinomycetes Potent Strain | Zone of Hydrolysis of Cellulase (mm) | Zone of Hydrolysis of Xylanase (mm) | Zone of Hydrolysis of Xylanase using SCB (mm) | |
|---|---|---|---|---|
| MMD1 | 36** | 34** | 32** | |
| MMD2 | 18 | 22 | 14 | |
| MMD3 | 16 | 11 | 14 | |
| MMD4 | 5 | 8 | 6 | |
| MMD5 | 6 | 9 | 5 | |
| MMD6 | 8 | 5 | 6 | |
| MMD7 | 0.4* | 0.5* | 0.3* | |
| MMD8 | 31 | 33 | 30 | |
| MMD9 | 6 | 8 | 7 | |
Footnote- **-highest zone of hydrolysis; * Lowest zone of hydrolysis.
Cellulase enzyme activity of potent actinomycetes strains
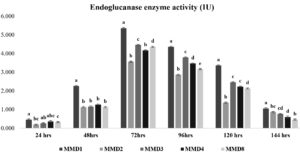
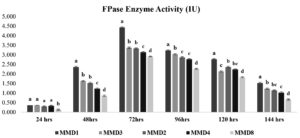
The endo—1,4-glucanase activity of five selected actinomycetes strains was quantified, and MMD1 was found to have the highest activity (5.4 IU), followed by MMD2 (4.5 IU), MMD4 (4.4 IU), MMD3 (4.2 IU), and MMD8 (3.6 IU) during the optimal 72 hours of incubation. The MMD1 enzyme activity profile revealed a rise in activity from 0.3 to 5.4 IU after 24 hours and a subsequent drop (Figure 2). The FPase at an optimal incubation time of 72 hours, the potent actinomycetes strains MMD1, MMD8, MMD2, MMD3, and MMD4 had displayed enzyme activities of 4.4 IU, 3.3 IU, 3.4, 3.9 IU, and 2.9 IU, respectively. While the most effective strain, MMD1, showed enhanced enzyme activity from 24 to 120 hours of incubation (Figure 3).
Figure 2. Endoglucanase enzyme activity of isolated potent Actinomycetes (MMD1 Streptomyces chartreusis) at 7 days of incubation period. Letters denote significant variations using Tukey’s test (P<0.05)
Figure 3. FPase enzyme activity of isolated potent Actinomycetes (MMD1 Streptomyces chartreusis) at 7 days of incubation period. Letters denote significant variations using Tukey’s test (P<0.05)
Screening of xylanase employing actinomycetes strain using beech wood xylan and extraction of treated xylan from SCB
The two potential actinomycetes strains (MMD1 & MMD2) displayed 34 mm and 22 mm, respectively, of xylanase catalysis during 72 hours of fermentation. Similar to this, using hemicelluloses (xylan) residue from treated sugarcane bagasse, MMD1 and MMD2 showed 32 mm and 14 mm of xylanase catalytic zone at 72 hours of fermentation duration, respectively.
Xylanase enzyme activity by selected actinomycete strains from beech wood xylan and extraction of treated Xylan from SCB
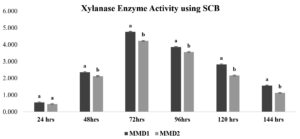
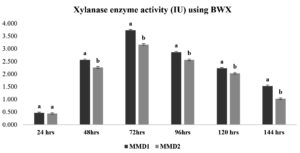
After treating sugarcane bagasse, 45% of the hemicellulose was removed (Figure 4). The quantitative experiment using treated SCB xylan fraction showed xylanase activity of 3.7 IU and 3.2 IU by MMD1 and MMD2, respectively, at an optimal 72-hour incubation period (Figure 5). While a quantitative analysis using beech wood xylan as a synthetic carbon source with MMD 1 and MMD 2 had found to be recorded with zone of 4.8 IU and 4.2 IU, respectively, at an optimal 72 hours of incubation period (Figure 6).
Figure 5. Xylanase enzyme activity using (Treated Sugarcane Bagasse) of isolated potent Actinomycetes (MMD1 Streptomyces chartreusis) at 7 days of incubation period. Letters denote significant variations using Tukey’s test (P<0.05)
Figure 6. Xylanase enzyme activity using (Beech wood xylan) of isolated potent Actinomycetes (MMD1 Streptomyces chartreusis) at 7 days of incubation period. Letters denote significant variations using Tukey’s test (P<0.05)
Morphological and biochemical characteristics of potential actinomycetes MMD1
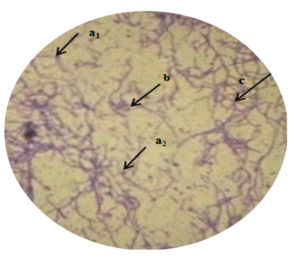
In the present study, selected potent strains Actinomycetes MMD1 had shown a preliminary characteristic property of the genus Streptomyces sp. as being represented in Table 2 and 3 by morphological and biochemical features. The colony characteristics were found to be recorded medium size, irregular in shape, round margin, powdery and rough consistency, and texture, raised elevation respectively. The aerial mycelia were whitish grey, and no reverse pigmentation observed. Light microscopic view of Gram staining of MMD1 in Figure 7 has shown the characteristic properties of Streptomyces sp. by the presence of branched aerial mycelia, sporophore in varied spirals and terminal of spore (Figure 8). Table 3 has exhibited the positive observation of different biochemical tests, like catalase, starch hydrolysis, gelatine liquification and nitrate reductions, while negative for Indole, MR, VP, and citrate utilization respectively.
Table (2):
Morphological Characteristics of Actinomycetes strain MMD1
Parameters |
Properties |
|---|---|
Size |
Medium |
Shape |
Irregular |
Margin |
Round |
Texture |
Powdery or Rough |
Elevation |
Raised |
Aerial mycelia colour |
Whitish grey |
Aerial mycelia morphology |
Loose spirals and tight spirals in sporophore morphology |
Substrate mycelia |
Yellowish white |
Spore/conidia |
Spore intercalary or terminal |
Gram staining |
Gram-Positive |
Table (3):
Biochemical Characteristics of Potent Strain MMD1
Biochemical tests |
Interpretations |
|---|---|
Indole |
Negative |
Methyl Red (MR) |
Negative |
Voges Proskauer (VP) |
Negative |
Citrate Utilization |
Negative |
Catalase test |
Positive |
Starch hydrolysis |
Positive |
Gelatine liquefaction |
Positive |
Nitrate reduction |
Positive |
Sugar Hydrolysis |
Positive |
Nitrogen utilization |
Positive |
Figure 7. Light microscopic image of Gram staining view of Actinomycetes strain MMD1 under 100x (a1, a2- Position of spore; b-Coiling of aerial mycelia; c- Branched aerial mycelia)
Figure 8. Scanning electron microscopy (SEM) of potent actinomycetes strain MMD1(A Magnification 5000x, Cell width 9.42 mm, B- Magnification 13000x, Cell width 9.02 mm)
Taxonomic characterization of Streptomyces sp. MMD1
In the present study of 16S rRNA sequence analysis a total data set of 16 nucleotide sequences with codon positions (1st+2nd+3rd+Noncoding) were recorded along with a total final dataset of 592 positions. The optimal tree with the sum of branch length = 4.91337812 is shown in Figure 9. The phylogenetic tree revealed that, the test isolate Streptomyces MMD1 had got 99.50% similarity with the type of strain Streptomyces chartreusis strain NBRC 12753 & confirmed the potential strain as Streptomyces chartreusis. NCBI GenBank accession number for the nucleotide sequence of the potential isolate is MT254830.
Streptomyces along with other actinomycetes, are commonly mentioned as efficient degraders of lignocellulosic biomass. In our study we have used Agro waste, broth culture, and synthetic medium-based rapid plate tests. The Two soil samples was collected from agricultural fields and dumping site for actinomycetes isolation and 30 actinomycetes colonies were isolated. Sukmawaty et al.20 reported the 15 actinomycetes colonies isolated from the soil of Malindo pine forest, South Sulawesi. MMD1 able to produce higher zone of hydrolysis (36 mm) of endoglucanase compare with the Lenni et al.21 reported the zone of hydrolysis 2.93 mm of endoglucanase by streptomyces spp. The formation of a clear zone surrounding the colony showed actinobacteria’s capacity to produce cellulase enzymes. Hydrolysing cellulose into glucose is possible with isolates that generate cellulase. High cellulase production was indicated by the large clear zone that appeared.22 The process of Quantitative screening of Cellulase from actinomycetes strain MMD1 able to produce much higher levels of endoglucanase (5.4 IU/ml) and FPase (4.4 IU/ml) similar results were also found in the Mohanta et al.23 they have reported 0.734±0.001 IU/mL for FPase and 1.381±0.024 IU/mL for CMCase. Budihal et al.24 have reported under optimized condition sorghum stover was found to be better production of cellulase under submerge 38 IU/ml and solid state 44 IU/ml bioprocess. According to the all-parameter Streptomyces spp having good efficiency to degrade cellulose.
The process of quantitative screening of both the artificial beech wood xylan and the agrowaste (treated sugarcane bagasse) xylan fraction had demonstrated quite satisfactory xylanase activity at 72 hours of incubation, which is also thought to be the beginning of the sporulation phase and the production of secondary metabolites (enzymes) in actinomycetes. The same isolate MMD1 has demonstrated positive confirmation for xylanase comparable cellulase and is also suggested by Brito et al.25, owing to the capacity of plant polymer (as a carbon source) degradation. Sporck et al.26 support the increased xylanase activity using xylan residue that has been recovered from heat- or alkali-treated agricultural wastes like rice straw, sugarcane bagasse, etc. Actinomycetes strains MMD1 and MMD2 were found to have 3.7 IU/ml and 3.2 IU/ml of xylanase activity, respectively, when hemicellulose (a source of xylan) was subjected to consecutive acid, alkali, and heat treatment in sequence in the current study. This is in stark contrast to beechwood xylan (a synthetic source), which had 4.8 IU/ml and 4.2 IU/ml. While the Xiuting et al.27 reported under optimized condition xylanase activity of 12 U/ml) with birch wood xylan and Streptomyces chartreusis strain L1105, which is quite higher than the present result of the study for Xylanase enzyme production.
Using a 100x magnification, isolates were examined, and Figure 7 shows the microscopic observations. Different forms of aerial mycelium were detected in isolates of Actinomycetes under a microscope reported Dhanasekaran et al.28 According to Maheswari et al.29, the aerial mycelium of the genus Streptomyces is flexible, spiral, straight, or open loops. Spores of Streptomyces are non-motile. Initially smooth, Streptomyces colonies can develop into aerial mycelium that can have a variety of appearances, including floccose, granular, powdery, or velvety.30 Our result shows the similarity with these Streptomyces chartreusis mycelium is straight, spores are non-motile, and colonies are powdery.
The isolate of actinomycete MMD1 was tentatively assigned to the genus Streptomyces based on morphological and biochemical characteristics for confirmation of genus identification by 16S rRNA Sequencing. current investigation to confirm the Streptomyces MMD1 species as Streptomyces chartreusis MMD1 matching (99.50) with Streptomyces chartreusis BRC12753T. In previous research by Wang et al.31, this strain was also investigated for drought tolerance in sugarcane fields using Streptomyces chartreusis WZS021. With this strain, laccase, endo alpha 1,3 glucanase, etc. have all been previously examined.
Actinomycetes are recognised as a powerful source of a number of significant commercial enzymes, including cellulase and xylanase. Streptomyces chartreusis MMD1 (NCBI GenBank submission number MT254830) has been identified and characterised as an effective cellulose and xylan degrader after a series of experiments exploring significant lignocellulose degraders that started with five of the best actinomycetes strains and culminated in one strain, MMD1. A series of treatments on sugarcane bagasse resulted in 45% hemicellulose yield, and the use of xylanase activity that was good but not as effective as Beechwood xylan (BWX) was discovered. Future applications for the lignocellulolytic degradation property include bioethanol generation, solid waste management, consortia formation for increased degradation, comparative omics level investigations and bioremediation of contaminated sites. application studies in pulp, paper, and petroleum industries and so on.
ACKNOWLEDGMENTS
The authors would like to acknowledge the Department of Education, Government of Gujarat, for providing SHODH Fellowship, Gene Explore Diagnostic lab Ahmedabad for genetic analysis and Junagadh Agricultural University for SEM analysis. The authors are also thankful to the Director, Research Innovation and Translation, Atmiya University and the Head of the Microbiology Department, Atmiya University Rajkot for providing Infrastructure and facilities. The authors are also thankful to Ms. Unnati B. Yagnik, Research Scholar, Department of Microbiology, for the manuscript revision.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
Both authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This study was supported by the Government of Gujarat SHODH fellowship under the UGC Scheme.
DATA AVAILABILITY
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
ETHICS STATEMENT
Not applicable.
- Limayem A, Ricke SC. Lignocellulosic biomass for bioethanol production: current perspectives, potential issues, and future prospects. Progress in Energy and Combustion Science. 2012;38(4):449-467.
Crossref - Semedo LTAS, Gomes RC, Bon EPS, Coelho RRR, Soares RMA, Linhares LF. Endocellulase and exocellulase activities of two Streptomyces strains isolated from a forest soil. Appl Biochem Biotechnol. 2000;84:267-276.
Crossref - Saini RK, Sivanesan I, Keum IY. Phytochemical of Moringa oleifera; a review of their nutritional, therapeutic and industrial significance. 3 Biotech. 2016;6(2):203.
Crossref - Shukla R, Kumar M, Chakraborty S, et al. Process development for the production of bioethanol from waste algal biomass of Gracilaria verrucosa. Bioresource Technol. 2016;220:584-589.
Crossref - Carolina S, Alcazar A Physiological properties, modification, and application of starches from different botanical sources. Food Sci Technol. 2015;35(2):215-236.
Crossref - Saitou N, Nei M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406-425.
- Cecchini L, Torquati B, Chiorri M. Sustainable agri-food products: A review of consumer preference studies through experimental economics. Agric Ecom-Czech. 2018 64(12):554-565.
Crossref - Kuster E, Williams ST. Selection of Media for Isolation of Streptomyces. Nature. 1964;202:928-929.
Crossref - Teather RM, Wood PJ. Use of Congo red-polysaccharide interactions in enumeration and characterization of cellulolytic bacteria from the bovine rumen. Appl Environ Microbiol. 1982;43(4):777-780.
Crossref - Bhagat S, Kokitkar S. Isolation, and identification of bacteria with cellulose-degrading potential from soil and optimization of cellulase production. J Appl Biol Biotechnol. 2021;9(6):154-161.
Crossref - Ghose TK. Measurement of cellulase activities. Pure Appl Chem. 1987;59(2):257-268.
Crossref - Maravi P, Kumar A. Isolation, screening, and identification of cellulolytic bacteria from soil. Biotechnol J Int. 2020;24(1):1-8.
Crossref - Gram C. The differential staining of schizomycetes in tissue sections and in dried preparations. Forischritte der Medicine. 1884;2:185-189.
- Shirling EB, Gottlieb D. Methods for characterization of Streptomyces sp. Int J Syst Bacterial. 1966;16(3):313-340.
Crossref - Marmur J. A procedure for the isolation of deoxyribonucleic acid from microorganisms. J Mol Biol. 1963;3(2):208-218.
Crossref - Cui XL, Mao PH, Zeng M, Li WJ, Zhang LP, Xu LH, Jiang CL. Streptimonospora salina gen. Nov., sp. Nov., a new member of the family Nocardiopsaceae. Int J Syst Evol Microbial. 2001;51(2):357-336.
Crossref - Nei M, Maruyama T, Chakraborty R. The bottleneck effect and genetic variability in populations. Evolution. 1975;29(1):1-10.
Crossref - Tamura K, Nei M, Kumar S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci U S A. 2004;101(30):11030-11035.
Crossref - Miller GL. Use of dinitro salicylic acid reagent for determination of reducing sugar. Anal Chem. 1959;31(3):426-428.
Crossref - Sukmawaty E, Sari SR, Masri M. Characterization of soil Actinomycetes from Malino pine forest rhizosphere of South Sulawesi. Elkawnie J Islam Sci Tech. 2020;6(2):315-328.
Crossref - Fitri L, Bessania MA, Septi N, Suhartono S. Isolation and characterization of soil actinobacteria as cellulolytic enzyme producer from Aceh Besar, Indonesia. Biodiversitas. 2021;22:5169-5180.
Crossref - Pesrita A, Linda TM, Devi S. Selection and activity of local Riau Actinomycetes cellulase enzymes on sugarcane bagasse lignocellulosic media. Jurnal Riau Biologia 2. 2017.
- Mohanta YK. Isolation of cellulose-degrading actinomycetes and evaluation of their cellulolytic potential. Bioeng Biosci. 2014;2(1):1-5.
Crossref - Budihal SR, Agsar D. Exploration of agrowastes for the production of cellulase by Streptomyces DSK29 under submerged and solid-state systems. Int J Curr Microbiol App Sci. 2015;4(11):681-689.
- Brito CCQ, Campos ITN, Faria FP, Bataus LAM. Screening and Xylanase production by Streptomyces sp. Grown on lignocellulosic Wastes. Appl Biochem Biotech. 2013;170(3):598-608.
Crossref - Sporck D, Reinoso FAM, Rencoret J, et al. Xylan extraction from pretreated sugarcane bagasse using alkaline and enzymatic approaches. Biotechnol Biofuels. 2017;10:296.
Crossref - Li X, Sun B, Zhao J, et al. Production and improved bleaching abilities of a thermostable xylanase from a newly isolated Streptomyces chartreusis strain. Afr J Biotechnol.2011;10(64):14132-14142.
Crossref - Dhanasekaran D, Yi J. Actinobacteria: Basics and Biotechnological Applications. IntechOpen, London. 2016.
Crossref - Maheswari MU, Chandra TS. Production and potential applications of a xylanase from a new strain of Streptomyces cuspidosporus. W J Microbiol Biotechnol. 2000;16(3):257-263.
Crossref - Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F. The Prokaryotes: Actinobacteria. Springer, Berlin. 2014.
Crossref - Wang Z, Solanki M, Yu Z, Yang L, Dong D, Yang L. Drft Genome analysis offers insights into the mechanism by which Streptomyces chartreusis WZS021 increases Drought Tolerance in Sugarcane. Front Microbio. 2019;9:3262.
Crossref
© The Author(s) 2024. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.