Diseases caused by Rickettsiales are often overlooked, although they pose important public health concerns. The Rickettsiales family comprises a broad range of intracellular bacteria with distinct evolutionary adaptations, making the development of treatment measures to combat infections, such as vaccines or antibiotics, a challenge. Interestingly, the outer membrane protein A (OmpA) was found to exist in the cell surface of most human pathogenic bacteria in the order Rickettsiales. However, knowledge about OmpA in each species and strain is scattered and ambiguous. In this study, we systematically compiled the existing information on OmpA and its relationship with human pathogenic rickettsiae to serve as a reference for future research. A comprehensive literature search was conducted using specific keywords across five databases. According to the literature, OmpA of spotted fever group rickettsia plays a crucial role as an adhesin and invasin that directly interacts with the surface of mammalian host cells to mediate bacterial localization in host cells. The presence of a premature stop codon in the amino acid sequence resulted in the secretion of non-functional OmpA, which is one of the main reasons for rickettsial strains or species to become avirulent. Similarly, OmpA also functions as an important adhesin in the Anaplasma family when it interacts with the sLex and sLex-like glycan of myeloid and endothelial cells, respectively. However, the OmpA of Anaplasma must be co-functional with the other two adhesins to promote bacterial internalization. Interestingly, certain sites in the amino acid residues of Ehrlichia and Orientia OmpA are predicted to be homologous to the binding domain region of Anaplasma OmpA. It is therefore suggested that OmpA is an important adhesin for bacteria to bind to their specific mammalian host cells.
Rickettsia, Outer Membrane Protein, Rickettsial Disease, Pathogenesis
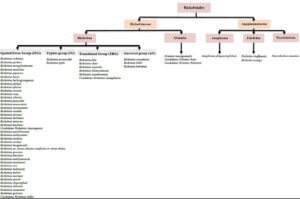
Based on the phylogenetic study, the Rickettsiales consist of three families, the Rickettsiaceae, Anaplasmataceae and Candidatus Midichloriaceae.1 While the Rickettsiaceae and Anaplasmataceae have been extensively studied due to their pathogenicity towards humans, Candidatus Midichloriaceae has been largely overlooked as it primarily affects corals and aquatic animals. Although two members of Candidatus Midichloriaceae (Candidatus Midichloria mitochondrii and Candidatus Lariskella arthropodarum) have been found in symbionts with ticks and in human blood, there is still insufficient evidence to suggest its pathogenicity towards humans.2,3 Therefore, Candidatus Midichloriaceae is often excluded from discussions on Rickettsial disease.
Rickettsiaceae is divided into two genera, Rickettsia and Orientia, which are differentiated by their distinct cell wall composition and structure. The bacteria in the genus Rickettsia are classified into four major antigenic groups: the spotted fever group (SFG), the typhus group (TG), the transitional group (TRG), and the ancestral group.1,4 SFG, TG, and TRG consist of pathogenic members of Rickettsia, which are the causative agents for the most frequently reported and deadly cases of Rickettsiosis.5 SFG represents the largest number of Rickettsia species, with more than 30 species having been discovered and new species emerging every year. Figure 1 illustrates all the known species of SFG that are potentially pathogenic to humans. However, following an extensive electronic literature search, we found numerous new species that have been reported under SFG and could potentially be harmful to humans. These include Candidatus Rickettsia mendelii,6,7 Candidatus Rickettsia andeanae,8,9 Candidatus Rickettsia tarasevichiae,10,11 Candidatus Rickettsia jingxinensis,12 Candidatus Rickettsia longicornii,12 Candidatus Rickettsia colombianensi, Rickettsia asembonensis, Rickettsia murinus, Candidatus Rickettsia barbariae,13 Candidatus Rickettsia wissemanii,14 Candidatus Rickettsia takensis, Candidatus Rickettsia laoensis, Candidatus Rickettsia Thierseensis, Candidatus Rickettsia Moyalensis, Candidatus Rickettsia uralica, Rickettsia sibirica mongolitimonae, Candidatus Rickettsia nicoyana,15 and Candidatus Rickettsia xinyangensis.16
A novel Orientia species, Candidatus Orientia chiloensis, has also been discovered, adding a new member to the existing Orientia genera, Orientia tsutsugamushi and Orientia chuto.17 As for the family Anaplasmataceae, there are four genera classified in this family which are Ehrlichia, Anaplasma, Neorickettsia, and Wolbachia. Anaplasma phagocytophilum, Ehrlichia chaffeensis, Ehrlichia ewingii, and Neorickettsia sennetsu are the species that have been shown to infect humans and cause the threatening human Anaplasmosis, Ehrlichiosis and Neoehrlichiosis.18 In addition, a new human pathogenic Ehrlichia species has also been discovered, namely Candidatus Ehrlichia erythraense.19 The emergence of various novel species under Rickettsiaceae and Anaplasmataceae is alarming, as there is still a lack of knowledge about developing effective vaccines that could provide heterologous protection for most of the current species in these families.
The antigenic heterogeneity of bacteria in the order Rickettsiales is the main reason that contributes to the difficulty in developing a vaccine, even though they are predicted to produce numerous virulence factors that may interact with host cells.20,21 Antigenic variation among strains of certain species is known to be one of the mechanisms for intracellular bacteria to establish persistent infection within the host cell.22 Therefore, identifying the most antigenic protein that is universally present throughout all the strains in a species and has immunogenicity properties is one of the early steps during vaccine development. While searching for antigenic protein in Rickettsiales in the literature databases, the OmpA protein always appeared in the search. The outer membrane protein A (OmpA) is a known virulence factor of gram-negative bacteria. It involves multiple stages of disease pathogenesis, including acting as the invasin and adhesin to facilitate the internalization of bacteria to the host cells through ligand-receptor interaction.23 However, the overall information on OmpA in certain genera in the order Rickettsiales is still unclear, especially in Orientia and Ehrlichia. It is expected that proteins annotated under similar “names” may not have similar homology between genera and should not be grouped. The DNA and protein sequence of OmpA varies greatly among different genera in Rickettsiales, as determined by bioinformatic analyses. However, in some manuscripts, certain sites in the amino acid sequence are discovered that show high similarity among different genera and these are used to predict the protein tertiary structure using in silico tools.24,25 This piques our curiosity to learn more about the OmpA protein in diverse types of bacteria classified as Rickettsiales. Therefore, the primary objective of this study is to compile all known information about the OmpA of bacteria in Rickettsiales. The information gathered in this review may be valuable for other researchers who intend to start research on the OmpA of any Rickettsiales member.
Survey methodology
Literature search strategy
An extensive literature search was conducted using five databases: Web of Science, Scopus, PubMed, Google Scholar, and Wiley Online Library. The first search occurred on August 9, 2022, the second search on July 31, 2023, and the third search on May 10, 2024. All databases’ alert has been enabled to receive any new linked publications. The search was undertaken to look for original research publications produced in English between 2000 and 2024 on the involvement of OmpA protein in the pathogenesis of all human Rickettsiales illnesses. The keywords used in the search are the combination of the word “Outer membrane protein A or OmpA” with all species and strains of Ricketsiales members including the disease names. A newly found or emerging genus, species, or disease may not be included in the keywords since its existence is unknown to the date this review was conducted. The duplicate manuscripts were removed by using EndNote 20 software.
Inclusion and exclusion criteria
The initial selection of potential manuscripts was based on the abstract and a quick view of the body of the manuscripts. All original research manuscripts published between 2000 and 2024 possess the keywords mentioned above were included in the review. The exclusion criteria: (i) manuscript used OmpA as a molecular detection for either identification of human infection, tick identification, geographical study, or phylogenetic tree construction; (ii) review manuscripts; (iii) non-relevant manuscript that does not have the keywords in the abstract and body of the manuscript; (iv) Books, letters to the editor, conference proceedings; and (v) non-English manuscripts.
Screening Process
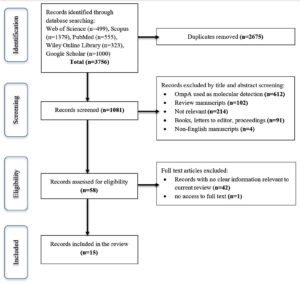
A flowchart of the study screening process is presented in Figure 2. A total of 3756 records were initially detected, resulting in a total of 1081 records after the duplicate was removed. Following this, 1023 records were removed after screening the title and abstract, as well as a cursory look at the body of the manuscripts. Of the 1023 records, 612 records showed OmpA was used as a marker for molecular detection, either for the identification of human infection or tick identification for geographical study or phylogenetic tree construction. While 102 records were review manuscripts, 91 were either books, letters to the editor, or conference proceedings; 4 were non-English manuscripts; and 214 were non-relevant as the manuscripts do not have the search keywords in the abstract and body of the manuscript. A total of 43 records were further eliminated, resulting in 15 records being included in this review.
Out of 15 records, three (3) manuscripts explained how the existence of variation in the OmpA DNA sequence of Rickettsia species or strains impacts their virulence. The avirulent Rickettsia OmpA (rOmpA) DNA sequence has either a deletion, an insertion, or a transversion, resulting in premature stop codons during translation and the lack of functional rOmpA protein synthesis (Table 1). While two (2) manuscripts discuss the post-translational process essential for the maturation of the rOmpA.26,27 Interestingly, two (2) records indicate the presence of substantial similarity in the OmpA-predicted tertiary structure of Orientia tsutsugamushi and Ehrlichia chaffeensis with Anaplasma phagocytophilium, implying a probable comparable function during disease pathogenesis.24,25 In addition, four (4) manuscripts show the known interaction of OmpA with the targeted human host cells and describe the particular region in OmpA that is involved in the interaction (Table 2). However, three (3) manuscripts suggested OmpA might not be involved in the virulence of Rickettsia during in vivo study or mice model.28-30 Similarly, one (1) manuscript explains that degradation of Ehrlichia chaffeensis’s OmpA does not affect the bacterial ability to adhere to its host cell, monocyte.31
Table (1):
Location of variation in OmpA gene sequence
| Origin of the OmpA use in comparison | Author and titles | Location of variation in the OmpA sequence | Effect of the variations in OmpA protein secretion | Further effects |
|---|---|---|---|---|
| Rickettsia peacockii vs Rickettsia rickettsii | Baldridge et al.32
|
i. Deletion of “G” at base pair 403 ii. Insertions of “GT” at base pair 4872 iii. Insertions of “A” at base pair 5828 |
• Presence of three identical premature stop codons • Absence translation of OmpA during SDS PAGE and immunoblot assay |
• Rickettsia peacockii unable to induce actin tail in the host |
| transversion of “G” to “C” nucleotide at base pair 61 | • Change the putative sequence “AAGG” (as in Rickettsia rickettsii OmpA) to “AAGC”. • Weakens the consensus sequence of ribosome binding, which disturbs the translation efficiency of OmpA protein in Rickettsia peacockii |
|||
| Rickettsia rickettsii strain Iowa vs strain Morgan, Sheila Smith and R | Clark et al.33 | i. Single nucleotide deletion at 660 bp ii. Deletion of downstream 891 bp nucleotide |
• Absence production of OmpA protein during detection by using western blot | Guinea pigs used as animal model show no fever response upon infection with Rickettsia peacockii |
| Rickettsia rickettsii strain Iowa vs strain Sheila Smith | Ellison et al.34 | i. Single nucleotide deletion at 660 bp ii. An 891bp in-frame deletion located within the repeat region of OmpA |
• Introduction of a stop codon at amino acid 184, resulting in a premature stop during translation
• Absence of OmpA analysed using Immunoblotting and immunofluorescence |
• No increment of temperature in the guinea pig infected with lowa strain, in comparison with Sheila Smith • Iowa strains show a significant lower release of total lactate dehydrogenase [LDH] into the medium, in comparison with R strains, at days 6 & 7 of infection, which indicates lowa has a deficiency in the lysis of host cells • Mutation in OmpA does not affect the ability of Ricketsia rickettsii Iowa to form actin tails |
Table (2):
Outer membrane protein A (OmpA) interaction with the human host cells
| Origin of the OmpA | Author and titles | Targeted human host cells | Location of OmpA involved in interaction | Host protein interacts with OmpA |
|---|---|---|---|---|
| Rickettsia rickettsii | Sahni et al.49 | Human Dermal Microvascular Endothelial cells | β-peptide region | Fibroblast growth factor receptors 1/ Heparan sulfate proteoglycans [FGFR1/HSPGs] |
| Rickettsia conorii | Hillman et al.48 | Human Microvascular Lung Endothelial | C-terminal region, amino acid position 954-1735, rOmpA(954-1735) | First analysis: filamin-A, epiplakin 1 and cytoplasmic dynein 1 heavy chain 1 Further analysis: α2β1 integrin |
| Anaplasma phagocytophilum | Ojogun et al.43 | Myeloid cells (HL-60) | N-terminal region, amino acid position 59-74, ApOmpA(59-74) | sialyl Lewis x (sLex-capped P-selectin glycoprotein ligand 1) |
| Endothelial cells (RF/6A) | N-terminal region, amino acid position 59-74, ApOmpA(59-74) | sLex-like molecule, 6-sulfo-sLex | ||
| Anaplasma phagocytophilum | Seidman et al.54 | Myeloid cells (HL-60) | K64 residue ApOmpA(59-74) G61 residue ApOmpA(59-74) |
α2,3-sialic acid residue of sLex α2,3-sialic acid & α1,3-fucose residue of sLex |
| Endothelial cells (RF/6A) | K64 residue ApOmpA(59-74) G61 residue ApOmpA(59-74) |
α2,3-sialic acid residue of sLex-like molecule, 6-sulfo-sLex α2,3-sialic acid & α1,3-fucose residue of sLex-like molecule, 6-sulfo-sLex |
The existence of OmpA in the human pathogenic Rickettsiales
In the genus Rickettsia belongs to the family Rickettsiaceae, the gene sequence of OmpA exists throughout all the members of SFGR, not only in the virulence species such as Rickettsia rickettsii and Rickettsia conorii but also found in the avirulence Rickettsia peacockii.32-35 For this reason, most genetic studies have used the OmpA gene region to distinguish different Rickettsia species.35 OmpA contains species-specific epitopes that provide the basis for rickettsial serotyping crucial for species identification and construction of phylogenetic trees.36 The Rickettsia OmpA (rOmpA), also known as surface cell antigen 0 (Sca0), is one of the five autotransporter proteins in Rickettsia. All the autotransporter proteins in this family will generally have three functional domains, which are (i) amino-terminal leader sequence, (ii) secreted mature (a) protein, and (iii) a carboxy-terminal (b) domain.37 The amino acid sequences of the b-peptides of all the autotransporters are predicted to be highly homologous and might undergo a similar post-translational process.27,37 rOmpA has been shown to have a similar post-translational process as the OmpB (Sca5), in which the proteolytic cleavage occurs at the b-peptide region of the carboxy-terminal. This explains why rOmpA usually appears as a 190 kDa protein during SDS-PAGE analysis. This differs from the predicted size calculated based on the amino acid sequence, 224 kDa protein precursor. The cleavage occurs at the carboxy-terminal side of Ser-1958, producing a 190 kDa protein and a small ~32 kDa protein fragment.27 This post-translational process is essential for the maturation of the rOmpA, and it is facilitated by aspartic protease or specifically APRc (Aspartic Protease from Rickettsia conorii) enzyme.26 The similarities in processing mature rOmpA with OmpB, a well-known autotransporter of SFGR, and its high abundance expression show the importance of rOmpA in the pathogenesis of SFGR. However, the typhus group, another member of the genus Rickettsia, does not possess rOmpA. Typhus group bacteria might have different mechanisms in their pathogenesis than SFGR.
In genus Orientia, another member of the family Rickettsiaceae, the OmpA sequence was found highly conserved in 51 isolates of Orientia tsutsugamushi originating from several Asia-Pacific geographical areas. Interestingly, the OmpA of Orientia tsutsugamushi (OtOmpA) is transcriptionally expressed in mice infected with Karp and Gilliam strains.24 The expression was detected in the mice’s blood and organs, such as the kidney, liver, lung and spleen, suggesting these are the target organs of Orientia tsutsugamushi. In vitro study also found OtOmpA protein being expressed by Orientia tsutsugamushi Ikeda during infection of mammalian host cells (HeLa cells), suggesting the involvement of OtOmpA in the pathogenesis of Scrub typhus.38 Interestingly, the non-translated upstream promoter region of the OtOmpA gene was also investigated. It is confirmed that OtOmpA is transcribed in human mammalian cells, specifically in epithelial (HeLa), endothelial (RF/6A), and monocytes (THP-1) at different stages of infection.39
Anaplasma phagocytophilum and Ehrlichia chaffeensis belong to the family of Anaplasmataceae and also have been found to express OmpA. The OmpA is expressed predominantly by the DCs form of the organisms.40 DCs stand for “dense-cored”, which indicates the infectious form of Anaplasma phagocytophilum and Ehrlichia chaffeensis.41,42 Both of these bacteria have a unique biphasic life cycle. It undergoes reversible transitioning between an infectious dense-cored (DC) form and a noninfectious, replicative reticulate cell (RC) form. During the early stage of infection, DC form will attach and invade the host cells. Then, this DCs form will differentiate into RCs form and start to replicate by binary fission. This resulted in the formation of Morula, a bacterium-filled organelle. Later, the RCs will transform back to DCs form, and a new cycle of infection will be initiated.41,42 In Anaplasma phagocytophilum, OmpA gene expression was found to be upregulated during binding and invasion of Anaplasma phagocytophilum DCs form to the myeloid cells (HL-60 cells) and during transmission to mice by ticks infected with Anaplasma phagocytophilum.43 In fact, the recombinant OmpA can also be recognized by the sera of patients with human granulocytic anaplasmosis (HGA).43 While, in Ehrlichia chaffeensis, both mRNA and protein of OmpA are expressed in the infected monocyte cells (THP-1 cells) and were upregulated at the late stage of the bacterium’s intracellular growth. Furthermore, the immunofluorescent labelling technique has confirmed the existence of OmpA as one of the surface-exposed proteins. The anti-OmpA bound to the surfaces of paraformaldehyde-prefixed nonpermeabilised host-cell free Ehrlichia chaffeensis, confirming the localisation of OmpA in the cell surface.31 Therefore, it is clear that OmpA has been found to exist in most of the human pathogenic bacteria in the order Rickettsiales, with the exception of the Rickettsia’s typhus group.
Functional rOmpA secretion is important for the pathogenesis of spotted fever group rickettsia
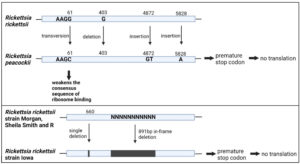
As discussed above, the rOmpA sequence exists throughout all the members SFGR. However, there are some interspecies variations in the rOmpA DNA sequence, which influence the conformation and functionality of the secreted rOmpA protein. Secretion of functional rOmpA is crucial in determining the virulency of rickettsia. A study by Baldridge et al. shown that, even though Rickettsia peacockii carry a high degree of similarity of rOmpA sequence with virulent Rickettsia rickettsii, there is no rOmpA translation product detected in its protein extract during in vitro infection of tick cells (Dermacentor andersoni cell line DAE100).32 This could be due to the presence of multiple premature stop codons within the rOmpA reading frame, that could interfere with the translation of the rOmpA (Table 1). Furthermore, sequence analysis by Baldridge et al. revealed a weakened ribosome binding site within the mRNA leader, adding another factor that interrupts the production of rOmpA protein.32 Similarly, there is also deficient expression of the rOmpA protein in the only reported avirulent strain of Rickettsia rickketsii, Iowa.33,34 This is believed to be due to the single nucleotide deletion at 660 bp upstream of rOmpA gene, which shifts the open reading frame of the rOmpA, introducing a stop codon, leading to protein truncation (Table 1). Apart from that, rOmpA sequence of Iowa also lacks downstream 891 bp sequence (in-frame insertion of 297aa) in comparison with the virulence Rickettsia rickettsii strain: (i) Morgan, (ii) Sheila Smith, and (iii) R.33,34
Figure 3 illustrates the location of interspecies variations in the OmpA DNA sequence. Immunoblotting assay using western blot confirmed the absence of the OmpA protein from Rickettsia rickettsii Iowa and Rickettsia peacockii, suggesting that the truncated product was either rapidly degraded or not recognized by monoclonal antibodies.
As a result, no polymerization of the host cell’s actin occurs during the infection, thus preventing the adhesion of the Rickettsia peacockii to the host cells.32 Actin-based motility is essential for rickettsia species to propel themselves from one host cell to another.34 Rickettsia movement disrupts the host cells’ actin scaffolding and facilitates rickettsia penetration of the internal and external membranes of the host cell.44 However, the mutation in rOmpA sequence in the Iowa strain of Rickettsia rickettsii does not defect the formation of its actin tails. But Iowa strain does have deficiency in lysis of the host cells.34 Iowa strains show a significantly lower release of total lactate dehydrogenase (LDH), compared to pathogenic R strains on days 6 & 7 of infection, indicating Iowa has a deficiency in the lysis of host cells. As a result, animal models (guinea pigs) infected with Iowa strains show no febrile response after the infection.34 However, there is no solid confirmation as to whether the deficiency of rOmpA has any correlation with the fever response and the ability to lyse the host cell.
rOmpA mediated the adherence and invasion of rickettsia to endothelial host cells
As an intracellular bacterium, it is crucial for rickettsia to invade and reside in the host cells in order to survive. Microvascular endothelial cells in the blood vessel lining have been known as the primary target of pathogenic rickettsia when infecting a human host.45 Localization of rickettsia in the vascular endothelial cells causes injury to the cell, leading to vasodilation by infiltering the perivascular mononuclear cells. This causes the increment of fluid in the interstitial space, which later leads to other serious complications depending on the targeted organs.46 Therefore, the ability of rickettsia to adhere and invade the host cells is very crucial during the early stages of rickettsia pathogenesis, which is mediated by specific ligand-receptor interaction.47 Functional rOmpA has been postulated to play the role of the ligand during the interaction. A study shown that the recombinant soluble C-terminal rOmpA (amino acid located at positions 954 to 1735) of Rickettsia conorii has the ability to bind to mammalian host cells, the human microvascular lung endothelial (HMVEC-L).48 The adherence ability of this soluble rOmpA(954 to 1735) has been confirmed, as it competitively inhibits Rickettsia conorii infection on HMVEC-L cells by 50%.
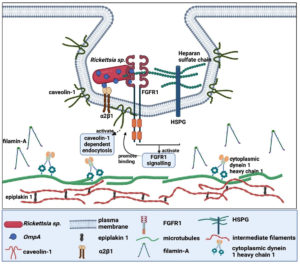
To fully understand the ligand-receptor interaction, two original articles have identified the host cell receptor that interacts with the OmpA. The most recent publication by Sahni et al., discovered the interaction between the β-peptide region of Rickettsia rickettsii OmpA with the fibroblast growth factor receptors 1/ heparan sulfate proteoglycans (FGFR1/HSPGs) complexes of the Human Dermal Microvascular Endothelial cells.49 This interaction activates FGFR1 signaling, leading to the binding of FGFR1 to caveolin-1, which could activate caveolin-1 dependent endocytosis and promote the invasion of Rickettsia rickettsii. The second manuscript found three potential proteins from HeLa cells that have interaction with rOmpA(954 to 1735) of Rickettsia conorii, namely filamin-A, epiplakin 1 and cytoplasmic dynein 1 heavy chain 1 (Table 2).48 However, all of these proteins are not directly associated with the plasma membrane, suggesting that they are not the potential candidate for the host cell receptor. The author later postulated that filamin-A, the most abundant species detected in the analysis, might form an adhesion complex with
b1 integrin. Further analysis of HMVEC-L suggests that a2b1 integrin is the mammalian receptor that interacts with OmpA of Rickettsia conorii.48 Figure 4 illustrates the possible ligand-receptor interaction between the OmpA of Rickettsia sp. and human cells.
Figure 4. Ligand-receptor interaction of Rickettsia sp. OmpA with human cells (Created with BioRender.com)
ApOmpA is the adhesin that facilitates the binding of Anaplasma phagocytophilum to mammalian host cells
Similar to rickettsia, Anaplasma phagocytophilum must invade host cells in order to survive. Neutrophil in peripheral blood and tissues are known to be the primary target of Anaplasma phagocytophilum and cause human granulocytic anaplasmosis (HGA).50 Internalization of Anaplasma phagocytophilum induces proinflammatory responses by increasing the secretion of pro-inflammatory cytokines (e.g., interleukin-10, IL-12, IL-1b and IFN-gamma).51 This promotes neutrophil degranulation and deactivation, leading to tissue injury. As a result, the neutrophils are unable to exert an effective antimicrobial response to the infection.52 In addition, Anaplasma phagocytophilum has also been found to affect the progenitors of myeloid, monocytic lineages and endothelial cells.
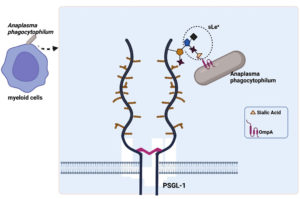
In vitro studies on myeloid (HL-60) and endothelial (RF/6A) cells have confirmed the adhesion capacity of ApOmpA. Interestingly, the N-terminal region of ApOmpA, which contains the protein’s extracellular domain, plays a role as a binding domain in the interaction with the host cells (Table 2). ApOmpA at amino acid position 59-74, ApOmpA(59-74) was found to interact with the a2,3-sialic acid of the sialyl Lewis x (sLex) tetrasaccharide that caps P-selectin glycoprotein ligand-1 (PSGL-1) on the surface of myeloid cells (HL-60) (Figure 5).43 It was known that Anaplasma phagocytophilum binds to and interacts with the sialic acid residue of neutrophils.53 Demolishment of sialic acid residues from myeloid cell surfaces by the use of sialidase has successfully inhibited the adhesion and infection of Anaplasma phagocytophilum.43 The molecular docking model prediction has identified three residues of ApOmpA(59-74), K64, G61 and K60, which interact with the sLex glycan. K64 was predicted to bind the a2,3-sialic acid, while G61 was predicted to bind the a2,3-sialic acid and the a1,3-fucose residue of sLex.
Figure 5. Binding of ApOmpA(59-74) with α2,3-sialic acid of the sialyl Lewis x (sLex) tetrasaccharide that caps P-selectin glycoprotein ligand-1 (PSGL-1) on myeloid cell (HL-60) surfaces (Created with BioRender.com)
However, the endothelial cell (RF/6A) does not express sLex and PSGL-1, but an sLex-like molecule, 6-sulfo-sLex. The a2,3-sialic acid and a1,3-fucose are also presented by the 6-sulfo-sLex, which becomes the site for ApOmpA(59-74) binding in the endothelial cell.54 The demonstrated adhesin properties of ApOmpA(59-74) in promoting the bacteria internalization to the host cell, make it a good candidate for vaccine development. However, anti-ApOmpA could only partially inhibit the infection of Anaplasma phagocytophilum in host cells.43 This is because Anaplasma phagocytophilum utilises multiple invasins to promote infection.55 To date, ApOmpA functions together with Asp14 (14-kDa Anaplasma phagocytophilum surface protein) and AipA (Anaplasma phagocytophilum invasion protein A) during bacterial infection. Interestingly, it has been shown that a mixture of antisera targeting ApOmpA, AipA and Asp14 can successfully clear the infection.55,56
OmpA of Orientia tsutsugamushi and Ehrlichia chaffeensis
As mentioned earlier, OmpA was also detected in Orientia tsutsugamushi and Ehrlichia chaffeensis. However, the actual role of OmpA in both these bacteria is still vague. What is known is that the predicted tertiary structure of the OmpA of Orientia tsutsugamushi (OtOmpA) shows high similarities with the OmpA of Anaplasma phagocytophilum (ApOmpA), particularly at the surface-exposed residue of the alpha-helix. Amino acid residues 103-118 of OtOmpA have been shown to be analogous to amino acid residues 59-74 of ApOmpA, which are functionally essential for the adhesion and invasion of Anaplasma phagocytophilum to its host cells.24 Interestingly, the amino acid sequence of Ehrlichia chaffeensis OmpA (EcOmpA) at positions 53-68 is also homologous to the ApOmpA59-74 receptor binding domain sequence.25 Since Orientia tsutsugamushi and Ehrlichia chaffeensis are originated from the same order as Anaplasma phagocytophilum, it is very likely that the OmpA of these bacteria play a similar role as adhesin and invasin in disease pathogenesis, even though they target different host cell surface proteins and cause different pathogenic diseases. In addition, antibodies develop against the EcOmpA53-68 fragments, successfully reducing the infection of the monocyte cell (THP-1) with Ehrlichia chaffeensis.25-40 However, another study found that protease-dependent degradation of EcOmpA by CDGA (2′-O-di(tert-butyldimethylsilyl)-c-di-GMP), a c-di-GMP antagonist, did not inhibit the adhesion of Ehrlichia chaffeensis to the monocyte cell (THP-1).31 It is suggested that other cell surface proteins are involved in facilizing the bacterial binding and internalization to the host cell, and that the presence of OmpA intensifies the process.31 In contrast, Cheng et al. (2011) suggests that the inhibitory effect of anti-OmpA IgG on the infection of Ehrlichia chaffeensis to the monocyte cell (THP-1) mentioned above is due to steric hindrance.40 These contradictory opinions need further investigation.
OmpA not involved in the virulence of Rickettsia in the in vivo study/mice model?
Based on all the information gathered and discussed above, the OmpA protein plays an important role in bacterial interaction with the host cell, especially for SFGR. The obstruction in the production of rOmpA by some of the members or strains in rickettsia is the key factor in repressing the virulence ability of these bacteria. However, unexpectedly, there are some manuscripts showing that the absence of rOmpA has no effect on the virulence of the rickettsia. This is mostly observed in the virulent strain of Rickettsia rickettsii. The knockout of the rOmpA sequence in the virulence strain Sheila Smith of Rickettsia rickettsii does not prevent the bacteria from infecting the animal model used, the guinea pig.28 The amount of viable rOmpA-knockout and wild-type Sheila Smith found in the spleen of the guinea pig after 6 days of infection shows no significant differences. This is consistent with no significant changes in the body temperature elevation of the guinea pig infected with both groups (wild-type and rOmpA knockout). Both groups induce a high fever response in the guinea pig, with the highest temperature occurring between day 5 and 7. In addition, both groups showed similar growth and entry rates when infecting the in vitro model, Vero cells. Furthermore, Riley et al. found that rOmpA mRNA expression in the in vivo infection of C3H/HeN mice was significantly lower than the expression in the in vitro study in Vero cells.30 This could be due to the fact that Rickettsia ricketsii uses different transcription strategies when infecting different host species. However, monoclonal anti-OmpA developed from Rickettsia conorii provided passive protection against future infection with Rickettsia conorii to C3H severe combined immunodeficiency mice (C3H SCID mice, a mouse model for spotted fever Rickettsiosis).29 As the result, the mice immunized with the anti-OmpA had a 100% survival rate compared to the 0% rate in the control group (normal serum/no serum). The author also investigated the kinetic production of antibodies in C3H/HeN mice after infection with Rickettsia conorii. It turns out that antibody reactive to OmpA do not appear during the early infection, but slowly appears on day 12, when the mice have already started to recover from the infection. Antibody production increased from day to day and was highest on day 30.29 It is suggested that rOmpA may provide protection against reinfection, but not during primary infection. The question of, whether or not OmpA is involved in the virulence of rickettsia during an in vivo study/mouse model is therefore in the grey area.
In this review, OmpA was shown to be present in most human pathogenic Rickettsiales, with the exception of the typhus group of rickettsia. In fact, the OmpA sequence was found to be highly conserved in various isolates of Orientia tsutsugamushi. This makes OmpA a good candidate for further investigation. Virulency of rickettsia has been shown to be highly dependent on the secretion of functional OmpA. The production of truncated OmpA due to the presence of a pre-mature stop codon in the OmpA sequence, has reduced the virulence of the rickettsia. OmpA plays an important role as an adhesin and invasin for the internalization of rickettsia and Anaplasma phagocytophilum into mammalian host cells. In rickettsia, the C-terminal region of OmpA has been shown to be the domain region for attachment and binding of Rickettsia to the endothelial host cell. In Anaplasma phagocytophilum, however, the N-terminal region acts as a ligand for binding to the myeloid and endothelial cells. Interestingly, OmpA from Orientia tsutsugamushi and Ehrlichia chaffeensis was computationally predicted to be homologous with the essential domain region of OmpA Anaplasma phagocytophilum. This suggests that the OmpA of Orientia tsutsugamushi and Ehrlichia chaffeensis may play a similar role in interacting with the host cell, even though they target different host cell surface proteins and cause different pathogenic diseases. Therefore, it can be concluded that OmpA plays an important role in the pathogenesis of human rickettsial diseases and is a potential candidate for vaccine development. However, further studies need to be carried out as some manuscripts showed contradictory results.
ACKNOWLEDGMENTS
All authors would like to thank the Institute for Medical Molecular Biotechnology, Faculty of Medicine, Universiti Teknologi MARA (UiTM), Selangor, Malaysia, for their support.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This work was supported by the Fundamental Research Grant Scheme from the Ministry of Higher Education Malaysia: FRGS/1/2019/STG03/UITM/02/2.
DATA AVAILABILITY
All datasets generated or analysed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies on human participants or animals performed by any of the authors.
- Driscoll TP, Verhoeve VI, Beier-Sexton M, Azad AF, Gillespie JJ. The Family Rickettsiaceae. In Green LH, Goldman E (eds.), Practical Handbook of Microbiology, 4th Ed. CRC Press, Boca Raton. 2021:511-526.
Crossref - Castelli M, McCarthy U, Petroni G, Bazzocchi C. Transmission of members of the “Candidatus Midichloriaceae” family to vertebrates and possible involvement in disease pathogenesis. In Thomas S (eds.). Rickettsiales: Biology, Molecular Biology, Epidemiology, and Vaccine Development, Springer, Cham. 2016:283-292.
Crossref - Sgroi G, Iatta R, Lovreglio P, et al. Detection of Endosymbiont Candidatus Midichloria mitochondrii and tickborne pathogens in humans exposed to tick bites, Italy. Emerg Infect Dis. 2022;28(9):1824-1832.
Crossref - Gillespie JJ, Williams K, Shukla M, et al. Rickettsia phylogenomics: unwinding the intricacies of obligate intracellular life. PloS one. 2008;3(4):e2018.
Crossref - Parola P, Paddock CD, Socolovschi C, et al. Update on tick-borne rickettsioses around the world: a geographic approach. Clin Microbiol Rev. 2013;26(4):657-702.
Crossref - Igolkina Y, Nikitin A, Verzhutskaya Y, et al. Multilocus genetic analysis indicates taxonomic status of “Candidatus Rickettsia mendelii” as a separate basal group. Ticks Tick Borne Dis. 2023;14(2):102104.
Crossref - Hajduskova E, Literak I, Papousek I, et al. ‘Candidatus Rickettsia mendelii’, a novel basal group rickettsia detected in Ixodes ricinus ticks in the Czech Republic. Ticks Tick Borne Dis. 2016;7(3):482-486.
Crossref - Fornadel CM, Zhang X, Smith JD, Paddock CD, Arias JR, Norris DE. High rates of Rickettsia parkeri infection in Gulf Coast ticks (Amblyomma maculatum) and identification of “Candidatus Rickettsia andeanae” from Fairfax County, Virginia. Vector Borne Zoonotic Dis. 2011;11(12):1535-1539.
Crossref - Arroyave E, Cornwell ER, McBride JW, Diaz CA, Labruna MB, Rodas JD. Detection of tick-borne rickettsial pathogens in naturally infected dogs and dog-associated ticks in Medellin, Colombia. Rev Bras Parasitol Vet. 2020;29(3):e005320.
Crossref - Lee MK, Kim HC, Choi YJ, et al. Detection of Candidatus rickettsia tarasevichiae from a tick collected from a human patient in South Korea. Syst Appl Acarol. 2019;24(2):193-197.
Crossref - Xue J, Ren Q, Jian R, et al. Molecular detection of “Candidatus Rickettsia tarasevichiae” by Loop-mediated Isothermal Amplification (LAMP) of the ompA gene. J Microbiol Methods. 2002;202:106601.
Crossref - Truong AT, Yun BR, Yoo MS, et al. Utility of ultra-rapid real-time PCR for detection and prevalence of Rickettsia spp. in ticks. BMC Vet Res. 2022;18(199):1-10.
Crossref - Abdelkadir K, Palomar AM, Portillo A, Oteo JA, Ait-Oudhia K, Khelef D. Presence of Rickettsia aeschlimannii, ‘Candidatus Rickettsia barbariae’ and Coxiella burnetii in ticks from livestock in Northwestern Algeria. Ticks Tick Borne Dis. 2019;10(4):924-928.
Crossref - Luz HR, Muooz-Leal S, de Carvalho WD, et al. Detection of “Candidatus Rickettsia wissemanii” in ticks parasitizing bats (Mammalia: Chiroptera) in the northern Brazilian Amazon. Parasitol Res. 2019;118(11):3185-3189.
Crossref - Moreira-Soto RD, Moreira-Soto A, Corrales-Aguilar E, Calderon-Arguedas O, Troyo A. ‘Candidatus Rickettsia nicoyana’: a novel Rickettsia species isolated from Ornithodoros knoxjonesi in Costa Rica. Ticks Tick Borne Dis. 2017;8(4):532-536.
Crossref - Li H, Li XM, Du J, et al. Candidatus Rickettsia xinyangensis as cause of spotted fever group rickettsiosis, Xinyang, China, 2015. Emer Infect Dis. 2020;26(5):985-988.
Crossref - Abarca K, Martםnez-Valdebenito C, Angulo J, et al. Molecular description of a novel Orientia species causing scrub typhus in Chile. Emerg Infect Dis. 2020;26(9):2148.
Crossref - Ismail N, Bloch KC, McBride JW. Human ehrlichiosis and anaplasmosis. Clin Lab Med. 2010;30(1):261-292.
Crossref - Lu M, Qin XC, Jiang YZ, et al. Emergence of ehrlichiosis by a new tick-borne Ehrlichia species in China. Int J Infect Dis. 2023;131, 32-39.
Crossref - Servin-Blanco R, Zamora-Alvarado R, Gevorkian G, Manoutcharian K. Antigenic variability: obstacles on the road to vaccines against traditionally difficult targets. Hum Vaccin Immunother. 2016;12(10):2640-2648.
Crossref - Valbuena G, Walker DH. Approaches to vaccines against Orientia tsutsugamushi. Front Cell Infect Microbiol. 2013;2:170.
Crossref - Futse JE, Buami G, Kayang BB, et al. Sequence and immunologic conservation of Anaplasma marginale OmpA within strains from Ghana as compared to the predominant OmpA variant. PLoS One. 2019;14(7):e0217661.
Crossref - Smith SG, Mahon V, Lambert MA, Fagan RP. A molecular Swiss army knife: OmpA structure, function and expression. FEMS Microbiol Lett. 2007;273(1):1-11.
Crossref - Evans SM, Adcox HE, VieBrock L, et al. Outer membrane protein a conservation among Orientia tsutsugamushi isolates suggests its potential as a protective antigen and diagnostic target. Trop Med Infect Dis. 2018;3(2):63.
Crossref - Green RS, Izac JR, Naimi WA, et al. Ehrlichia chaffeensis EplA interaction with host cell protein disulfide isomerase promotes infection. Front Cell Infect Microb. 2020;10:9.
Crossref - Cruz R, Huesgen P, Riley SP, et al. RC1339/APRc from Rickettsia conorii is a novel aspartic protease with properties of retropepsin-like enzymes. PLoS Pathog. 2014;10:e1004324.
Crossref - Noriea NF, Clark TR, Mead D, Hackstadt T. Proteolytic cleavage of the immunodominant outer membrane protein rOmpA in Rickettsia rickettsii. J Bacteriol. 2017;199(6):e00826-16.
Crossref - Noriea NF, Clark TR, Hackstadt T. Targeted knockout of the Rickettsia rickettsii OmpA surface antigen does not diminish virulence in a mammalian model system. MBio. 2015;6(2):e00323-15.
Crossref - Feng HM, Whitworth T, Olano JP, Popov VL, Walker DH. Fc-dependent polyclonal antibodies and antibodies to outer membrane proteins A and B, but not to lipopolysaccharide, protect SCID mice against fatal Rickettsia conorii infection. Infect Immun. 2004;72(4):2222-2228.
Crossref - Riley SP, Pruneau L, Martinez JJ. Evaluation of changes to the Rickettsia rickettsii transcriptome during mammalian infection. PloS One. 2017;12(8):e0182290.
Crossref - Kumagai Y, Matsuo J, Hayakawa Y, Rikihisa Y. Cyclic di-GMP signaling regulates invasion by Ehrlichia chaffeensis of human monocytes. J Bacteriol. 2010;192(16):4122-4133.
Crossref - Baldridge GD, Burkhardt NY, Simser JA, Kurtti TJ, Munderloh UG. Sequence and expression analysis of the ompA gene of Rickettsia peacockii, an endosymbiont of the Rocky Mountain wood tick, Dermacentor andersoni. Appl Environ Microb. 2004;70(11):6628-6636.
Crossref - Clark TR, Noriea NF, Bublitz DC, et al. Comparative genome sequencing of Rickettsia rickettsii strains that differ in virulence. Infect Immun. 2015;83(4):1568-1576.
Crossref - Ellison DW, Clark TR, Sturdevant DE, Virtaneva K, Porcella SF, Hackstadt T. Genomic comparison of virulent Rickettsia rickettsii Sheila Smith and avirulent Rickettsia rickettsii Iowa. Infect Immun. 2008;76(2):542-550.
Crossref - Xiao Y, Beare PA, Best SM, Morens DM, Bloom ME, Taubenberger JK. Genetic sequencing of a 1944 Rocky Mountain spotted fever vaccine. Sci Rep. 2023;13(1):4687.
Crossref - Parola P, Paddock CD, Raoult D. Tick-borne rickettsioses around the world: emerging diseases challenging old concepts. Clin Microbiol Rev. 2005;18(4):719-756.
Crossref - Walker DH, Yu XJ. Progress in rickettsial genome analysis from pioneering of Rickettsia prowazekii to the recent Rickettsia typhi. Ann N Y Acad Sci. 2005 1063(1):13-25.
Crossref - Beyer AR, Rodino KG, VieBrock L, et al. Orientia tsutsugamushi Ank9 is a multifunctional effector that utilizes a novel GRIP like Golgi localization domain for Golgi to endoplasmic reticulum trafficking and interacts with host COPB2. Cell Microbiol. 2017;19:e12727.
Crossref - Hunt JR, Carlyon JA. Analysis of Orientia tsutsugamushi promoter activity. Pathog Dis. 2021;79(7):ftab044.
Crossref - Cheng Z, Miura K, Popov VL, Kumagai Y, Rikihisa Y. Insights into the CtrA regulon in development of stress resistance in obligatory intracellular pathogen Ehrlichia chaffeensis. Mol Microbiol. 2011;82(5):1217-1234.
Crossref - Troese MJ, Carlyon JA. Anaplasma phagocytophilum dense-cored organisms mediate cellular adherence through recognition of human P-selectin glycoprotein ligand 1. Infect Immunity. 2009;77(9):4018-4027.
Crossref - Zhang JZ, Popov VL, Gao S, Walker DH, Yu XJ. The developmental cycle of Ehrlichia chaffeensis in vertebrate cells. Cell Microbiol. 2007;9(3):610-618.
Crossref - Ojogun N, Kahlon A, Ragland SA, et al. Anaplasma phagocytophilum outer membrane protein A interacts with sialylated glycoproteins to promote infection of mammalian host cells. Infect Immun. 2012;80(11):3748-3760.
Crossref - Kurtti TJ, Simser JA, Baldridge GD, Palmer AT, Munderloh UG. Factors influencing in vitro infectivity and growth of Rickettsia peacockii (Rickettsiales: Rickettsiaceae), an endosymbiont of the Rocky Mountain wood tick, Dermacentor andersoni (Acari, Ixodidae). J Invertebr Pathol. 2005;90(3):177-186.
Crossref - Sahni SK, Rydkina E. Host-cell interactions with pathogenic Rickettsia species. Future Microbiol. 2009;4(3):323-339.
Crossref - Valbuena G, Walker DH. Infection of the endothelium by members of the order Rickettsiales. Thromb Haemost. 2009;102(6):1071-1079.
Crossref - Sahni A, Fang R, Sahni SK, Walker DH. Pathogenesis of Rickettsial Diseases: Pathogenic and Immune Mechanisms of an Endotheliotropic Infection. Annu Rev Pathol. 2019;14:127-152.
Crossref - Hillman Jr RD, Baktash YM, Martinez JJ. OmpA mediated rickettsial adherence to and invasion of human endothelial cells is dependent upon interaction with a2b1 integrin. Cell Microb. 2013;15(5):727-741.
Crossref - Sahni A, Patel J, Narra HP, Schroeder CL, Walker DH, Sahni SK. Fibroblast growth factor receptor-1 mediates internalization of pathogenic spotted fever rickettsiae into host endothelium. PloS One. 2017;12(8):e0183181.
Crossref - Dumler JS, Choi KS, Garcia-Garcia JC, et al. Human granulocytic anaplasmosis and Anaplasma phagocytophilum. Emerg Infect Dis. 2005;11(12):1828-1834.
Crossref - Schotthoefer AM, Schrodi SJ, Meece JK, Fritsche TR, Shukla SK. Pro-inflammatory immune responses are associated with clinical signs and symptoms of human anaplasmosis. PLoS One. 2017;12(6):e0179655.
Crossref - Guzman N, Yarrarapu SNS, Beidas SO. Anaplasma Phagocytophilum. [Updated 2023 Aug 8]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing. 2023. https://www.ncbi.nlm.nih.gov/books/NBK513341/
- Carlyon JA, Akkoyunlu M, Xia L, et al. Murine neutrophils require a1,3-fucosylation but not PSGL-1 for productive infection with Anaplasma phagocytophilum. Blood. 2003;102(9):3387-3395.
Crossref - Seidman D, Hebert KS, Truchan HK, et al. Essential domains of Anaplasma phagocytophilum invasins utilized to infect mammalian host cells. PLoS Pathog. 2015;11(2):e1004669.
Crossref - Seidman D, Ojogun N, Walker NJ, et al. Anaplasma phagocytophilum surface protein AipA mediates invasion of mammalian host cells. Cell Microbiol. 2014;16(8):1133-1145.
Crossref - Kahlon A, Ojogun N, Ragland SA, et al. Anaplasma phagocytophilum Asp14 is an invasin that interacts with mammalian host cells via its C terminus to facilitate infection. Infect Immun. 2013;81(1):65-79.
Crossref
© The Author(s) 2024. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.