ISSN: 0973-7510
E-ISSN: 2581-690X
This study has been undertaken for the media optimization of pectinase producing strain, Bacillus subtilis BKDS1 isolated from dump yards of vegetable wastes. The ideal concentration of the substrate pectin was optimized by One–factor-at-a-time (OFAT) optimization method and found to be 0.25%. Optimization of the rest of fermentation variables (yeast extract, CaCl2, NaCl, (NH4)2SO4, KH2PO4, K2HPO4, MgSO4.7H2O, NaNO3, inoculum volume and pH) was carried out by Plackett-Burman design (PBD) followed by response surface methodology (RSM). Among the ten fermentation variables, three variables (yeast extract, CaCl2 and inoculum size) were selected to significantly affect pectinase production by PBD. The optimum levels of these variables were further determined using response surface methodology (RSM) based on central composite design (CCD). The optimal medium compositions were determined as follows (g/L): yeast extract 7.6g, CaCl2 0.81g, inoculum size of 1.5% and pectin 2.5g. The pectinase activity reached the maximum value of 1065.95U/mL in the optimized medium and it showed many fold increase in enzyme production compared to other pectinase production media tested in shake flask experiments.
Bacillus subtilis BKDS1, CCD, PBD, pectinase, RSM.
Pectinases constitute an exclusive group of enzymes which catalyze the degradation of pectic polymers present in the plant cell walls. They are classified based on the mechanism used to “attack” the galacturonan backbone as; the de-esterifying enzymes and the depolymerizing enzymes. The former catalyses the de-esterification of pectins and the later break the glycosidic a-(1- 4) bonds between GalA residues either by hydrolysis (hydrolases) or by trans elimination (lyases) 1. Today pectinases are one of the contemporary enzymes with prodigious industrial and commercial applications. The production of pectinase shares about 10% of the overall manufacturing of enzyme preparations and constitutes 25% of the global food enzyme market 2, 3. As many of the industries are purely depending on pectinases, their demand has increased progressively.
Among the wide group of microorganisms comprising fungi, bacteria, yeast and actinomycetes, filamentous fungi are commonly recognized as the best natural sources for the production of pectinase enzyme. But bacterial strains producing commercial enzymes are always preferred over fungal strains because of various reasons 4-7. Pectinases from fungal sources produce best under acidic pH and low temperature conditions and can therefore not be used in various industrial bioprocesses which utilize neutral to alkaline pH with high temperatures exceeding 45 °C. It has been shown that bacteria produce pectinase that withstands high pH and temperature 8, 9. Also, it is easy to harvest pectinase than fungus as it is an extracellular product in bacterial culture 10.
Medium optimization is one of the prime requirements for quantitative enhancement leading to overproduction of the enzymes. Factors like carbon and nitrogen sources and their concentrations have always been of great interest to the researchers in the industry for the low-cost media design. It is also known that 30-40% of the production cost of industrial enzymes is projected to be the cost of growth medium, which demands optimization for cost-efficient enzyme production 11. The conventional system for optimizing enzyme production by OFAT involves varying a single independent variable while maintaining the others at a constant level. This one-dimensional approach is arduous and time-consuming particularly for a large number of variables and does not consider interactions among variables. The limitations of this approach are overcome by the use of techniques like PBD and RSM. The PBD is an effective technique for the screening of significant factors influence the production and eliminate the insignificant components in order to obtain a smaller, more manageable set of factors 12. The selected significant factors were further optimized with the help of RSM that enables the study of interaction effects among different variables. It usually involves an experimental design such as CCD to fit a second-order polynomial by the least squares technique. Nowadays, RSM is used in wide range of scientific fields including production media optimization 13.
In view of the potential industrial applications of pectinases, it is essential to identify a new source of microorganisms capable of producing the enzyme at a cheap rate. In our earlier study, it was found that B. subtilis BKDS1 was selected as the predominant pectinolytic bacterial strain isolated from dump yards of market vegetables and fruits 6. The aim of this study was to identify key medium components and optimize suitable medium components for pectinase production by B. subtilis BKDS1 using a statistical design.
Culture maintenance and bacterial strain
Bacillus subtilis BKDS1, the pectinase producing strain, isolated from dump yards of market vegetables and fruits 6 was used for the study. The strain was identified by sequencing the 1500 bases of 16S rRNA gene followed by BLAST homology search. The nucleotide sequences have been deposited with NCBI database under accession number KT004506.1. The culture was maintained on nutrient agar slants at 4 °C and also stored as glycerol stocks at -20 °C. Media and chemicals were purchased from Hi- Media, Sigma and SRL chemicals. Overnight grown cultures (1% inoculum volume) of the bacterial isolate in yeast extract pectin (YEP) were used to inoculate the pectinase production medium.
Optimization of growth medium for pectinase production
Optimal pectin concentration (as the sole source of carbon) for the enzyme
YEP media with different concentrations (0.05 – 1%) of citrus pectin (Pectin from citrus peel-sigma) was used to study the pectinase production in the conventional OFAT method. Sterile media was inoculated with 1% overnight culture of B. subtilis BKDS1 and incubated in a shaker incubator (40 °C and 150 rpm) for 48 h. The pectinase activity was measured in the culture supernatant using DNS method adapted from Miller, 1959 14. The standard curve was prepared using D-galacturonic acid as a reducing sugar. One unit (U) of polygalacturonase activity was defined as the amount of enzyme that releases 1µmol of galacturonic acid per min under the assay conditions.
Optimization of media components using statistical softwares
Plackett-Burman design (PBD)
The variables that significantly influence the pectinase production were screened by PBD using the statistical software Minitab (Release 17, PA, USA). Ten independent variables were evaluated at two levels (high and low) which were designated as; coded values + 1 and -1 respectively. The factors and levels are depicted in Table: 1.
Table (1):
Experimental levels of 10 factors tested in PBD
No. |
Factors code |
Factors |
Min. value [-1](g/L) |
Max. value [+1] (g/L) |
|---|---|---|---|---|
1 |
A |
Yeast extract (YE) |
0.20 |
1.0 |
2 |
B |
Calcium chloride (CaCl2) |
0.01 |
0.11 |
3 |
C |
Sodium Chloride (NaCl) |
0.02 |
0.1 |
4 |
D |
Ammonium sulphate (NH4)2SO4 |
0.05 |
0.15 |
5 |
E |
potassium dihydrogen phosphate (KH2PO4) |
0.03 |
0.15 |
6 |
F |
Dipotassium phosphate (K2HPO4) |
0.01 |
0.05 |
7 |
G |
Magnesium sulfate (MgSO4•7H2O) |
0.01 |
0.11 |
8 |
H |
Sodium Nitrate (NaNO3) |
0.01 |
0.07 |
9 |
J |
Inoculum size (%) |
0.5 |
2.5 |
10 |
K |
(pH) |
6 |
8 |
PBD is based on the first order polynomial model;
![]() Where Y is the response (pectinase activity), β0 is the model intercepts, βi is the linear coefficient, and xi is the level of the independent variable. The ten factors chosen were; yeast extract (YE), NaCl, (NH4)2SO4, KH2PO4, K2HPO4, CaCl2, MgSO4·7H2O, NaNO3 and pH. As per the design, a set of twenty runs were performed. In all experiments, the concentration of pectin (0.25%) was kept as constant. Pectinase assays were carried out in duplicates and the averages of enzyme activity were taken as the response. From the regression analysis, the variables which were significant at or above 95 % level (P < 0.05), were considered to have a greater impact on pectinase activity and were further optimized by CCD.
Where Y is the response (pectinase activity), β0 is the model intercepts, βi is the linear coefficient, and xi is the level of the independent variable. The ten factors chosen were; yeast extract (YE), NaCl, (NH4)2SO4, KH2PO4, K2HPO4, CaCl2, MgSO4·7H2O, NaNO3 and pH. As per the design, a set of twenty runs were performed. In all experiments, the concentration of pectin (0.25%) was kept as constant. Pectinase assays were carried out in duplicates and the averages of enzyme activity were taken as the response. From the regression analysis, the variables which were significant at or above 95 % level (P < 0.05), were considered to have a greater impact on pectinase activity and were further optimized by CCD.
Central composite design (CCD)
RSM is a medley of statistical and mathematical methods used to select the best experimental conditions requiring the lowest number of experiments in order to get the applicable result. The CCD approach based on RSM was used for determining optimum levels of critical variables (identified by PBD) for enhanced enzyme production. CCD has been widely used as a statistical method based on the multivariate nonlinear model for the optimization of process and production variables. The statistical software ‘Design Expert 6.0’ was used to generate and analyze the experimental design. The CCD was used for fitting a second-order model which requires only a minimum number of experiments for modelling. Each significant parameter was assessed at five levels (-2, -1, 0, +1, +2), with six replicates at the centre points. Experimental range and levels of independent process variables are shown in Table: 2.
Table (2):
Ranges of variables used in RSM
No. |
Variables |
code |
-2 |
-1 |
0 |
1 |
+2 |
|---|---|---|---|---|---|---|---|
1 |
YE |
A |
0.20 |
0.4 |
0.60 |
0.8 |
1.0 |
2 |
Cacl2 |
B |
0.01 |
0.035 |
0.06 |
0.085 |
0.11 |
3 |
Inoculum |
C |
0.05 |
1.0 |
1.5 |
2.0 |
2.5 |
A total of 20 experimental runs were conducted, and the results were used to fit the second order polynomial model as shown in below equation;
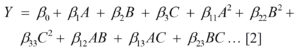 Where Y is the predicted response, β0 model constant; A, B and C are independent variables; β1, β2 and β3 are linear coefficients; b12, b13 and b23 are cross product coefficients; β11 β22 and β33 are the quadratic coefficients. With the help of Design Expert Software, the 3-dimensional surface plots were generated. The quality of the fit of the polynomial model was expressed by the value of correlation coefficient (R2). The model F-value (Fisher variation ratio), probability value (Prob > F) and adequate precision are the main indicators showing the significance and adequacy of the employed model.
Where Y is the predicted response, β0 model constant; A, B and C are independent variables; β1, β2 and β3 are linear coefficients; b12, b13 and b23 are cross product coefficients; β11 β22 and β33 are the quadratic coefficients. With the help of Design Expert Software, the 3-dimensional surface plots were generated. The quality of the fit of the polynomial model was expressed by the value of correlation coefficient (R2). The model F-value (Fisher variation ratio), probability value (Prob > F) and adequate precision are the main indicators showing the significance and adequacy of the employed model.
Verification Experiments
In order to validate the optimization of medium compositions, the result predicted by the design was compared with the actual values from the fermentations conducted using predicted medium compositions.
Comparison of enzyme production in optimized medium with various other pectinase production media
The pectinase production attained in the optimized medium was compared with previously reported pectinase production media viz. (i). YEP15, (ii). Czapek’sDox pectin medium 16 production media used by (iii) Soares et al., (1999 )17 and (iv) Jayani et al., (2010)18.
Effect of Incubation time and temperature on enzyme production
The optimized medium was used for analysing the effect of incubation time and temperature on enzyme activity. The inoculum was prepared and incubated in a rotary shaker at 150 rpm in different temperatures (30 °C, 40 °C, and 50 °C). The enzyme assay was performed at 12 h intervals.
The optimization of the culture medium for enhanced pectinase production using the strain B.subtilis BKDS1 was carried out by a combination of non-statistical and statistical based experimental designs.
Optimum concentration of pectin
Considering the commercial importance of pectinase, studies have been carried out to assess the optimum conditions for enhanced enzyme production. The concentration of pectin was optimized using OFAT method. For this, YEP broth was prepared with varying concentrations (0.05-1%) of pectin and the enzyme activity is calculated from the culture supernatant. The result obtained was plotted in Figure:1.
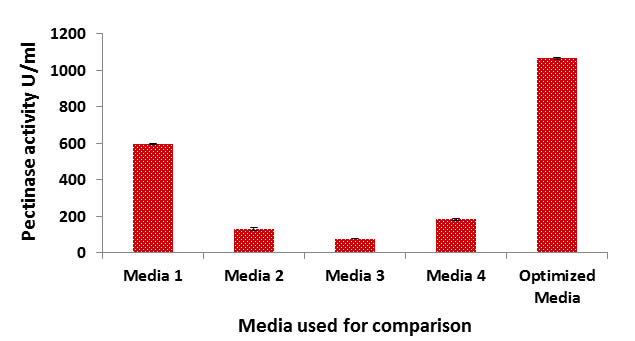
Fig. 1. Pectinase activity with different concentration of pectin
Maximum enzyme activity (599.38 U/ml) was found in medium containing 0.25% of pectin. This result clearly indicates that enzyme activity is decreased with increasing concentration of pectin. The similar type of observations were previously noted in B. subtilis by Kashyap
et al., (2000)15 where maximum pectinase activity was in 0.25% of pectin also in Streptomyces sp. RCK-SC 19. Decreased enzyme production in a higher concentration of pectin can be accredited to the phenomenon of catabolite repression, where galacturonic acid or one of the metabolites produced is undergoing self catabolite repression 20, 21 and also because of increase in viscosity of the broth 11.
Screening of the most significant medium components by PBD
Application of statistical method in media optimization for pectinase production was reported previously by many researchers 22. This step was initialized with PBD to screen some vital factors that have an immense role in the pectinase production by B. subtilis BKDS1. The result of PBD analysis is given in Table: 3 and 4. The corresponding pareto chart is shown in Figure: 2.
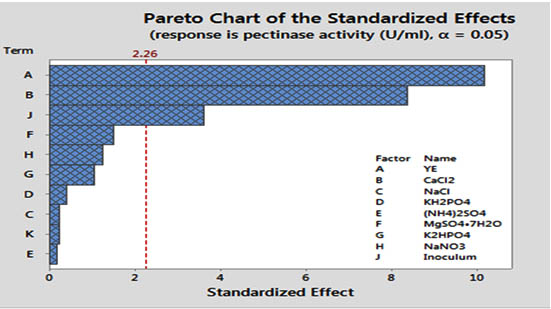
Fig. 2. Pareto chart showing the effect of ten media components on pectinase activity
Table (3):
PBD generated for nine variables.
| Run order | Coded Factors | Pectinase activity (U/ml) |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| A | B | C | D | E | F | G | H | J | K | ||
| 1. | -1 | -1 | -1 | -1 | -1 | -1 | -1 | -1 | -1 | -1 | 137.247 |
| 2. | -1 | -1 | -1 | 1 | -1 | 1 | -1 | 1 | 1 | 1 | 231.686 |
| 3. | 1 | -1 | -1 | 1 | 1 | -1 | 1 | 1 | -1 | -1 | 295.162 |
| 4. | 1 | -1 | 1 | 1 | -1 | -1 | -1 | -1 | 1 | -1 | 487.911 |
| 5. | 1 | 1 | -1 | -1 | 1 | 1 | -1 | 1 | 1 | -1 | 812.256 |
| 6. | -1 | -1 | 1 | 1 | -1 | 1 | 1 | -1 | -1 | -1 | 191.433 |
| 7. | 1 | -1 | -1 | -1 | -1 | 1 | -1 | 1 | -1 | 1 | 377.990 |
| 8. | -1 | 1 | 1 | -1 | -1 | -1 | -1 | 1 | -1 | 1 | 285.873 |
| 9. | -1 | 1 | 1 | 1 | 1 | -1 | -1 | 1 | 1 | -1 | 440.552 |
| 10. | 1 | -1 | 1 | 1 | 1 | 1 | -1 | -1 | 1 | 1 | 519.333 |
| 11. | -1 | -1 | -1 | -1 | 1 | -1 | 1 | -1 | 1 | 1 | 284.324 |
| 12. | -1 | 1 | -1 | 1 | 1 | 1 | 1 | -1 | -1 | 1 | 424.435 |
| 13. | -1 | 1 | -1 | 1 | -1 | 1 | 1 | 1 | 1 | -1 | 439.917 |
| 14. | -1 | 1 | 1 | -1 | 1 | 1 | -1 | -1 | -1 | -1 | 375.667 |
| 15. | 1 | 1 | -1 | 1 | 1 | -1 | -1 | -1 | -1 | 1 | 616.410 |
| 16. | 1 | 1 | 1 | 1 | -1 | -1 | 1 | 1 | -1 | 1 | 688.401 |
| 17. | 1 | 1 | -1 | -1 | -1 | -1 | 1 | -1 | 1 | -1 | 679.112 |
| 18. | 1 | -1 | 1 | -1 | 1 | 1 | 1 | 1 | -1 | -1 | 439.285 |
| 19. | 1 | 1 | 1 | -1 | -1 | 1 | 1 | -1 | 1 | 1 | 886.569 |
| 20. | -1 | -1 | 1 | -1 | 1 | -1 | 1 | 1 | 1 | 1 | 206.915 |
Table (3):
PBD generated for nine variables
| Run order |
Coded Factors | Pectinase activity (U/ml) |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| A | B | C | D | E | F | G | H | J | K | ||
| -1 | -1 | -1 | -1 | -1 | -1 | -1 | -1 | -1 | -1 | 137.247 | |
| -1 | -1 | -1 | 1 | -1 | 1 | -1 | 1 | 1 | 1 | 231.686 | |
| 1 | -1 | -1 | 1 | 1 | -1 | 1 | 1 | -1 | -1 | 295.162 | |
| 1 | -1 | 1 | 1 | -1 | -1 | -1 | -1 | 1 | -1 | 487.911 | |
| 1 | 1 | -1 | -1 | 1 | 1 | -1 | 1 | 1 | -1 | 812.256 | |
| -1 | -1 | 1 | 1 | -1 | 1 | 1 | -1 | -1 | -1 | 191.433 | |
| 1 | -1 | -1 | -1 | -1 | 1 | -1 | 1 | -1 | 1 | 377.990 | |
| -1 | 1 | 1 | -1 | -1 | -1 | -1 | 1 | -1 | 1 | 285.873 | |
| -1 | 1 | 1 | 1 | 1 | -1 | -1 | 1 | 1 | -1 | 440.552 | |
| 1 | -1 | 1 | 1 | 1 | 1 | -1 | -1 | 1 | 1 | 519.333 | |
| -1 | -1 | -1 | -1 | 1 | -1 | 1 | -1 | 1 | 1 | 284.324 | |
| -1 | 1 | -1 | 1 | 1 | 1 | 1 | -1 | -1 | 1 | 424.435 | |
| -1 | 1 | -1 | 1 | -1 | 1 | 1 | 1 | 1 | -1 | 439.917 | |
| -1 | 1 | 1 | -1 | 1 | 1 | -1 | -1 | -1 | -1 | 375.667 | |
| 1 | 1 | -1 | 1 | 1 | -1 | -1 | -1 | -1 | 1 | 616.410 | |
| 1 | 1 | 1 | 1 | -1 | -1 | 1 | 1 | -1 | 1 | 688.401 | |
| 1 | 1 | -1 | -1 | -1 | -1 | 1 | -1 | 1 | -1 | 679.112 | |
| 1 | -1 | 1 | -1 | 1 | 1 | 1 | 1 | -1 | -1 | 439.285 | |
| 1 | 1 | 1 | -1 | -1 | 1 | 1 | -1 | 1 | 1 | 886.569 | |
| -1 | -1 | 1 | -1 | 1 | -1 | 1 | 1 | 1 | 1 | 206.915 | |
Table (4):
Statistical analysis of PBD showing coefficient value, standard error coefficient value, t and p value for each variable
Term |
Factors |
Effect |
Coef |
SE Coef |
T-Value |
P-Value |
|---|---|---|---|---|---|---|
A |
YE |
282.4 |
141.2 |
13.8 |
10.20 |
0.000 |
B |
CaCl2 |
231.8 |
115.9 |
13.8 |
8.37 |
0.002 |
C |
NaCl |
6.3 |
3.2 |
13.8 |
0.23 |
0.824 |
D |
KH2PO4 |
-11.0 |
-5.5 |
13.8 |
-0.40 |
0.700 |
E |
(NH4)2SO4 |
4.8 |
2.4 |
13.8 |
0.17 |
0.866 |
F |
MgSO4•7H2O |
41.7 |
20.8 |
13.8 |
1.50 |
0.167 |
G |
K2HPO4 |
29.1 |
14.5 |
13.8 |
1.05 |
0.321 |
H |
NaNO3 |
-34.4 |
-17.2 |
13.8 |
-1.24 |
0.245 |
J |
Inoculum size (%) |
99.7 |
49.8 |
13.8 |
3.60 |
0.006 |
K |
pH |
6.3 |
3.2 |
13.8 |
0.23 |
0.824 |
The PBD analysis of ten factors indicated a significant variation in pectinase activity from 191.43 U/ml to 886.569 U/ml in twenty experimental runs. This variation confirmed the impact of all the factors on pectinase activity. The analysis of regression coefficients and t-value of ten ingredients are depicted in Table: 4. Generally, a large t-value associated with a low P-value of a variable indicates a high significance of the corresponding model term. From Table: 4 and the corresponding pareto chart in Figure: 2, it is clear that variables YE, CaCl2, and inoculum displayed a high positive significant effect for enzyme production with ‘0’ p-value whereas NaCl, (NH4)2SO4, MgSO4·7H2O, K2HPO4 and pH showed non-significant positive effects. Factors such as KH2PO4 and NaNO3 displayed a non-significant negative effect. None of the tested factors showed significant negative effect.
Yeast extract and CaCl2 were the prime media components selected in PBD screening with lowest (0.00) p-value. Yeast extract is verified to be the principal nitrogen source probably because it provided other stimulatory components such as vitamins. Previous research reports are available to authorize this result. Among various nitrogen sources tested, yeast extract (7.5 g/L) is selected as most effective for pectinase production by B.subtilis 23. Similarly, pectinase production by marine B.subtilis is enhanced with the presence of yeast extract in the production medium 21. Furthermore, B. sphaericus MTCC 7542 presented maximal polygalacturonase production when grown on mineral medium containing yeast extract as sole nitrogen source 18.
PBD selected CaCl2 the metal ion, as the next factor that significantly affects pectinase production. Previous reports are available that indicating the impact of CaCl2 in pectinase production in various microbial genera24-26. Metal ions like Ca2+ might play a vital role in maintaining the active conformation of pectinases to stimulate the activity 2, 27. The third factor selected by the PBD to affect the pectinase production significantly was inoculum size with (with p-value 0.006). The optimization of inoculum size is a well-accepted criterion in microbial fermentation because higher inoculum densities may cause lesser enzyme production. This result was supported by studies of various researchers28-31.
Optimization of significant variables using CCD
At the end of screening experiment by PBD, three factors were found to play a significant role in pectinase production. The optimal levels of these three most significant factors and effect of interaction on pectinase production were determined by CCD of RSM in the Design Expert software. The critical factors selected by the PBD were studied at five different levels (-2, -1, 0, +1,+2) as shown in Table: 2 and a set of 20 experiments with a different combination of these factors were carried out. Quadratic regression analysis using ANOVA was used to estimate the significance of model coefficients and the p values indicated the significance of each coefficient, which also showed the interaction strength between each independent variable. The actual yield of pectinase and the yield predicted by the model equation are given in Table: 5. The ANOVA analysis of the optimization study is given in Table: 6. The three-dimensional response surface plots for pectinase production showing the interactive effects between the selected media components were shown in Figure: 3 (i-iii).
Table (5):
CCD matrix of four variables (in coded & actual units) with experimental and predicted response
| Run | Yeast extract ( A ) |
Inoculum (B ) |
CaCl2 ( C ) | Pectinase activity (U/ml) | ||||
|---|---|---|---|---|---|---|---|---|
| Response | Observed | Predicted | ||||||
| Coded | Actual | Coded | Actual | Coded | Actual | |||
| 1 | 0 | 0.6 | +2 | 2.5 | 0 | 0.06 | 798.2293 | 1055.02 |
| 2 | 0 | 0.6 | 0 | 1.5 | 0 | 0.06 | 1065.229 | 1027.42 |
| 3 | 0 | 0.6 | 0 | 1.5 | 0 | 0.06 | 972.3383 | 1027.42 |
| 4 | 0 | 0.6 | 0 | 1.5 | 0 | 0.06 | 1026.525 | 1027.42 |
| 5 | +2 | 1 | 0 | 1.5 | 0 | 0.06 | 948.4962 | 1085.66 |
| 6 | +1 | 0.8 | -1 | 1 | -1 | 0.035 | 782.0662 | 935.7 |
| 7 | 0 | 0.6 | 0 | 1.5 | 0 | 0.06 | 1044.484 | 1027.42 |
| 8 | 0 | 0.6 | 0 | 1.5 | +2 | 0.11 | 972.6479 | 1109.7 |
| 9 | -1 | 0.4 | +1 | 2 | -1 | 0.035 | 853.7472 | 973.72 |
| 10 | 0 | 0.6 | 0 | 1.5 | 0 | 0.06 | 1026.525 | 1027.42 |
| 11 | -1 | 0.4 | -1 | 1 | +1 | 0.085 | 860.8689 | 1018.02 |
| 12 | -2 | 0.2 | 0 | 1.5 | 0 | 0.06 | 831.763 | 969.18 |
| 13 | 0 | 0.6 | 0 | 1.5 | -2 | 0.01 | 800.7992 | 945.14 |
| 14 | -1 | 0.4 | -1 | 1 | -1 | 0.035 | 865.2038 | 1014.12 |
| 15 | +1 | 0.8 | +1 | 2 | +1 | 0.085 | 1058.495 | 1182.24 |
| 16 | 0 | 0.6 | -2 | 0.5 | 0 | 0.06 | 810.0884 | 999.82 |
| 17 | +1 | 0.8 | -1 | 1 | +1 | 0.085 | 933.9432 | 1086.64 |
| 18 | 0 | 0.6 | 0 | 1.5 | 0 | 0.06 | 1042.007 | 1027.42 |
| 19 | +1 | 0.8 | +1 | 2 | -1 | 0.035 | 906.0759 | 1021.58 |
| 20 | -1 | 0.4 | +1 | 2 | +1 | 0.085 | 868.3002 | 987.34 |
From the ANOVA analysis, the model F-value of 18.8 implies the model is significant. Values of “Prob> F” less than 0.05 indicate model terms are significant. In this case A,C,AB,AC,A2,B2, C2 are significant model terms. The regression equation coefficients were calculated and the data was fitted to a second order polynomial equation. Thus the response (Y), (in terms of coded factors) i.e. pectinase production by B. subtilis BKDS1 can be expressed in terms of the following regression equation;![]() Where A is yeast extract, B is CaCl2, and C is inoculum.
Where A is yeast extract, B is CaCl2, and C is inoculum.
Table (6):
Analysis of variance (ANOVA table for response surface quadratic model-CCD)
Source |
Sum of Squares |
df |
Mean Square |
F- Value |
p-value Prob > F |
|
|---|---|---|---|---|---|---|
Model |
171203.9 |
9 |
19022.655 |
18.802 |
significant |
|
A-YE |
13567.97 |
1 |
13567.975 |
13.411 |
0.0044 |
|
B-inoculum |
3047.531 |
1 |
3047.531 |
3.012 |
0.1133 |
|
C-CaCl2 |
27077.63 |
1 |
27077.633 |
26.764 |
0.0004 |
|
AB |
7974.996 |
1 |
7974.996 |
7.883 |
0.0186 |
|
AC |
10810.23 |
1 |
10810.230 |
10.685 |
0.0084 |
|
BC |
47.18933 |
1 |
47.189 |
0.047 |
0.8334 |
|
A2 |
32398.28 |
1 |
32398.281 |
32.023 |
0.0002 |
|
B2 |
82808.86 |
1 |
82808.863 |
81.850 |
||
C2 |
33953.55 |
1 |
33953.547 |
33.560 |
0.0002 |
|
Residual |
10117.18 |
10 |
1011.718 |
|||
Lack of Fit |
5174.495 |
5 |
1034.899 |
1.05 |
0.4806 |
not significant |
Pure Error |
4942.686 |
5 |
988.537 |
|||
Cor Total |
181321.1 |
19 |
The ‘lack of fit F-value’ of 1.05 implies the lack of fit is not significant relative to the pure error. The p-value of lack of fit in in this model is 0.4806 (>0.05) means the model fits well. The design predicted an ‘R-squared value’ of 0.7241, which is in reasonable agreement with the ‘adjacent R-squared’ value of 0.8940. For a good statistical model, the R2 value should be in the range of 0 – 1.0, and the value as obtained in the data analysis indicates that the model is good. The interaction effects and optimal levels of the factors were determined by plotting the response surface curves. The three-dimensional response surface plots for pectinase production showing the interactive effects between the selected media components were shown in Figure: 3 (i-iii).
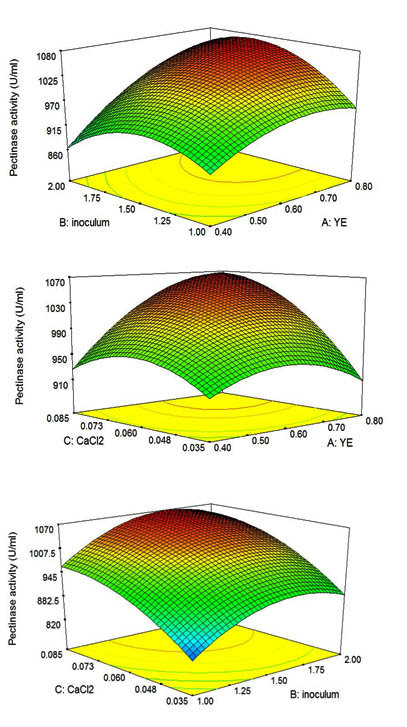
Fig. 3. 3D-Response surface plot for pectinase production showing the interactive effects of; i). YE (A) & inoculum size (B), ii). YE (A) & CaCl2 (C), and iii). inoculum size (B) & CaCl2 (C)
The interaction effects and optimal levels of the factors were determined by plotting the response surface curves. Figure: 3(i) depicts the interactive effects between factors yeast extract and inoculum and this is a significant interaction as the p value is 0.0186 (from Table: 8). From the figure, it is evident that the pectinase activity is maximum when the concentration of yeast extract reach 7.6 g/L at inoculum volume 1.5%. The enzyme activity tends to decrease above and below this range. The significant interactive effect between yeast extract and CaCl2 is presented in Figure: 3(ii). The enzyme activity rises with increasing concentration of CaCl2 and reaches maximum at 0.81g/L of CaCl2 at this point the concentration of yeast extract is 7.6 g/L. From table: 8, it is visible that the interaction between inoculum and CaCl2 is a non-significant interaction as the p value is 0.8334 which is <0.05. The response surface plot of this interaction is shown in 3(iii).
Validation of the Model
Validation of the experimental model was tested by carrying out the experiments under optimized conditions: (g/L) yeast extract 7.6g, CaCl2 0.81g and inoculum size of 1.5%, Citrus pectin 2.5g at pH 7. The experiments were performed in triplicates and the results were compared. The pectinase activity (1065.95 U/ml) obtained from experiments was very close to the actual response (1069.84 U/ml) predicted by the regression model, which proved the validity of the model.
Comparison of enzyme production in optimized medium with various other pectinase production media
The pectinase production levels attained in the optimized medium was compared with culture media previously used by various researchers. The data obtained were plotted in Figure: 4.
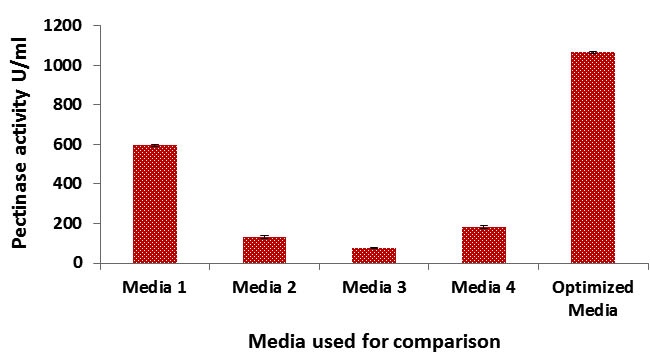
Fig. 4. Comparison of pectinase production in various media with RSM optimized medium
From the comparison result, it is clear that the optimized medium showed many fold increase compared to other pectinase production media tested. The corresponding fold of increase was; 1.78, 8.08, 13.74, 5.82 folds in media1, 2, 3 and 4 respectively. This implies a good optimization result.
Optimum temperature and incubation time for maximum enzyme activity
Effect of incubation time and temperature on enzyme production was studied in 12 h time interval at a temperature range of 30 – 50 °C at150 rpm. The enzyme assay was performed at every 12 h incubation period and the results obtained were illustrated in Figure: 5.
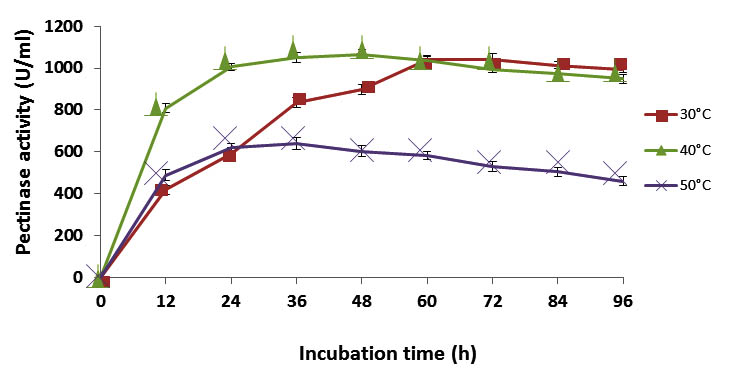
Fig. 5. Enzyme activity (U/ml) in different incubation temperature
From the graph (Figure: 5), it is clear that incubation temperature and time had an impact on enzyme yield. The optimal incubation time for maximal pectinase activity in 30 °C was found to be 72 h. At the point when the temperature is increased from 30 °C to 40 °C, the optimum incubation period diminishes from 72 h to 48 h. The incubation period again decreased to 24- 36 h at a temperature of 50 °C but the level of enzyme production was very low compared to other temperature ranges. So 40°C is taken as the optimum temperature for maximal enzyme production. Despite the fact that, the enzyme production achieved its peak at 48 h (1066.255 U/ml), the level of enzyme production increases even from 24 h (1006.398 U/ml) of incubation period. Reports have shown that many Bacillus species produce pectinase maximally at an incubation time of 72 h and above18, 32, 33.
Medium optimization is one of the most critically investigated process that is carried out before any large-scale metabolite production. As a preferred statistical experimental method, RSM is suitable for describing a near optimum region and thus identifying the exact criterion for a multifactorial system, which reduces the number of experiments without neglecting the interaction among the parameters. The present study has been attempted to optimize the growth-promoting factors for the enhanced production of pectinase by the isolated strain B.subtilis BKDS1 using statistical methods. The study began with optimizing the substrate pectin (0.25%) by OFAT method. Then, PBD was used to determine the relative importance of ten variables on pectinase production and found that yeast extract, CaCl2 and inoculum size were the major factors. The optimal concentration ranges of the three factors were optimized successively by CCD. In the optimized fermentation broth that contain yeast extract (7.6g/L), CaCl2 (0.81g/L) and pectin (2.5g/L) at an inoculum size of 1.5%, the pectinase activity reached 1065.95U/mL compared with the predicted value of 1069.84U/mL. Further, the incubation temperature and incubation period were also optimized and found to be 40°C and 42 h respectively. The optimized media showed many fold increase in enzyme production compared to various other production media tested. So, this study demonstrates the prospects of the new strain B.subtilis BKDS1 for pectinase production and applicability of statistical media optimization for augmented enzyme production.
ACKNOWLEDGMENTS
The authors express their deep sense of gratitude to the Department of Life Sciences, University of Calicut for providing the necessary facilities. We also thank UGC-RGNF for the Ph.D. fellowship and our colleagues who have supported us in every way towards the completion of this work.
Authors’ contributions
This work was carried out in collaboration between both the authors. Author DS conceived and designed the study. Author BK carried out the experiments, performed the data analysis, wrote the first draft of the manuscript and managed the literature searches. Author DS edited and proofread the final manuscript. Both the authors read and approved the final manuscript.
- Alkorta, I., Garbisu, C., Llama, M.J. & Serra, J.L. Industrial applications of pectic enzymes: a review. Process Biochem, 1998; 33, 21-28.
- Jayani, R.S., Saxena, S. & Gupta, R. Microbial pectinolytic enzymes: a review. Process Biochem 2005; 40: 2931-2944.
- Pedrolli, D.B., Monteiro, A.C., Gomes, E. & Carmona, E.C. Pectin and pectinases: production, characterization and industrial application of microbial pectinolytic enzymes. Open Biotechnol J, 2009; 3, 9-18.
- Prathyusha, K. & Suneetha, V. Bacterial pectinases and their potent biotechnological application in fruit processing/juice production industry: a review. J Phytol, 2011; 3, 16-19.
- Joshi, M., Nerurkar, M. & Adivarekar, R. Characterization, kinetic, and thermodynamic studies of marine pectinase from Bacillus subtilis. Preparative biochemistry & biotechnology 2015′ 45, 205-220.
- Kavuthodi, B., Thomas, S.K. & Sebastian, D. Co-production of Pectinase and Biosurfactant by the Newly Isolated Strain Bacillus subtilis BKDS1. Br Microbiol Res J, 2015; 10.
- Kavuthodi, B. & Sebastian, D. Review on bacterial production of alkaline pectinase with special emphasis on Bacillus species. Biosci Biotechnol Res Commun, 2018; 11, 18-30.
- Hoondal, G.S., Tiwari, R.P., Tewari, R., Dahiya, N. & Beg, Q.K. Microbial alkaline pectinases and their industrial applications: a review. Appl Microbiol Biot, 2002; 59, 409-418.
- Andrade, M.V.V.d., Delatorre, A.B., Ladeira, S.A. & Martins, M.L.L. Production and partial characterization of alkaline polygalacturonase secreted by thermophilic Bacillus sp. SMIA-2 under submerged culture using pectin and corn steep liquor. Food Sci Technol (Campinas), 2011; 31, 204-208.
- Sohail, M. & Latif, Z. Phylogenetic Analysis of Polygalacturonase producing Bacillus and Pseudomonas isolated from plant waste material. Jundishapur J Microbiol, 2016; 9: e28594.
- Palaniyappan, M., Vijayagopal, V., Viswanathan, R. & Viruthagiri, T. Screening of natural substrates and optimization of operating variables on the production of pectinase by submerged fermentation using Aspergillus niger MTCC 281. Afr J Biotechnol, 2009; 8: 682-686.
- Montgomery, D.C. Design and analysis of experiments, Edn. 7th ed. 2009; (Wiley, Hoboken, N.J.).
- Ghaffari-Moghaddam, M., Yekke-Ghasemi, Z., Khajeh, M., Rakhshanipour, M. & Yasin, Y. Application of response surface methodology in enzymatic synthesis: A review. Russ J Bioorgan Chem, 2014; 40: 252-262.
- Miller, G.L. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 1959; 31, 426-428.
- Kashyap, D.R., Chandra, S., Kaul, A. & Tewari, R. Production, purification and characterization of pectinase from a Bacillus sp DT7. World J Microbiol Biotechnol, 2000; 16, 277-282.
- Reda A et al. Production of Bacterial Pectinase(s) from Agro-Industrial Wastes Under Solid State Fermentation Conditions J Appl Sci Res, 2008; 4 1708-1721.
- Soares, M.M., Silva, R.d. & Gomes, E. Screening of bacterial strains for pectinolytic activity: characterization of the polygalacturonase produced by Bacillus sp. Revista de Microbiologia, 1999; 30, 299-303.
- Jayani, R.S., Shukla, S.K. & Gupta, R. Screening of bacterial strains for polygalacturonase activity: its production by Bacillus sphaericus (MTCC 7542). Enz res, 2010.
- Kuhad, R.C., Kapoor, M. & Rustagi, R. Enhanced production of an alkaline pectinase from Streptomyces sp. RCK-SC by whole-cell immobilization and solid-state cultivation. World J Microbiol Biotechnol, 2004; 20, 257-263.
- Tsuyumu S. Self-catabolite repression” of pectate lyase in Erwinia carotovora. J Bacteriol, 1979; 137, 1035.
- Joshi, M., Nerurkar, M. & Adivarekar, R. Use of Citrus limetta Peels for Pectinase Production by Marine Bacillus subtilis. Innov Rom Food Biotechnol 2013; 12, 75.
- Yu, P. & Xu, C. Production optimization, purification and characterization of a heat-tolerant acidic pectinase from Bacillus sp. ZJ1407. Int J Biol Macromol, 2018; 108: 972-980.
- Qureshi, A.S. et al. Production of pectinase by Bacillus subtilis EFRL 01 in a date syrup medium. Afr J Biotechnol, 2012; 11, 12563-12570.
- Ortiz, G.E. et al. Pectinase production by Aspergillus giganteus in solid-state fermentation: optimization, scale-up, biochemical characterization and its application in olive-oil extraction. J ind Microbiol Biotechnol 2017; 44, 197-211.
- Munir, N., Asad, J.M. & Haidri, S. Production, Purification and Characterization of Endopolygalacturonase by Bacillus subtillus. Biochem Anal Biochem 2015; 4: 1.
- Bennamoun, L. et al. Production and Properties of a Thermostable, pH—Stable Exo-Polygalacturonase Using Aureobasidium pullulans Isolated from Saharan Soil of Algeria Grown on Tomato Pomace. Foods 2016; 5: 72.
- Oumer, O.J. & Abate, D. Characterization of Pectinase from Bacillus subtilis Strain Btk 27 and Its Potential Application in Removal of Mucilage from Coffee Beans. Enzyme Res 2017.
- Reddy, L.V.A., Wee, Y.-J., Yun, J.-S. & Ryu, H.-W. Optimization of alkaline protease production by batch culture of Bacillus sp. RKY3 through Plackett–Burman and response surface methodological approaches. Bioresour Technol, 2008; 99, 2242-2249.
- Shabbiri, K. et al. Medium optimization of protease production by Brevibacterium linens DSM 20158, using statistical approach. Braz J Microbiol, 2012; 43: 1051-1061.
- Gupta, A., Emmanuel, S. & Lakshminaras, M. Protease production by newly isolated Bacillus Sp.: Statistical media optimization. Arch Appl Sci Res, 2010; 2: 109-123.
- Ghazala, I., Sayari, N., Romdhane, M.B., Ellouz-Chaabouni, S. & Haddar, A. Assessment of pectinase production by Bacillus mojavensis I4 using an economical substrate and its potential application in oil sesame extraction. Journal of food science and technology, 2015; 52: 7710-7722.
- Kumar, A. & Sharma, R. Production of alkaline pectinase by bacteria (Cocci sps.) isolated from decomposing fruit materials. J Phytol, 2012; 4: 01-05.
- Paudel, Y.P., Lin, C., Shen, Z. & Qin, W. Characterization of pectin depolymerising exo polygalacturonase by Bacillus sp. HD2 isolated from the gut of Apis mellifera L. Microbiol Discov, 2015; 3: 1-8.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


