ISSN: 0973-7510
E-ISSN: 2581-690X
Colistin is considered the last option for treatment of infections caused by Multidrug Resistant (MDR) Gram negative pathogens. The mcr-1 gene could transfer the resistance to colistin between different species. Therefore, screening of this resistance mechanism will help greatly in the control of further spread of colistin resistance and enhancing the outcomes of patients. The current work aimed to study the frequency of colistin resistance among MDR Gram negative pathogens isolated from clinical samples at Ain Shams University Hospitals and to explore if the mcr-1 gene was the responsible mechanism for this resistance. These pathogens were isolated from various samples including; blood, pleural fluid, sputum, urine, swabs from surgical and burn wounds, that were submitted from wards and intensive care units to the central microbiology laboratory at Ain Shams University Hospital, Cairo, Egypt, during the period from June to December 2019. Culture and bacterial identification were done by conventional microbiologic techniques. Eighty Gram negative bacterial pathogens were assayed for antimicrobial susceptibility by disc diffusion test. Sixty MDR Gram negative isolates were identified and further studied for colistin susceptibility by E- test, as well as real- time PCR to detect mcr-1 gene. Totally four isolates (6.7%) were phenotypically resistant to colistin. We found mcr-1 gene in three of the obtained isolates (5%); 1 Pseudomonas aeruginosa ( 5.3%) and 2 Acinetobacter baumannii (14.3%). In conclusion, although we detected a low prevalence of mcr-1 positive isolates from human clinical samples, continuous monitoring and implementation of infection control precautions are greatly required, to interfere with the further occurrence and transfer of bacterial species carrying the mcr-1 gene.
Colistin resistance, mcr-1 gene, Multidrug resistant Gram negative pathogens, Real- time PCR
The emergence of multidrug-resistance among Enterobacteriaceae, Acinetobacter baumannii (A. baumannii ) as well as Pseudomonas aeruginosa (P. aeruginosa) is considered a critical issue in different healthcare settings. Infections that are caused by such antimicrobial resistant pathogens usually result in much higher morbidity and mortality rates1. Treatment options are limited because MDR Gram negative bacteria are resistant to most classes of antimicrobials2. The high incidence of these microorganisms and the shortage of suitable newly developed antimicrobials are the leading causes for the re-introduction of polymyxins, a previously known class of cyclic polypeptide antimicrobials, again as a helpful therapeutic option3 .
Clinicians used colistin, which is known as polymyxin E, in the time period between 1950s and 1980s. However, they stopped using it because of its noticed toxic effects on the kidneys and nervous system. This antibiotic was limited for treatment of chest infections in patients with cystic fibrosis and decontamination of intensive care patients’ guts4. Colistin acts mainly by binding to lipid A within the cell wall of Gram negative bacteria, displacing calcium and magnesium ions, resulting in lysis of the bacterial cell wall and death. Other suggested mechanisms include possible endotoxin effect and inhibition of respiratory enzymes located in the bacterial membrane5.
Polymyxin resistance has been attributed to chromosomal mutations. Until late 2015, when a plasmid-mediated polymyxin resistance gene named mcr-1 was first reported in China. The frequency of positive isolates for this gene from human patients’ samples was 1%, while higher rates were found in raw meat samples and animals of 15% and 21% respectively6. Several countries reported an increasing detection rate of isolates harbouring mcr-1 gene7. A transferase enzyme is encoded by this gene adds a phosphoethanolamine molecule to lipid A, thus decreasing the negative charge of the target and hence reducing its affinity for ploymyxins8. Moreover, the mcr-1 gene has been observed on plasmids containing other antimicrobial resistance genes9.
The prevalence of colistin resistant bacterial isolates varies among different countries and changes over the eras. Conducting surveillance programs for mcr-1 gene detection and applying proper infection control precautions are the two main pillars to minimize the spread of isolates positive for this gene. Thus the current work aimed to study the frequency of colistin resistance among MDR Gram negative pathogens isolated from various clinical samples at the central microbiology laboratory of Ain Shams University Hospital and to explore if mcr-1 gene was the responsible mechanism for this resistance.
Study setting
This work was performed on different clinical samples, submitted from different wards and Intensive Care Units (ICUs) to the central microbiology laboratory at Ain Shams University Hospital, during the period from June to December 2019. The studied samples included sputum, pleural fluid, blood, urine, and swabs from surgical and burn wounds, collected under complete aseptic precautions.
Identification and antibiotic susceptibility of the isolated organisms
Identification of the isolated bacterial species was done based on colony morphology, microscopic examination of Gram stained films and conventional biochemical reactions of the isolates in accordance with Wilson et al.10 Eighty Gram negative pathogens were randomly selected and assayed for their susceptibility to different antimicrobial classes, by disc diffusion test, following the CLSI guidelines11. The following antibiotics (Oxoid, England) were included: amoxicillin/clavulanic acid, ampicillin/ sulbactam, piperacillin /tazobactam, cefepime, cefotaxime, ceftazidime, imipenem, meropenem, tobramycin, amikacin, sulfamethoxazole / trimethoprim, doxycycline, ciprofloxacin, and levofloxacin. Nitrofurantoin was used only in urine samples. MDR organism was defined as the isolate was non-susceptible to at least one drug in three or more classes of antimicrobial12.
Determination of colistin minimal inhibitory concentration by E- test
Minimal inhibitory concentration (MIC) of colistin was determined for the different MDR bacterial species, by the E-test method (Himedia laboratories, India) according to the manufacturer’s recommendations. The breakpoints used here were: ≤2 mg/l considered as sensitive to colistin, while ≥4 mg/l considered as resistant 11. Escherichia coli (E. coli) ATCC 25922 and P. aeruginosa ATCC27853 served as a quality control for the antimicrobial susceptibility testing.
Detection of mcr-1 gene by Real time PCR
DNA Extraction from the studied MDR bacterial isolates was performed by QIAamp DNA Mini Kit (Qiagen, Germany) following the manufacturer’s instructions. Purified DNA was preserved at -20°C until used. Amplification and detection of mcr-1 were performed using SYBR Green quantitative PCR (qPCR) assay by Rotor Gene Q5 plex (Qiagen, Germany). The used primers were CLR5-F 5’ CGGTCAGTCCGTTTGTTC- 3’; CLR5-R 5’-CTTGGTCGGTCTGTAGGG-3’, according to previous report 6. The final volume of PCR reaction was 25 μl; composed of 12.5 μl QuantiTec SYBR Green qPCR Master Mix (2X), 5 μl template DNA, 0.7 μl for each primer, 6 μl RNase- free water. Positive and negative controls were also included. The real-time thermal cycler was programmed as: initial denaturating step for 15 min at 95°C, then 45 cycles of denaturation at 95°C for 15 sec, annealing at 55°C for 30 sec and extension at 72°C for 30 sec. This program was followed immediately by a melting program consisting of 1 minute at 95°C, 30 sec at 55°C, lastly to 95°C for 30 sec.
Data analysis
All statistical procedures were carried out using SPSS version 22.0 for Windows (SPSS Inc, Chicago, IL, USA). Descriptive statistics, frequencies and percentages were calculated.
Our study included 60 MDR Gram negative bacterial isolates; 76.67% (46/60) of which were obtained from ICUs and 23.3% (14/60) obtained from different wards at Ain Shams University Hospital. The studied isolates were identified as follows: 26 Klebsiella pneumoniae (K. pneumoniae), 19 P. aeruginosa, 14 A. baumannii, and 1 E. coli. The majority of isolates were from wound swabs (25/60, 41.67%) (Table 1).
Table (1):
The distribution of MDR Gram negative bacterial isolates in different wards and different samples.
| MDR (N0.=60) | |||||
|---|---|---|---|---|---|
| Enterobacteriaceae | P. aeruginosa | A. baumannii | Total | ||
| No.(%) | No.(%) | No.(%) | No.(%) | ||
| Department | Surgical ICU | 10 (37.04%) | 6 (31.58%) | 2 (14.29%) | 18 (30%) |
| Surgical ward | 2 (7.41%) | – | 2 (14.29%) | 4 (6.67%) | |
| Medical ICU | 4 (14.81%) | 4 (21.05%) | 3 (21.43%) | 11 (18.32%) | |
| Medical ward | – | – | 2 (14.29%) | ||
| 2 (3.33%) | |||||
| Pediatric ICU | 2 (7.41%) | 3 (15.79%) | 1 (7.14%) | 6 (10%) | |
| Neonatal ICU | – | 2 (10.53%) | – | 2 (3.33%) | |
| Out patients | 2 (7.41%) | 1 (5.26%) | – | 3 (5%) | |
| Neurology ICU | 1 (3.70%) | – | – | 1 (1.67%) | |
| Oncology unit | – | 2 (10.53%) | 1 (7.14%) | 3 (5%) | |
| Burn ICU | 5 (18.52%) | 1 (5.26%) | 1 (7.14%) | 7 (11.67%) | |
| Chest ward | – | – | 1 (7.14%) | 1 (1.67%) | |
| Orthopedic ward | 1 (3.70%) | – | – | 1 (1.67%) | |
| Geriatric ICU | – | – | 1 (7.14%) | 1 (1.67%) | |
| Sample | Wound swabs | 14 (51.85%) | 9 (47.37%) | 2 (14.29%) | 25 (41.67%) |
| Respiratory samples* | 8 (29.65%) | 3(15.79%) | 7 (50%) | 18 (30%) | |
| Urine | 4 (14.80%) |
5 (26.31%) |
3 (21.42%) | 12 (20%) | |
| Blood | 1 (3.70%) |
2 (10.53%) |
2 (14.29%) | 5 (8.33%) | |
| Total | 27(45%) | 19(31.7%) | 14(23.3%) | 60(100%) | |
*Sputum and pleural fluid.
As regards the resistance pattern of the studied MDR bacterial isolates, Enterobacteriaceae had the highest rate of resistance (100%) against cephalosporins. P. aeruginosa displayed the highest resistance (94.74%) against ceftazidime, imipenem, and ciprofloxacin. All the studied A. baumannii isolates (100%) were resistant to ampicillin/sulbactam, cephalosporins, ciprofloxacin, and levofloxacin (Table 2).
Table (2):
Resistance pattern of the MDR bacterial isolates to different antimicrobials.
Antimicrobial agent |
Enterobacteriaceae (No.= 27) |
P. aeruginosa (No.= 19) |
A. baumannii (No.= 14) |
|---|---|---|---|
Amoxicillin- clavulanic acid |
25 (92.59%) |
NA |
NA |
Ampicillin-sulbactam |
26 (96.30%) |
NA |
14 (100%) |
Piperacillin-tazobactam |
23 (85.19%) |
17 (89.47%) |
13 (92.86%) |
Cefotaxime |
27 (100%) |
NA |
14 (100%) |
Ceftazidime |
27 (100%) |
18 (94.74%) |
14 (100%) |
Cefepime |
27 (100%) |
17 (89.47%) |
14 (100%) |
Imipenem |
25 (92.59%) |
18 (94.74%) |
12 (85%) |
Meropenem |
26 (96.30%) |
18 (94.74%) |
12 (85%) |
Amikacin |
26 (96.30%) |
17 (89.47%) |
13 (92.86%) |
Tobramycin |
26 (96.30%) |
17 (89.47%) |
13 (92.86%) |
Ciprofloxacin |
25 (92.59%) |
18 (94.74%) |
14 (100%) |
Levofloxacin |
25 (92.59%) |
16 (84.21%) |
14 (100%) |
Trimethoprim-sulfamethoxazole |
26 (96.30%) |
NA |
12 (85%) |
Doxycycline |
NA |
NA |
11 (78.57%) |
Nitrofurantoin (Additional for urine samples) |
3 (75%) |
NA |
NA |
NA: not applied
As for the results of colistin MIC by the E- test method (Fig.1), a total of four isolates (4/60, 6.7%) were colistin resistant, their MICs ranged from 4 to 64µg/ml. Colistin-resistant isolates included 2 P. aeruginosa (2/19, 10.5%) and 2 A. baumannii (2/14, 14.3%). They were isolated from urine (2/4,50%), blood (1/4, 25%), and sputum samples (1/4, 25%).
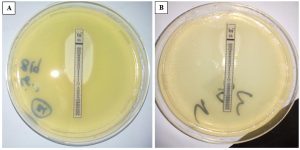 Fig. 1. Colistin susceptibility by E test. A: sensitive isolate (MIC = 1.5µ g/ml). B: resistant isolate (MIC = 64µ g/ml)
Fig. 1. Colistin susceptibility by E test. A: sensitive isolate (MIC = 1.5µ g/ml). B: resistant isolate (MIC = 64µ g/ml)
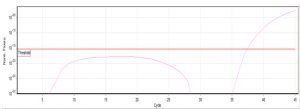
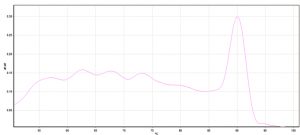
Real time PCR was done to detect mcr-1 expression (Fig.2 and 3). The gene existed in 3 of the studied isolates (3/60, 5%). The positive isolates were 1 P. aeruginosa (1/19, 5.3%) and 2 A. baumannii (2/14, 14.3%). Only 1 P. aeruginosa isolate was phenotypically non-susceptible to colistin by E- test but was negative for mcr1 gene by real time PCR.
Colistin is usually used as the last option of antimicrobials in treatment of multiresistant infections13. Transfer of plasmid -mediated colistin-resistance between different Gram negative pathogens requires frequent surveillance to limit the spread of this global threat 14.
As previously reported by Somily et al.15, most of the MDR isolates in this study were obtained from ICU patients (76.67%). This can be explained by several factors that predispose ICU patients to bacterial infections such as the use of invasive devices, severity of diseases, prolonged duration of hospitalization, comorbidities, and the higher use of broad-spectrum or multiple antibiotics.
In this study phenotypic colistin susceptibility was carried out by E- test. Several studies found the high agreement between E-test and dilution methods, which have made it a reliable alternative to these tests16. All Enterobacteriaceae isolates obtained in this study were sensitive to colistin, while 12.12% (4/33) of the non-fermenters were resistant to colistin; representing 6.7% of the studied MDR isolates. One previously reported study in India found a total of 24 colistin resistant isolates (8.5%) among 281 Enterobacteriaceae and non-fermenters isolates17. In Kuwait, a study reported that 12% of studied Acinetobacter isolates were colistin- resistant18. Several studies conducted in other Egyptian hospitals also reported low rates of colistin resistance ranging from 0 to 5% among A. baumannii19-21. Kandeel et al.16, found 8% and 3% rates of colistin resistance among MDR P. aeruginosa and A. baumannii, respectively, in Saudi Arabia. Khedr et al.22 found that 4.5% of Enterobacteriaceae and 7.9% of non-fermenters isolated from patients’ samples in the pediatric oncology department at the national cancer institute, Egypt, were resistant to colistin. Other investigators reported higher rates of colistin resistance. Shawky et al.23 detected 13.8% of 65 carbapenem resistant K. pneumoniae were colistin resistant. Another report studied MDR and extensively- drug resistant Gram negative bacteria isolated from Tanta University Hospital, and found 10 isolates among 61 Gram- negative bacilli (16.4%) were resistant to colistin. Among these colistin resistant isolates, 8 were K. pneumoniae and only 1 E. coli isolate and 1 P. aeruginosa24. The difference in the rates of resistance to colistin reported in the above studies is mostly attributed to differences in the studied populations and variability in the infection control measures applied. Also the low level of colistin resistace observed in our study may be due to the limited use of this antibiotic in our hospital.
In this study colistin resistant isolates were most obtained from urine samples (50%), followed by blood and sputum samples (25%) each. This finding agreed with previous reports14,25, where colistin resistant isolates where most frequently cultured from urine (46%), and blood samples (30%). However, Emara et al.24, found respiratory samples as the most common source of colistin resistant isolates (50%), followed by urine (30%).
Regarding the results of the real time PCR, 3 of the studied isolates (5%) were positive for mcr-1 expression; including 1 P. aeruginosa (1/19; 5.3%) and 2 A. baumannii (2/14; 14.3%). This report confirms previously reported findings that alarmed the spread of this gene. El nahriry et al.26 firstly reported mcr-1 gene in single E. coli isolate out of 241 Gram-negative clinical bacterial isolates, from Egypt. Zaki et al.25 revealed this gene in 2 bacterial strains (4%); “1 E. coli and 1 K. pneumoniae” out of 50 colistin resistant Enterobacteriaceae spp., that were recovered from health-care associated infections in Mansoura. However, Osama et al.27 detected one E. coli isolate (3.3%) harbouring mcr-1 gene, while Emara et al.24 didn’t detect mcr-1 gene among any of the phenotypically colistin resistant K. pneumoniae isolates.
The four colistin resistant isolates recovered in this study exhibited a high level of resistance against tested antimicrobials; as 75% were resistant to piperacillin/ tazobactam, fluoroquinolones, cephalosporins, carbapenem, and aminoglycosides. These findings also agrees with Zaki et al.25, who reported marked resistance to the third generation cephalosporin (60%), fourth generation cephalosporin (78%), and carbapenem antibiotics (50%). Another study found resistance to ciprofloxacin, piperacillin/tazobactam and nitrofurantoin was 90% for each one, 30% to carbapenems, and 20% to aminoglycosides24. This variability in resistance pattern can be explained by difference in antibiotics policy applied in health care settings within different geographical regions.
Worldwide the prevalence of mcr-1 in animal isolates is much higher compared with human clinical isolates, which may be related to the misuse of colistin in agriculture and the animal production28. Genes for colistin resistance are excreted in the feces of animals, and infection can be transmitted to humans when animal feces are used as manure, or contaminate the water bodies29.
In this study, we found one phenotypically colistin resistant P. aeruginosa isolate (MIC=4μg/ml). This could be explained by either the presence of chromosomal-mediated resistance, or the presence of other mcr gene variants. About 13 mcr-1 subgroups were already described in several countries, differing from mcr-1 by only one nucleotide. In addition other nine mcr variants have been described30.
Ongoing transfer of mcr-1 may lead to higher rates of poor treatment outcomes and consequently greater morbidity and mortality rates31. Thus surveillance for colistin resistance mediated by this gene should be conducted and studies should involve greater collection of isolates.
Although we detected a low prevalence of mcr-1 positive isolates from human clinical samples, continuous monitoring and implementation of infection control precautions are greatly required, to interfere with the further occurrence and transfer of bacterial species carrying mcr-1 gene.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have contributed to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Theuretzbacher U. Global antimicrobial resistance in Gram-negative pathogens and clinical need. Curr Opin Microbiol. 2017;39:106-112.
Crossref - Bakthavatchalam YD, Pragasam AK, Biswas I, Veeraraghavan B. Polymyxin susceptibility testing, interpretative breakpoints and resistance mechanisms: An update. J Glob Antimicrob Resist. 2018;12:124–136.
Crossref - Elbediwi M, Li Y, Paudyal N, et al. Global Burden of Colistin-Resistant Bacteria: Mobilized Colistin Resistance Genes Study (1980-2018). Microorganisms. 2019;7(10):461.
Crossref - Nang SC, Li J, Velkov T. The rise and spread of mcr plasmid-mediated polymyxin resistance. Crit Rev Microbiol, 2019;45:131–161.
Crossref - Kai J, Wang S. Recent progress on elucidating the molecular mechanism of plasmid-mediated colistin resistance and drug design. Int Microbiol. 2019.
Crossref - Liu YY, Wang Y, Walsh TR, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16(2):161-168.
Crossref - Dalmolin TV, de Lima-Morales D, Barth AL. Plasmid-mediated Colistin Resistance: What Do We Know? J Infect. 2018; 1(2):16-22.
Crossref - Sun J, Zhang H, Liu YH, Feng Y. Towards Understanding MCR-like Colistin Resistance. Trends Microbiol. 2018; 26: 794–808.
Crossref - Biswas S, Elbediwi M, Gu G, Yue M. Genomic Characterization of New Variant of Hydrogen Sulfide (H2S)-Producing Escherichia coli with Multidrug Resistance Properties Carrying the mcr-1 Gene in China. Antibiotics. 2020;9:80.
Crossref - Wilson ML, Weinstein MP, Rellerv LB. Laboratory detection of bacteremia and fungemia, pp. 15-28. In : Jorgensen JH, Pfaller MA, Carroll KC, Funke G, Landry M, Richter S, Warnock DW (eds.), Manual of Clinical Microbiology, 11th Ed.American Society for Microbiology, Washington, DC, USA. 2015.
Crossref - Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 27th edition. CLSI document M100. Clinical and Laboratory Standards Institute, Wayne, PA; 2017.
- Magiorakos AP, Srinivasan A, Carey R, et al.Multidrug resistant, extensively drug resistant and pandrug resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol and Infect. 2012;18(3):268-281.
Crossref - Lima Barbieri N, Nielsen DW, Wannemuehler Y, et al. Mcr-1 identified in Avian Pathogenic Escherichia coli (APEC). PLoS ONE. 2017;12(3): e0172997.
Crossref - Arjun R, Gopalakrishnan R, Nambi PS, et al. A Study of 24 Patients with Colistin-Resistant Gram-negative Isolates in a Tertiary Care Hospital in South India. Indian J Crit Care Med. 2017;21:317-321.
Crossref - Kandeel A. Detection of Colistin Susceptibility in Multi- drug Resistant Pseudomonas Aeruginosa and Acinetobacter Baumannii by four Different Methods. Egypt J Med Microbiol. 2016;25(2):17-23.
Crossref - Somily A. Comparison of E-test and disc diffusion methods for the in vitro evaluation of the antimicrobial activity of colistin in multi-drug resistant Gram Negative Bacilli. Saudi Med J. 2010;31(5):507-511.
- Behera B, Mathur P, Das A, et al. Evaluation of susceptibility testing methods for polymyxin. Int J Infect Dis. 2010;14:e596–e601.
Crossref - Al-Sweih NA, Al-Hubail MA, Rotimi VO. Emergence of tigecycline and colistin resistance in Acinetobacter species isolated from patients in Kuwait hospitals. J Chemother. 2011;23(1):13–16.
Crossref - Al-Agamy MH, Khalaf NG, Tawfick MM, Shibl AM, El-Kholy AA. Molecular characterization of carbapenem-insensitive Acinetobacter baumannii in Egypt. Int J Infect Dis. 2014;22:49-54.
Crossref - Amer WH. What remains against carbapenem-resistant Enterobacteriaceae and Acinetobacter spp? Evaluation ofTigecycline and Colistin. Int J Curr Microbiol Appl Sci. 2015;4 (9):613-24.
- El-Kohly AA, Sayed AM, Soliman MS, Reda NM, Attia AS . Investigation of the Colistin Resistance among Acinetobacter Baumannii from Patients at Cairo University Hospitals. Med J Cairo Univ. 2017; 85(8):3037-3042
- Khedr RA, Hussein M, Elswify M, El-Mahallawy HA, Rafeh N. Current prevalence of Colistin resistance and its impact on mortality among pediatric cancer patients in Egypt. 2016; Abstr. ECCMID, Amsterdam – Netherlands.
Crossref - Shawky SM, Abdallah A, Khouly M . Antimicrobial activity of Colistin and Tiegecycline against carbapenem resistant Klebsiella pneumoniae clinical isolates in Alexandria, Egypt. Int J Curr Microbiol App Sci. 2015;4(2):731-742
- Emara MM, Abd-Elmonsef MM, Abo Elnasr LM, Elfeky AA . Study of mcr-1 Gene-Mediated Colistin-Resistance in Gram-Negative Isolates in Egypt. Egypt J Med Microbiol. 2019;28 (3):9-16
- Zaki ME, AbouElKheir N, Mofreh M. Molecular study of colistin resistant clinical isolates of Enterobacteriaceae species. J Clin Mol Med. 2018;1(1):2-4.
Crossref - Elnahriry SS, Khalifa HO, Soliman AM, et al. Emergence of plasmid-mediated colistin resistance gene mcr-1 in a clinical Escherichia coli isolate from Egypt. Antimicrob Agents and Chemother. 2016;60(5):3249–3250.
Crossref - Osama R, Bakeer W, Fadel S, Amin M.Association of Carbapenem and Colistin Resistance in Pathogenic Gram Negative Bacteria. J Pure Appl Microbiol. 2019;13(2):733-739.|
Crossref - Poirel L, Jayol A, Nordmann P. Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin Microbial Rev. 2017;30(2):557-596.
Crossref - Bitrus AA, Chuanchuen R, Luangtongkum T . Emergence of colistin resistance in extended-spectrum beta lactamase producing Enterobacteriaceae isolated from food animals and its public health implication: A review. J Adv Vet Anim Res. 2018;5(1):1-11.
Crossref - Wang C, Feng Y, Liu L, Wei L, Kang M, Zong Z. Identification of novel mobile colistin resistance gene mcr-10. Emerging Microbes & Infections. 2020; 9(1):508-516.
Crossref - Caniaux I, van Belkum A, Zambardi G, Poirel L, Gros MF. MCR modern colistin resistance.Eur. J Clin Microbiol Infect Dis. 2017;36(3):415-420.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.