ISSN: 0973-7510
E-ISSN: 2581-690X
Phytopathogenic diseases are a major concern in modern agriculture, and for decades, pesticides have been used to prevent potential damage. Bacillus megaterium is proposed as a biological controlling agent, and gene expression of the lipopeptide genes FEND and ITUDI was assessed using RT-qPCR. Inhibition effects of B. megaterium on Alternaria sp. and Botrytis sp. were examined over a period of nine days, which confirmed the potential use of this bacterium to counteract these two pathogens. In addition, expression of FEND and ITUDI genes was assessed over nine days in the aforementioned dual cultures and inhibition tests. FEND expression in B. megaterium increased 20.16-fold in response to Alternaria sp., and ITUDI expression increased 3.20-fold in response to Botrytis sp. on day five of incubation. These results were corroborated by gene expression data obtained from B. megaterium during fermentation, where FEND and ITUDI gene expression increased 95.14- and 18.70-fold, respectively. In conclusion, B. megaterium can increase lipopeptide synthesis when exposed to these particular phytopathogens and can significantly increase the respective expression during fermentation.
Gene expression, Antagonism, Antimicrobial genes, Plant pathogens
Agriculture is one of the most important economic fields. It is presented with serious problems, such as economic and production losses owing to diseases affecting numerous types of crops during growth and after harvest, most of which are caused by phytopathogenic fungi1. A large number of pathogens that cause severe plant diseases in the most important staple crops belong to the fungal genera, Penicillium, Botrytis, Diplodia, Mucor, Phomopsis, Fusarium, Rhizopus, Monilinia, Alternaria, and Colletotrichum2. In Ecuador, two of the genera that cause the greatest number of diseases in crops of great production and consumption value are Botrytis and Alternaria. For example, Botrytis can infect 596 vascular plants, of which strawberry plants are particularly affected. This fungus occurs in various geographical areas, ranging from tropical to temperate and cold zone environments3,4. Alternaria sp. causes diseases in tomato plants and in plants of the genus Brassica5.
Damage caused by these pathogens is a severe concern for crop production; therefore, controlling their spread is an urgent matter. To control these pathogens, different techniques and compounds have been used, the most common of which are synthetic pesticides owing to their considerable efficiency; however, long-term use of such chemical agents has caused collateral damage to the environment and human health. Novel alternatives for the control of pathogens are therefore required, and biological agents that may help mitigate or eliminate fungal pathogen-induced damage of crop plants are a promising option6. Bacillus megaterium is a growth promoter in plants and also shows pathogen-controlling effects as it produces lipoproteins, antibiotics, and hydrolytic enzymes. B. megaterium may, therefore, be a promising and ecologically versatile alternative for controlling the spread and reproduction of various pathogens7.
Molecular methods may help elucidate certain mechanisms and factors, such as the optimal time point of applying biocontrolling agents, the optimal concentration, and the required conditions for optimal effects, all of which are crucial aspects and should be subsequently determined using in vivo tests. In this regard, quantification of gene expression allows us to understand the underlying mechanisms and to design targeted studies on the efficiency of biological controllers8-10.
B. megaterium has been shown to contain antifungal genes, which can inhibit growth of Alternaria sp. and Botrytis sp.11; however, it is important to examine the respective expression patterns and to assess how they change over time and under certain conditions so as to determine the factors that induce expression and thereby increase the pathogen-controlling potential. Therefore, using RT-qPCR, we aimed to assess expression of FEND and ITUDI antifungal lipopeptide genes in B. megaterium in response to Alternaria sp. and Botrytis sp. and during fermentation.
Inoculum preparation
Microbes that had been identified using molecular genetic means as B. megaterium, Alternaria sp. and Botrytis sp. were used in this study. The cultures, cryopreserved at -80°C, were obtained from the Laboratories of Life Sciences of the Universidad Politécnica Salesiana, Quito, Ecuador12. First, a culture medium was prepared using 22 g of Nutritional Agar and 1 L distilled water, in which the bacteria were incubated at 36°C. For fungal cultures, potato dextrose agar (PDA) culture medium was prepared at the same concentration, which was then used for incubation at 26°C.
Dual culture assays to test antagonistic activity
In order to test inhibitory effects of B. megaterium on Alternaria sp. and Botrytis sp., a dual culture method was used, in which each pathogenic fungus was exposed to B. megaterium in PDA culture medium where the fungus was placed at the center of the box, and the battery was inoculated through a 3-mm striation between bacteria and fungus. Incubation was continued at 36°C for nine days13,14. Genetic material and data for calculating the proportion of inhibition were obtained after this step. Data were collected on days 1, 5, and 9. Bacteria cultured without fungi were used as a control. The proportion of radial growth inhibition was calculated using the following equation15:
PICR = [R1 – R2/R2]X 100
Where R1 is the diameter of radial fungus growth and R2 is the diameter of radial growth of fungus exposed to B. megaterium.
Liquid Fermentation
Preparation of the inoculum and fermentation
Nutritional Agar medium was used to reactivate cryopreserved B. megaterium for incubation at 35°C for 24 h. Next, the bacterial concentration was recorded as a starting point. For this purpose, a suspension was prepared from normal saline that was set to a standard of 0.5 using spectrophotometry and the McFarland scale, thus representing a concentration of 105 CFU mL-1. After preparing the initial inocula, the fermentation process was initiated, starting with the preparation of 100 mL bacterial suspension (105 CFU mL-1) per Erlenmeyer flask using Triptocasa Soja Broth as the liquid culture medium. B. megaterium was incubated at 25°C under agitation at 100 rpm for 120 h.
RNA extraction
Extraction and quantification of Bacillus megaterium RNA from dual cultures
RNA was extracted from dual cultures of Botrytis sp. and Alternaria sp. exposed to B. megaterium on days 1, 5, and 9 using the PureLink RNA Mini-Kit (Thermo Fisher Scientific, Waltham, MA, USA). Subsequently, RNA was quantified in µg/µL using a Qubit RNA BR Assay Kit (Thermo Fisher Scientific).
Extraction and quantification of RNA from Bacillus megaterium in fermentation
RNA was extracted from bacterial cultures fermented in broth for 24, 48, 72, and 120 h, as specified above.
Reverse transcription and quantification
A Superscript® III First-Strand Synthesis SuperMix for RT-qPCR (Thermo Fisher Scientific) kit was used to obtain cDNA from the RNA templates, according to the manufacturer’s instructions. Briefly, 5 µg/µL RNA were used, and the reaction was incubated in a thermal cycler at 25°C for 10 min, 50°C for 30 min, and 85°C for 5 min before cooling to 4°C. Then, 1 µL E. coli RNase H was added, and the reaction mixture was incubated at 37°C for 20 min. cDNA was quantified using a Qubit ssDNA Assay Kit (Thermo Fisher Scientific). RT-qPCR
The ITUDI fragment was amplified using primers F: 5′-GATGCGATCTCCTGGATGT-3′ and R: 5′-ATCGTCATGGCTGAG-3′, and the FEND fragment was amplified using primers F: 5′-TTTGGCAGCAGGAGAAGTT-3′ and R: 5′-GCTGTCCGTGCTTT-3′ at a concentration of 10 µM. The 16S rRNA housekeeping gene was used for data normalization.
A LightCycler 2.0 (Roche, Basel, Switzerland) and 2 µg/µL cDNA were used to perform RT-qPCR with primers and Fast Sybr Green Master Mix (Thermo Fisher Scientific) at a final volume of 20 µL, according to the manufacturer’s instructions. Thermocycling was performed as follows: 95°C for 20 s, followed by 40 cycles of 95°C for 3 s, 60°C for 30 s, and 72°C for 30 s, after which a melting curve was generated at 95°C for 0 s, 65°C for 1 min, and 95°C for 0 s. The reaction was stopped by cooling to 4°C. To calculate expression of FEND and ITUDI, the initial concentration of gene copies (N0) was required, including the respective efficiencies (E) of each reaction, which was computed using LinRegPCR version 11.0, 16 which uses the 2−∆∆ct method17,18. Data were standardized with the 16S housekeeping gene as a reference, according to the following equation19:
N0 (normalizado)= [N0 gen problema / N0 gen normalizador]
Gene expression was calculated from standardized data by comparing all treatments with the control, which in this case was cultivation of B. megaterium without fungi, using the following equation19:
Relative expression level= [ N0 gene day n / N0 gene day 1 ]
Statistical analyses
FEND and ITUDI gene expression in B. megaterium were considered the response variables, and presence of the phytopathogenic fungi Botrytis sp. and Alternaria sp. and incubation time (1, 5, or 9 days) were used as independent variables. A completely randomized design was used for a 3 X 3 factorial analysis. The 27 experimental units were categorized according to incubation time and growth using three replicates of exposure of bacteria to each pathogen, plus the respective controls. Gene expression data obtained from dual cultures were compared with data collected at 24, 48, 72, and 120 h of bacterial fermentation under optimal growth conditions. For this, an analysis of variance (ANOVA) was performed, followed by Tukey’s post hoc test at 5%, using the statistical software, Infostat (2018).
Radial growth inhibition
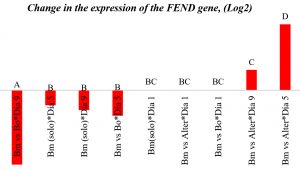
ANOVA showed a highly significant difference for both growth and incubation time (p < 0.0001), and the interaction between these variables was significant (p = 0.0395). Therefore, radial growth inhibition differed between the two fungi and depended on incubation time. Tukey’s test (Fig. 1) showed three ranges of significance, A, B, and C, with range C representing antagonism of Alternaria sp. on day 5 of incubation with an average inhibition of 76.45% and day 9, with inhibition averaging 98.48%, cultures that presented the highest inhibition, which indicated that B. megaterium inhibits Alternaria sp. growth more effectively, with inhibition levels of almost 99%. In previous studies, Bacillus sp. produced inhibition effects on Alternaria sp. ranging from 11% to 31% on day 5 20, 11, which are comparable to the results of the current study. Moreover, B. subtilis was previously reported to show a 60% inhibition of Botrytis sp. and 80% inhibition of Alternaria sp. 21, which is consistent with our results.
Note: Alter: Alternaria sp.; Bo: Botrytis sp.; Bm: Bacillus megaterium.
Fig. 1. Change in FEND gene expression (logarithmic scale, base 2).
Kurniawan et al. (2018)20 and García (2016)22 established that several species of the genus Bacillus, such as B. megaterium, can potentiate their inhibitory effects if they are grown under exposure to a pathogen. Moreover, B. megaterium, A. alternata on biopreparations when applied to broccoli crops infected with A. japonica kept values below 15.56 of severity23.
Table (1):
Percentage of Radial Growth Inhibition.
| Growth | Day | RIPP % | |
|---|---|---|---|
| B. megateirum vs. | 1 | 4,55 ± 5,25 | a |
| Botrytis sp. | 5 | 10,71 ± 0,00 | a |
| 9 | 30,00 ± 0,00 | ab | |
| B. megateirum vs. | 1 | 43,33 ± 5,62 | b |
| Alternaria sp. | 5 | 76,45 ± 5,22 | c |
| 9 | 98,48 ± 2,14 | c |
Note: RPI: Percentage of Radial Growth Inhibition.
Table (2):
Change in relative gene expression (RGE) of the FEND gene in Bacillus megaterium alone and against pathogens.
| Growth | Incubation days |
Average EGR change (Linear) |
Average EGR change (Log2) |
|---|---|---|---|
| B. megaterium | 1 | 1,00 ± 0,00 | 0,00 ± 0,00 |
| 5 | 0,56 ± 0,29 | -0,97 ± 0,78 | |
| 9 | 0,53 ± 0,35 | -1,31 ± 1,49 | |
| B. megaterium vs. Botrytis sp. | 1 | 1,00 ± 0,00 | 0,00 ± 0,00 |
| 5 | 0,32 ± 0,06 | -1,66 ± 0,27 | |
| 9 | 0,04 ± 0,01 | -4,89 ± 0,60 | |
| B. megaterium vs. Alternaria sp. | 1 | 1,00 ± 0,00 | 0,00 ± 0,00 |
| 5 | 20,16 ± 0,18 | 4,33 ± 0,02 | |
| 9 | 2,55 ± 0,47 | 1,34 ± 0,27 |
FEND gene expression in B. megaterium
When B. megaterium was not challenged with pathogenic fungi, no increase in relative expression of FEND by more than 2 was observed, which was not considered significant. Lengyel (2018)24 reports that molecules involved in gene expression are an inevitable source of chance because they are found under certain conditions in the cells, which causes proteins to be produced randomly and at minimal quantities, which is termed as the “noise” of expression. However, when B. megaterium was exposed to Alternaria sp., a significant increase in gene expression was evident on day 5 of incubation. ANOVA showed a highly significant difference in growth between pathogens and incubation times, and the interaction between the two variable was also significant (p < 0.0001, each); thus, relative expression of FEND depended on the fungus and time of incubation.
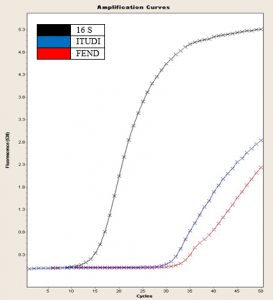
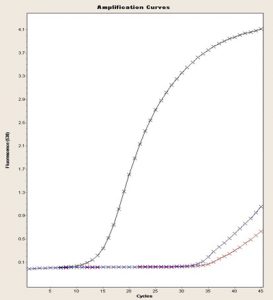
Fig. 2. 16S gene amplification curve, FEND, ITUDI of Bacillus megaterium vs Alternaria sp. at day 5 (LightCycler®, Roche).
In Fig. 2, according to Tukey’s test, four ranges of significance are presented: A, B, C, and D; within range A, the test presented the lowest expression, and range D includes the effect of Alternaria sp. on B. megaterium on day 5, which showed the greatest increase in FEND expression since day 1. FEND expression was down-regulated over time in the control. A similar phenomenon was observed when B. megaterium was exposed to Botrytis sp.
FEND expression was upregulated in B. megaterium exposed to Alternaria sp. with a 20-fold increase until day 5. On day 9, FEND expression decreased, which is in line with the results of Dragovoz et al. (2016), who reported that different strains of Bacillus exposed to A. alternata showed a 20- to 100-fold increased lipopeptide gene expression from day 1, with FEND as the highest expressed gene. They confirmed that fengicins are lipopeptides that are produced in large quantities. A different study found that fengicines are specific inhibitors of fungi25; however, surfactins more specifically inhibit bacteria.
The FEND gene is largely expressed as a defense against specific pathogens that are considered targets. Velho et al. (2015)10 suggested that biocontrollers increase the production of specific and effective antimicrobial proteins as a defense against certain microorganisms. The results of the current study showed that B. megaterium may be a promising biocontroller of Alternaria sp. as it showed upregulation of FEND expression when challenged with this phytopathogenic fungus.
Knight et al. (2018) reported that B. subtilis, a species very similar to B. megaterium, is an effective controller of the phytopathogens, Alternaria sp. and Fusarium sp., with upregulated FEND expression on day 3 of incubation. A different study using in vitro assays found that B. subtilis inhibited the growth of Alternaria sp. and Fusarium sp., and on day 4 of incubation confirmed the presence of genes such as ituA, bamC, and sfp which encode iturine A, bacilomycin D, and surfactin, respectively26.
ITUDI gene expression in B. megaterium
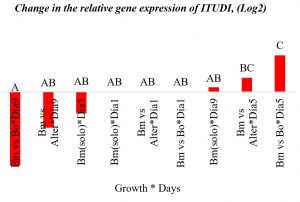
No significant changes in ITUDI expression over time were observed; however, B. megaterium exposed to Botrytis sp. and Alternaria sp. showed ITUDI upregulation on day 5. Exposure to Botrytis sp. led to a 3-fold increased expression, compared to that on day 1 (Table 3).
Table (3):
Change in relative gene expression (RGE) of ITUDI in Bacillus megaterium alone and against pathogens.
| Growth | Incubation days |
Average EGR change (Linear) |
Average EGR change (Log2) |
|---|---|---|---|
| B. megaterium | 1 | 1,00 ± 0,00 | 0,00 ± 0,00 |
| 5 | 0,81 ± 0,94 | -0,97± 1,65 | |
| 9 | 1,22 ± 0,61 | -0,17 ± 0,71 | |
| B. megaterium vs. Alternaria sp | 1 | 1,00 ± 0,00 | 0,00 ± 0,00 |
| 5 | 1,73, ± 1,02 | 0,64 ± 0,78 | |
| 9 | 033, ± 0,08 | -1,61 ± 0,36 | |
| B. megaterium vs. Botrytis sp | 1 | 1,00 ± 0,00 | 0,00 ± 0,00 |
| 5 | 3,20 ± 0,64 | 1,66 ± 0,31 | |
| 9 | 0,10 ± 0,01 | -3,31 ± 0,15 |
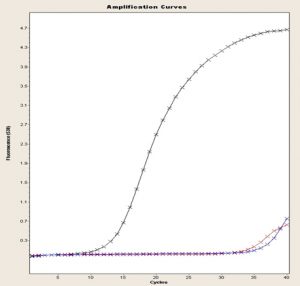
Fig. 4. 16S gene amplification curve, FEND, ITUDI of Bacillus megaterium versus Botrytis sp., at day 5 (LightCycler®, Roche).
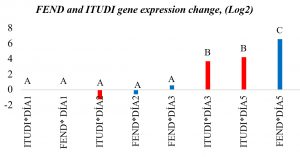
Fig. 5. Representation of relative gene expression change in logarithmic scale base 2 of FEND and ITUDI in Bacillus megaterium under fermentation.
Fig. 6. 16S gene amplification curve, FEND, ITUDI of Bacillus megaterium versus Botrytis sp., at day 5 (LightCycler®, Roche).
Only moderate changes in ITUDI expression were observed; however, the strength of inhibitory effects depends on several metabolites, including other lipopeptides (fengicins, surfactins, bacilliomycins, etc.), suggesting that each metabolite is required. Thus, both fengicins and surfactins depend on the expression of iturins, as they play an important role for effective inhibitory activity27,28.
Table (4):
16S gene amplification curve, FEND, ITUDI of Bacillus megaterium versus Botrytis sp., at day 5 (LightCycler®, Roche).
| Genes | Fermenting days | Average EGR change (Linear) |
Average EGR change (Log2) |
|---|---|---|---|
|
FEND |
1 | 1,00 ± 0,00 | 0,00 ± 0,00 |
| 2 | 0,85 ± 0,05 | -0,24 ± 0,08 | |
| 3 | 1,16 ± 0,04 | 0,21 ± 0,05 | |
| 5 | 95,14 ± 6,59 | 6,57 ± 0,10 | |
|
ITUDI |
1 | 1,00 ± 0,00 | 0,00 ± 0,00 |
| 2 | 0,42 ± 0,10 | -1,27 ± 0,37 | |
| 3 | 12,96 ± 1,02 | 3,96 ± 0,12 | |
| 5 | 18,70 ± 0,38 | 4,23 ± 0,03 |
Even so, gene expression differed between incubation conditions. For this purpose, an ANOVA was carried out considering each variable of interest separately, as well as the interactions between them. No significant difference was found regarding growth, whereas the effect of incubation time showed 1.91-fold increased expression until day 5, compared to that on day 1. In Fig. 4, data are shown as a logarithm to the base 2, and in Fig. 5, amplification curves until day 5 are shown. These results indicated that ITUDI expression by B. megaterium exposed to Botrytis sp. increased by 3.20-fold until day 5, compared to that on day 1. Such a response was expected as B. megaterium can reach the exponential growth phase after 48 h; thus on day 9, the bacteria would have already entered the dying phase29. In addition, production of lipopeptides by several species of the genus Bacillus to which inhibitory effects of Botrytis sp. are attributed suggest the presence of iturines, among other lipopeptides. Stable production occurs from 120 h to 168 h. Thus, on day 5, it increases by up to three fold30. Moreover, fengicines31 can also be produced during the stationary phase which occurs from 48–72 hours of incubation; however, inhibitory effects of B. megaterium are attributed to the synthesis of lipopeptides, as observed previously in microbiological test including exposure to Alternaria sp. and Botryris sp.32.
FEND and ITUDI gene expression in fermenting B. megaterium
FEND and ITUDI expression during fermentation of B. megaterium was evaluated using RT-qPCR on days 1, 3, 5, and 9 (Table 4), and we observed that expression of both genes significantly increased until day 5 of fermentation; however, it was necessary to perform an ANOVA. Tukey’s post hoc test showed a significant difference in gene expression between days of incubation. Gene expression of both lipopeptides increased 56.93-fold until day 5. Regarding differences in expression between genes, the results showed stronger upregulation of fengicines with a 95.14-fold increase, and an 18.71-fold increase in iturines. This indicates that lipopeptide synthesis increases during fermentation. Fig. 6 shows data of logarithmic phase 2 and Fig. 7 shows the amplification curve during fermentation.
The fermentation environment is more appropriate for bacterial growth33. In addition, with respect to time, the production of iturines is associated with the early stationary phase, and fengicines accumulate during the later stationary phase, which should occur after 72 hours, as reflected in the observed changes in gene expression34. During fermentation, the results were more promising, regardless of whether liquid or solid culture media are used35. Large commercial producers use this route of production because of its strong performance and high yield of metabolites such as antibiotics, bioregulators, and toxins for use in different fields including agriculture. B. subtilis, which is closely related to B. megaterium, showed considerable production of iturines and fengicines from days 2–5 of fermentation in our study, and similar results have been obtained previously36. Therefore, B. megaterium apparently increases the expression of both lipopeptides independently when exposed to pathogens and during fermentation.
We conclude that B. megaterium inhibits Alternaria sp. growth by 98.48% during 9 days of incubation. FEND gene expression increased 20.16-fold until day 5 of incubation when B. megaterium was exposed to Alternaria sp., which was the maximum of upregulation, suggesting that B. megaterium is a promising controlling agent to counteract specifically Alternaria sp. Regarding iturines, gene expression in B. megaterium increased three times more when exposed to Botrytis sp., and incubation time is a key factor for the synthesis of this lipopeptide, as evidenced by our results. Fermentation was associated with a significant increase in expression of both genes as under optimal growth conditions, this bacterium could produce metabolites at higher yields. Both genes were upregulated rapidly, but upregulation of the FEND gene was stronger. B. megaterium can be considered an promising candidate for agricultural use as it produced excellent results in in vitro microbiological and molecular tests; specifically, it may be used as a biological controlling agent to counteract Alternaria sp. and Botrytis sp.
ACKNOWLEDGMENTS
Thank you To Engineer Johana Meneses, for her support in the Research.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
VC authorship and direction of the project, management of resources for research and publication, review and translation of the manuscript. KM execution of laboratory tests, analysis, processing of results and writing of the manuscript. AC execution of laboratory tests, analysis, processing of results and writing of the manuscript. RA donation of microbiological material, support in statistical analysis and review of the manuscript.
FUNDING
This study was supported by grants $ 5000 from the Universidad Politécnica Salesiana as an internal project of the institution.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Agrios G. Plant Pathology. Elsevier Acadmic Press. 2005.
- FHIA. Deterioro Poscosecha de las frutas y hortalizas frescas por hongos y bacterias. Apartado Postal. 2007.
- Tomas-Grau RH, Requena-Serra FJ, Hael-Conrad V, Martinez-Zamora MG, Guerrero-Molina MF, Diaz-Ricci JC. Soft mechanical stimulation induces a defense response against Botrytis cinerea in strawberry. Plant Cell Rep. 2018;37(2):239-250.
Crossref - Elad Y, Pertot I, Cotes Prado AM, Stewart A. Plant Hosts of Botrytis spp. in Botrytis – the Fungus, the Pathogen and its Management in Agricultural Systems. Springer International Publishing. 2016:413-486.
Crossref - AyukeFO,LagerlofJ,JorgeG,.Söderlund S, Muturi JJ, Sarosh BR, Meijere J. Effects of biocontrol bacteria and earthworms on the severity of Alternaria brassicae disease and the growth of oilseed rape plants (Brassica napus). Appl Soil Ecol. 2017;117-118:63-69 (2017).
Crossref - Martínez-Padrón HY, Osorio-Hernández EO, Estrada-Drouaillet B, López-Santillán JA, Varela-Fuentes SE, Torres-Castillo JA. Control biológico de Fitopatógenos mediante aislados de Trichoderma spp. Agro Product. 2017;10(3):9-14.
- Knight CA, Bowman MJ, Frederick L, Day A, Lee C, Dunlap CA. The first report of antifungal lipopeptide production by a Bacillus subtilis subsp. inaquosorum strain. Microbiol Res. 2018;216:40-46.
Crossref - Grabova AY, Dragovoz IV, Zelena LB, Tkachuk DM, Avdeeva LV. Antifungal activity and gene expression of lipopeptide antibiotics in strains of Bacillus genus. Biopolym Cell. 2016;32(1):41-48.
Crossref - Sajitha KL, Dev SA. Quantification of antifungal lipopeptide gene expression levels in Bacillus subtilis B1 during antagonism against sapstain fungus on rubberwood. Biol Control. 2016;96:78-85.
Crossref - Leaes FL, Velho RV, Caldas DGG, Ritter AC, Tsai SM, Brandelli A. Expression of essential genes for biosynthesis of antimicrobial peptides of Bacillus is modulated by inactivated cells of target microorganisms. Res Microbiol. 2015;167(2):83-89.
Crossref - Saravanakumar D, Thomas A, Banwarie N. Antagonistic potential of lipopeptide producing Bacillus amyloliquefaciens against major vegetable pathogens. Eur J Plant Pathol. 2019;154:319-335.
Crossref - Chiluisa-Utreras V, Campaña M, Acurio R. Determinación microbiológica y molecular mediante PCR en tiempo real de dos bacterias del género Bacillus de interés agro biotecnológico. Bionatura. 2020;5.
Crossref - DeFilippi S, Groulx E, Megalla M, Mohamed R, Avis TJ. Fungal Competitors Affect Production of Antimicrobial Lipopeptides in Bacillus subtilis Strain B9-5. J Chem Ecol. 2018;44:374-383.
Crossref - Zalila-Kolsi I, Mahmoud AB, Ali H, Sellami S, Nasfi Z, Tounsi S, Jamoussi K. Antagonist effects of Bacillus spp. strains against Fusarium graminearum for protection of durum wheat (Triticum turgidum L. subsp. durum). Microbiol Res. 2016;192:148-158.
Crossref - Meza CLS, Barbosa RJF, Valero NO, Carrillo RMG, Redondo ARP. Antagonismo in vitro de Trichoderma harzianum Rifai sobre Fusarium solani (Mart.) Sacc., asociado a la marchitez en maracuyá. Rev Colomb Biotecnol. 2008;10(2):35-43.
- Ramakers C, Ruijter JM, Deprez RHL, Moorman AFM. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett. 2003;339(1):62-66.
Crossref - Livak KJ, Schmittgen TD. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods. 2001;25:402-408.
Crossref - Morales Segovia J. Expresión génica de flavanona 3-hidroxilasa y p-cumaril ester 3-hidroxilasa en el fruto de Solanum caripense dunal mediante RT-PCR y RT-qPCR. 2019.
- Evrard A, Boulle N, Lutfalla G. Nanoscience: Nanobiotechnology and nanobiology. in Real-Time PCR 1-1200. 2010.
- Kurniawan O, Wilson K, Mohamed R, Avis TJ. Bacillus and Pseudomonas spp. provide antifungal activity against gray mold and Alternaria rot on blueberry fruit. Biol Control. 2018;126:136-141.
Crossref - Hang NTT, Oh SO, Kim GH, Hur JS, Koh YJ. Bacillus subtilis S1-0210 as a biocontrol agent against Botrytis cinerea in strawberries. Plant Pathol J. 2005;21:59-63.
Crossref - García, M. Nuevas estrategias analíticas de extracciones selectivas en continuo acopladas a la cromatografía de líquidos para la determinación de antibióticos en muestras bioológicas. UNED. 2015.
- Acurio Vásconez RD, Tenorio Moya EM, Collaguazo Yépez LA, Chiluisa-Utreras VP, Vaca Suquillo IA. Evaluation of Bacillus megaterium strain AB4 as a potential biocontrol agent of Alternaria japonica, a mycopathogen of Brassica oleracea var. itálica. Biotechnol Reports. 2020;26:e00454.
Crossref - Lengyel I. Oscilaciones y ruido en la expresión genética: diálogos entre teoría y experimentos. Universidad de Buenos Aires. 2018.
- Jha SS, Joshi SJ, Geetha SJ. Lipopeptide production by Bacillus subtilis R1 and its possible applications. Brazilian J Microbiol. 2016;47(4):955-964.
Crossref - Mejía-Bautista MA, Alejo J, Tum-Suárez JM, Reyes-Ramírez A. Actividad in vitro de Bacillus spp. en la inhibición de crecimiento micelial de Fusarium equiseti Y Fusarium solani aislado de chile habanero (Capsicum chinense Jacq.). Agrociencia. 2016;50:1123-1135 .
- Maget-Dana R, Thimon L, Peypoux F, Ptak M. Surfactin/iturin A interactions may explain the synergistic effect of surfactin on the biological properties of iturin A. Biochimie. 1992;74(12):1047-1051.
Crossref - Romero D, de Vicente A, Rakotoaly RH, et al. The iturin and fengycin families of lipopeptides are key factors in antagonism of Bacillus subtilis toward Podosphaera fusca. Mol Plant-Microbe Interact. 2007;20(4):430-440.
Crossref - Tortolo-Cabanas K, Bell-García A. Producción de proteinas recombinantes en Bacillus megaterium: estado del arte. ICIDCA. 2015;49:0138-6204.
- Juárez Y. Detección de Botrytis cinerea mediante PCR en cultivos de vid y evaluación del biocontrol con Bacillus subtilis Q11. 2017.
- Jacques P, Hbid C, Destain J, et al. Optimization of Biosurfactant Lipopeptide Production from Bacillus subtilis S499 by Plackett-Burman Design. Appl Biochem Biotechnol. 1999;77:223-234.
Crossref - Ma Y, Kong Q, Oin C, et al. Identification of lipopeptides in Bacillus megaterium by two-step ultrafiltration and LC-ESI-MS/MS. AMB Express. 2016;6.
Crossref - Beltrán-García E, Macedo-Raygoza G, Villafana- Rojas J, et al. Production of Lipopeptides by Fermentation Processes: Endophytic Bacteria, Fermentation Strategies and Easy Methods for Bacterial Selection. in Fermentation Processes. InTech. 2017.
Crossref - Acurio R, Ñacato C, Valencia, M. Cepas autóctonas de Bacillus subtilis como agente de biocontrol in vitro de Alternaria spp. en Brassica oleracea var. itálica. Bionatura. 2018;3(2).
Crossref - Marinelli F, Molinari F. Las fermentaciones en la producción de metabilitos secundarios de interés farrmacéutico. Monogr la Real Acad Nac Farm. 2012;12:20-25.
- Ariza Y, Sánchez L. Determinación de metabolitos secundarios a partir de Bacillus subtilis efecto biocontrolador sobre Fusarium sp. Nova-Publicación Científica en Ciencias Biomédicas. 2012;10:149-155.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.