ISSN: 0973-7510
E-ISSN: 2581-690X
The present study describes the protease purification from halophilic Bacillus sp. MTCC 9558 by a single step hydrophobic interaction chromatographic using Phenyl-Sepharose CL – 4B column. After the final purification steps, protease was purified to 59.62-fold with a specific activity of 165.75 U/mg protein and overall recovery of 5.89%. The homogeneity of the enzyme preparation was confirmed by procedures such as SDS – PAGE, zymogram analysis and MALDI-TOF techniques. The molecular weight was determined to be 80.7 KDa and casein zymogram analysis of purified protease showed a single band indicating that the protease is active as a monomer. This paper also describes comparison of purified protease with crude in order to determine the effect of impurities or contaminating proteins on its characteristics. The purified protease exhibited improved thermostability and stability in presence of organic solvents in comparison with crude enzyme. The alkaline protease was determined to be halotolerant and Zn2+ activated metalloprotease. The purified protease was characterized to determine its kinetic constants such as Km, Vmax, kcat and kcat/Km values. The purified enzyme exhibited promising characteristics of potential biotechnological interest. In this paper, one of the applications of this protease in protein hydrolysis for bioactive peptide production with emphasis on antibacterial studies is discussed.
MALDI-TOF, Thermostability, Metalloprotease, Kinetic constants, Bioactive peptides.
Protein engineering techniques are extensively useful to modify or design proteins to meet the industrial requisites. This target can be achieved by purification of proteins of interest so that their characteristics, conformations, reactions with other ligands, substrate specificities, and specific activities can be understood. The selection of suitable purification steps is dependent on several properties such as molecular size, solubility, charge, and other features of the target protein.1,2 Efficient purification of enzymes through downstream processes is an essential prerequisite for subsequent commercial exploitation. Majority of the purification procedures for extracellular enzymes involves concentration of the culture filtrate either with salt or solvent precipitation or by ultrafiltration followed by purification by various chromatographic procedures.3 Several proteases from Bacillus sp. have been purified4 and the characteristics of purified proteases are determined5 for further exploration commercially. Many types of purified protease are subjected to sequence analysis to determine their amino acid sequence.6 Several enzymes are extensively studied for its applications in production of biologically active peptides – “bioactive peptides”, as they have great potential for commercial applications.7 The biopeptides liberated from animal and plant proteins, may exhibit various bioactivities such as angiotensin I converting enzyme (ACE) inhibitory activity8, antioxidative9, antimicrobial10 and several more. The most commonly used proteases are those from Bacillus sp, lactic acid bacteria, and marine yeasts.11
This paper describes the protease purification from halophilic Bacillus sp. by a single step chromatographic procedure by the test organism and the homogeneity of the enzyme preparation was confirmed by procedures such as sodium dodecyl sulphate polyacrylamide gel electrophoresis (SDS – PAGE), zymogram analysis and Matrix assisted laser desorption ionization- time of flight (MALDI-TOF) techniques. The comparison of purified protease with crude enzyme in order to determine the effect of impurities or contaminating proteins on its characteristics was determined. The purified protease was characterized to for its properties, kinetic constants and application prospective. Thus purified protease was used to hydrolyze several protein rich sources for the formation of bioactive peptides and its antibacterial property was determined.
Bovine Serum Albumin (BSA), Phenyl-Sepharose CL – 4B, acrylamide, bisacrylamide, dialysis tubing and Diethylaminoethyl cellulose (DEAE – cellulose) were procured from Sigma Chemical Co. (St Louis, MO, USA). Protein molecular weight marker (broad range) was purchased from Bangalore Genei India Private Limited. Amicon Ultra-4 3K centrifugal filter device was procured from Millipore, USA. All other chemicals used were of analytical grade and were purchased from SD Fine Chemicals (Mumbai, India).
Microorganism and culturing
The bacterium was isolated from poultry soil and was identified as Bacillus sp. MTCC 9558 belonging to B. cereus cluster was found to be a good producer of protease. It has been reported by us earlier for its potential for protease production in bioreactor12. The protease enzyme was produced by wheat bran an agro waste as protein source13 and Bacillus sp. was routinely sub-cultured and maintained on nutrient agar slants.
Protease assay
The protease activity was assayed according to procedure of Kunitz14 with some modifications. The absorbance was recorded spectrophotometrically at 280 nm. One unit of protease activity (U) was defined as the amount of enzyme which releases 1 ìmole of tyrosine per minute under the assay conditions.
Single step protease purification by hydrophobic interaction chromatography
The enzyme supernatant concentrated by ammonium sulphate precipitation was ultrafiltered using Amicon 3 kDa cut off centrifugal filter unit (Millipore, USA) and the amount of protein was estimated using BSA as standard.15 The protease activity was observed in the retentate fraction and was purified by hydrophobic interaction chromatography on Phenyl-Sepharose CL-4B column (1.5 cm X 10 cm) equilibrated with Tris-HCl buffer (50 mM pH 7.0) containing 1.0 M ammonium sulphate.16 The retentate with added ammonium sulphate (final concentration 1.0 M) was loaded onto the column. The bound proteins were eluted with a decreasing gradient of ammonium sulphate (1.0 – 0.0 M in the equilibrating buffer) and analyzed for protease activity. The protease did not elute at this stage and still remained bound and it was further eluted by a linear gradient of ethylene glycol (0 – 30%, v/v in 0.1 M Tris-HCl buffer pH 7.0). Active fractions were pooled together and used as purified protease for further characterization.
Determination of purity of protease
Sodium dodecyl sulphate polyacrylamide gel electrophoresis was carried out to examine the fractions as per the method of Laemmli17 using 5% (w/v) stacking gel and 12% (w/v) resolving gel. Protein molecular weight marker containing proteins covering the molecular weight range between 205 to 14.4 kDa were used to determine the molecular weight. Protein bands on the gel were detected by silver staining method and casein zymogram analysis was done to confirm the proteolytic activity. The gel was stained with Coomassie Brilliant Blue (CBB) R-250 followed by until clear zone appeared on a blue background indicating protease activity. The purified enzyme was analyzed with Matrix-assisted Laser Desorption Ionization-Time of Flight (MALDI-TOF) mass spectrometer (Shimadzu, Kyoto, Japan).
Characterization of crude and purified protease
The enzyme activity in each was studied and compared with crude protease characteristics to determine the effect of contaminating proteins on protease characteristics. Both crude and pure protease characteristics were determined with casein as substrate. The crude and purified protease from Bacillus sp. were subjected to characterization by studying the effects of pH (5.0 – 11.0 for 24 hours at 37°C) , temperature (30, 35, 37, 40, 45, 50, 55, 60, 65 and 70°C for 1 hour), NaCl (5 M – 0.5 M for 1 hour at 37°C), stability in organic solvents ratio (enzyme: solvent 3: 1 with shaking of 120 rpm for one hour at 37°C), various metal ions as activators (10 mM concentration) and inhibitors (Pepstatin, Iodoacetamide, Phenylmethylsulphonyl fluoride and EDTA at 37°C for 1 hour).
Determination of enzyme kinetics constants
The kinetic constants (Vmax, Km, kcat and kcat/Km) were determined using Lineweaver-Burk double reciprocal plot by assaying protease activity at a fixed enzyme concentration with varying concentrations of substrates 1 – 10 mg/ml at 37°C.
Application studies of purified protease from Bacillus sp. MTCC 9558 in synthesis of bioactive peptides with antibacterial activity
The purified protease from Bacillus sp. MTCC 9558 was used for hydrolysing different proteins (Spirulina, Yeast, casein, soy protein and â-lactoglobulin) resulting in low molecular weight peptides. The suspensions of different protein sources were mixed with purified protease from Bacillus sp. with activity of 5.1 U/ml and the mixtures were incubated and the reaction was terminated. The mixtures were centrifuged to remove the precipitate and the resulting protein hydrolysates were ultrafiltered using Amicon 3 kDa cut off centrifugal filter unit to remove proteins greater than 3 kDa. The filtrate thus obtained was used as peptides sample11.
The potential of the bioactive peptides to exhibit antibacterial activity was determined by well diffusion method against different pathogens such as Escherichia coli, Pseudomonas aeruginosa, Staphylococcus aureus, Proteus mirabilis and Salmonella typhimurium. The different microorganisms were uniformly spread on the solidified agar and five different peptide samples were added into the wells with protein concentration of 0.7 mg/well and examined for zone of clearance.
The crude enzyme supernatant was concentrated by ammonium sulphate precipitation and the maximum specific activity of 3.73 U/mg was obtained in 40 – 60% pellet fraction. The Bacillus sp. being halophilic has another issue to be addressed i.e., problem associated with using normal purification techniques. To achieve purity of halophilic enzymes has been difficult and in most of the cases, it is required to maintain high salt concentration of 5 – 20% throughout the purification procedure to retain protein activity. But this interrupts chromatographic procedures18 and therefore, purification of halophilic enzymes needs different protocols.
Protease purification by hydrophobic interaction chromatography
The protease from halophilic bacterium Bacillus sp. MTCC 9558 was concentrated from the cell-free supernatant by precipitation, subjected to ultra filtration and purified by Phenyl-Sepharose CL-4B hydrophobic interaction chromatography. The impurities were initially removed from matrix with decreasing gradient of ammonium sulphate and hence many protein peaks were observed. The protein of interest eluted in the ethylene glycol gradient and this protein peak coincided with the protease activity peak. The elution profile (Figure 1) and results of purification are summarized (Table 1). After the final purification steps, protease was purified to 59.62-fold with a specific activity of 165.75 U/mg protein and overall recovery of 5.89%.
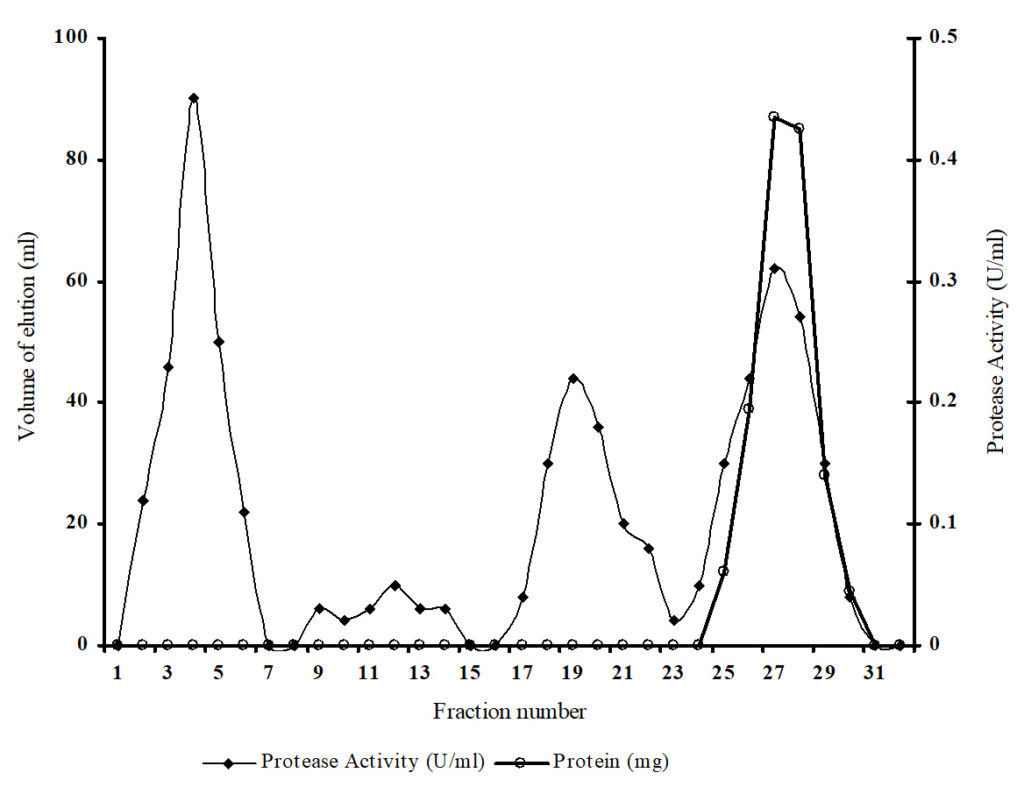
Fig. 1. Elution profile of protease from Bacillus sp. on phenyl Sepharose CL-4B column
Table (1):
Summary of purification of protease from Bacillus sp.
Purification Steps |
Total Activity (U) |
Total Protein (mg) |
Specific Activity (U/mg) |
Purification fold |
Yield (%) |
|---|---|---|---|---|---|
Crude |
11250 |
4050 |
2.78 |
1 |
100 |
Ammonium sulfate precipitation |
5600 |
1500 |
3.73 |
1.34 |
49.78 |
Ultrafiltration (5 kDa) |
3880 |
438 |
8.86 |
3.19 |
34.49 |
Phenyl-Sepharose CL-4B hydrophobic interaction chromatography |
663 |
4 |
165.75 |
59.62 |
5.89 |
Several researchers have used the Phenyl Sepharose column, such as Karan and Khare19 for purification of protease from Geomicrobium sp. EMB 2 with 22.6 fold of purification and yield of 51.2%. Joo and Chang16 purified oxidant and SDS-stable alkaline protease from Bacillus clausii I-52 on Phenyl-Sepharose matrix with 2.7 fold of purification and 85.2% yield.
SDS-PAGE and zymogram analysis of protease
The purity and relative molecular weight were determined by SDS-PAGE in 12% gels (Figure 2). The SDS-PAGE of the eluted protease showed a high level of purity and the enzyme purified from hydrophobic interaction chromatography appeared as a single band corresponding to a single polypeptide change of large molecular weight of 80.7 kDa as shown in the Figure 2. The casein zymogram analysis of purified protease after column purification showed a single band indicating that the protease is active as a monomer (Figure 3). These results indicated that the protease is a monomeric enzyme.
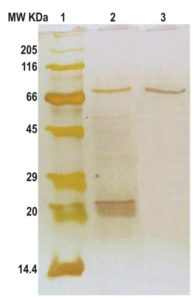
Fig. 2. SDS-PAGE analysis of protease purification and determination of molecular weight upon silver staining; Lanes indicate:
1. Molecular range protein marker: Myosin, Rabbit Muscle (205 kDa); âglucosidase (116 kDa); Bovine Serum Albumin (66 kDa); Ovalbumin (45 kDa); Carbonic Anhydrase (29 kDa); Soyabean Trypsin Inhibitor (20 kDa) and Lysozyme (14.4 kDa);
2. Culture supernatant and
3. Purified protease
![]()
Fig. 3. Casein zymogram analysis of purified protease
MALDI-TOF
MALDI-TOF analysis of purified protease showed a major peak corresponding to the molecular weight of 80.77 kDa (Figure 4). This corresponded to the result obtained by SDS-PAGE analysis. The presence of a single peak demonstrated that a single charged protein species of average mass of 80.77 kDa was present. The absence of any other significant peaks in the mass spectrum also demonstrated the homogeneity of the purified enzyme.
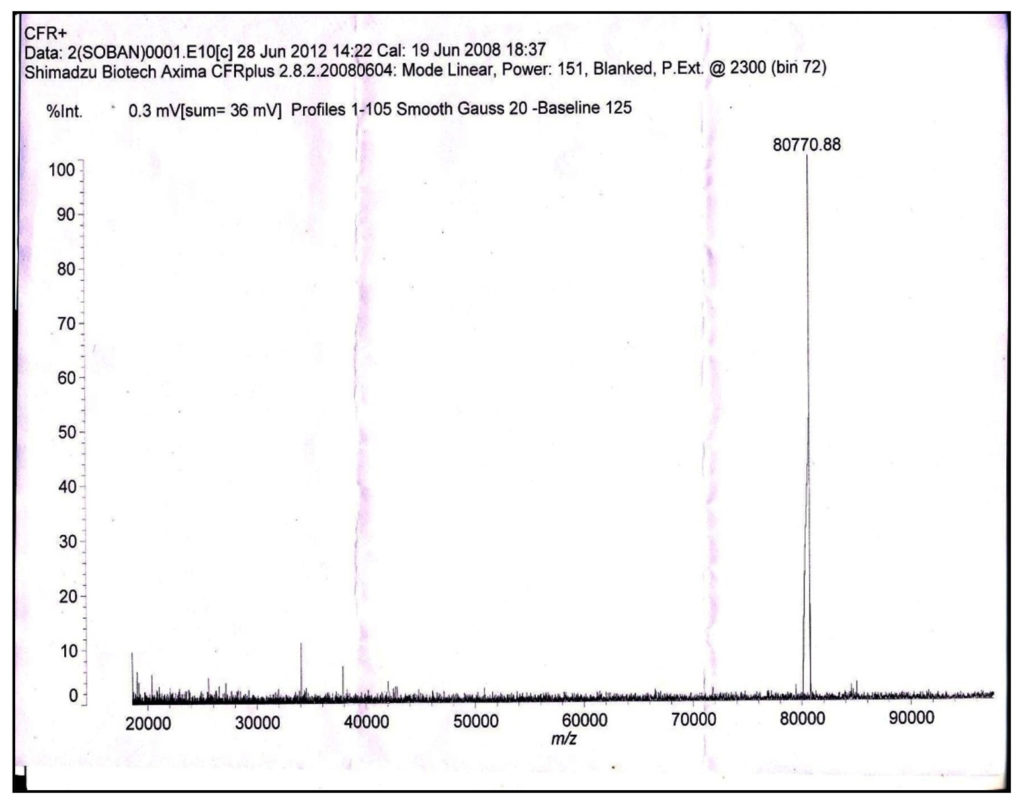
Fig. 4. MALDI-TOF of purified protease from Bacillus sp.
Characterization of purified protease
Effect of pH on protease activity
The proteolytic activity of the pure and crude protease was determined at several pH values (Figure 5). The crude protease was active in the pH range of 6.0 – 9.0, with an optimum at pH 7.0, whereas the purified protease was active in the pH range of 7.0 -10.0, with an optimum at pH 9.0 and showed 56 and 9% activity at pH 7.0 and 10.0 respectively. From this comparative study it was clear that the purified protease is an alkaline protease whereas the impure protease with other proteins exhibited neutral proteolytic activity. Several reports are available for optimum pH of majority of proteases from Bacillus sp. in the range of 8.0 – 10.0 and the optimum pH of serine alkaline protease from Bacillus sp. was reported as pH 8.20
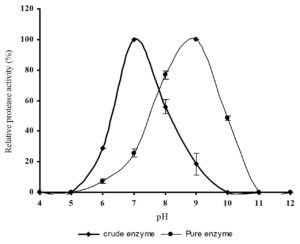
Fig. 5. Effect of pH on crude and purified protease activity
Effect of temperature on protease activity
The temperature dependence of the crude and purified protease for 1 hour assay at pH 9.0 is shown in the Figure 6. Both the crude and purified enzymes were most active at 37°C. Purified enzyme exhibited 70% of the initial activity at 60°C whereas the crude enzyme had almost lost its casein hydrolyzing ability. These studies proved that through purification and removal of contaminating proteins improved the desirable thermostability of the enzyme. It was reported that purified thermostable serine protease of Streptomyces tendae retained 70% of the initial activity after being treated at 60°C for 30 minutes.21
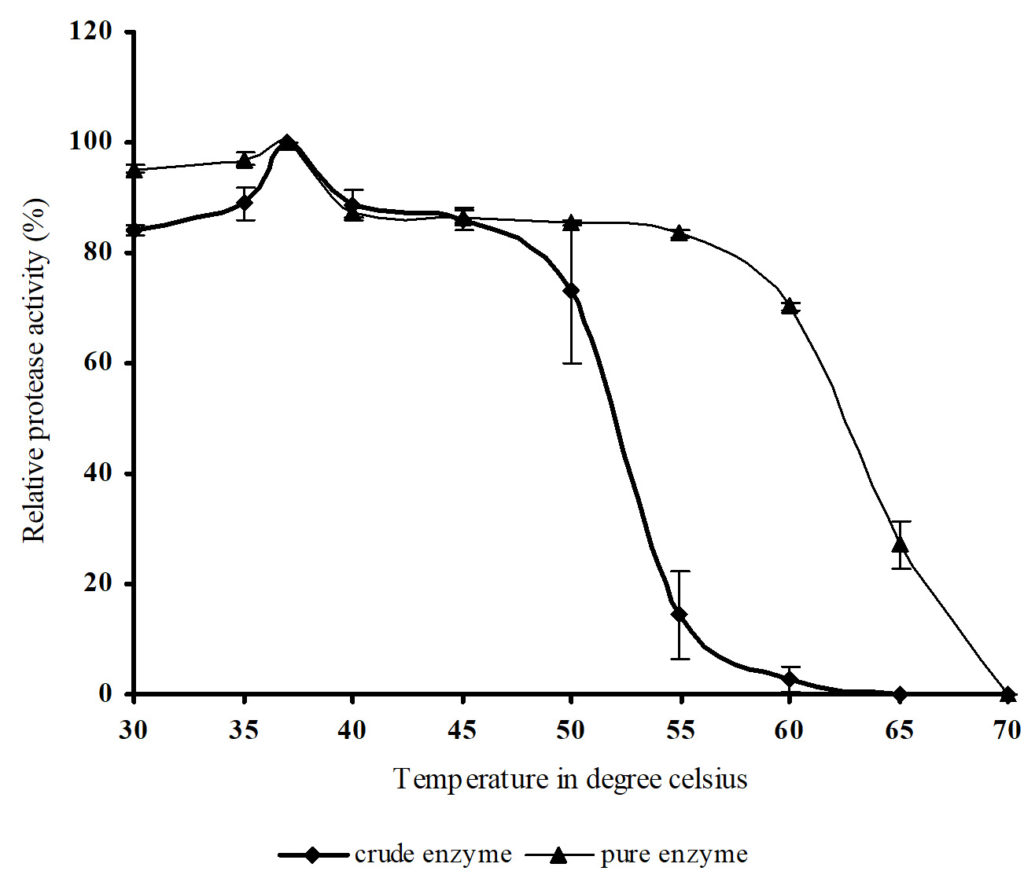
Fig. 6. Effect of temperature on crude and purified protease activity
Protease stability in organic solvents
Enzymes are usually denatured or inactivated in the presence of organic solvents as they have a tendency to strip essential water from enzyme molecules.22 Solvent stability of the crude and purified protease was investigated by incubating it with organic solvents of varying logarithm of the partition coefficient of a particular solvent between n-octanol and water (log P) at 37°C (Figure 7). Tris-HCl buffer of pH 9.0 was used as control and the effects of various organic solvents ethanol (log P -0.24), acetone (log P -0.23), butanol (log P 0.8), benzene (log P 2.0), toluene (log P 2.5), xylene (log P 3.1) and hexane (log P 3.5) were investigated. The crude protease exhibited lower activity in all organic solvents compared to control. But the purified protease was able to tolerate hydrophobic organic solvents having log P of 2.0 and above such as benzene, toluene, xylene and hexane and retained the same residual activity as the control. The purified protease was more stable in presence of xylene and hexane by nearly double than that in control in case of xylene.
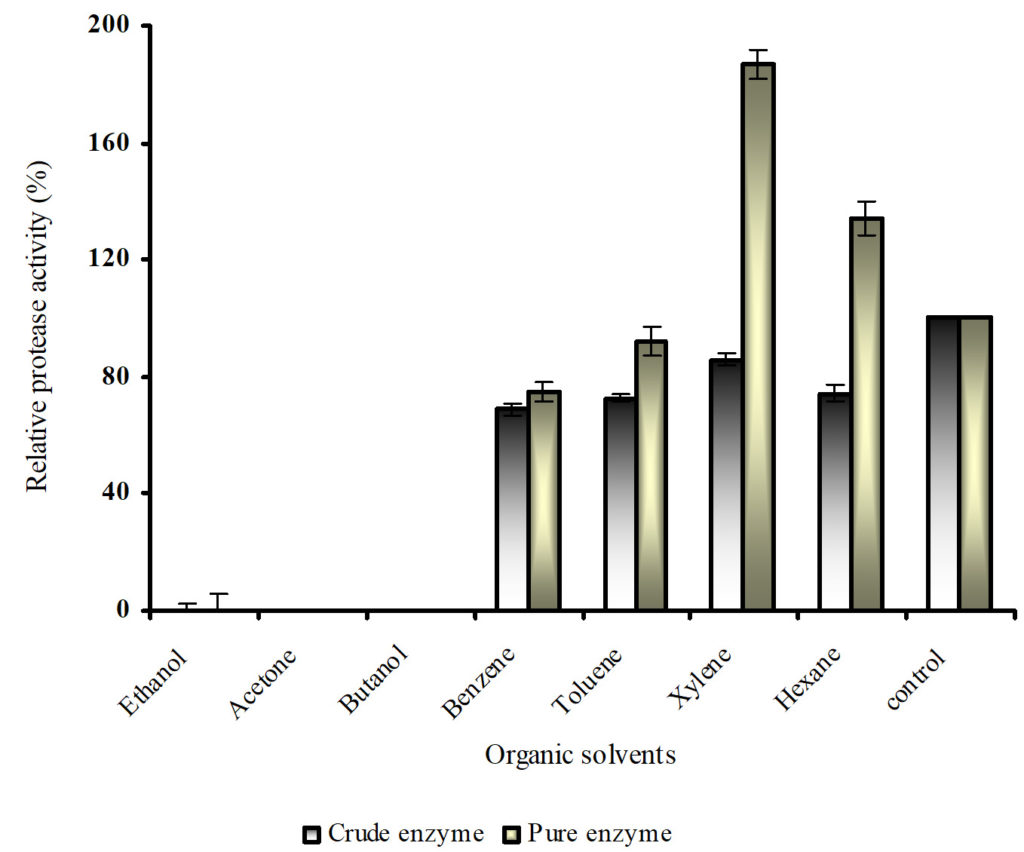
Fig. 7. Organic solvent stability of crude and purified protease
Similar results are reported by Karbalaei-Heidari.23 However, it did lose the activity in the presence of hydrophilic solvents; for instance, it suffered a 25% loss in activity in presence of butanol. Even in this study the purified protease devoid of contaminating proteins had better tolerance towards organic solvents. Based on these results, we conclude that the protease produced by Bacillus sp. is stable in hydrophobic solvents.
Effect of NaCl concentration on protease activity
The crude and purified enzyme retained 37% and 64% of the residual protease activity respectively at 5M NaCl concentration (Figure 8). The halophilic nature of the protease confers several unique properties such as stability to solvents, pH and temperature denaturation as the enzyme undergoes structural adaptations to halophilic environments.19
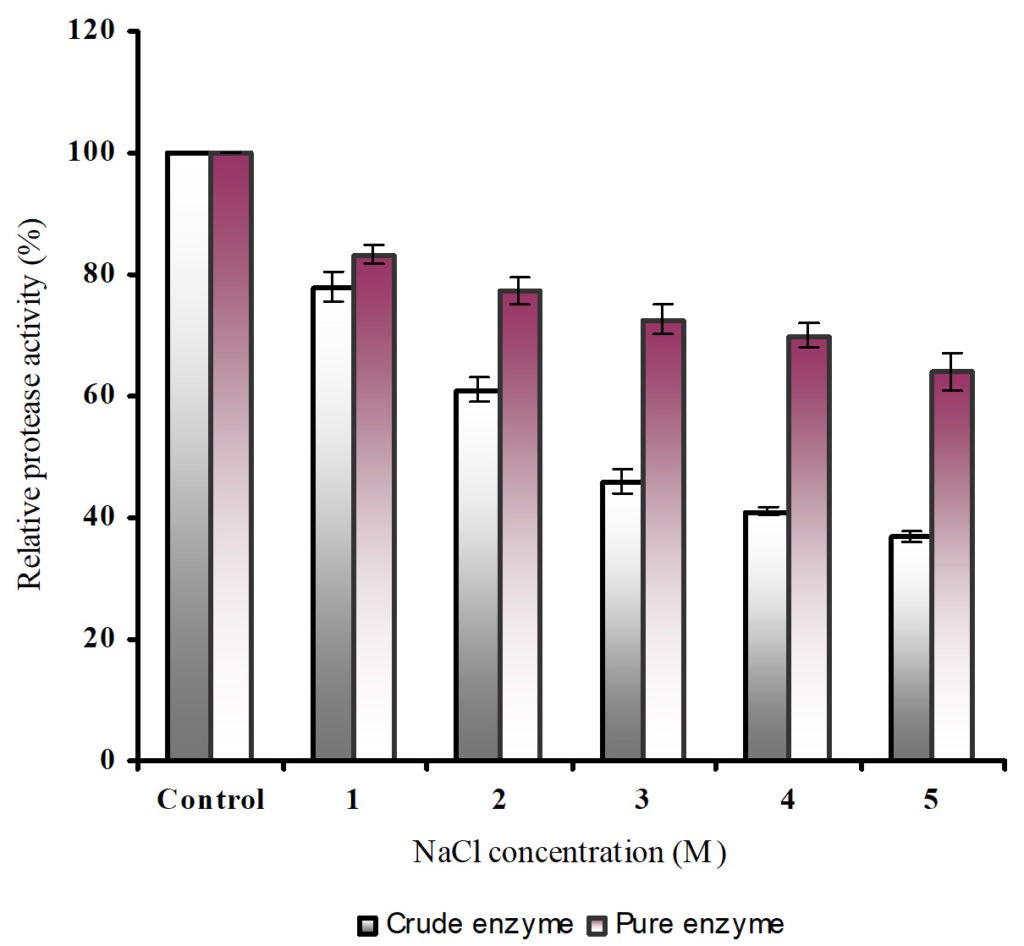
Fig. 8. Effect of NaCl concentration on crude and purified protease activity
Effect of inhibitors on protease activity
Different inhibitors are used to establish the identity and type of protease. North24 has classified proteases based on the activities in presence of various inhibitors. The alkaline protease activity was inhibited by PMSF from a facultative methylotrophic strain,11 where as our results revealed that the crude protease was completely inhibited by EDTA and iodoacetamide indicating that the preparations contained both metallo and cysteine protease. But the purified protease was completely inhibited even at 1 mM EDTA, a chelator of divalent cations suggesting that the purified protease from Bacillus sp. belongs to metalloproteases group (Table 2).
Table (2):
Effect of inhibitors on protease activity.
| Inhibitors |
Concentration (mM) | Residual Activity (%) | Inhibitor class | |
|---|---|---|---|---|
| Crude enzyme | Purified enzyme | |||
| Pepstatin | 0.05 | 73.33 ± 3.79 | 95.67 ± 2.08 | Aspartic |
| Iodoacetamide | 10 | 4.33 ± 1.53 | 51.67 ± 2.52 | Cysteine |
| Iodoacetamide | 1 | 32.33 ± 0.98 | 68.89 ± 2.76 | Cysteine |
| PMSF | 10 | 104.33 ± 3.06 | 102.67 ± 2.08 | Serine |
| PMSF | 1 | 102.78 ± 2.67 | 101.45 ± 1.57 | Serine |
| EDTA | 1 | 0 | 0 | Metallo |
Effect of metal ions on protease activity
To further characterize the Bacillus sp. halotolerant alkaline protease, the effects of divalent metals on crude and purified protease activity were examined. The divalent ions such as Ca2+, Mg2+, Zn2+ and Na+ exhibited positive regulation on enzyme activity and maximum relative activity (%) was observed in presence of Zn2+. The protease activity was inhibited by addition of FeCl2 and completely inhibited in presence of BaCl2, EDTA and HgCl2 (Table 3).
Table (3):
Effect of metal ions on protease activity.
| Metal ions (10mM) | Relative Activity (%) | |
|---|---|---|
| Crude enzyme | Purified enzyme | |
| CaCl2 | 91.67 ± 1.53 | 451.33 ± 5.69 |
| MgSO4 | 104 ± 3.00 | 216.33 ± 2.52 |
| ZnSO4 | 34.33 ± 2.52 | 552.33 ± 4.16 |
| NaCl | 95 ± 2.00 | 138 ± 2.65 |
| FeCl2 | 54.67 ± 2.52 | 39.67 ± 1.53 |
| BaCl2 | 0 | 0 |
| EDTA | 0 | 0 |
| HgCl2 | 0 | 0 |
| Control | 100 | 100 |
Similar results of Zn2+, Mg2+ ions activation and Hg2+ inhibition on protease activity are reported25. In addition, protease inactivated by EDTA treatment could only be activated by Zn2+, indicating that Zn2+ is the main metal ion to catalyze chemical reactions suggesting that zinc was essential for protease activity. These studies confirmed that the protease of this study was a Zn2+ activated metalloprotease.
Determination of kinetic parameters
The effect of substrate concentration on the initial velocity of the reaction was studied (Figure 9). The kinetic parameters – maximum reaction rate (Vmax) and Michaelis-Menten constant (Km) for the purified protease was estimated from the Lineweaver-Burk equation plot26. The kinetic studies of the pure enzyme obtained from Lineweaver-Burk plot of 1/v versus 1/[S]; demonstrate that the purified protease showed Km value of 15 µM, with casein as substrate. Vmax value of 0.485 U/ml for the maximum substrate concentration, kcat value of 3.23 s-1 and kcat/ Km value of 0.215 s-1/µM was obtained.
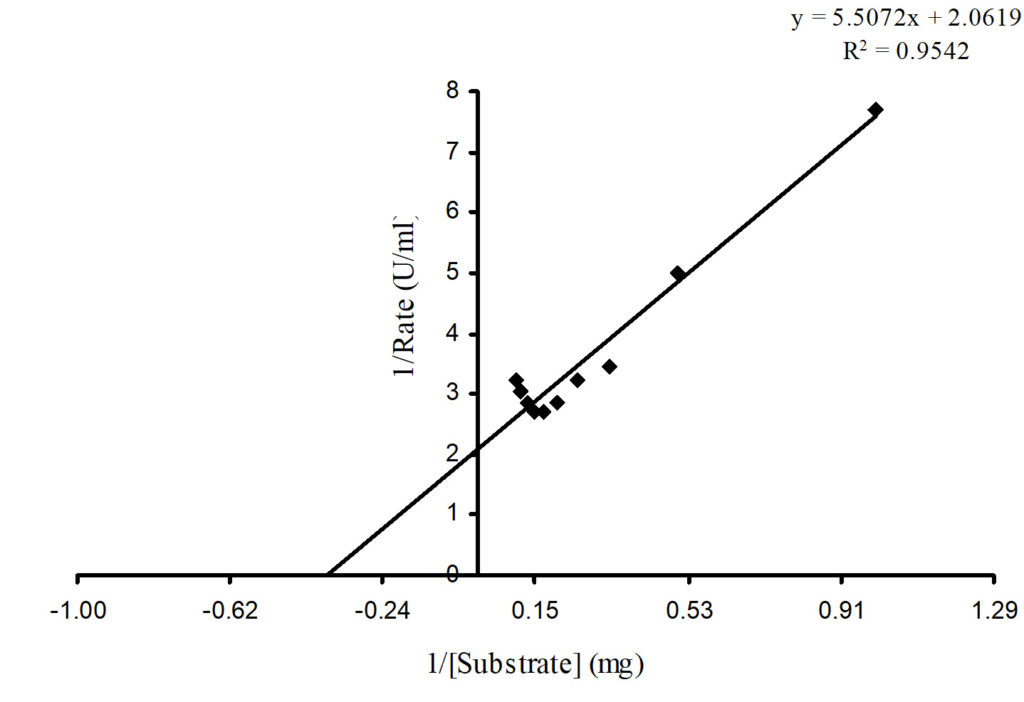
Fig. 9. Lineweaver-Burk plot
Table (4):
Kinetic parameters of purified protease from Bacillus sp.
Substrate |
Vmax (U/ml) |
Km (µM) |
kcat (s-1) |
kcat/Km (s-1/µM) |
|---|---|---|---|---|
Casein |
0.485 |
15 |
3.23 |
0.215 |
Determination of antibacterial activity – well diffusion method
The aim of the present study was to investigate the potential of the protease from Bacillus sp. MTCC 9558 to generate antibacterial peptides from different natural protein sources. It was observed that the peptides from bovine casein as protein source exhibited clearing zone (0.15 cm from the edge of the well) against Staphylococcus aureus, Pseudomonas aeruginosa and Salmonella typhimurium at protein concentration of 0.7 mg/well (Figure 10). A number of bioactive peptides have been identified in milk proteins, such as casein and whey proteins, where they are present in an encrypted form, stored as propeptides or mature C-terminal peptides that are only released upon proteolysis.10
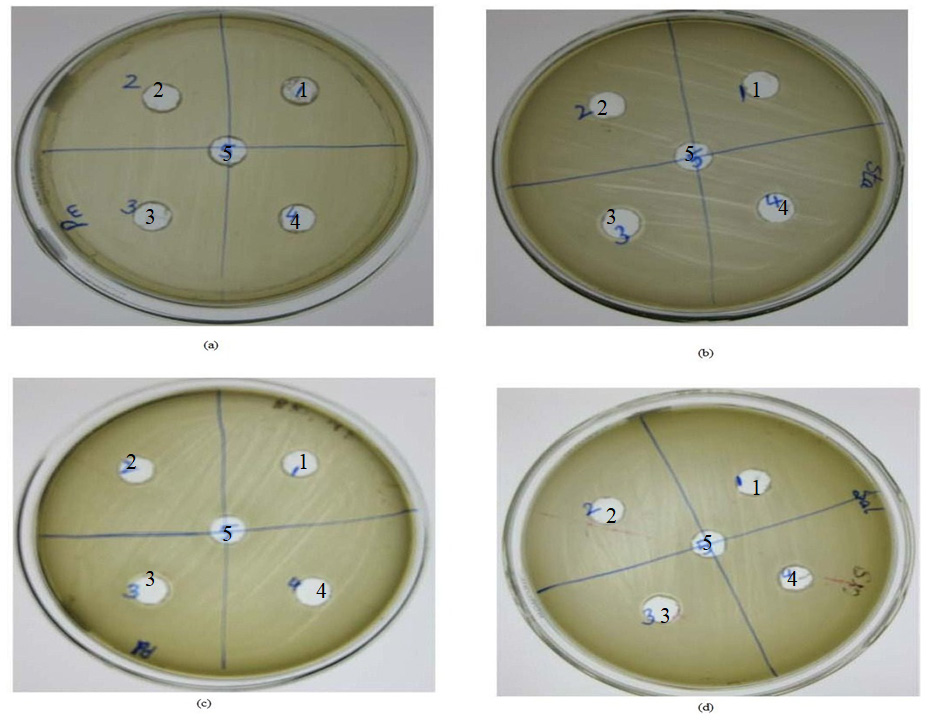
Fig. 10. Well diffusion studies for antibacterial activity of bioactive peptides
Table (5):
Characteristics of purified protease from Bacillus sp.
Parameter |
Characteristics |
|---|---|
Protease class |
Metalloprotease |
Molecular Mass |
80.77 kDa |
Optimum pH |
9.0 |
Optimum temperature |
37°C |
Thermostability |
70% (for 60 minutes at 60°C) |
Kinetic value |
Vmax (U/ml): 0.485; Km (µM): 15; kcat (s-1): 3.23; kcat/ Km (s-1/µM): 0.215 |
Metal ion activators |
Ca2+, Mg2+, Zn2+ and Na+ |
Proteolytic nature |
Endoprotease |
Commercialization of alkaline proteases depends on their stability during isolation, purification and storage as well on their robustness against solvents, surfactants and oxidants. Purification improved certain desired characteristics of enzyme such as temperature stability, organic solvent tolerance and 5.0 M NaCl. The alkaline protease was inhibited by 10 mM EDTA and significantly activated by Zn2+ indicating that it was a Zn2+ activated metalloprotease. The kinetic constants Vmax value 0.485 U/ml, Km value 15 µM, kcat value 3.23 s-1 and kcat/Km value 0.215 s-1/µM were determined. After the studies on the purified protease characteristics, we believe that this protease is novel and have the potential to be developed as an enzyme for industrial or medical applications. With this insight, the applications of purified protease in protein hydrolysis for bioactive peptide synthesis were studied. The potential of the protease from Bacillus sp. MTCC 9558 to generate antibacterial peptides from different natural protein sources was determined.
Thus, the significant emphasis has been given to the development of safe and effective food supplements and serving various bioactivities peptides derived from hydrolyzed food proteins. Moreover, the utilization of proteins or their hydrolysates for food and cosmetic applications not only provides additional advantages but also confers nutritional and functional properties.
ACKNOWLEDGMENTS
The authors are grateful to Department of Biotechnology, Kerala University for all the support in availing facilities.
- Amersham Pharmacia Biotech, Protein Purification Handbook, AB ed., Amersham Pharmacia Biotech Inc. New Jersey, USA, 1999.
- G. Zubay, Biochemistry, second ed., Macmillan Publishing Co., New York, USA, 1988.
- Huang, Q., Peng, Y., Li, X., Wang, H., Zhang, Y. . Purification and characterization of an extracellular alkaline serine protease with dehairing function from Bacillus pumilus. Curr. Microbio., 2003; 46: 0169 – 0173.
- Kim, S.S., Kim, Y.J., Rhee, I.K. Purification and characterization of a novel extracellular protease from Bacillus cereus KCTC 3674. Arch. Microbiol. 2001; 175: 458 – 461.
- Kobayashi, T., Hakamada, Y., Adachi, S., Hitomi, J., Yoshimatsu, T., Koike, K., Kawai, S., Ito, S. Purification and properties of an alkaline protease from alkalophilic Bacillus sp. KSM-K16. Appl. Microbiol. Biotechnol. 1995; 43(3): 473 – 481.
- Miyaji, T., Otta, Y., Shibata, T., Mitsui, K., Nakagawa, T., Watanabe, T., Niimura, Y., Tomizuka, N. Purification and characterization of extracellular alkaline serine protease from Stenotrophomonas maltophilia strain S-1. Lett. Appl. Microbiol. 2005; 41: 253 – 257.
- Rao, M.B., Tanksale, A.M., Ghatge, M.S., Deshpande, V. Molecular, biotechnological aspects of microbial protease. Microbiol. Mol. Biol. R. 1998; 62: 597 – 635.
- Cha, M., Park, J.R., Yoon, K.Y. Purification and characterization of an alkaline serine protease producing angiotensin I-converting enzyme inhibitory peptide from Bacillus sp. SS103. J Med. Food. 2005; 8(4): 462 – 468.
- Kim, S.K., Kim, Y.T., Byun, H.G., Park, P.J., Ito, H. Purification and characterization of antioxidative peptides from bovine skin. J. Biochem. Mol. Biol. 2001; 34: 214 – 219.
- Kamysu, W., Okroj, M., Lukasiak, J. Novel properties of antimicrobial peptides. Acta Pol. Biochim. 2003; 50: 236 – 239.
- Ma, C.L., Ni, X.M., Chi, Z.M., Ma, L.Y., Gao, L.M. Purification and characterization of an alkaline protease from the marine yeast Aureobasidium pullulans for bioactive peptide production from different sources. Mar. Biotechnol. 2007; 9: 343 – 351.
- Prasad, R., Abraham, T.K., Nair A.J. Scale up of production in a bioreactor of a halotolerant protease from moderately halophilic Bacillus sp. isolated from soil. Braz. Arch. Biol. Technol. 2014; 57(3): 448 – 455.
- Prasad, R., Nair A.J. Immobilization of Bacillus sp. protease on different matrices and its enzymatic characterization. Int. J. Pharm. Bio. Sci. 2013; 4(4): 155 – 161.
- Kunitz, M. Crystalline soybean trypsin inhibitor. J. Gen. Physiol. 1947; 30: 291 – 310.
- Lowry, O.H., Rosebrough N.J., Farr A.L., Randall R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 1951; 193: 265 – 275.
- Joo, H.S., Chang, C.S. Oxidant and SDS-stable alkaline protease from a halotolerant Bacillus clausii I-52: enhanced production and simple purification. J. Appl. Microbiol. 2005; 98: 491 – 497.
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nat. 1970; 227: 680 – 685.
- Gupta, M.N., Roy, I. Bioseparation of proteins: Some cutting edge technologies. Chem. World. 2002; 1: 30 – 31.
- Karan, R., Khare, S.K. Purification and characterization of solvent-stable protease from Geomicrobium sp. EMB 2. Environ. Technol. 2010; 31: 1061 – 1072.
- Aysha K., Haneef, U.R., Shah Ali, U.Q., Abdul, H.B., Mustafa, K. Purification and characterization of thiol dependent, oxidation-stable serine alkaline protease from thermophilic Bacillussp. J. Genet. Eng. Biotechnol. 2015; 13(1): 59 – 64.
- Seong, C.N., Jo, J.S., Choi, S.K., Kim, S.W., Kim, S.J., Lee, O.H., Han, J.H., Yoo, J.C. Production, purification & characterization of a novel thermostable serine protease from soil isolate Streptomyces tendae. Biotechnol. Lett. 2004; 26: 907 – 909.
- Ghorbel, B., Kamoun, A.S., Nasri, M. Stability studies of protease from Bacillus cereus BG1. Enzyme Microb. Technol. 2003; 32: 513 – 518.
- Karbalaei-Heidari, H.R., Ziaee, A.A., Amoozegar, M.A. Purification and biochemical characterization of a protease secreted by the Salinivibrio sp. strain AF-2004 and its behaviour in organic solvents. Extremophiles. 2007; 11: 237 – 243.
- North, M.J. Comparative biochemistry of the proteinases of eukaryotic microorganisms. Microbiol. Rev. 1982; 46: 308 – 340.
- Yuan, Q., Hayashi, A., Kitamura, Y., Shimada, T., Na, R., Jin, X. Purification and characterization of cold-adapted metalloprotease from deep sea water lactic acid bacteria Enterococcus faecalis TN-9. Int. J. Biol. 2009; 1(2): 12 – 21.
- Lineweaver, H., Burk, D. The Determination of Enzyme Dissociation Constants. J. Am. Chem. Soc. 1934; 56(3): 658 – 666.
© The Author(s) 2017. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


