ISSN: 0973-7510
E-ISSN: 2581-690X
Milk and dairy products form a significant part of the human diet. Temperature control is considered as the key to extend shelf life for dairy products. Psychrotrophs grow at temperature such as 7oC or lower but had higher optimal growth temperature. Most psychrotrophs produce extracellular enzymes while growing in milk and deteriorate its quality. Six types of samples constituting buffalo milk, flavored milk, curd, shrikhand, lassi and butter were collected in different seasons to study the psychrotrophic microbiota. Sixty psychrotrophic bacteria were grouped into fifteen clusters of very similar isolates. One isolate from each of the fifteen clusters of closely related isolates was identified by phylogenetic analysis namely Stenotrophomonas maltophilia, Pseudomonas putida, Pseudomonas stutzeri, Enterobacter hormaechei, Macrococcus caseolyticus, Exiguobacterium acetylicum, Exiguobacterium indicum, Micrococcus luteus, Kocuria rosea, Staphylococcus equorum, Ornithinmicrobium humiphilum, Planococcus psychrotoleratus, Bacillus cereus and two isolates upto genus level namely Exiguobacterium spp. and Kocuria spp. Fatty acid profile of milk sample inoculated with lipolytic psychrotrophic bacterial isolate was determined using Hydrolytic Extraction Gas Chromatographic method.
Psychrotrophs, dairy products, spoilage, gas chromatography.
Psychrotrophs grow in cold environments due to unique features like cold shock proteins, short and unsaturated fatty acids in membranes, enzymes with high specific activity, thermolability and genetic changes to thermal shifts (Margesin et al., 2007). Milk is an ideal and perfect medium for growth of bacteria and therefore it gets contaminated very easily and readily. It is extremely perishable in nature and its shelf life is limited to 3 to 4 hours depending upon the temperature of storage (Kumar, A. and Seth, R., 2008). Extension of shelf life from hours to months has been a prime objective of the dairy industry for many years to meet the demands for increasing distribution times and distances (Goff, H.D. and Griffiths, M.W., 2006). The growth of microorganisms in milk causes disintegration of fat, protein and lactose and will soon make the product unsuitable for drinking. The microbial changes in milk and milk products are produced as a result of fermentation of one or more of the milk constituents by the causative micro-organisms. These milk fermentations may be of normal or abnormal type. Prolonging the storage of raw milk before pasteurization results in increasing the psychrotrophic, lipolytic and proteolytic bacteria that produce enzymes and cause the milk to change, such that there will be problems in processing the milk and quality of the milk products. Law, B.A. reported that Farm refrigeration of cow’s milk has led to increased levels of psychrotrophic Gram negative rods (Law, B.A., 1979). Proteolysis by the psychrotrophic microorganisms results in coagulation of the milk and unclean and bitter flavor of the milk (Burdova et al., 2002). Psychrotrophic bacteria grow luxuriantly in dairy products and multiply at refrigeration temperature (Xin et al, 2017). The problem of maintaining good shelf life or achieving extended keeping quality has always been one of great concern to milk processors. Keeping quality of milk products depends on initial quality of the products, on care and techniques employed in processing and distribution and on storage temperatures.
Collection and coding of samples
Forty different samples of milk and milk products constituting 10 samples of pasteurized milk, 5 samples of flavored milk, 5 samples of curd, 9 samples of shrikhand, 9 samples of lassi and 2 samples of butter were collected from ten different dairies of Nasik district.
Isolation and identification
Isolation of psychrotrophs was carried out by streak plate technique using sterile milk agar plates. Plates were incubated at 7oC for 10 days. Colonies developed on milk agar were picked up, purified repeatedly and preserved on nutrient agar slants and sub-cultured after every two months. Isolates of different morphotypes were appropriately coded and studied for morphological and biochemical characters.
Physiological studies of the isolates
The effect of pH, temperature and salt concentration on growth of isolates was studied using the method given by Cruickshank et al. (1975). Log phase cultures of each isolate of a cell density 106 cells per ml were inoculated in 0.01 ml quantity in to 5 ml of sterile nutrient broth in triplicates adjusted to different pH values such as 4, 5, 6, 7, 8, 9, 10, 11 and 12. To study the effect of different temperatures on growth, cells were inoculated in 0.01 ml quantity in to 5 ml of sterile nutrient broth in triplicates. The sets were incubated at different temperatures such as 7oC for ten days, 15oC, 25oC, 37oC, 45oC and 55oC for 24 hrs. Each of isolates were inoculated in 0.01 ml quantity in to 5 ml of sterile nutrient broth in triplicates having different concentrations of NaCl such as 0.5%, 1%, 1.5%, 2%, 2.5%, 3%, 3.5%, 4%, 4.5%, 5%, 5.5%, 6% 6.5%, 7%, 7.5% and 8% for observation of growth at different salt concentrations. All the sets were incubated at 7oC for ten days. After incubation tubes were observed for the growth.
Molecular identification and phylogenetic analysis
The Microseq 16S rRNA gene kit (Applied Biosystems Division) was used for PCR and sequencing. The amplification of the genomic DNA was carried out by PCR, upon optimization. (ABI 9700 geneamp PCR). The facility was availed from Molecular Diagnostic Centre, Pune. The sequence was pasted in the BLAST tool and program was run using default parameters against non-redundant databases to avoid redundancy in the obtained hits. Hits obtained were analyzed and used for identification of the isolates.
Analysis of fatty acids profile
The 500 ml tryptone dextrose yeast extract broth media was inoculated at 1.0% (v/v) with cultures of lipolytic isolate and incubated at 7oC for 10 days along with the control flask (uninoculated pasteurized milk). After incubation of milk samples (Control and test) fatty acid profile was determined using Hydrolytic Extraction Gas Chromatographic method (AOAC Official Method 996.06).
Isolation of psychrotrophic bacteria and coding of isolates
A total of sixty psychrotrophic bacterial isolates were obtained from six different types of milk and fermented milk products which constitutes nineteen isolates from milk namely RAtM-1, RAtM-2, RAtM-3, RAtM-4, WSM-1, WSM-2, WSM-3,WTM-1, WTM-2, RRM-1, SPM-1, SShM-1, SNM-1, WAM-1, WVM-1, WVM-2, WVM-3, RGM-1 and RGM-2, five isolates from flavored milk namely SShFM-1, RShFM-1, RRFM-1, RPFM-1 and WPFM-1, five isolates from curd namely SAtC-1, SGC-1, SSC-1, WSC-1and RSC-1, fourteen isolates from shrikhand namely RRS-1, RTS-1, WTS-1, WTS-2, WShS-1, WShS-2, WAtS-1, RVS-1, RVS-2, WVS-1, WVS-2, RSS-1,WSS-1and WSS-2, fifteen isolates from lassi namely WTL-1, RShL-1, SShL-1,SNL-1, RGL-1, RGL-2, SGL-1, SGL-2, SSL-1, WRL-1, WRL-2, RAL-1, RAL-2, RAL-3 and RAL-4 and two isolates from butter samples namely RVB-1 and RSB-1.
Table (1):
Characterization of psychrotrophic bacteria
Sr. No. |
Gram nature, motility, morphology |
No. of Isolates |
|---|---|---|
1 |
Gram negative motile rods |
19 |
2 |
Gram negative motile short rod |
1 |
3 |
Gram positive nonmotile cocci |
22 |
4 |
Gram positive motile short rods |
9 |
5 |
Gram positive motile rods |
5 |
6 |
Gram positive nonmotile short rods |
3 |
7 |
Gram positive motile Cocci |
1 |
Microbiological examination of milk, flavored milk, curd, shrikhand, butter and lassi
Santana et al. (2004) reported that the psychrotrophic bacteria in the refrigerated milk include 69% Gram negative bacilli that means 45% of the total proteolytic microorganisms in the dairy process.
Table (2):
Utilization of Carbohydrates by isolates
Sr.
No. |
Sugar |
Positive Isolates |
|---|---|---|
1 |
Glucose |
36 |
2 |
Lactose |
01 |
3 |
Mannitol |
21 |
4 |
Sucrose |
34 |
5 |
Arabinose |
29 |
6 |
Xylose |
15 |
7 |
Galactose |
20 |
8 |
Cellobiose |
17 |
9 |
Maltose |
41 |
10 |
Trehalose |
31 |
11 |
Mannose |
21 |
12 |
Raffinose |
03 |
13 |
Rhamnose |
08 |
14 |
Ribose |
18 |
15 |
Sorbitol |
12 |
16 |
Fructose |
34 |
17 |
Dulcitol |
05 |
During cold storage after milk collection, psychrotrophic bacterial populations dominate the microflora and their extracellular enzymes, mainly proteases and lipases contribute to the spoilage of dairy products (Elionora, H.Z. and Malka, H., 2007). Bashir et al. (2010) used citrate test, methyl red test and Voges-Proskauer’s test for identification of psychrotrophic bacteria isolated from freezer (-20°C).
Table (3):
Enzymatic characteristics
Sr. No. |
Enzyme |
Positive Isolates |
|---|---|---|
1 |
Catalase |
60 |
2 |
Oxidase |
44 |
3 |
Gelatinase |
28 |
4 |
Amylase |
14 |
5 |
Protease |
29 |
6 |
Lipase |
50 |
7 |
Lecithinase |
05 |
8 |
Phosphatase |
18 |
9 |
Urease |
09 |
10 |
Nitrate Reductase |
24 |
11 |
Arginine Hydrolase |
03 |
12 |
Ornithin Decarboxylase |
07 |
13 |
Indole test |
00 |
14 |
Methyl Red test |
12 |
15 |
Voges-Proskauer’s test |
08 |
16 |
Citrate Utilization test |
31 |
Physiological characteristics of the isolates
The isolates showed variable response of growth towards temperature, pH and salt concentrations. All sixty isolates showed maximum growth at pH 7. None of the sixty isolates showed growth at pH 4. Only one isolate showed growth at a very high pH value i.e. pH 12. All sixty isolates showed growth at 7oC, 15oC and 25oC. Forty one isolates showed growth at 37oC. Out of sixty isolates, only five isolates showed growth at 45oC. None of the sixty isolates grew at 55oC temperature. All sixty isolates showed growth at mesophilic temperature as well as at 7oC, an important distinguishing characteristic feature of psychrotrophic bacteria. Psychrotrophic bacterial isolates showed variable response of the growth to different salt concentrations ranging from 0.5% to 8%. All sixty isolates showed growth at a salt concentration in a range of 0.5% to 3%. None of the isolates grew at 8% salt concentration. It can also be seen that twelve isolates tolerated maximum 3.0% salt concentration, eleven isolates tolerated maximum 3.5% salt concentration, five isolates tolerated maximum 4% salt concentration, eight isolates tolerated maximum 4.5% salt concentration, five isolates tolerated maximum 5% salt concentration, two isolates tolerated maximum 5.5% salt concentration, six isolates tolerated maximum 6% salt concentration, two isolates tolerated maximum 7% salt concentration and nine isolates tolerated maximum 7.5% salt concentration.
Table (4):
Clustering of isolates
Cluster |
Isolates |
Sample |
|---|---|---|
1 |
RAtM-1, WVM-1, WTM-2, WPFM-1, WTS-2, SGL-2, RAL-4 (07) |
Milk, Flavored milk,
Shrikhand, Lassi |
2 |
RAtM-4 , WAM-1, RVS-2, RAL-1 (04) |
Milk, Shrikhand, Lassi |
3 |
SPM-1, WTM-1, SShM-1, WSC-1, WRL-2, RAL-2, RAL-3, RSB-1 (08) |
Milk, Curd, Lassi, Butter |
4 |
SAtC-1 (01) |
Curd |
5 |
SGL-1 (01) |
Lassi |
6 |
WSM-2, WShS-1, WRL-1, WVM-2 (04) |
Milk, Shrikhand, Lassi |
7 |
WSM-1, RRFM-1 (02) |
Milk, Flavored Milk |
8 |
WVM-3, WSM-3, WShS-2 (03) |
Milk, Shrikhand |
9 |
RAtM-2, RRM-1, RGM-1, SGC-1, WVS-2, SNL-1, RGL-2, RVB-1 (08) |
Milk, Curd, Shrikhand,
Lassi, Butter |
10 |
RGM-2, WSS-2, WAtS-1 (03) |
Milk, Shrikhand |
11 |
SShFM-1 (01) |
Flavored Milk |
12 |
RSC-1 , SSC-1, WSS-1, RSS-1, RRS-1, SShL-1, RGL-1, SSL-1, WTL-1 (09) |
Curd, Shrikhand, Lassi |
13 |
RAtM-3, RTS-1, RShL-1 (03) |
Milk, Shrikhand, Lassi |
14 |
WVS-1 (01) |
Shrikhand |
15 |
SNM-1, RPFM-1, RShFM-1, RVS-1, WTS-1 (05) |
Milk, Flavored Milk, Shrikhand |
Forming clusters of isolates based on morphological, cultural, biochemical and physiological characters
On the basis of maximum similarities of characteristics studied, sixty isolates of psychrotrophic bacteria were grouped into fifteen clusters of very similar isolates and labelled as 1 to 15. Isolates of each of the fifteen clusters were obtained from different sources.
Molecular identification and phylogenetic analysis
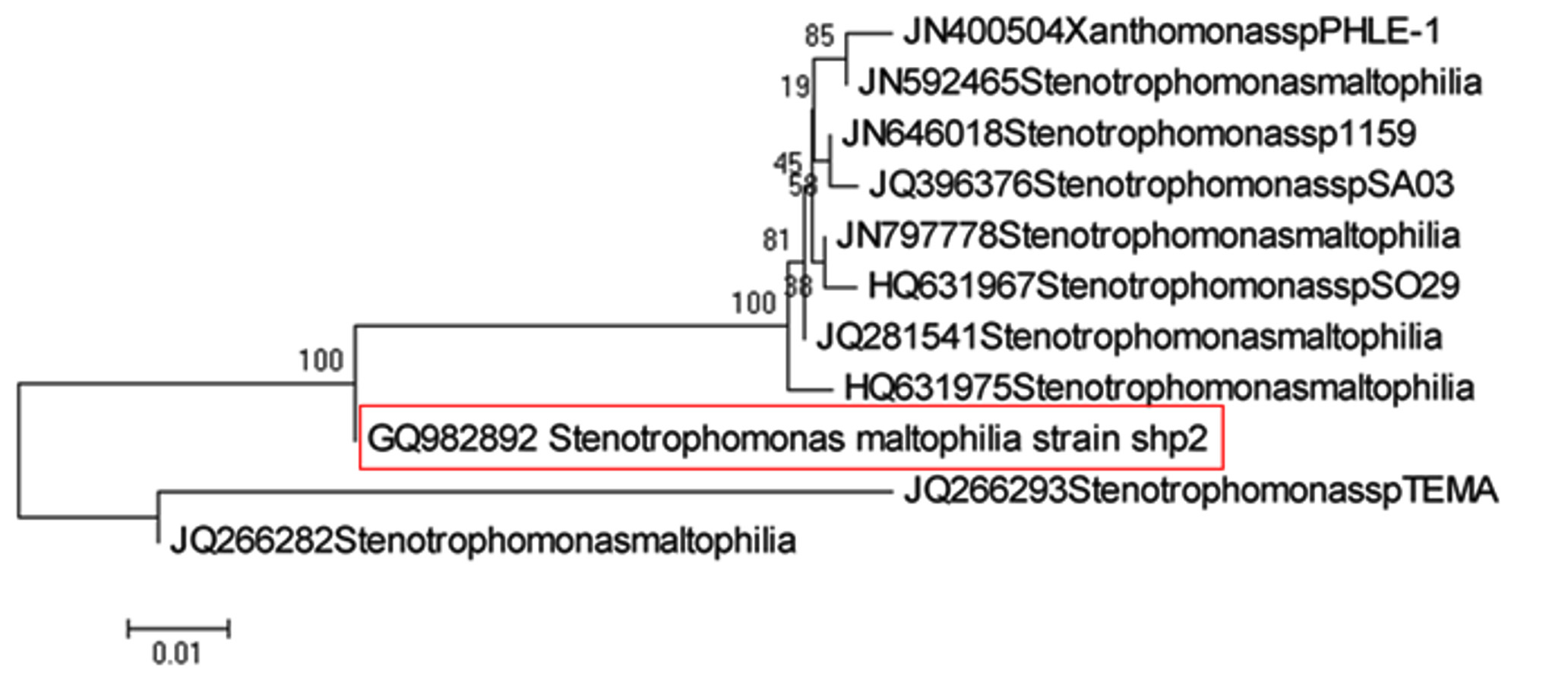
One isolate from each of the fifteen clusters of closely related isolates was identified by phylogenetic analysis and sequencing 16S rRNA gene. The results of molecular identification of such fifteen representative isolates are presented in Table 5 and Fig. 1 to 11.
Table (5):
Identification of bacterial isolates
Code of Isolate |
Phylum |
Order |
Family |
Genus |
Species |
|---|---|---|---|---|---|
RAL-4 |
Proteobacteria |
Xanthomonadales |
Xanthomonadaceae |
Stenotrophomonas
|
maltophilia
|
WAM-1 |
Proteobacteria |
Pseudomonadales |
Pseudomonadaceae |
Pseudomonas |
putida
|
SPM-1 |
Proteobacteria |
Pseudomonadales |
Pseudomonadaceae |
Pseudomonas |
stutzeri |
SAtC-1 |
Proteobacteria |
Enterobacteriales |
Enterobacteriaceae |
Enterobacter |
hormaechei
|
SGL-1 |
Firmicutes |
Bacillales |
Staphylococcaceae |
Macrococcus |
caseolyticus
|
WSM-2 |
Firmicutes |
Bacillales |
Bacillaceae |
Exiguobacterium |
acetylicum |
RRFM-1 |
Firmicutes |
Bacillales |
Bacillaceae |
Exiguobacterium |
indicum |
WShS-2 |
Firmicutes |
Bacillales |
Bacillaceae |
Exiguobacterium spp. |
– |
RRM-1 |
Actinobacteria |
Actinomycetales |
Micrococcaceae |
Micrococcus |
luteus |
WSS-2 |
Actinobacteria |
Actinomycetales |
Micrococcaceae |
Kocuria |
rosea |
SShFM-1 |
Actinobacteria |
Actinomycetales |
Micrococcaceae |
Kocuria spp. |
– |
SSL-1 |
Firmicutes |
Bacillales |
Staphylococcaceae |
Staphylococcus |
equorum |
RAtM-3 |
Actinobacteria |
Actinomycetales |
Intrasporangiaceae |
Ornithinmicrobium |
humiphilum |
WVS-1 |
Firmicutes |
Bacillales |
Planococcaceae |
Planococcus |
psychrotoleratus |
SNM-1 |
Firmicutes |
Bacillales |
Bacillaceae |
Bacillus |
cereus |
As shown in Fig. 1, isolate namely RAL-4 which was identified as Stenotrophomonas maltophilia (Acc. No.: GQ 982892) (100% similarity) forms cluster with the clade of Stenotrophomonas maltophilia (Acc. No.: HQ631975, JQ281541 and JQ266282) indicating that this strain (RAL-4) is identified as Stenotrophomonas maltophilia.
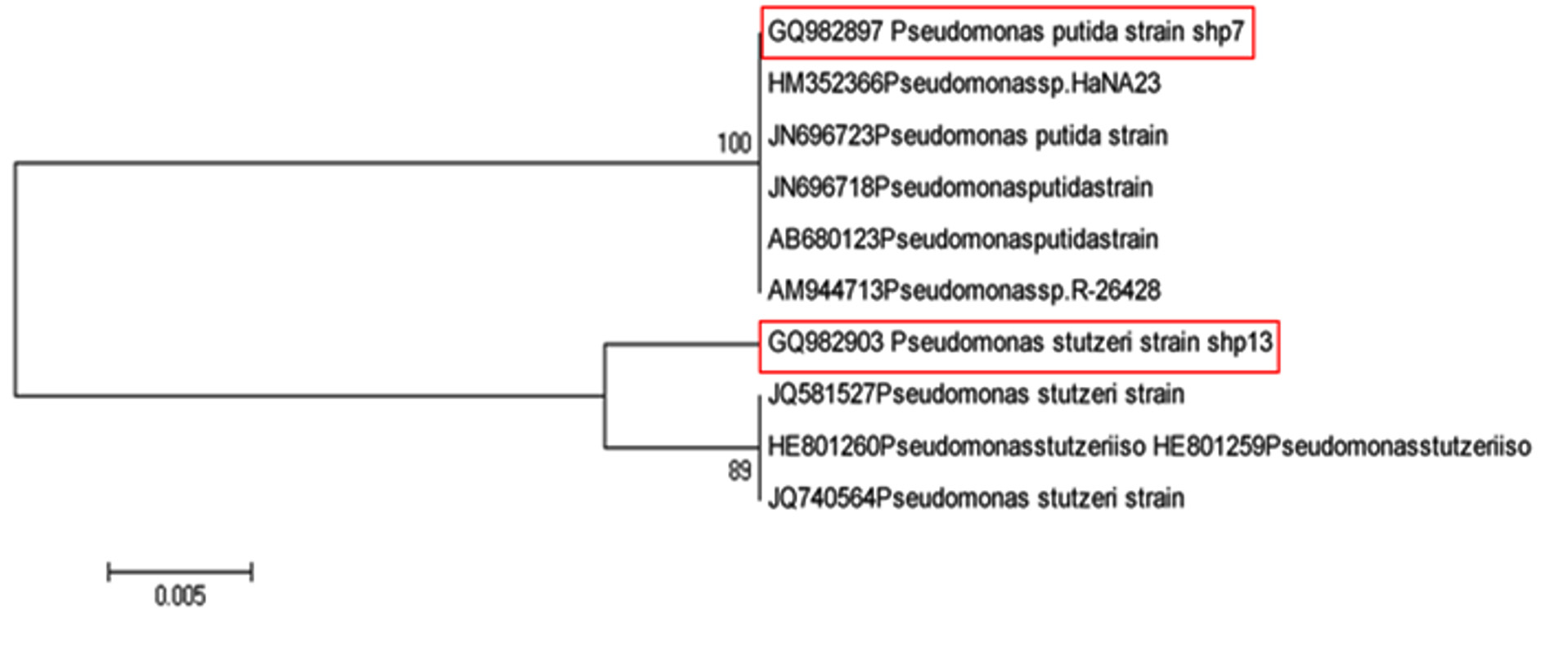
As shown in Fig. 2, isolate namely WAM-1, which was identified as Pseudomonas putida (Acc. No.: GQ982897) (100% similarity) forms cluster with the clade of five isolates namely Pseudomonas spp. HaNA23 (Acc. No.: HM352366), Pseudomonas putida strain (Acc.No. JN696723), Pseudomonas putida strain (Acc. No.: JN696718), Pseudomonas putida strain (Acc. No.: AB680123) and Pseudomonas spp. R-26428 (Acc. No.: AM944713). It can be also seen from the phylogenetic analysis that strain SPM-1, which was identified as Pseudomonas stutzeri (Acc. No.:GQ982903) (99% similarity) by analysis of 16S rRNA gene sequence analysis forms cluster with the clade of isolates of Pseudomonas stutzeri.
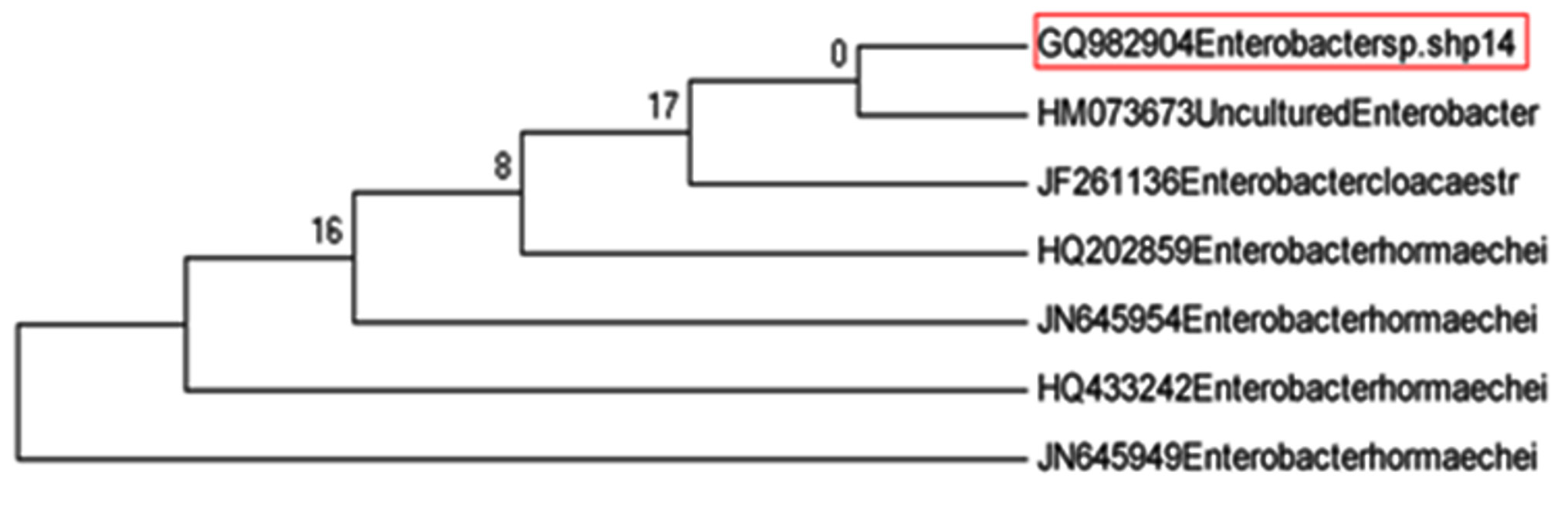
As shown in Fig. 3, isolate SAtC-1 (GQ982904) identified as Enterobacter hormaechei (100% similarity).
As shown in Fig. 4, isolate SGL-1 (GQ982900) was identified as Macrococcus caseolyticus (100% similarity) forms clade with Macrococcus caseolyticus.
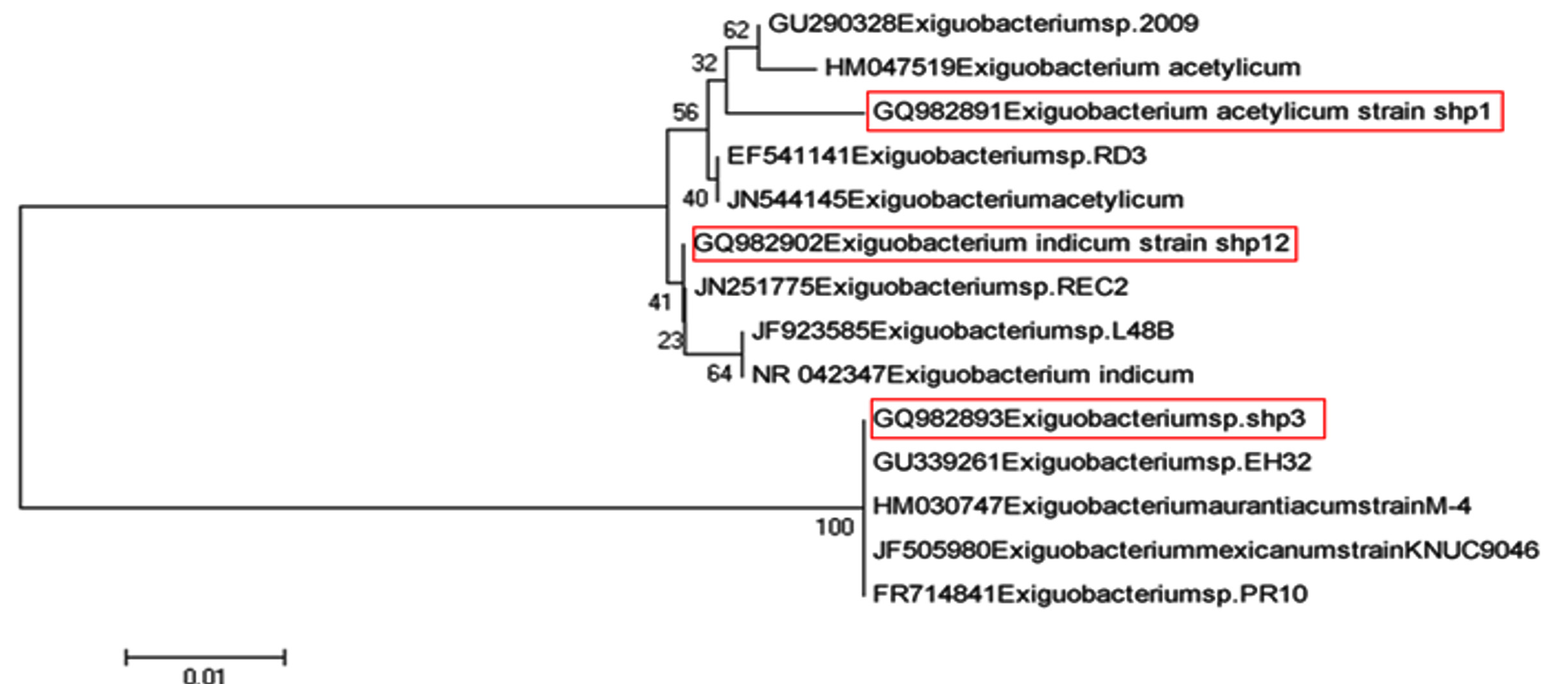
Phylogenetic analysis revealed that an isolate namely WSM-2 (Acc.No.: GQ982891) which was identified as Exiguobacterium acetylicum (98% similarity) by analysis of 16S rRNA gene sequence did not cluster with clades of Exiguobacterium acetylicum (HM047519 and JN544145) instead forms separate clade indicating that probably this strain might be a different species of Exiguobacterium acetylicum on the basis of genotypic analysis. It can be seen from the phylogenetic analysis that an isolate namely RRFM-1 (Acc. No.: GQ982902) which was identified as Exiguobacterium indicum (99% similarity) by analysis of 16S rRNA gene sequence did not cluster with clade of Exiguobacterium indicum (Acc. No.: NR042347). The strain found to be very distant from the other strains of Exiguobacterium indicum in the phylogenic tree. Phylogenetic analysis revealed that Exiguobacterium spp. ((Acc. No.: GQ982893) formed a separate cluster from that of the known Exiguobacterium type strain viz. Exiguobacterium acetylicum (Acc. No.: GQ982891) and Exiguobacterium indicum ((Acc. No.: GQ982902). It was also seen from the phylogenetic analysis that strain WShS-2 which was identified as Exiguobacterium spp. ((Acc. No.: GQ982893) by analysis of 16S rRNA gene sequence did not cluster with the clade of Exiguobacterium acetylicum (Acc. No.: GQ982891) and Exiguobacterium indicum ((Acc. No.: GQ982902) but forms clade with Exiguobacterium aurantiacum (Acc. No.: HM030747) and Exiguobacterium mexicanum ((Acc. No.: JF505980) indicating that probably this strain is complex of Exiguobacterium aurantiacum and Exiguobacterium mexicanum (Fig. 5).
As shown in Fig. 6, isolate RRM-1(GQ 982894) identified as Micrococcus luteus (100% similarity).
Phylogenetic analysis reveals that an isolate namely WSS-2, which was identified as Kocuria rosea (Acc. No.: GQ 982895) (100% similarity) by analysis of 16S rRNA gene sequence did not forms cluster with clade of Kocuria rosea (Acc. No.: JQ684249) indicating that probably this strain is different from the Kocuria rosea (Acc. No.: JQ684249). Phylogenetic analysis also reveals that an isolate namelySShFM-1 identified as Kocuria spp. (Acc. No.: GQ982898) (100% similarity) forms cluster with clade of different species of Kocuria (Fig. 7).
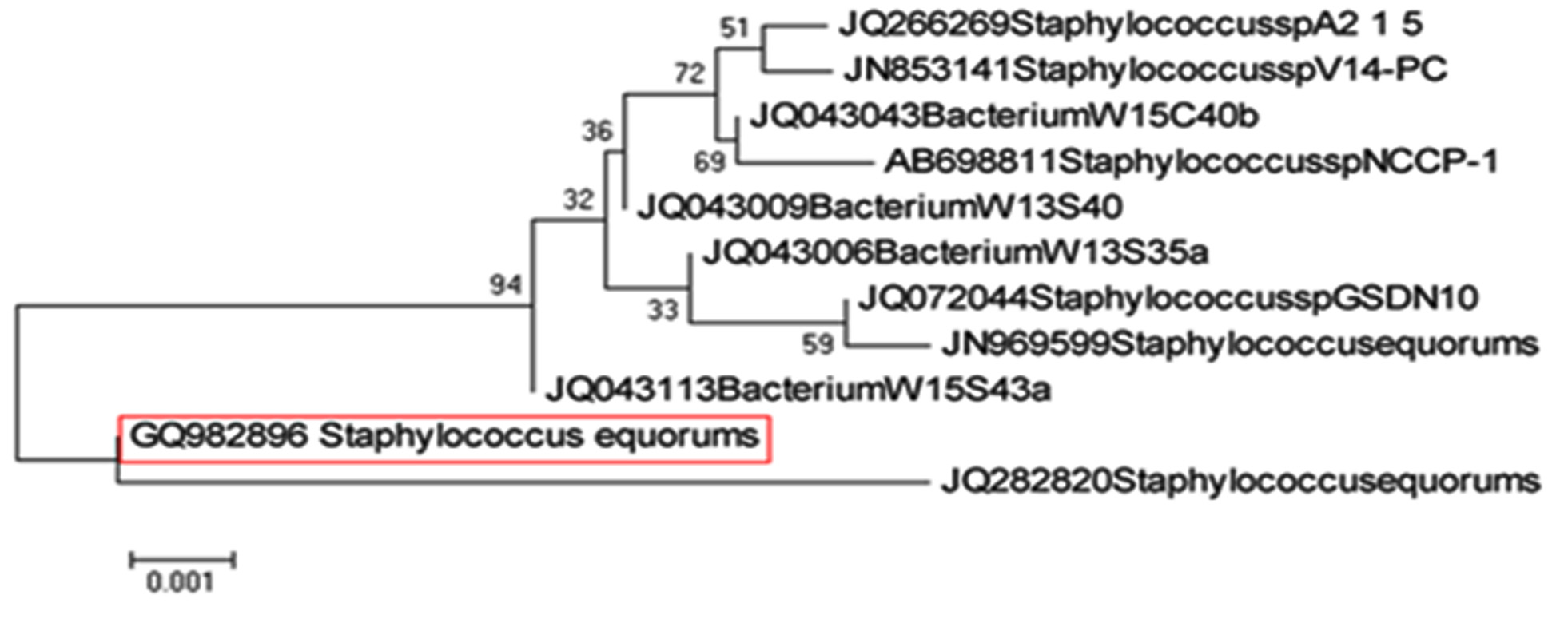
As shown in Fig. 8, SSL-1 (GQ982896) identified as Staphylococcus equorum (100% similarity) and forms cluster with clade of Staphylococcus equorum.
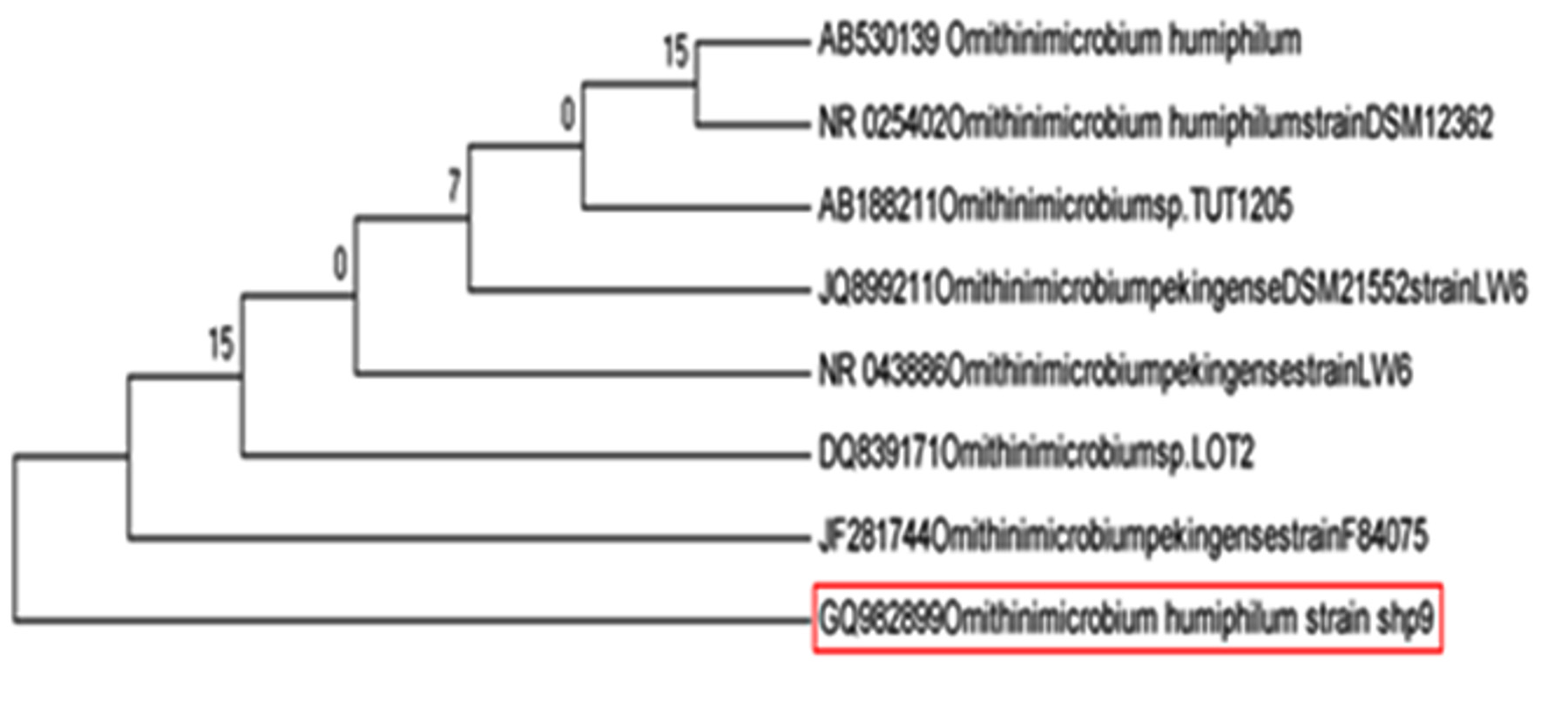
As shown in Fig. 9, RAtM-3 (GQ982899) was identified as Ornithinmicrobium humiphilum (100% similarity).
Phylogenetic analysis revealed that an isolate namely WVS-1 which was identified as Planococcus psychrotoleratus (Acc. No.: GQ982901) (96% similarity) indicating that this strain (WVS-1) is identified as Planococcus psychrotoleratus. It can be also seen from phylogenetic analysis that strain WVS-1 did not cluster with the clade of Planococcus psychrotoleratus, indicating that probably this strain might be a different or new species of Planococcus (Fig. 10)
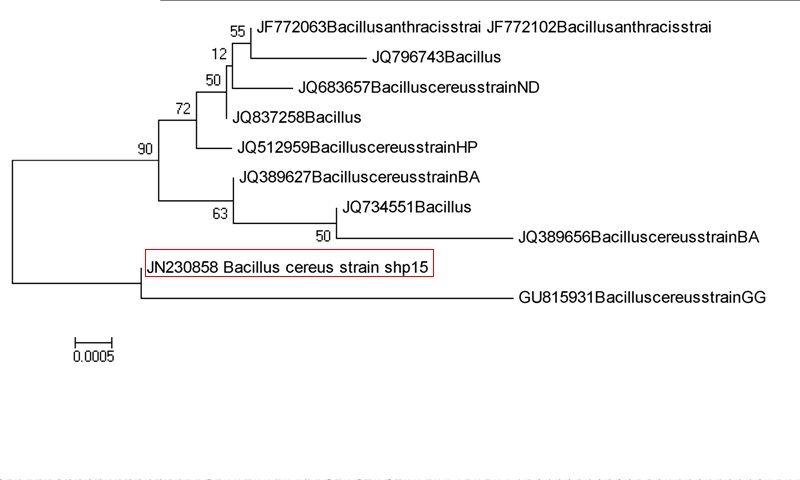
As shown in Fig. 11, isolate namely SNM-1 (Acc. No.: JN230858) is identified as Bacillus cereus and this strain SNM-1shown 99% similarity with Bacillus cereus (Acc.No.: GU81593).
Delbes et al. (2007) using the 16S rRNA gene, shown that culturable bacterial flora in raw milk were highly diversified. These organisms are one of the most prevalent groups found in the form of biofilms on dairy equipments causing spoilage of conventionally pasteurized liquid milk. Stenotrophomonas maltophilia (formerly Pseudomonas and Xanthomonas maltophilia) is a ubiquitous bacterium. It is increasingly prevalent in hospitals as an opportunistic human pathogen causing nosocomial infections (Berg et al., 1999). It is important to note that some heat resistant strains of B.cereus exhibiting psychrotrophic characteristics have been reported to grow and produce toxins in milk and dairy products at refrigeration temperatures (Guinebretiere et al., 2010). B.cereus causes problems to the food industry both by deteriorating the products and by endangering people’s life upon consuming them (Ghelardi et al., 2002). The storage and transportation of raw milk at low temperatures promote the growth of psychrotrophic bacteria and the production of thermo-stable enzymes, which pose great threats to the quality and shelf-life of dairy products (Yuan et al. 2017)
Analysis of fatty acids profile
Analysis of fatty acid profile of milk sample inoculated with Stenotrophomonas maltophilia (RAL-4) showed variations in saturated fatty acids, monounsaturated fatty acids, polyunsaturated fatty acids and trans fatty acids. Increase in short chain as well as middle length chain saturated fatty acids was observed which are responsible for development of rancid, soapy, unclean or bitter flavors. According to Chen et al., (2003) and Huis Veld J (1996), the hydrolysis of as little as 1% of milk triglycerides could lead to rancid off flavors. Jaeger et al., (1994) reported that increased levels of short chain fatty acids (C4 – C8) give rise mainly to rancid flavors, while the middle length chains (C10 – C12) give rise to most of the soapy, unclean or bitter flavors.
Although the milk and milk products are preserved primarily by refrigeration, yet the Psychrotrophic bacteria can responsible for their spoilage. On the basis of morphological, cultural, biochemical and physiological characteristics, sixty psychrotrophic bacterial isolates were grouped into fifteen clusters of closely related isolates. Psychrotrophic bacteria producing protease and lipase enzymes emphasizes the spoilage potential from them in dairy products because proteolysis is a main factor limiting shelf life of milk and milk products due to flavour and texture changes. Spoilage of milk and milk products may be avoided or prevented by controlling proteolytic and lipolytic psychrotrophic organisms or protease or lipase enzymes produced by them in milk and milk products.
Acknowledgements
The author is grateful to Dr. Sonali Sanghavi of Molecular Diagnostic Center, Pune, for base sequencing of the bacterial cultures & TUV Lab., Pune for Hydrolytic Extraction Gas Chromatography.
Funding
None
Data Availability
All datasets generated or analyzed during this study are included in the manuscript.
Ethics Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
- Bashir A., Imran J., Aamer A.S., Abdul H., Fariha H. Psychrotrophic bacteria isolated from -20°C freezer. African Journal of Biotechnology, 2010; 9: 718-724.
- Berg G., Roskot N., Smalla K. Genotypic and phenotypic relationships between clinical and environmental isolates of Stenotrophomonas maltophilia. Journal of Clinical Microbiology, 1999; 37: 3594–3600.
- Burdova O., Baranova M., Laukova A., Rozanska, H. and Rola J.G. Hygiene of pasteurized milk depending on psychrotrophic microorganisms. Bulletin of Veterinary Institute in Pulawy. 2002; 46: 325-329.
- Chen L., Daniel R.M. and Coolbear T. Detection and impact of protease and lipase activities in milk and milk powders. International Dairy Journal, 2003; 13: 255-275.
- Cruickshank R., Dugid J.P., Shwain R.H.A. ‘Medical Microbiology’- A guide to the laboratory diagnosis and control of infection. 1975 11th edn.
- Delbes C.I., Mandjee A. and Montel M.C. Monitoring bacterial communities in raw milk and cheese by culture-dependent and independent 16S rRNA gene-based analyses. Applied and Environmental Microbiology, 2007; 73: 1882-1891.
- Elionora H.Z. and Malka H. Culturable psychrotrophic bacterial communities in raw milk their proteolytic and lipolytic traits. Applied and Environmental Microbiology, 2007; 73: 7162-7168.
- Ghelardi E., Celandroni F., Salvetti S., Barsoti C., Baggiani A., Senesi S. Identification and characterization of toxigenic Bacillus cereus isolates responsible for two food-poisoning outbreaks. FEMS Microbiology Letters, 2002; 208: 129-134.
- Goff H.D. and Griffiths M.W. Major advances in fresh milk and milk products. Journal of Dairy Science, 2006; 89: 1163-1173.
- Guinebretiere M.H., Velge P., Couvert O., Carlin F., Debuyser M.L., Christophe N.T. Ability of Bacillus cereus group to cause food poisoning varies according to phylogenetic affiliation (Groups I to VII) rather than species affiliation. Journal of Clinical Microbiology, 2010; 48(9): 3388-3391.
- Huis Veld J.H.J. Microbial and biochemical spoilage of foods: an overview. International Journal of Food Microbiology, 1996; 33: 1-18.
- Jaeger K.E., Ransac S., Dijkstra B.W., Colson C., Hauvel M.V. and Misset O. Bacterial lipases. FEMS Microbiology Reviews, 1994; 15: 29-63.
- Kumar A. and Seth R. Chemical Quality of Spray Dried Skim Milk Powder. Indian Journal of Dairy Science, 2008; 61: 27-30.
- Law B.A. Enzymes of psychrotrophic bacteria and their effects on milk and milk products. Journal of Dairy Research, 1979; 46: 573-588.
- Margesin R., Neuner G. and Storey KB. Cold loving microbes, plants and animals fundamental and applied aspects. Naturwissenschaften, 2007; 94: 77-99.
- Santana E., Beloti V., Muller E., Ferreira M., Moraes L., Pereira M. and Gusmao V. Milk contamination in different points of the dairy process. Ciencias Agrarias Londrina, 2004; 5: 349-358.
- Xin L., Meng Z., Zhang L., Cui Y., Han X., Yi H. The diversity and proteolytic properties of psychrotrophic bacteria in raw cows’ milk from North China. Int. Dairy J., 2017; 66: 34–4110.1016/j.idairyj.2016.10.014.
- Yuan L., Sadiq F.A., Liu T.J., et al. Psychrotrophic bacterial populations in Chinese raw dairy milk. LWT, 2017; 84: 409-418.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.