ISSN: 0973-7510
E-ISSN: 2581-690X
Circulating immune complexes (CIC’s) are associated with disease progression in Tuberculosis (TB) though their role in pathogenesis is still unclear. Hence the present study was undertaken to identify proteins of diagnostic potential in tuberculosis by proteomic profiling of CIC’s. Serum samples from tuberculosis patients (n=28), latent TB (n=10) and healthy (n=15) individuals were collected and CIC’s levels were estimated by ELISA. CIC’s were isolated by 7% Polyethylene Glycol precipitation and were subjected to proteomic analysis. Bioinformatic analysis and functional annotation of identified proteins was performed using Mascot search engine and PANTHER respectively. Identified protein was validated by ELISA. Statistical analysis was performed with SPSS version 16 and Graph pad prism 5. The mean CIC concentration in TB, latent TB and healthy individuals was found to be 38.23±11.45, 24.43±15.09, and 8.61±2.47µg/ml respectively. A total of 74, 48 and 60 proteins were identified in CIC’s from of TB, latent TB and healthy individuals respectively. Among identified proteins the sensitivity, specificity, positive predictive value, negative predictive value and accuracy of C1qC to distinguishing TB patients from controls (with respect to both latent TB and healthy controls) was found to be 87.18% , 93.33%, 94.44%, 84.85% and 89.86% respectively. Our exploratory analysis suggests that immune-complex based assays might provide better alternate to invasive diagnostic techniques especially in diagnosis of extra pulmonary TB. However, further elaborate studies are required.
Immune complexes, Mass Spectrometry, Proteomics, Tuberculosis
Early diagnosis and appropriate treatment of active pulmonary tuberculosis (PTB) patients is essential for reducing the morbidity, mortality and incidence of tuberculosis (TB). Traditionally for the majority of TB patient’s diagnosis still depends mainly on sputum smear microscopy, chest radiography, tuberculin test and culture. However, these methods have suboptimal sensitivity or specificity and culture requires 6-8 weeks, whereas test based on DNA amplification (PCR, Gene Xpert MTB/RIF) and interferon-g release (Quntiferon TB Gold) are expensive and need particular expertise1,2. The aforementioned limitations of available diagnostic tools highlight an urgent need of competent, affordable and accessible biomarkers for diagnosis and therapeutic monitoring of TB3. Authors in the past have associated circulating immune complexes with disease progression in tuberculosis4,5. A recent study monitoring the transition of latent TB to active TB also reported activation of complement pathway genes and increased levels of antigen/antibody complexes in the early stage of Human Immunodeficiency Virus (HIV) associated TB6. An earlier study also reported the possible role of CIC’s in generating an immune response and regulating variety of immune complexes mediated inflammatory events in tuberculosis7. The clinical role of the CIC present in the serum of tuberculosis patients is still unclear, although several studies have tried to demonstrate the specificity of these immune complexes. Major antigenic components of Mycobacteria are lipoarabinomannan, arabinogalactan, peptidoglycan, heat shock protein and lipoprotein but which of these antigens are responsible for the immune complex formation and plays a role in patho-physiology of tuberculosis is still unknown. A recent dot blot assay based study of dissociated immune complexes demonstrated the presence of Mycobacterium antigens in all TB patients8.
Thus, in the present study, we made an attempt to explore the proteomic profile of CIC’s from tuberculosis patients in comparison with latent TB and healthy controls to identify the novel proteins that can be used as biomarkers for TB diagnosis.
A cross sectional, prospective study was undertaken in the Department of Microbiology at a tertiary care centre in India after obtaining approval from the Institutional Ethics Committee. An informed consent was obtained from all participants before enrolling them in the present study.
Selection of Subjects
Group-1: Pulmonary Tuberculosis Patients
Sputum samples from all patients suspected of pulmonary Tuberculosis attending the OPD of Department of Pulmonary Medicine & Tuberculosis and Chest Diseases were collected in a sterile container and was transported to the Microbiology Department maintaining the cold chain. On receiving the sample, AFB smears were prepared and were stained with 0.3% Auramine phenol O for 10 minutes, then decolorized with 1% acid alcohol solution for 2 minutes and finally counter stained with 0.1% potassium permanganate for 1 minute. Stained smears were examined using LED-FM as per RNTCP guidelines 9. All smear positive patients were enrolled for the present study after obtaining consent.
Group-2: QuantiFERON-TB Gold ELISA Positive (Latent Tuberculosis) and Group 3- QuantiFERON-TB Gold ELISA Negative (Healthy Controls)
To distinguish latent tuberculosis from healthy control household contacts of smear positive pulmonary tuberculosis patients were screened using the QuantiFERON-TB Gold ELISA test. Briefly 1 ml of whole blood was collected in a NIL tube (negative control tube coated with saline), a TB antigen tube (coated with ESAT-6, CFP-10 and TB7.7) and Mitogen tube (positive control coated with Phyto-hemagglutinin, PHA) from each participant using a sterile blood collection needle. The contents in the tubes were mixed thoroughly with the blood by shaking firmly for 10 times and incubated at 37°C in an upright position for 16-24 hours. After incubation harvesting of plasma was performed by centrifuging the tubes for 15 minutes at 3000 g and the plasma was collected in sterile 2 ml tubes and stored at -80°C until use. Human IFN-g ELISA was performed using QuantiFERON-TB Gold ELISA kit following manufacturer’s guidelines and Optical Density (O.D) was measured at 450 nm using a micro plate reader. The raw data were recorded and analyzed using the QFT Analysis Software (Cellestis). The samples showing <0.35 IU/ml of IFN- g or ³0.35 IU/ml of IFN-g and <25% of the negative control value were considered negative, whereas the samples showing ³0.35 IU/ml and ³25% of the negative control value were considered positive. When the mitogen-negative control value was <0.5 IU/ml the test results were considered inconclusive. All patients found positive by QuantiFERON-TB Gold ELISA were enrolled as latent TB, whereas negative were enrolled as healthy controls.
Collection of Blood Samples
Blood samples (2-5 ml) from each subject from aforementioned three groups was collected in vacutainers and were kept at 37°C for 1 hour, followed by 1 hour at 4°C and centrifuged at 1600 g for 5 minutes and the supernatant was collected as serum and stored at -80°C until use.
Detection of Levels of Circulating Immune Complexes in Serum from Different Groups Enrolled for the Study
The level of circulating immune complexes in serum samples was determined by a commercially available kit from DiaMetra (Italy) and optical density (O.D) was read at 450 nm against a reference wavelength at 620 nm. The O.D was tabulated in a Microsoft Excel spreadsheet and concentration in µg/ml of circulating immune complexes in each sample was calculated by generating a standard calibrator curve.
Isolation of Circulating Immune Complexes
Three hundred microliters of serum was centrifuged at 10,000 rpm for 5 minutes and 200µl upper layer sera was mixed with an equal volume of 7% Polyethylene Glycol (PEG) and incubated at 4°C overnight for precipitation of immune complexes. The precipitated immune complexes were centrifuged at 13000 rpm for 10 minutes at 4°C. The resulting pellet was washed twice with 3.5% PEG and the pelleted circulating immune complexes were stored at -80°C until use10.
Proteomic Profiling of Circulating Immune Complexes
For proteomics analysis, the disulfide reduction was carried out by resuspending the pelleted circulating immune complexes in 100µl dithiothreitol (10 mM, Sigma-Aldrich) and incubation at 56°C for 45 minutes. Alkylation of sulfhydryl groups of cysteine residues was performed by addition of 100µl iodoacetamide (55 mM, Sigma-Aldrich) and incubation at room temperature for 30 minutes in the dark. Digestion of proteins was performed by the addition of 0.5 mg/ml trypsin (Sigma-Aldrich) followed by overnight incubation at 37°C. Digestion was stopped by the addition of 5µl of 10% Trifluoroacetic acid and the supernatant was collected after brief centrifugation at 13000 rpm11. The supernatant was lyophilised and mixed with 10µl of solution containing 5% acetonitrile (ACN) and 0.1% formic acid and finally 8µl was used for Liquid Chromatography coupled to mass spectrometry (LCMS) analysis using an Agilent 6520 Accurate mass Q-TOF LC/MS system (Agilent Technologies, Santa Clara, USA). Chromatographic separation was performed using Aligent eclipse plus C18 column (4.6×150 mm, 5µM). The mobile phase contained 0.1% formic acid in water (solvent-A) and 90% ACN and 0.1% formic acid (solvent-B). The gradient profiling was Tmin/A%: B% used was as follows T0 /97%:3%; T40/30%:70, T45 /70%:30%; T50 /97%:3% with a flow rate of 0.4 ml/min. The column temperature and sample temperature were maintained at 37°C and 4°C respectively. Electrospray ionization (ESI) was used in positive mode with spray voltage for positive ion injection at 3.5 kV. Instrument control and data acquisition was performed using Agilent Mass Hunter Workstation Data Acquisition software B.05.01 (Agilent Technologies, Santa Clara, USA). Agilent Mass Hunter Qualitative Analysis software (version B.06.00) was used for processing the LC-MS data.
Bioinformatic Analysis
The LC-MS data in Mascot generic format (MGF) were searched against the Swiss-Prot database (human and Mycobacterium tuberculosis) using the MASCOT search engine (Matrix Science) with the following search parameters, enzyme: trypsin, fixed modifications: carbamidomethylation (C), variable modifications: oxidation (M), mass values: monoisotopic, protein mass: unrestricted, peptide mass tolerance: ± 1.2 Da, fragment mass tolerance: ± 0.5 Da , maximum missed cleavages: 1 ,Instrument type: ESI-QUAD-TOF and significant threshold p<0.05.For optimal identity or extensive homology (p<0.05) only protein with peptide score >25 were considered for analysis12. The Venn diagrams were generated using a web-based tool (http://bioinformatics.psb.ugent.be/webtools/Venn/).
Gene Ontology for Functional Annotation of Identified Proteins
To further understand the functions of identified proteins in each group, we classified proteins by Gene ontology categories which included molecular functions, protein class, biological processes, cellular components and pathway analysis using functional annotation tool available at PANTHER13,14.
Validation of Proteomic Result
Based on biological functions associated with immunity and other infectious diseases Complement C1q subcomponent subunit C (C1QC_HUMAN) was selected and evaluated for distinguishing TB from latent TB and healthy controls using ELISA kit from Biospes Co., Ltd (Chongqing, China) following manufacturers protocol. The O.D was read at 450nm and concentration of C1qC in each sample was calculated by generating a standard calibrator curve.
Statistical Analysis
Statistical analyses were performed using SPSS version 16 for Windows (SPSS, Inc., Chicago, IL, USA) and Graph Pad prism 5 (Graph Pad, San Diego, CA).Continuous variables were presented as the mean ± standard deviation (SD) or median (range); categorical variables are presented as percentages. For all statistical analysis the level of significance was set at p< 0.05.
Demographic of Enrolled Participants
A total of 53 serum samples from participants comprising of 28 smear positive tuberculosis patients, 10 latent TB patients and 15 healthy controls were used for CIC’s estimation and proteomic profiling in the present study. Thirty four (64.15%) participants were male, whereas 19 (35.85%) were female. The mean ± SD age of the participants was found to be 39.22±12.89. A detailed description of the demographic characteristics of the patient enrolled in the present study is shown in Table 1.
Table (1):
Demographic Characteristic of the Participants
Characteristic |
Tuberculosis |
Latent TB |
Healthy |
|---|---|---|---|
Sample Size, n (%) |
28 (52.83%) |
10 (18.87%) |
15 (28.30%) |
Age
Mean± SD (in years) Range ( in years) |
39.142 ± 14.8
17- 60 |
35.9 ±13.35
13 – 56
|
41.6 ± 8.11 32- 60 |
Gender, Male
Female |
22 (41.50%)
6 (11.32%) |
7 (13.20%)
3 (5.66%) |
5 (9.43%)
10 (18.86%) |
BCG vaccination status |
28/28 |
10/10 |
15/15 |
Levels of Circulating Immune Complexes
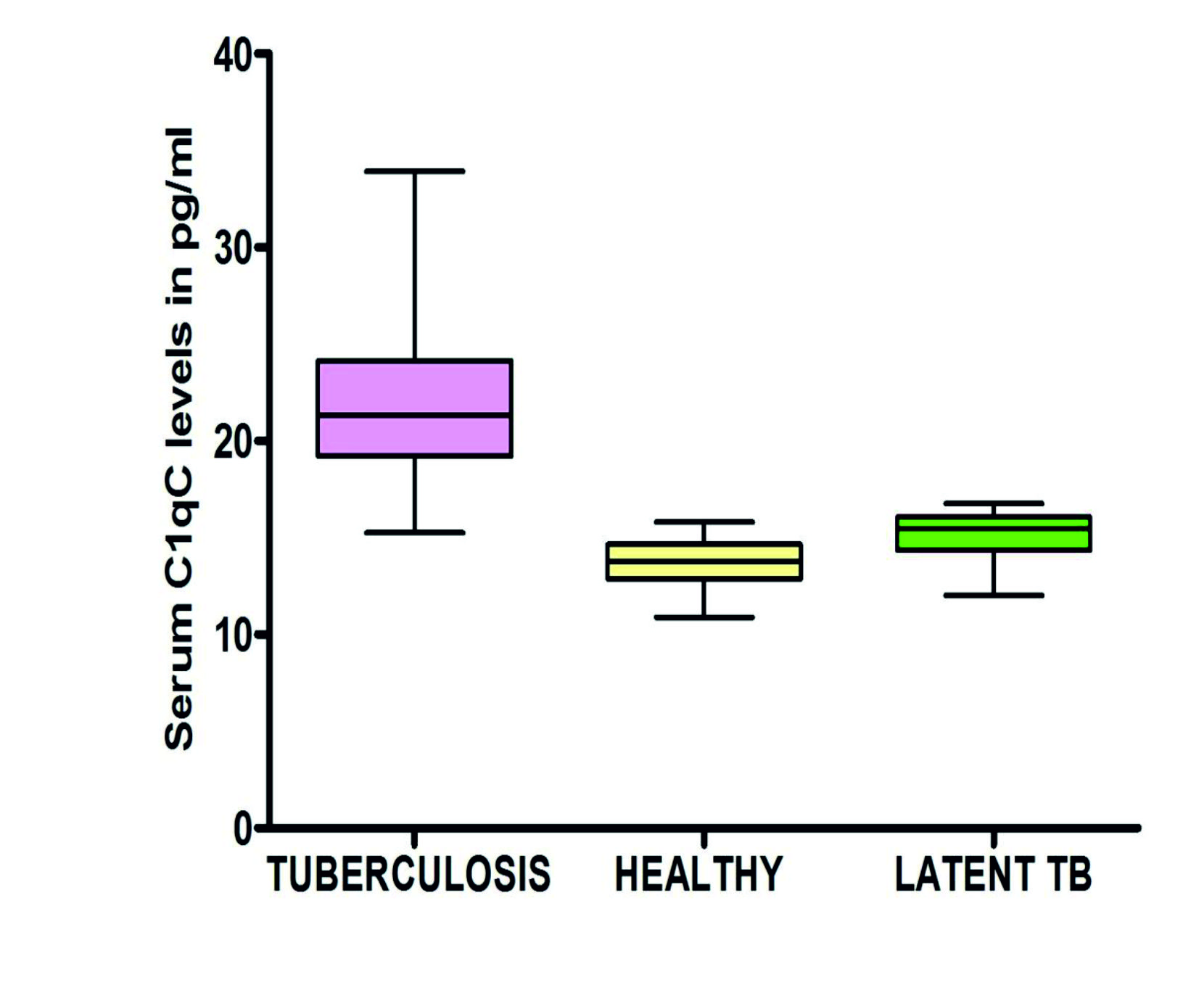
The CIC concentration in TB patients was found significantly higher (p<0.0001) when compared to both latent TB and healthy individuals (Fig 1). The mean CIC concentration in TB, latent TB and healthy individuals was found to be 38.23 (SD, ± 11.45), 24.43 (SD, ±15.09), and 8.61(SD, ±2.47) µg/ml respectively.
 Fig. 1. Scatter dot plot showing concentration of circulating immune complexes in Healthy (n= 15), Latent TB (n= 10) and Tuberculosis patients (n = 28). The mean and standard error of mean in each group is shown by the horizontal bars.
Fig. 1. Scatter dot plot showing concentration of circulating immune complexes in Healthy (n= 15), Latent TB (n= 10) and Tuberculosis patients (n = 28). The mean and standard error of mean in each group is shown by the horizontal bars. Proteomic Profiling of Circulating Immune Complexes
On analyzing the LCMS data on the MASCOT search engine a total of 74, 48 and 60 proteins were identified in circulating immune complexes of TB, latent TB and healthy individuals respectively. Venn diagram analysis (Fig. 2) showed that 26 proteins were overlapped in all three groups. Further, there were 37, 10 and 19 proteins which were specifically seen in TB, latent TB and healthy individuals respectively. Among observed proteins in TB, latent TB and healthy individuals, 89.20% (66/74), 95.83% (46/48) and 90% (54/60) proteins were human proteins respectively. Further 8 (10.81%), 2(4.16%) and 6 (10%) mycobacterial proteins in TB, latent TB and healthy individuals were also observed though the majority of these identified proteins were from M. bovis. A detailed description of proteins identified in different study groups is shown in Table: 2. Further, Kruskal-Wallis test of the identified proteins showed a statistically significant difference between the three groups with p-value 0.00831 (H= 9.5803).
Table (2):
Detailed Descriptions of Identified Proteins in Different Study Groups.
Study Groups
(n=total proteins) |
Identified Proteins ( Symbol) |
|---|---|
Latent TB, Healthy
& Tuberculosis (n=26) |
Apolipoprotein B-100 (APOB_HUMAN), C4b-binding protein alpha chain (C4BPA_HUMAN), Complement C1q subcomponent subunit C (C1QC_HUMAN), Ig gamma-3 chain C region (IGHG3_HUMAN), C4b-binding protein beta chain (C4BPB_HUMAN), Suppressor of cytokine signaling 6 (SOCS6_HUMAN), Ig mu chain disease protein (MUCB_HUMAN), Compliment C3 (CO3_HUMAN), Immunoglobulin J chain (IGJ_HUMAN), Nidogen-1 (NID1_HUMAN), Titin (TITIN_HUMAN), Ig gamma-1 chain C region (IGHG1_HUMAN),Ig alpha-1 chain C region (IGHA1_HUMAN), Apolipoprotein(a) (APOA_HUMAN), Compliment C4-A (CO4A_HUMAN),Ig mu chain C region (IGHM_HUMAN), Fibronectin (FINC_HUMAN) Protein CBFA2T3 (MTG16_HUMAN), Apolipoprotein C-III (APOC3_HUMAN), Iglamda chain C region (LAC_HUMAN), Serum albumin (ALBU_HUMAN), Complement C1s subcomponent (C1S_HUMAN),Ig kappa chain C region (IGKC_HUMAN), Complement factor H (CFAH_HUMAN), Apolipoprotein A-I (APOA1_HUMAN),Ig gamma-2 chain C region (IGHG2_HUMAN) |
Latent TB & Tuberculosis
(n=04) |
Ig lambda chain V-IV region HiL (LV403_HUMAN), Putative membrane protein mmpS 5 (MMPS5_MYCBO), Fibrocystin-L (PKHL1_HUMAN), Complement C1q subcomponent subunit B (C1QB_HUMAN) |
Healthy & Tuberculosis
(n=07) |
Neurabin-2 (NEB2_HUMAN), Apolipoprotein E (APOE_HUMAN), Exodeoxyribonuclease V gamma chain (EX5C_MYCTU), 26S proteasome non-ATPase regulatory subunit 11 (PSD11_HUMAN), DNA polymerase IV 2 (DPO42_MYCBO), Apolipoprotein C-II (APOC2_HUMAN), Protein SON ( SON_HUMAN) |
Healthy & Latent TB
(n=08) |
DENN domain containing protein 2C (DEN2C_HUMAN), LisH domain and heat repeat containing protein KIAA1468 (K1468_HUMAN, Endoplasmic reticulum metallopeptidase 1 (ERMP1_HUMAN), Fibrinogen gamma chain (FIBG_HUMAN), Fibrinogen beta chain (FIBB_HUMAN), Hemoglobin subunit beta (HBB_HUMAN), Apolipoprotein A-II (APOA2_HUMAN), Fibrinogen alpha chain (FIBA_HUMAN) |
Tuberculosis
(n=37) |
Translational activator GCN1 (GCN1L_HUMAN), Exopolysaccharide phosphotranferase cpsY (CPSY_MYCBO), C-reactive protein (CRP_HUMAN), Vasohibin-2(VASH2_HUMAN), Bromodomain-containing protein 1 (BRD1_HUMAN), Complement factor H related protein1 (FHR1_HUMAN), Serum amyloid A-1 protein (SAA_HUMAN), Putative helicase MOV-10 (MOV10_HUMAN), Haptoglobin (HPT_HUMAN),Ig gamma-4 chain C region (IGHG4_HUMAN),Ig kappa chain V-III region SIE (KV302_HUMAN), Haptoglobin related protein (HPTR_HUMAN), Laminin subunit alpha-5 (LAMA5_HUMAN), Apoptotic chromatin condensation inducer in the nucleus (ACINU_HUMAN), Ig kappa chain V-III region B6 (KV301_HUMAN),Tubulin polyglutamylase TTLL11 (TTL11_HUMAN), Complement C5 (CO5_HUMAN), 30S ribosomal protein S2 (RS2_MYCBO),Ras-related protein Rab-21(RAB21_HUMAN), Polypeptide N-acetylgalactosaminaly transferase 4 (GALT4_HUMAN), Serine protease inhibitor Kazal type-5 (ISK5_HUMAN), Vitamin K-dependent protein S (PROS_HUMAN), Dynein heavy chain 10 (DYH10_HUMAN), Tyrosine-protein kinase Yes (YES_HUMAN),Ig lambda chain V-III region LOI (LV302_HUMAN), Sin3 histone deacetylase corepressor complex component SDS3 (SDS3_HUMAN), Uncharacterized HTH-type transcriptional regulator Rv0474 (Y474_MYCTU), Complement factor I (CFAI_HUMAN), Putative maltooligosyl trehalose synthase (TREY_MYCTU), S phase cyclin A-associated protein in the endoplasmic reticulum (SCAPE_HUMAN), Prothrombin (THRB_HUMAN), Complement C4-B (COB4_HUMAN), Voltage dependent T-type calcium channel subunit alpha-1I (CAC1I_HUMAN), Glycine–tRNA ligase (SYG_MYCBO), Alpha-2-HS-glycoprotein (FETUA_HUMAN), Complement C1q subcomponent subunit A (C1QA_HUMAN), Sorbin and SH3 domain-containing protein 2 (SRBS2_HUMAN) |
Latent TB
(n=10) |
Tau-tubulin kinase 2 (TTBK2_HUMAN), pH-sensitive adenylate cyclase Rv1264 (Y1264_MYCTU), Probable ATP dependent RNA helicase DDX28 (DDX28_HUMAN), UPF0687 protein C20orf27 (CT027_HUMAN), La-related protein 7 (LARP7_HUMAN), Polycomb protein SCMH1 (SCMH1_HUMAN), Vascular endothelial growth factor receptor 3 (VGFR3_HUMAN), Neuralized-like protein 4 (NEVL4_HUMAN), Uncharacterized protein C3orf43 (CC043_HUMAN), Hemoglobin subunit alpha (HBA_HUMAN) |
Healthy
(n=19) |
Dystrophin (DMD_HUMAN), Major centromere autoantigen B (CENPB_HUMAN),Glycosyltransferase 6 domain-containing protein 1 (GL6D1_HUMAN), Regulating synaptic membrane exocytosis protein 2 (RIMS2_HUMAN) , Cell division protein FtsZ (FTSZ_MYCBO), Steroid C26-monooxygenase (CP125_MYCBO), Annexin A7 (ANXA7_HUMAN) , PDZ domain-containing protein 2 (PDZD2_HUMAN), Putative 2-dehydropantoate 2-reductase (PANE_MYCTU), Reticulon-4 (RTN4_HUMAN), Probable ATP-dependent Clp protease ATP-binding subunit (CLPC_MYCBO), Formin-binding protein 1-like (FBP1L_HUMAN), Ligand-dependent corepressor (CJO12_HUMAN), Nuclear pore complex protein Nup205 (NU205_HUMAN), Estrogen sulfotransferase (ST1E1_HUMAN), RING finger protein 32 (RNF32_HUMAN), Immunoglobulin heavy variable 3-48 (HV302_HUMAN), AT-rich interactive domain-containing protein 2 (ARID2_HUMAN), Methylmalonate-semialdehyde dehydrogenase[acylating], mitochondrial (MMSA_HUMAN) |
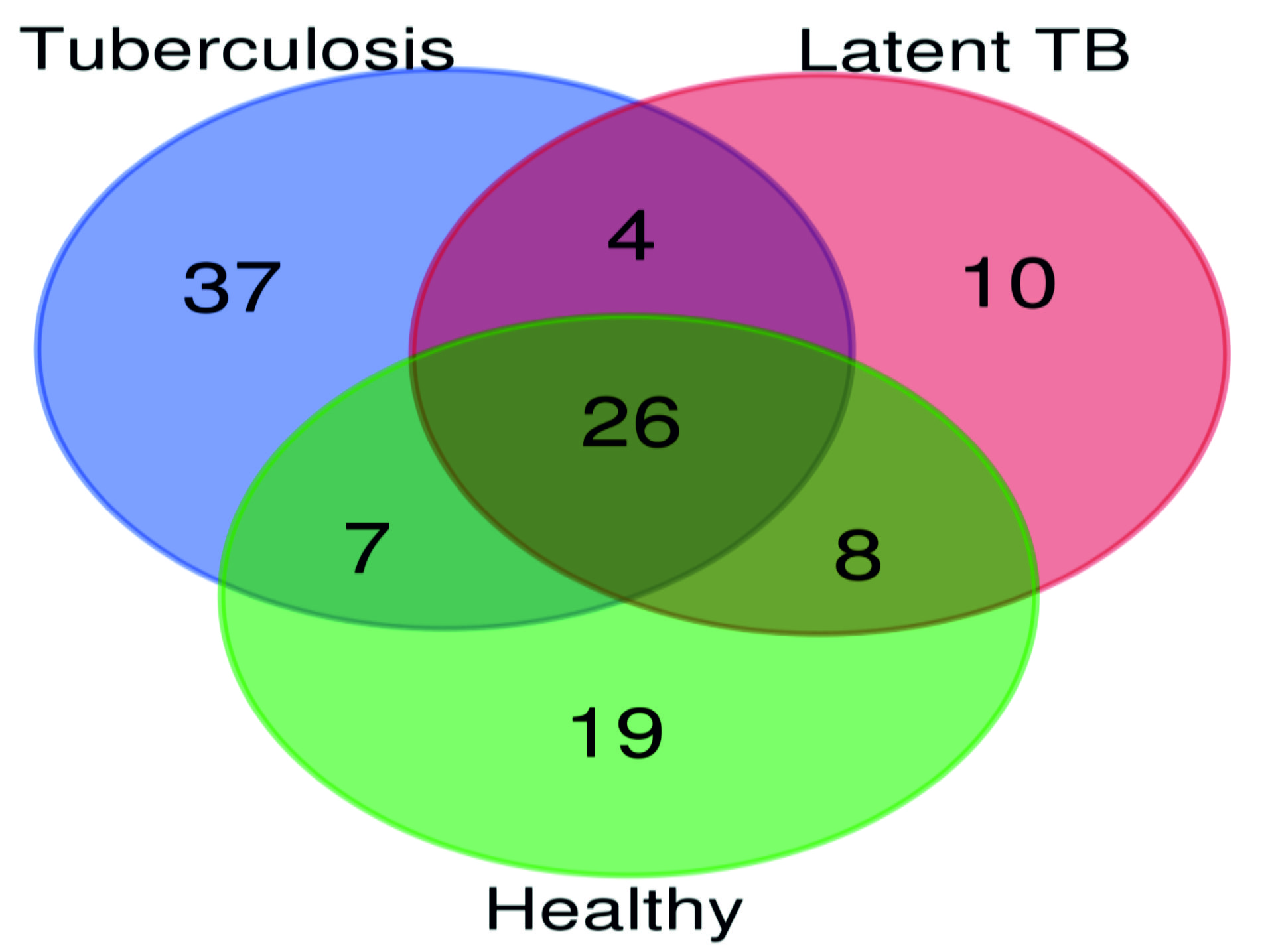 Fig. 2. A venn diagram showing total proteins identified in at least one individual from each group of Tuberculosis, Latent TB and Healthy. Numbers of identified proteins are shown in different overlaps between groups.
Fig. 2. A venn diagram showing total proteins identified in at least one individual from each group of Tuberculosis, Latent TB and Healthy. Numbers of identified proteins are shown in different overlaps between groups.Gene Ontology for Functional Annotation of Identified Proteins
Upon analyzing molecular functions of identified proteins, we observed that proteins associated with binding and catalytic activity were dominant in all study groups. On analyzing protein classes’ protein associated with defence/ immunity and enzyme modulation were more dominant in TB patients as compared to latent TB and healthy individuals. Among cellular components proteins associated with extracellular region and macromolecular complex were dominant in all three study groups. A detail description of classification of proteins according to molecular function, protein class, biological processes, cellular component and pathway analysis is shown in Figure: 3.
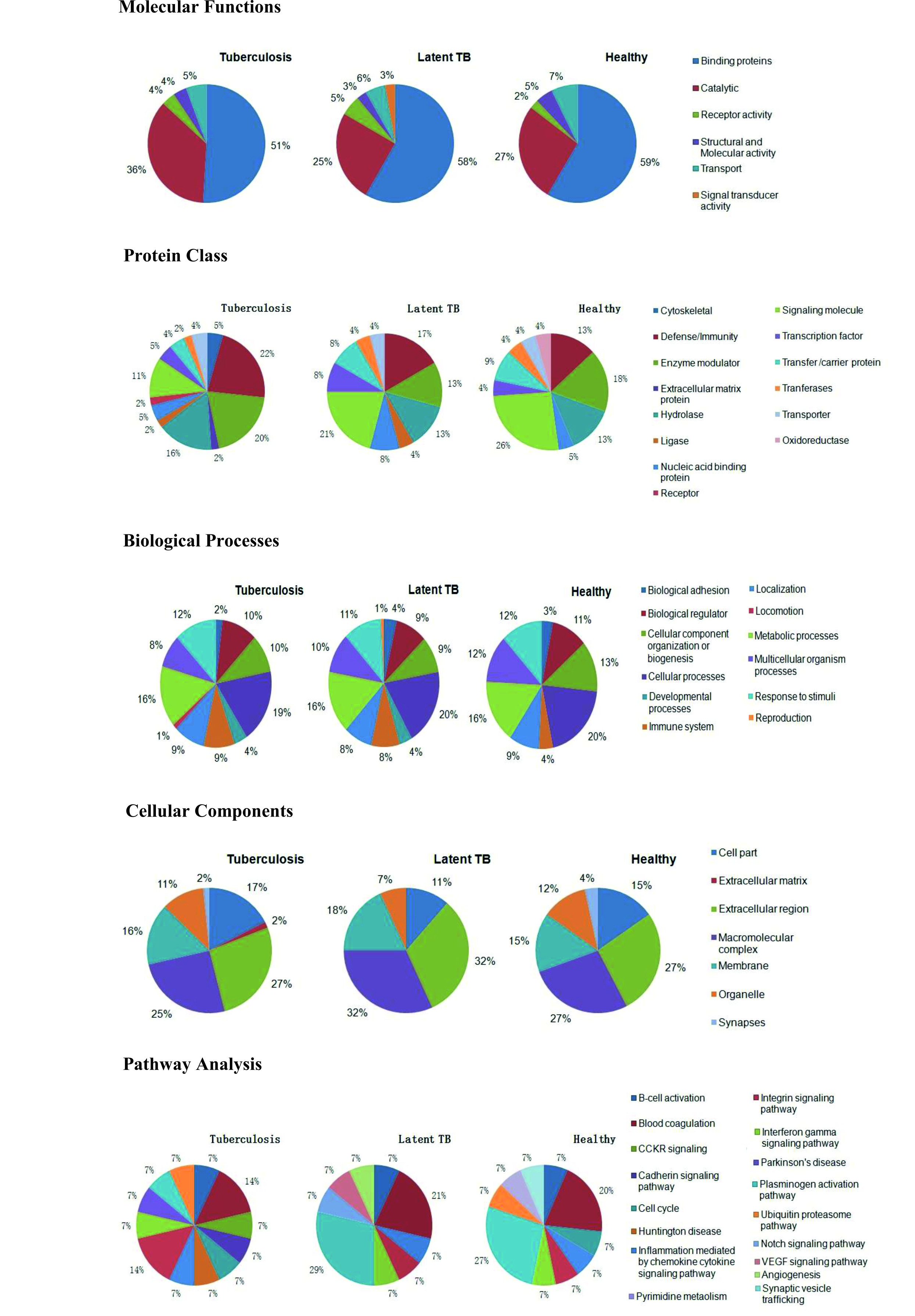 Fig. 3. Classification of proteins according to molecular function, protein class, biological processes, cellular component and pathway analysis.
Fig. 3. Classification of proteins according to molecular function, protein class, biological processes, cellular component and pathway analysis.Validation of Complement C1q Subcomponent Subunit C (C1QC_HUMAN)
Mean (± SD) of C1qC levels in Tuberculosis, latent TB and healthy controls was found to be 21.8±4.39 pg/ml, 15.18±1.3 pg/ml and 13.64±1.32 pg/ml respectively (Fig 4). On the receiver operator characteristic curve (ROC) analysis the area under curve was found to be 0.903 (95%CI, 0.822-0.983) with a standard error 0.041. The sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV) and accuracy of C1qC for distinguishing tuberculosis patients from controls (latent TB and healthy controls) was found to be 87.18% (95%CI, 72.57%-95.70%), 93.33%(95%CI,77.93%-99.18%), 94.44%(95%CI, 81.59%-98.49%), 84.85% (95%CI, 71.07%-92.74%) and 89.86%(95%CI, 80.21%-95.82%) respectively.
Immune complexes are formed when a specific antibody combines with an antigen in different proportions. An equal proportion of antigen and antibody results in precipitation of immune complexes whereas when there is an excess of antigen these immune complexes may remain soluble and circulate in the blood. Comprehensive analysis of these circulating immune complexes might be useful in identifying antigens incorporated in CIC’s that may be useful in developing diagnostic and treatment strategies for various circulating immune complex associated diseases. Presence of IC’s is established in both circulation and in tissues in TB7. Thus, in the present study, we made an attempt to explore the proteomic profile of circulating immune complexes from tuberculosis patients in comparison with latent TB and healthy controls. Earlier studies have reported high levels of CIC’s in active TB patients and its correlation with the severity of the disease which was similar to our findings in the present study and also the CIC’s concentration in TB patients was observed significantly higher (p<0.05) as compared to both latent TB and healthy controls8. The majority of studies have reported Immunoglobulin G in all of the CICs from the patients with tuberculosis15-22, whereas Bhattacharya et al.16 and Samuel et al.22 reported the presence of IgM and IgA. May et al.19 confirmed mycobacterial antigen specific antibodies (IgG and IgM), whereas isotypes of IgG and IgM were reported by Raja et al.20 and Jhonson et al.17. In the present proteomic study, we also observed Ig alpha-1 chain C region (IGHA1_HUMAN), Ig gamma-1 chain C region (IGHG1_HUMAN),Ig gamma-2 chain C region (IGHG2_HUMAN), Ig gamma-3 chain C region (IGHG3_HUMAN), Ig gamma-4 chain C region (IGHG4_HUMAN),Ig lambda chain C region (LAC_HUMAN) and Ig mu chain C region (IGHM_HUMAN) which showed the presence of IgA, IgG, and IgM antibodies in CIC’s which is in concordance with the findings of Bhattacharya et al.16 and Samuel et al 22. May et al.19 reported the presence of C1q, C3 and C4 components in CIC’s while Carr et al.15 and Jhonson et al.17 reported only C1q, where as Bhattacharya et al.16 reported the presence of C3 and C4 in CIC isolated from the serum of TB patients. Though our study confirmed the presence compliment components C1q, C1s, C1s, C1qC, C3, C4 we also observed complement factor H (CFAH) in 9 and C5 in 2 TB patients. In a recent study, C1qC was found to be the most differentially expressed protein among the components of C1q component and reported a sensitivity of 83% and specificity of 89% to distinguish active TB from controls in the present study also the overall sensitivity and specificity of C1qC to distinguish active TB from controls was found to be 87.18% and 93.33% respectively which is in concordance with earlier findings23.Studies exploring novel serum biomarkers for TB diagnosis showed C4BPA, SAA24, APOA1, APOB l 25 to be promising biomarkers for TB, the presence of all these proteins in immune-complexes in all three groups in present study warrants further exploration of these proteins for establishing the diagnosis of tuberculosis. A study reported significantly high CRP level and low levels of Fibrinogen in active TB patients in comparison with healthy controls and on similar lines, we also observed CRP in 8 TB patients and interestingly, fibrinogen was found (Fibrinogen alpha chain, Fibrinogen beta chain and Fibrinogen gamma chain) in latent TB and healthy controls only26.
A label free quantitative proteomics study27 of plasma from TB, latent TB and healthy samples have reported the presence of MTB secretory proteins prpC, pbp, rpsN, pyrR, mntH, and Rv0102 among active TB samples, relA, vapB18, clpP2, kasA, Rv0064, and Rv2633c among LTB samples, aceAa and dxr among healthy samples. The spectral data search for MTB proteins in the present study also identified 8 (CPSY_MYCBO, MMPS5_MYCBO, RS2_MYCBO, SYG_MYCBO, TREY_MYCTU, Y474_MYCTU, DPO42_MYCBO, EX5C_MYCTU), 2 (MMPS5_MYCBO, Y1264_MYCTU) and 6 (CLPC_MYCBO, PANE_MYCTU, CP125_MYCBO, FTSZ_MYCBO, DPO42_MYCBO, EX5C_MYCTU) mycobacterial proteins in TB, latent TB and healthy respectively. Since all these proteins were detected in a limited number of samples further targeted proteomics approaches are required to fully explore the role of these proteins in pathogenesis of TB. Protein class analysis showed a majority of proteins in TB belonged to defence/immunity, enzyme modulator and hydrolyses which suggests infection status. Further, majority of identified proteins were associated with protein binding and catalytic activities. Protein associated with cadherin signaling pathway and CCKR signaling pathway were also observed in CIC’s from Tuberculosis patients. Though present study was the first comprehensive study on proteomic profiling of circulating immune complexes in tuberculosis in comparison with latent TB and healthy individuals, one of the major limitations of this study was its limited sample size and validating these findings in larger cohorts might provide better insights of these proteins for their diagnostic potential.
Present study provides the first insight into the composition of circulating immune complexes in TB, latent TB and healthy controls and. Identification of TB specific biomarkers in Immune-complexes provides better alternate to invasive diagnostic techniques, especially in diagnosis of extra pulmonary TB and paediatric TB, thus further elaborate study with larger number of subjects is desirable.
Acknowledgments
We are grateful to all the participants enrolled in the present study for their consent and cooperation during the study. We also thank the physicians, nursing staff and technical staff for their help during the course of study.
Conflict Of Interest
The authors declare that there is no conflict of interest.
Authors’ Contribution
All authors have made substantial, direct and intellectual contribution to the work and approved it for publication.
Funding
None.
Data Availability
All datasets generated or analyzed during this study are included in the manuscript.
Ethics Statement
This study was undertaken after obtaining approval from the Institutional Ethics Committee. An informed consent was obtained from all participants before enrolling them in the present study.
- Drobniewski F.A., Caws M., Gibson A. & Young D. Modern laboratory diagnosis of tuberculosis. Lancet Infec Dis., 2003; 3: 141.
- Dorman S.E. New diagnostic tests for tuberculosis: bench, bedside, and beyond. Clin Infect Dis., 2010; 50 (Supplement_3): S173-177.
- Yerlikaya S, Broger T, MacLean E, Pai M, Denkinger CM. A tuberculosis biomarker database: the key to novel TB diagnostics. Int J Infect Dis, 2017; 56:253-257.
- Simonney N., Bourrillon A., Lagrange P.H. Analysis of circulating immune complexes (CICs) in childhood tuberculosis: levels of specific antibodies to glycolipid antigens and relationship with serum antibodies. Int. J. Tuberc. Lung Dis., 2000; 4(2):152-160.
- Gupta I., Jain A., Singh N.B., Chaturvedi V., Agarwal S.K. Role of antigen specific circulating immune complexes in diagnosis of tuberculosis. Int. J. Tuberc. Lung Dis., 1998; 2(6):456-461.
- Esmail H., Lai R.P., Lesosky M., Wilkinson K.A., Graham C.M., Horswell S., Coussens A.K., Barry C.E., O’Garra A., Wilkinson R.J. Complement pathway gene activation and rising circulating immune complexes characterize early disease in HIV-associated tuberculosis. Proc. Natl. Acad. Sci., 2018; 115(5): E964-73.
- Senbagavalli P., Hilda J.N., Ramanathan V.D., Kumaraswami V., Nutman T.B., Babu S. Immune complexes isolated from patients with pulmonary tuberculosis modulate the activation and function of normal granulocytes. Clin. Vaccine Immunol., 2012; 19(12): 1965-71..
- Przybylski G., Golda R. Research on the occurrence of Mycobacterium tuberculosis antigens in the circulating immune complexes, isolated from serum of patients with tuberculosis. Med. Sci. Monit., 2014; 20: 6.
- Reza L.W., Satyanarayna S., Enarson D.A., Kumar A.M., Sagili K., Kumar S., Prabhakar L.A., Devendrappa N.M., Pandey A., Wilson N., Chadha S. LED-fluorescence microscopy for diagnosis of pulmonary tuberculosis under programmatic conditions in India. PLoS ONE, 2013; 8(10): e75566.
- Ranganathan U.D., Bethunaickan R., Raja A. Isolation of Circulating Immune Complexes from TB Patient Serum for Serodiagnosis. Bio-protocol, 2012; 2(11): e188.
- Ohyama K., Huy N.T., Yoshimi H., Kishikawa N., Nishizawa J.E., Roca Y., Revollo Guzman R.J., Velarde F.U., Kuroda N., Hirayama K. Proteomic profile of circulating immune complexes in chronic Chagas disease. Parasite Immunol., 2016; 38(10):609-17.
- Huy N.T., Trieu H.T., Okamoto K., Ninh T.T., Ha T.T., Morita K., Huong V.T., Lan N.T., Thuy T.T., Nga C.T., Kikuchi M. Proteomic profile of circulating immune complexes in dengue infected patients. J. Trop. Dis., 2013; 1: 109.
- Mi H., Thomas P. Panther pathway: an ontology-based pathway database coupled with data analysis tools. Protein Networks and Pathway Analysis, Humana Press; 2009; 123-140.
- Thomas P.D., Kejariwal A., Guo N., Mi H., Campbell M.J., Muruganujan A., Lazareva-Ulitsky B. Applications for protein sequence–function evolution data: mRNA/protein expression analysis and coding SNP scoring tools. Nucleic Acids Res., 2006; 34(suppl_2): W645-50.
- Carr R.I., Charkraborty A.K., Brunda M.J., et al. Immune complexes and antibodies to BCG in sera from patients with mycobacterial infections. Clin. Exp. Immunol., 1980: 39: 562-569
- Bhattacharaya A., Ranadive S.N., Kale M., Bhattacharaya S.: Antibody based Enzyme Linked Immunoabsorbent Assay for determination of immune complexes in clinical tuberculosis. Am. Rev. Respir. Dis., 1986; 134: 205-209
- Johnson NmcI, McNicol M.W., Burton-Kee E.J., Mowbray J.F.: Circulating immune complexes in tuberculosis. Thorax., 1981; 36: 610-617
- Gatner E.M.S., Anderson R.: An in vitro assessment of cellular and humoral immune function in pulmonary tuberculosis; correction of defective neutrophli motility by ascorbate, levamisole, metoprolol, pronalonol. Clin. Exp. Immunol., 1980; 40: 327-336
- May J.J., Katilus J., henson P.M., Dreisin R.B.: The purification and identification of circulating immune complexes in tuberculosis. Am. Rev. Respir. Dis., 1983; 128: 920-925
- Raja A., Narayanan P.R., Mathew R., Prahakar R.: Characterization of mycobacterial antigens and antibodies in circulating immune complexes from pulmonary tuberculosis. J. Lab. Clin. Med., 1995; 125: 581-587
- Radhakrishnan V.V., Mathai A., Sundaram P.: Diagnostic significance of circulating immune complexes in patients with pulmonary tuberculosis. J. Med. Microbiol., 1992; 36: 128-131
- Samuel A.M., Ashtekar M.D., Ganatra R.D.: Significance of circulating immune complexes in pulmonary tuberculosis. Clin. Exp. Immunol., 1984; 58: 317-324
- Cai Y., Yang Q., Tang Y., Zhang M., Liu H., Zhang G., Deng Q., Huang J., Gao Z., Zhou B., Feng C.G. Increased complement C1q level marks active disease in human tuberculosis. PloS one, 2014; 9(3): e92340.
- Jiang T.T., Shi L.Y., Wei L.L., Li X., Yang S., Wang C., Liu C.M., Chen Z.L., Tu H.H., Li Z.J., Li J.C. Serum amyloid A, protein Z, and C4b-binding protein chain as new potential biomarkers for pulmonary tuberculosis. PloS one, 2017; 12(3): e01734.
- Wang C., Wei L.L., Shi L.Y., Pan Z.F., Yu X.M., Li T.Y., Liu C.M., Ping Z.P., Jiang T.T., Chen Z.L., Mao L.G. Screening and identification of five serum proteins as novel potential biomarkers for cured pulmonary tuberculosis. Sci. Rep., 2015; 5: 15615.
- Adigun K., Edem V.F., Ige O.M., Arinola O.G. Plasma levels of C – reactive protein and Fibrinogen in Pulmonary Tuberculosis Patients in Ibadan, Southwest, Nigeria. Afr. J. Biomed. Res., 2017; 20(2): 127-30.
- Sun H., Pan L., Jia H., Zhang Z., Gao M., Wang J., Sun Q., Wei R., Du B., Xing A., Zhang Z. Label-Free Quantitative Proteomics Identifies Novel Plasma Biomarkers for Distinguishing Pulmonary Tuberculosis and Latent Infection. Front Microbiol., 2018; 9: 1267.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.