‘COVID-19’ the recent virulent viral infection had influenced the lives of millions globally leading to both loss of life, economic and financial crisis. Coronavirus belongs to family coronaviridae with four genus viz. a/b and g-coronavirus, infecting both aves and mammals. The SARS-Cov-2 emerged in Wuhan, China in Dec, 2019 and since then had spread to 213 countries. Its origin is debatable with both natural origin and conspiracy theory providing no conclusive evidences. Coronavirus have ‘+’ive RNA and encodes for 29 proteins, which carries out its life cycle including infection and disease progression. The study of its proteome organization could illustrate the proteins which act as the key molecular players in the infection cycle of the virus. These proteins can also act as important drug targets in combating COVID-19 infection. Majority of the drugs have been formulated in order to act as agonist to spike proteins inhibiting infection by binding to ACE2 receptors. Proteome analysis has also revealed the critical mutated proteins that are responsible for COVID-19 pathogenesis and virulence. mRNA based vaccines (mRNA-1273, BNT162) also targets these spike proteins. Although DNA vaccine has also been attempted using RDT, but the high rate of mutation associated with COVID-19 have made such vaccines ineffective even before use. Thus evolutionarily conserved proteins have been the best candidature for vaccine development. Similarly phylogenetic analysis of its proteins could help us to understand the evolutionary pattern of COVID-19. It could be used to develop a predictable model for such pathogenic infections, preparing ourselves to take preventive action against its reoccurrence.
COVID-19, Coronavirus, phylogenetic analysis, mRNA-1273, BNT162, ACE2 receptors, CHAdOx1 vaccine
This novel coronavirus (covid-19) has been classified in the order Nidovirales under family Coronaviridae1-2. Family Coronaviridae can be classified into four genus viz. Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus. Out of these, mammals are infected by α-coronavirus and β-coronavirus, avian species were infected by γ-coronavirus while both mammalian and avian species by δ-coronavirus3. a-coronaviruses consist of human coronavirus NL63 (HCoV-NL63), porcine respiratory coronavirus (PRCV) and porcine transmissible gastroenteritis coronavirus (TGEV), PEDV. b-coronavirus consist of SARS-CoV, MERS-CoV, bat coronavirus HKU4, mouse hepatitis coronavirus (MHV), bovine coronavirus (BCoV), and human coronavirus OC43, while the γ & δ-coronaviruses consist of avian infectious bronchitis coronavirus (IBV) and porcine δ coronavirus (PdCV), respectively, thus corona virus has the ability to cause infections in both animas as well as humans. Moreover data suggest that 80% or more infections are mild or asymptomatic, thus these infected people serves as a breeding ground for corona virus4. Although the mortaility rate is small (3.4% latest WHO estimate)5 yet immune compromised individual’s especially senior citizens (above 60 years) and young ones (below 5) are much more susceptible for the disease. Globally till date (22nd May, 2020) more than 50 million cases of COVID-19 has been reported and 328000 deaths have been reported, and infection has spread to more than 213 countries6.
In the year 2003-2004, about 8000 people infected with severe acute syndrome coronavirus (SARS-CoV) having mortality rate of about 10%7-10. Then in 2012, more than 1700 people become infected with Middle East respiratory syndrome coronavirus (MERS-CoV) along with the mortality rate of about 36%11-12. In 2013, United States was chronically suffered by porcine epidemic diarrhea coronavirus (PEDV) in piglets that cause 100% mortality; as a result their population has decreased by more than 10% within a year13-15. From all these findings, it has now become evident that coronavirus pose serious threat to human health by causing infections related to respiratory, central nervous system and gastrointestinal system along with economic losses1-2. It has also been reported through modern research that coronavirus has the potential to undergo mutations and recombination processes, so become capable of surviving in diverse environmental conditions along with wide host range and tissue tropism efficiency16–18. Due to these unique characteristics, coronavirus pose very stable and long term infections in humans and animals globally disabling the global healthcare system and economy.
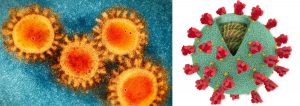
Fig. 1. Corona Virus (Covid-19) structural analysis. (a) SEM image of the COVID-19 coronavirus. (b) Structure image of corona virus. *(Photo source: Scripps Research)
Structural Organization
Coronaviruses have positive-stranded RNA as the genetic material along with an envelope (Fig. 1). It has been found that coronavirus posses the largest genome amongst all the RNA viruses (ranges from 27 to 32 kb). The genome of coronavirus is surrounded by an outermost envelope, inside which helical capsid is present that is made up of nucleocapsid protein (N). Three structural proteins are associated with the viral envelope viz. membrane protein (M), envelope protein (E) and spike protein (S)19. Among these, envelope protein (E) and membrane protein (M) are responsible for assembly of the virus, while the spike protein (S) helps in the virus entry into host cell. In addition to these three structural proteins, some coronavirus have additional envelope associated hemagglutinin-esterase protein (HE)20. Large protrusions emerged out from the spike of viral surface, which forms a crown like appearance, that’s why given the name ‘coronavirus’ (Latin word means crown) (Fig. 1 (a) & (b)). The spike of the coronavirus is not only associated with entry of virus inside the host cell but also act as a vital determinant for tissue tropism, viral host range and a key stimulator of host immune responses.
The spike of coronavirus posses three segments viz. big ectodomain, a single-pass transmembrane anchor, and a small intracellular tail (Fig. 1b). The ectodomain segment contains a S1 subunit that binds to the receptors and a S2 subunit that helps in the membrane-fusion. It has become evident from the electron microscopy that spike is in the form of clove-shaped trimer consisting of three S1 subunits as heads and a trimetric S2 subunit as stalk21-24 (Fig. 1 (a) & (b)). During the entry of virus inside the host cell, S1 subunit binds to the receptors present on the surface of the host cell and S2 subunit merged with the membranes of host cell and viral cell, allowing the entry of viral genomes inside the host cell. The important steps in the infection cycle of coronavirus are the receptor mediated binding and membrane fusion that act as a key targets for drug development. In this review article, we have focused on the structural, functional aspect of coronavirus proteome along with the identification of key targets for vaccine development.
Prevalence and COVID-19
Molecular phylogenetic analysis across different geographical isolates has been carried out to identify viral proteins those are under positive selection pressures, i.e. accumulating mutations that enhance pathoadaptivity of virus over time. These proteins have potential to act as vaccine candidates as these also act as B cell and T cell epitopes against which host mounts an immune response.
It further helps to track the global outbreak of coronavirus pandemic. The first case of COVID-19 was observed in the Wuhan city in China, on December 2019 and has since then it had acquired the form of pandemic, disseminating in 213 countries and 26 cruise ships across the globe10.
The origin and evolution of COVID-19 has been speculated through two different hypotheses including both natural origin of COVID-19 and conspiracy hypothesis. The hypothesis for the natural origin of COVID-19 assumes that it is the result of natural evolution, according to conclusion published by scientists from Australia, United Kingdom and United States in Nature Medicine25. The investigation of data in public genome sequence from SARS-CoV-2 and MRSA viruses have not initiate any substantial evidence that the virus was constructed in the laboratory or engineered otherwise. Although the origin of the virus is still debatable but the recent mutational studies about its evolution had suggested it to be of natural origin rather then been produced through genetic engineering. Scientists in China have successfully sequenced the genome of SARS-CoV-2 and made the data has been submitted to NCBI, USA26.
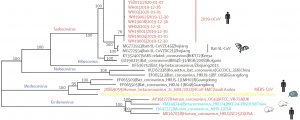
Fig. 2. Phylogentic analysis of full length genome of 2019-n CoV and its interconnection with animal viruses. MERS-Co V is Middle East respiratory syndrome coronavirus and SARS-CoV is severe acute respiratory syndrome27.
Several independent sequencing studies on SARS-Co V-2, through phylogenetic sequence similarity plot clearly revealed that major similarity in nucleotide regions with members of subgenus Sarbecovirus27-29 (Fig. 2). In accordance to the results of the phylogenetic analysis 2019 nCoV had clustered around SARS-COV, Bat-SL-CoVZ45 and bat-SL-CoVZXC21 indicating that it could be a recombinant in between the two28-30 (Fig. 2).
Although with recent identified strains of the Covid-19 it is clear that the virus is capable of self evolving and crossing the host range, yet some research workers still believe that the recombination could have been possible in a BSL-4 facility using the tools and techniques of RDT31-34. Scientists believing in the Conspiracy hypothesis have provided circumstantial evidences for the indicating that the virus could be a potential bioweapon31-35. Firstly the origin of virus and its report occurred in an area just 30 km away from the Wuhan Institute of Virology, Chinese Academy of Sciences, located in Jiangxia District, Wuhan31-34,36.
Secondly since 2005 a Coronavirus research group was actively involving in research at the Wuhan Institute of Virology reported that horse shoe bats were the natural reservoir for the same37. In 2015 the research group also published scientific findings on infection of Homo sapiens HeLa cell lines with coronavirus38-39. In Feb, 2020 New York Time reported that in Feb, 2020 a patent claim was filed by the Chinese authorities for the drug remdesivir which was found to contain the infection of Corona virus rising smoke indicating dubious role of the Chinese in containing the viral infection40. Although considerable evidence can never be made about the virus being genetically engineered but the roles of Chinese agencies will always remain doubtful31-36.
Another key target protein to illustrate the evolution of COVID-19 are the spike proteins, which is used by the virus to capture and pierce the plasma membrane of animal cells. Spike protein have two important features that includes the receptor-binding domain (RBD), which helps the viral particles in attachment onto the host cells and the cleavage site which helps in viral integration into the host cell.
Recent research have identified that RBD portion of the SARS-CoV-2 spike proteins has evolved to successfully bind to a specific target protein of human cells called ACE2, a receptor involved in blood pressure regulation41. The binding efficiency of the virus to the human cell has also illustrated that the virus had evolved through natural evolution.
The spike protein present in SARS-CoV-2 helps in its effective binding to the human cells. However, the scientists have reached to a conclusion that it was the product of natural selection and not the creation of genetic engineering. Coronavirus belongs to large family of viruses that results in severe infections of wide variety. In 2003, coronavirus has caused a chronic infection i.e. Severe Acute Respiratory Syndrome (SARS) which has caused severe illness in China. Another epidemic of such severe illness occurred in 2012 with the Middle East Respiratory Syndrome (MERS) in Saudi Arabia42-43. In January, 2020 scientists deciphered the genome of the virus that causes COVID-19 i.e. SARS-CoV-2. The first cluster of cases came from workers from the seafood market in Wuhan44,45. Researchers and scientists are now engaged in recognizing the morphology of this virus, which could inspire new drugs, vaccines and other tools to fight the ongoing pandemic.
It is now confirmed that the genome of the new coronavirus is less than 30,000 nucleotide long. Scientists have identified genes coding for 29 proteins, which carries out its life cycle and are responsible for all sought of infections, progressions, disease and ultimately death. The coronavirus genome and its associated proteins encodes are depicted in Table 1. Inside the host cell the first viral protein produced is a chain of 16 proteins that are joined together. Among them two work like scissors, nicking the links between different proteins and releasing them apart from each other in order to carry out their activities including the three structural proteins.
THE SARS-CoV-2 GENOME ORF1ab PROTEIN STRUCTURAL PROTEINS
Spike S, E, M, N
30,000 RNA letters ▶
NONSTRUCTURAL PROTEINS (NSPs)
1, 2, 3, 4, 5, 6, 7, 7a, 7b, 12, 9, 10, 8, 11, 13, 14, 15, 16
ACCESSORY PROTEINS
ORF3a, ORF6, ORF7a, ORF8, ORF 9b, ORF 9c
Researches on other corona viruses have indicated that some of the proteins are structural components while other serves as assessor proteins and there are also others whose function and properties are still mysterious. The key important features and function of each protein has been summarized in table 1.
Table (1):
Comparative analysis of the key important features, function, active amino acid residues of the structural and non structural proteins constituting the proteome of COVID-19.
| S. No. | Category | Protein name | Key structural feature | Probable function | Active site Active Amino acid residues | Reference |
|---|---|---|---|---|---|---|
| 1. | Spike proteins | Spike S | Spike protein (S protein) is a large type I transmembrane protein. It is highly glycosylated, contains 21-35 N-glycosylation sites | It binds to ACE2 receptors over pharynx cells then fuses viral and host membranes through its S2 subunit. Thus is the main protein responsible for viral invasion into the host cell. | Receptor protein (PDB ID: 6LZG) | 42, 43 |
| 2. | Spike E | Structural components still under investigation | The corona virus genome encodes four major structural spike proteins (S, E, M, N) all of which can produce structurally complete viral particles. | |||
| 3. | Spike M | |||||
| 4. | Spike N | |||||
| 5. | Non Structural proteins (NSPs) | NSP1 | b-barrel fold and two disordered regions at the N and C terminal | Triggers the infection cycle by suppressing host gene expression | K164; H165 (PDB ID: 2HSX) | 44 |
| 6. | NSP2 | Four transmembrane helical segments | Acts as a docking site for proteins and stabilizing mutations in endoscope associated proteins makes it contagious | S543; Q321 (Modelled structure) | 45 | |
| 7. | NSP3 | Six transmembrane helical segments | It helps in replication of viral particles in the infected cells, acts as protease to cleave and activate viral proteins | P192 (Modelled structure) | ||
| 8. | NSP4 | Transmembrane tetrapass protein with hydrophobic stretch of amino acids | It helps to anchor the viral replication transmembrane complex (RTC) tomodified endoplasmic reticulum membrane | H120; F121 (EMBL:PUB 00052994) | 46, 47 | |
| 9. | NSP5 | A three domain structure with a chymotrypsin like fold formed by domains Ist and IInd. | Acts as a proteinase to cleave c-terminus of NSP4 protein through NSP 16 for all Corona virus | H41; C145 (Modelled structure) | 46,48 | |
| 10. | NSP6 | Transmembrane multipass protein | It restricts autoplagosomes expansion | (Modelled structure) | 49 | |
| 11. | NSP7 | Three helical (α2 to α4) forms a flat up-down-up antiparallel α-helix sheet. | Protein NSP7 and NSP8 aids NSP12 protein in replication and transcription of coronavirus. NSP12 plays a major role in corona virus replication/ transcription and is the primary target for drugs. |
D618; D760-1 (NSP12) (PDB ID: 6NUR) |
50, 51 | |
| 12. | NSP12 | It has a novel b-hairpin domain at its N-terminal | ||||
| 13. | NSP9 | Four crystallographic structures with variety of dimeric interfaces are reported. NSP9 shows 7 b-strand and a single α-helix. | NSP9 infiltrates nuclear pore apparatus, influencing transcription and translation of human genome. Influences nuclear localization signals and transport. | G100; G104 (Modelled structure) | 52, 53 | |
| 14. | NSP10 | 148 AA NSP10 containing two zinc figure motifs. | It interacts with both NSP14 and NSP16 stimulating their 3’-5- exoribonuclease and 2’-O-methylatransferase activity. | Y126 (PDB ID: 2FYG) | 54 | |
| 15. | NSP8 | Two NSP8 acts as co-factors and binds to N-terminal extension of NSP12. | The NSP-12-NSP7-NSP8 complex is essentially required for nucleotide polymerization. | (PDB ID: 6NUR) | 51 | |
| 16. | NSP13 | It is also a Upf1-like helicase. It has four domains, an N-terminal CH domain and two helicase core domain RecA; RecA2 and 1B. | Exact function unknown. Scientist predicts it could act as a topoisomerase, alter RNA conformation relaxing it to biosynthesize new viral particles | K288; S289; D374; E375; Q404; R567 (PDB ID: 6JYT) |
55 | |
| 17. | NSP14 | It is bimodular; amino acids 287 folds into ExoN domain and 288-527 form N7-MTase domain, a loop of amino acids 288-301. | It acts as a proofreader and proofreads replicated genome as NSP12 lacks a proofreading activity, thus its defective expression leads to high mutational rate in covid-19 | D355; E357; D424; Y497; D501; E191 (PDB ID: 5C8U) |
56 | |
| 18. | NSP15 | Its monomeric structure has a novel protein fold, two timeric NSP15 forms hexamer with active sites | It acts as a RNAse and cleaves off any leftover Viral RNA to protest cell from antiviral defense | H262; H277; K317 (PDB ID: 2GTI) |
57, 58 | |
| 19. | NSP16 | It is only active in the presence of an activating partner NSP10. It binds to NSP10 through ≈930 A°2 surface area bindin pocket | It works along with NSP10 and provides protection to viral genome from antiviral proteins of the cell | (PDB ID: 2XYV) | 59, 60 |
The study of the spike proteins S, E, M and N has revealed that these proteins forms a timeric structure which is responsible for binding to ACE2 receptors over the pharynx cells that causes viral invasion. The genes for these spike proteins are responsible for the evolution of zoonotic corona virus crossing its host range into a disease causing virus for Homo sapiens.
The proteome analysis of the COVID19 proteins has illustrated some key important drug targets. As NSP12 protein in conjugation with NSP7 and NSP8 initiates the replication and transcription of coronavirus COVID-19. So it is the primary target of majority of drugs in order to halt replication and transcription. Recently the first FDA approved drug remdesivir has been shown to exhibit promising effect over combating COVID-19 infection65. Similarly, SARS-NSP13 has been identified as an ideal target for development of anti-viral drugs due to its sequence conservation and indispensability across all CoV species56,57.
The study of COVID-19 proteome is of special significance as it could help in the identification of critically mutated proteins responsible for its pathogenesis and virulence as compared to its non virulent corona viral strains66. Spike proteins (Spike S, Spike E, Spike M and Spike N) of COVID-19 are critical for the invasion of human cell as these binds to the ACE2 receptors on the pharynx cell, have been targeted as a vaccine candidate46-47,66. Another vaccine mRNA-1273, coding for the “spike protein” is under human clinical trials67, while Pfizer vaccine BNT162 is also exploring similar mRNA technology to combat COVID-19. Oxford University research project on the vaccine CHAdOx1 nCOV-19 is based upon the use of genetic engineering to ligate segments of the COVID-19 genome into a non pathogenic modified viral host, hoping to provoke an immune response in humans68.
Although several attempts are also made to synthesize DNA vaccines using recombinant DNA technology, but the high rate of mutations in COVID-19 would make such vaccines ineffective even before the actual treatment begins69. Thus the entire scientific world is left with the choice of developing a protein based vaccine in order to ensure long term protection against COVID-19.
Some other accessory proteins have also been reported that influences the pathogenesis and infection of COVID-19, these are referred to as accessory proteins. ORF3a refers to a group of accessory protein that helps in altering the internal environment of the infected cell making it effective to replicate. It causes the appearance of lessons in the membrane of the infected cell making it easier for new viruses to escape. Another protein encoding gene ORF3b overlaps the same RNA although its function is not well defined70. The ORF6 protein another accessory protein inhibits the signals that the infected cell would produce to the immune system for response. It inhibits the cell antiviral proteins making it more prone to viral infection.
The ORF7a accessory protein triggers an increase in viral load as it enhances movement of viral particles out of the cell. It also triggers auto suicide of the cell causing damage to alveolar tissues that could lead to death. The function of the other accessory protein ORF7b is unknown. The ORF8 mystery protein drastically varies in different corona viruses strains and is responsible for influencing viral pathogenesis. The accessory proteins ORF9b and ORF9c overlap this same stretch of RNA in COVID-19 genome. ORF9b blocks interferon and other cytokines which are an effective molecules of the immune system in combating the virus. Majority of the closed related viruses such as SARS/ MARS do not encode the genes for this protein, thus causing the protein to facilitate the infection of COVID-19.
The corona virus genome ends with a snippet of RNA that stops the cell’s protein-making machinery. The comparative analysis of the COVID-19 protein can help us to identify key important viral proteins which could serve as the targets for vaccine development in combating COVID-19 infection.
Although the origin of Corona virus COVID-19 remains dubious and the matter will always remain debatable among scientific communities but the containment of the virus is the cause of concern. The study of the proteome of the virus could be useful in identifying key target proteins that could serve as the candidature for vaccine development, as well as the phylogenetic analysis of these proteins could help us in predicting the evolutionary pattern of COVID-19 and its transformation from a Zoonotic virus to an infection one. This could help us in predicting the pattern of viral evolutions and taking preventive actions against the reoccurrence of such pandemics in near future.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declares that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets analyzed during this study are included in the manuscript.
- Enjuanes L, Almazan F, Sola I and Zuniga S. Biochemical aspects of coronavirus replication and virus-host interaction. Annu Rev Microbiol. 2006;60:211–30.
Crossref - Perlman S and Netland J. Coronaviruses post-SARS: update on replication and pathogenesis. Nat Rev Microbiol. 2009;7:439–50.
Crossref - National Center for Immunization and Respiratory Diseases (NCIRD), Division of Viral Diseases. February 15, 2020.( https://www.cdc.gov/coronavirus/types.html).
- https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200306-sitrep-46-covid-19.pdf?sfvrsn=96b04adf_2.
- https://www.sfgate.com/bayarea/article/WHO-global-mortality-rate-death-World-Health-Org-15104402.php
- https://coronavirus.jhu.edu/map.html.
- Ksiazek TG, Erdman D, Goldsmith CS, Zaki SR, Peret T. A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med. 2003;348:1953-66.
Crossref - Peiris JSM, Lai ST, Poon LLM, Guan Y, Yam LYC. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet. 2003;361:1319-25.
Crossref - Marra MA, Jones SJM, Astell CR, Holt RA, Brooks-Wilson A. The genome sequence of the SARS-associated coronavirus. Science. 2003;300:1399-404.
Crossref - Rota PA, Oberste MS, Monroe SS, Nix WA, Campagnoli R. Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science. 2003;300:1394-99.
Crossref - Zaki AM, van-Boheemen S, Bestebroer TM, Osterhaus A, Fouchier RAM. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med. 2012;367:1814–20
Crossref - De-Groot RJ, Baker SC, Baric RS, Brown CS, Drosten C. Middle East respiratory syndrome coronavirus (MERS-CoV): announcement of the Coronavirus Study Group. J Virol. 2013;87:7790–92.
Crossref - Mole B. Deadly pig virus slips through US borders. Nature. 2013;499:388.
Crossref - Stevenson GW, Hoang H, Schwartz KJ, Burrough ER, Sun D. Emergence of porcine epidemic diarrhea virus in the United States: clinical signs, lesions, and viral genomic sequences. J Vet Diagn Invest. 2013;25:649–54.
Crossref - Chen Q, Li G, Stasko J, Thomas JT, Stensland WR. Isolation and characterization of porcine epidemic diarrhea viruses associated with the 2013 disease outbreak among swine in the United States. J Clin Microbiol. 2014;52:234–43.
Crossref - Graham RL, Baric RS. Recombination, reservoirs, and the modular spike: mechanisms of coronavirus cross-species transmission. J Virol. 2010;84:3134–46.
Crossref - Li F. Receptor recognition and cross-species infections of SARS coronavirus. Antiviral Res. 2013;100:246–54.
Crossref - Li WH, Wong SK, Li F, Kuhn JH, Huang IC. Animal origins of the severe acute respiratory syndrome coronavirus: insight from ACE2–S-protein interactions. J Virol. 2006;80:4211–19.
Crossref - Perlman S. Another decade, another coronavirus. The N Eng J Med. 2020. doi: 10.1056/NEJMe2001126. [Epub ahead of print] No abstract available.
Crossref - Wu F, Zhao S, Yu B, Chen YM, Wang W, Hu Y. Complete genome characterisation of a novel coronavirus associated with severe human respiratory disease in Wuhan, China. bioRxiv. 2020.
Crossref - Kirchdoerfer RN, Cottrell CA, Wang N, Pallesen J, Yassine HM. Pre-fusion structure of a human coronavirus spike protein. Nature. 2016;531:118–21
Crossref - Walls AC, Tortorici MA, Bosch BJ, Frenz B, Rottier PJ. Cryo-electron microscopy structure of a coronavirus spike glycoprotein trimer. Nature. 2016;531:114–17.
Crossref - Beniac DR, Andonov A, Grudeski E, Booth TF. Architecture of the SARS coronavirus prefusion spike. Nat Struct Mol Biol. 2006;13:751–52.
Crossref - Li F, Berardi M, Lim WH, Farzan M, Dormitzer PR, Harrison SC. Conformational states of the severe acute respiratory syndrome coronavirus spike protein ectodomain. J Virol. 2006;80:6794–800.
Crossref - Kristian G, Andersen AR, WIL, Edward C, Holmes RFG. The proximal origin of SARS-CoV-2. Nature Medicine. 2020;
Crossref - Zhao WM, Song SH, Chen ML. The 2019 novel coronavirus resource. Yi Chuan. 2020;42(2):212–221. doi:10.16288/j.yczz.20-030 [PMID: 32102777].
- Lu R, Zhao X, Li J, et al. Genomic characterization and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. The Lancet. 2020;395:565-574.
Crossref - Hu D, Zhu C, Ai L. Genomic characterization and infectivity of a novel SARS-like coronavirus in Chinese bats. Emerging Microbes and Infections. 2018;7:154.
Crossref - Lu G, Wang Q, Gao GF. Bat-to-human: spike features determining‘host jump’ of coronaviruses SARS-CoV, MERS-CoV, and beyond. Trends in Microbiology. 2015;23:468–78.
Crossref - Wang Q, Wong G, Lu G, Yan J, Gao GF. MERS-CoV spike protein: targets for vaccines and therapeutics. Antiviral Res. 2016;133:165–77.
Crossref - Cohen J. “Scientists ‘strongly condemn’ rumors and conspiracy theories about origin of coronavirus outbreak”. Science. 2020, April 14.
Crossref - Fisher M. “Why Coronavirus Conspiracy Theories Flourish. And Why It Matters”. The New York Times. (ISSN 0362-4331)., 2020, April 20.
- Noor P. “A third of Americans believe Covid-19 laboratory conspiracy theory – study”. The Guardian. (ISSN 0261-3077)., 2020, April 20.
- Holland K. “Sorry, conspiracy theorists. Study concludes COVID-19 ‘is not a laboratory construct’”. ABC News, 2020. April 20.
- Taylor J. “Bat soup, dodgy cures and ‘diseasology’: the spread of coronavirus misinformation”. The Guardian, 2020, February 3.
- Taylor A. “Experts debunk fringe theory linking China’s coronavirus to weapons research”. Washington Post., 2020, February 3.
- Li W, Shi Z, Yu M, et al. “Bats Are Natural Reservoirs of SARS-Like Coronaviruses”. Science. 2020;310(5748):676–679.
Crossref - Vineet DM, Boyd LY, Kari D, et al. “A SARS-like cluster of circulating bat coronaviruses shows potential for human emergence”. Nature Medicine. 2015;21(12):1508–1513.
Crossref - Butler D. “Engineered bat virus stirs debate over risky research: Lab-made coronavirus related to SARS can infect human cells”. Nature News, Nature. 2015.
Crossref - “China Wants to Patent Gilead’s Experimental Coronavirus Drug”. Bloomberg News. 2020, February 5.
- Li F, Li W, Farzan M, Harrison SC. Structure of SARS coronavirus spike receptor-binding domain complexed with receptor. Science. 2005;309:1864–68.
Crossref - Peiris JS, Guan Y, Yuen KY. Severe acute respiratory syndrome. Nature Medicine. 2004;10(suppl 12):S88–97.
Crossref - Chan-Yeung M, Xu RH. SARS: epidemiology. Respirology. 2003;8(suppl):S9–14.
Crossref - Tan W, Zhao X, Ma X. A novel coronavirus genome identified in a cluster of pneumonia cases—Wuhan, China 2019−2020. China CDC Weekly. 2020;2:61–62.
- Zhu N, Zhang D, Wang W. A novel coronavirus from patients with pneumonia in China. N Eng J Med. 2020; published online Jan 24.
Crossref - Almeida MS, Johnson MA, Herrmann T, Geralt M, Wuthrick K. Novel beta-barrel fold in the nuclear magnetic resonance structure of the replicase nonstructural protein 1 from the severe acute respiratory syndrome coronavirus. J Virol. 2007;81:3151-3161.
Crossref - Silvia A, Domenico B, Martina B, Marta G, Stefano P, Massimo C. COVID-2019: the role of the nsp2 and nsp3 in its pathogenesis. J Med Virol. 2020.
Crossref - Graham RL, Sparks JS, Eckerle LD, Sims AC, Denison MR. SARS corona virus replicase proteins in pathogenesis. Virus Res. 2008;133(1):88-100.
Crossref - Fung TS, Liu DX. Post-translational modifications of coronavirus proteins: roles and function. Future Virology. 2018;13(6):405-430.
Crossref - Stobart CC, Sexton NR, Munjal H, et al. Chimeric exchange of coronavirus nsp5 proteases (3CLpro) identifies common and divergent regulatory determinants of protease activity. J Virol. 2013;87(23):12611-8.
Crossref - Cottam EM, Whelband MC, Wileman T. Coronavirus NSP6 restricts autophagosome expansion. Autophagy. 2014;10(8):1426-41.
Crossref - Wolfgang P, Margaret A, Johnson TH, et al. Structural Genomics of the Severe Acute Respiratory Syndrome Coronavirus: Nuclear Magnetic Resonance Structure of the Protein nsP7. J Virol. 2005;79(20):12905-12913.
Crossref - Robert NK, Andrew BW. Structure of the SARS-CoV nsp12 polymerase bound to nsp7 and nsp8 co-factors. Nature Communications. 2019;10:2342.
Crossref - Zachary J, Miknis EF, Donaldson TC, Umland RA, Rimmer RS, Baric L, Wayne S. Severe Acute Respiratory Syndrome Coronavirus nsp9 Dimerization Is Essential for Efficient Viral Growth. J Virol. 2009;83(7):3007-3018.
Crossref - Littler DR, Gully BS, Colson RN, Rossjohn J. Crystal structure of the SARS-CoV-2 non-structural protein 9, Nsp9. BioRxiv. 2020;
Crossref - Joseph JS, Saikatendu KS, Subramanian V, et al. Crystal structure of nonstructural protein 10 from the severe acute respiratory syndrome coronavirus reveals a novel fold with two zinc-binding motifs. J Virol. 2006;80(16):7894-7901.
Crossref - Zhihui J, Liming Y, Zhilin R, et al. Delicate structural coordination of the Severe Acute Respiratory Syndrome coronavirus Nsp13 upon ATP hydrolysis. Nucleic Acids Res. 2019;47(12):6538-6550.
Crossref - Yuanyuan M, Lijie W, Neil S, et al. Structural basis and functional analysis of the SARS coronavirus nsp14–nsp10 complex. PNAS. 2015;112(30):9436-9441.
Crossref - Xiaoling X, Yujia ZFS, Zhiyong L, et al. New Antiviral Target Revealed by the Hexameric Structure of Mouse Hepatitis Virus Nonstructural Protein nsp15. J Virol. 2006;80(16):7909-7917.
Crossref - Decroly E, Debarnot C, Ferron F, et al. Crystal Structure and Functional Analysis of the SARSCoronavirus RNA Cap 29-O-Methyltransferase nsp10/ nsp16 Complex. PLoS Pathogens. 2011;7(5):1-14.
Crossref - Kim Y, Jedrzejczak R, Maltseva N, et al. Crystal Structure of NSP15 Endoribonuclease from SARS CoV-2. Center for Structural Genomics of Infectious Diseases (CSGID)., 2020, April 1.
Crossref - Qihui W, Yanfang Z, Lili W, et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell. 2020;181:1-11.
Crossref - Guangwen L, Qihui W, George FG. Bat-to-human: spike features determining ‘host jump’ of coronaviruses SARS-CoV, MERS-CoV, and beyond. Trends in Microbiology. 2015;23(8):468-478.
Crossref - Minasov G, Shuvalova L, Rosas-Lemus M, et al. 1.80 Angstrom Resolution Crystal Structure of NSP16-NSP10 Complex from SARS-CoV-2. Center for Structural Genomics of Infectious Diseases (CSGID). 2020, April 1.
Crossref - Wang M, Cao R, Zhang L, et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel corona virus (2019-nCoV) in vitro. Cell Research. 2020;30:269-271.
Crossref - Kim E, Erdos G, Huang S, et al. Microneedle array delivered recombinant coronavirus vaccines: Immunogenicity and rapid translational development, EBioMedicine. 2020.
Crossref - Negahdaripour M. The Battle Against COVID-19: Where Do We Stand Now? Iranian Journal of Medical Sciences. 2020;45(2):81-82. doi: 10.30476/ijms.2020.46357.
- Doremalen NV, Haddock EFeldmann F, White KM, et al. A single dose of ChAdOx1 MERS provides protective immunity in rhesus macaques. Science Advances. 2020; 1-15.
Crossref - Yu J, Tostanoski LH, Peter1 L, et al. DNA vaccine protection against SARS-CoV-2 in rhesus macaques. Science, 2020; 1-12.
Crossref - The New York Times. Accessed April 21, 2020. https://ww.nytimes.com/2020/04/13/ opinion/coronavirus-immunity.html?utm_source=Nature+Briefing&utm_campaign
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.