ISSN: 0973-7510
E-ISSN: 2581-690X
The diverse microbial populations in milk produce a range of antimicrobial compounds that confer preservative action. Goat milk characterized by high nutritional value, medicinal properties and hypoallergenic in nature, constitutes the most prevalent non-bovine milk consumed globally. Lactic acid bacteria (LAB), renowned for their probiotic properties, are important constituents of goat milk microflora. In this study, bacterial strains with probiotic potential were isolated and characterized from milk samples of indigenous Indian goat breeds. On MRS medium, Gram-positive rods were observed after anaerobic culture. Based on 16s rRNA technique LAB from goat milk were identified to belong to Enterococcus durans and Lactobacuillus plantarum. Antimicrobial activities were observed against known pathogens (Staphylococcus aureus, Klebsiella pneumoniae, Escherichia. coli and Salmonella typhimurium) and minimum inhibitory concentration was found to be low (6.5 mg/ml) for most of the isolates. High resistance to both acidic condition (pH 2.0) and 0.3% bile salts was observed along with marginal increase in cell counts in some isolates. Adherence to Caco-2 cell lines was observed in all the four identified LABs and was higher in case of Enterococcus durans compared to Lactobacillus plantarum. LC-MS analysis revealed the presence of lacticin as one of the key component of cell free extracts (CFE) of selected isolate. Goat milk as a source of possible LAB probiotics opens up a whole new avenue of probiotics from non-bovine milk sources. LAB strains isolated in this study are potential probiotic candidates that can be employed as antimicrobial agents in food and pharmaceutical industries.
Goat milk, Lactic acid bacteria, Probiotics, Antimicrobial activity, Cell surface characteristics, Bacteriocins
Goat milk production accounts for 13.5 % percent of total non-bovine milk production, making it one of the most widely produced non-bovine milk.1,2 The demand and production is on the rise considering its potential to replace bovine milk consumption due to its non-allergenic nature, high digestibility, nutritional superiority and medicinal properties.3 Similar to bovine milk, goat milk harbors rich and complex microbiota, and detailed knowledge of microbial composition of this milk is essential for the production of fermented products.4,5 Microorganisms from the Lactic Acid Bacteria (LAB) group are primarily responsible for the natural fermentation of dairy products, and they are widely studied due to their potential use as adjuvants and bio preservatives in food products.4,6,7 Several studies have shown that LAB have considerable inhibitory activity against pathogenic bacteria such as methicillin-resistant Staphylococcus aureus (MRSA) and microorganisms (Listeria monocytogenes and Salmonella spp) causing food spoilage.8-10 This is attributed to a class of compounds called bacteriocins which form an important class of bioactive compounds in milk.11
Bacteriocins are ribosomally synthesized antibacterial peptides. These peptides have gained considerable interest in the recent past owing to their inhibitory action against a wide range of pathogenic microorganisms.12,13 These peptides are largely produced by Gram-positive bacteria which include species like Lactobacillus, Leuconostoc and Enterococcus that constitute the dominant microflora of milk and other dairy products. Bacteriocins produced by LABs are generally inhibitory to Gram-positive bacteria, many of which are precarious human pathogens such as MRSA, vancomycin intermediate S. aureus, vancomycin resistant enterococci, Streptococcus pneumoniae and Clostridium difficile, amongst others.14 This feature makes them suitable for use in human and veterinary medicine and also in the pharmaceutical industry.15
The natural nutrient rich composition of goat milk makes it an excellent medium to support survival of microorganisms. Because of its richness and the presence of various bacteriocinogenic LAB bacteria, the native microbiota in raw goat milk is particularly noteworthy.2,16 It has been reported that bacteria with known probiotic and bacteriocinogenic properties such as Lactobacillus, Lactococcus, Leuconostoc, Enterococcus and Streptococcus species form an integral part of the microbiota in goat milk.17,18 Different substances, such as lactic acid, hydrogen peroxide, and bacteriocins, have been associated to the antibacterial activity of bacteria isolated from goat milk.19 However, there is lack in information on possible bacteriocins that the strains are capable of producing.18,20 Data on inhibitory activity of LAB strains and the tolerance of LAB from goat milk to gastric conditions along with adherence to gut epithelium are also important to evaluate the probiotic potential of these strains and products derived from goat’s milk. This information will be essential to justify further research with pure bacteriocins required to evaluate a variety of features that allow them to be used as biopreservatives in the food applications.
The aim of this study was to characterize the diversity of the major LAB strains that make up the microflora of raw goat milk from goat breeds indigenous to India and their probiotic potential, although previous studies have isolated and identified LAB from bovine milk and non-bovine milk in other countries, this study was taken up to in order to identify new strains capable of producing bacteriocin from goat breeds in India that could be used as bio preservatives.
Samples and isolation of lactic acid bacteria (LAB)
Raw goat milk samples were gathered from five Indian goat species (Beetal, Osmanabadi, Boer, Jamunapari and Barbari goats). Samples were diluted ten times in 0.85 % NaCl (w/v), and selected dilutions were spread plated onto sterile de Man, Rogosa, and Sharpe (MRS) media (Oxoid, India) containing 1% CaCO3. The plates were incubated both under aerobic and anaerobic conditions at 37 °C for 48 h. After incubation, colonies were enumerated and representative colonies from each sample were selected and subjected to morphological evaluation (Supplementary Fig. ). Gram staining and catalase tests. Gram-positive bacterial strains which tested negative for catalase enzyme were selected for further studies.
Production of lactic acid by selected LAB
Lactic acid production was estimated by standard FeCl3 method, LAB isolates (24 hrs old) were centrifuged at 3500 rpm for 15 min at 4 °C and 25 µl of the cell free supernatant was added to 1ml of 0.2% FeCl3.6H2O. Absorbance was measured at 390 nm and standard plot of absorbance v/s concentration was prepared. Solutions of standard lactic acid was used to plot calibration curve.
Acid and bile tolerance properties of LAB isolates
LAB isolates were cultured in MRS broth at 37 °C for 24 hours. Active cultures were aliquot (0.1 ml) and pH was adjusted to 2.5 with 1N HCl followed by incubation at 37 °C for 3hours for estimating tolerance of isolates to acidic condition. For analyzing bile tolerance, filter sterilized bile solution was added to active cultures to achieve a final concentration of 0.3 % with a duplicate set without bile solution serving as control. All LAB isolated were taken from 0, 1 and 2 hours cultured MRS broth and directly subjected to each of the two stress factors: low pH, bile salts and viable cells were enumerated and growth estimated spectrophotometrically by measuring absorbance at 600 nm. All the experiments were carried out in triplicates, mean standard deviation was calculated.
In-vitro adhesion assay using Caco-2 cell lines
Preparation of LAB isolates for adhesion assay
Freshly cultured isolated colonies of each LAB strain were inoculated into 5ml MRS broth and grown for 18-hours at 37 °C. Bacterial cells were harvested after 18 h from MRS cultures by centrifugation (3500 rpm for 10 min) followed by one wash with PBS buffer (pH=7.3) and one with the assay buffer (DMEM without FBS and antibiotic, pH=7.4). Cell density was adjusted approximately to the desired levels with assay buffer.
Preparation and maintenance of Caco-2 cell lines
Caco-2 cell line was obtained from American Type Culture Collection (ATCC). The cells were routinely cultured in Dulbecco’s modified Eagle’s medium (DMEM) with L-glutamine, 10% fetal bovine serum and 100µg/mL Penicillin-Streptomycin. Incubation was carried out at 37 °C under 5% CO2 atmosphere.
For adhesion assay, grown Caco-2 cells were washed in the flask once with warm Dulbecco’s phosphate buffered saline. The cells were incubated with 0.05% trypsin – EDTA for 5 min followed by addition of warmed complete medium. Detached cells were resuspended and the cell suspension was centrifuged at 2,000 rpm for 3min. Cell pellet was then resuspended in DPBS followed by centrifugation under same condition. The cell pellet (2×105 cells/ml) was resuspended in medium supplemented with 10% serum and without antibiotics.
Adhesion assay
Caco-2 cells (104 cells/well) were seeded in 6-well standard tissue culture plates incubated at 37 °C under 5% CO2 atmosphere, further medium was changed every alternate day. Cells were inspected under an inverted microscope to rule-out contamination. The spent medium was completely removed 24-hours before adhesion assay and cells were washed twice with warm DPBS.1 ml of DMEM (only with 10% serum) was added to each well and incubated at 37 °C for 30 min. Overnight LAB cultures (at 1 × 106 CFU) suspended in 1 ml DMEM medium (only with antibiotics) were added to different wells, to represent a multiplicity of infection (MOI) of 5:1 (bacteria: cells). The plates were incubated at 37 °C in 5% CO2 for 6 hours. Medium was decanted and cell monolayer was washed thrice with warm DPBS. Adhesion was measured by enumerating adhered bacteria per 10 different microscopic fields under 100X objective and mean adherence estimated.
Detection of antimicrobial activity
The inhibitory effect of isolated LAB strains of LAB was determined by disc diffusion method21. LAB isolates were grown in MRS broth for 24 h at 37 °C and cell-free supernatant (CFS) was prepared by centrifugation at 4000 rpm for 30 min followed by filtration through a 0.22 μm filter for sterilisation. Antimicrobial substance from CFS were concentrated by modified solvent extraction method using ethyl acetate. 24 h cultures of the indicator strains (K. pneumoniae, E. coli, S. aureus and S. typhi) in Peptone broth at 37 °C were inoculated on Soyabean Casein Digested agar media at a concentration of 106 cells/ml. Filter paper disks impregnated with CFS (15µl) and Ciprofloxacin (15µl, as standard) were placed on the media with the help of forceps followed by incubation at 37 °C for 24h. The plates were observed for zone of inhibition around the discs.
Identification of selected isolates by 16S rRNA sequencing
Genomic DNA was isolated by using the InstaGeneTM Matrix Genomic DNA isolation kit (Catalog # 732-6030). Ribosomal RNA16S Region Universal primers 27F (AGAGTTTGATCATGGCTCAG) and 1492R (TACGGCTACCTTGTTACGACTT) were used for amplification and sequencing. DNA fragments are amplified using 1 µl of template DNA in 20 µl of total PCR reaction mixture using 50 pmol27F/1492Rprimers and 35 amplification cycles with following program: denaturation: 94 °C for 45 sec; annealing: 52 °C for 60 sec; chain elongation: 72 °C for 60 sec. Montage PCR Clean up kit was used to remove unincorporated PCR primers and dNTPs from PCR products. Sequencing reactions were performed using a ABI PRISM® Big Dye TM Terminator Cycle Sequencing Kits with AmpliTaq®DNA polymerase (FS enzyme) (Applied Biosystems). 16S rRNA gene 27F/1492R universal primers were used to perform single-pass sequencing on each template. Using an ethanol precipitation technique, the fluorescent-labeled fragments were separated from the unincorporated terminators. The samples were resuspended in distilled water and subjected to electrophoresis in an ABI 3730XL Sequencer (Applied Bio systems, USA).
Identification of bacteriocin from LAB isolate by LS/MS
Sample preparation and pre-treatment
Selected LAB isolate was grown in MRS broth for 24-hours and CFS was collected post centrifugation at 4000 rpm at 4 °C for 30 min. CFS was sterilized by filtration through a 0.22 μm filter. Ethyl acetate extracts were obtained from CFS by liquid-liquid partition chromatography in a separating funnel. The ethyl acetate extracts were evaporated to dryness and reconstituted in 1ml of ethyl acetate for LCMS analysis.
LCMS analysis
LCMS was carried out on HPLC system (Shimadzu) coupled with TSQ Quantum Access MAX Triple Quadruples LC-MS system (Finnigan). Liquid chromatography was carried on aLuna C18 (4.6 x 100mm, 5µm) column (Phenomenox) maintained at 35 °C. Acetonitrile: 0.1% formic acid in H2O, 60:40v/v was used as mobile phase at 0.5 ml/min. Mass spectrophotometer was operated in positive ion and negative ion mode with precursor ion mass scan from 50 – 1050 Daltons. Curtain gas was set to 10 psi, gas1 at 20 psi and gas2 at 0 psi. Needle voltage operated at 5000 V and declustering potential at 100 V.
Microbiological characteristics of LAB strains from goat milk and other samples
Microbial analysis of probiotics bacteria from milk samples of 5 different indigenous goat breeds. Total of 12 different bacterial isolates were obtained after growth under anaerobic conditions. Colonies on MRS medium were small, raised, circular and appeared white, yellow, creamish-white of pale yellow in color. All isolates were Gram-positive and were rod shaped or cocco-bacillary in morphology when examined microscopically (Table 1). The morphological and microscopic characteristics of the isolates were consistent with the features of LAB. Isolates from goat milk sample (12 in number) were used in further studies.
Table (1):
Microbiological characteristics and identification of bacterial strains from samples.
| Source | Isolates | Color | Morphology | Gram stain | Identification by 16s rRNA sequencing |
|---|---|---|---|---|---|
| Goat milk samples | Strain 1 | White | Cocci shaped | Positive | Enterococcus durans |
| Strain 2 | Yellow | Cocci shaped | Positive | – | |
| Strain 3 | Cream | Rod shaped | Positive | – | |
| Strain 4 | White | Rod shaped | Positive | – | |
| Strain 5 | Light yellow | Cocci shaped | Positive | — | |
| Strain 6 | Cream | Rod shaped | Positive | Lactobacillus plantarum | |
| Strain 7 | White | Rod shaped | Positive | – | |
| Strain 8 | White | Cocci shaped | Positive | ||
| Strain 9 | White | Rod shaped | Positive | Lactobacillus plantarum | |
| Strain 10 | Light yellow | Rod shaped | Positive | Lactobacillus plantarum | |
| Strain 11 | Light cream | Rod shaped | Positive | – | |
| Strain 12 | white | Cocci shaped | Positive | – |
Probiotic properties of LAB isolates from goat milk
Production of lactic acid by selected LAB strains
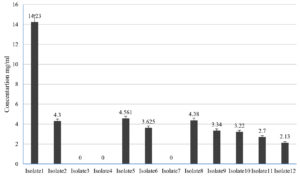
Lactic acid produced by LAB isolates was estimated by comparison with lactic acid standard curve. Of the 12 isolates lactic acid production was observed in 9 isolates. The concentration of lactic acid ranged from 2.0 mg/ml to 14.23 mg/ml among the isolate, however, isolates LAB 3, 4 and 7 were negative for lactic acid production (Fig. 1).
Fig. 1. Concentration of lactic acid produced by LAB isolates from goat milk (Error bars in the figure indicates standard deviation).
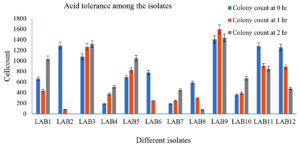
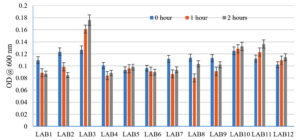
Acid and bile tolerance properties of LAB isolates
LAB 2 and LAB 6 isolates were not resistant to the acidic environment; however, LAB 3 and LAB 9 had good resistance as there was no decrease in cell count was observed (Fig. 2a). In bile tolerance study it was observed that LAB 2 and 4 were not resistant to bile. However, LAB 3, 6,9 10,11 and 12 were tolerant to bile (Fig. 2b).
Adhesion of LAB isolates to Caco-2 cells lines
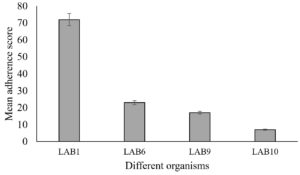
The selected LAB isolates exhibited good adhesion to Caco-2 cell lines after 6 hours of incubation with the Caco-2 cells. LAB 1, 6, 9 and 10 had adherence property. LAB 01 exhibited the highest adherence score of 88 from 10 microscopic fields (Fig. 3).
Fig. 3. Mean adherence scores of selected probiotic LAB isolated from goat milk (Error bars in the figure indicates standard deviation).
Antimicrobial activity of LAB isolates and minimum inhibitory concentration of LAB isolates towards standard pathogenic bacterial strains
All 12 LAB isolates obtained from goat milk samples showed antimicrobial activities towards standard pathogens viz. S. aureus, E. coli, K. pneumonia and S. typhi. Inhibition was estimated by measuring zone of inhibition around wells charged with LAB isolates on plates seeded with pathogens. Maximum inhibition was observed in case of LAB 9, LAB 10, LAB 1 and LAB 12 against all tested pathogens (Table 2). The inhibitory effect was close to that observed with standard ciprofloxacin, indicating potential of these isolates to serve as probiotics. Minimum inhibitory concentration was also found to be low (6.25 mg/ml) for these isolates against the pathogens (Table 2) further corroborating the probiotic potential of these strains.
Table (2):
Zone of inhibition and minimum inhibitory concentration of LAB isolates against standard pathogenic strains.
| K. pneumoniae | E. coli | S. aureus | S. typhi | |||||
|---|---|---|---|---|---|---|---|---|
| Inhibition (mm) | MIC (mg/ml) | Inhibition (mm) | MIC (mg/ml) | Inhibition (mm) | MIC (mg/ml) | Inhibition (mm) | MIC (mg/ml) | |
| Ciprofloxacin standard | 23 | NA | 25 | NA | 23 | NA | 15 | NA |
| LAB1 | 14 | 6.25 | 11 | 25 | 12 | 12.5 | 9 | 50 |
| LAB2 | 11 | 25 | 12 | 12.5 | 11 | 25 | 11 | 25 |
| LAB3 | 11 | 25 | 11 | 25 | 12 | 12.5 | 12 | 12.5 |
| LAB4 | 12 | 12.5 | 12 | 12.5 | 12 | 12.5 | 11 | 25 |
| LAB5 | 12 | 12.5 | 9 | 50 | 10 | 50 | 10 | 50 |
| LAB6 | 14 | 6.25 | 9 | 50 | 10 | 50 | 11 | 25 |
| LAB7 | 10 | 50 | 10 | 50 | 12 | 12.5 | 13 | 6.25 |
| LAB8 | NA | 50 | 11 | 25 | 10 | 50 | 12 | 12.5 |
| LAB9 | 13 | 6.25 | 12 | 12.5 | 13 | 6.25 | 13 | 6.25 |
| LAB10 | 13 | 6.25 | 8 | 50 | 11 | 12.5 | 13 | 6.25 |
| LAB11 | 12 | 12.5 | 9 | 50 | 11 | 12.5 | 12 | 12.5 |
| LAB12 | 10 | 50 | 11 | 25 | 14 | 6.25 | 11 | 25 |
NA: Not available; MIC: Minimum inhibitory concentration expressed as mg/ml of 108 CFU/ml cells in PBS
Molecular identification of selected LAB isolates
Potential LAB strains were identified by 16S rRNA amplification and sequencing. The 16S rRNA sequences were obtained after amplification using 27F/1492R universal primers and were searched by BLAST followed by sequence homology analysis for generation of phylogenetic tree for species identification. Considering morphological characteristics and other microbiological aspects along with data generated by 16s rRNA sequencing, LAB 6, LAB 9, LAB 10 isolates were identified as L. plantarum, one isolate as E. durans (Table 1).
Characterization of bacteriocin form selected LAB isolates by LC-MS
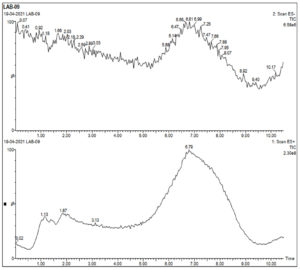
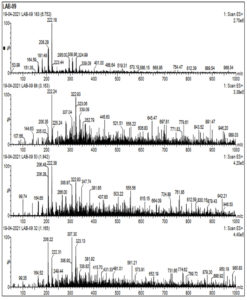
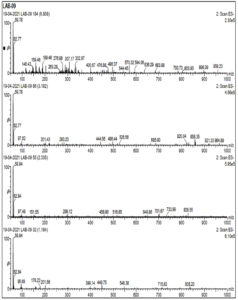
Cell free supernatant for LAB 09, identified as L. plantarum, was analyzed by LC-MS to identify key extracellular compounds. Total ion chromatogram of sample in both negative and positive mode is shown in Fig 4a. In positive mode, 3 peaks were observed at retention time of 6.79, 1.87 and 1.13 min. In negative mode single peak at 6.87min was recorded. LCMS extracted ion chromatogram in positive mode is shown in Fig. 4. In LAB-08 183 fraction the positive mode mass spectrums of compounds show abundant (M + H) +1mass of 222.24, 206.22 and 322.93. LCMS extracted ion chromatogram in negative mode is shown in Fig. 4b. In LAB-08 86 fraction the positive mode mass spectrums of compounds show abundant (M + H) +1mass of 59.78, 189.46 and 159.46. Positive mode m/z of 968.54 at retention time of 6.75 matches with m/z 968.54 reported for Lacticin, which is one of the identified compounds in the CSF of LAB 09 isolate.
LAB has been tested for potential probiotic characteristics from a range of sources, including dairy products, fermented foods and intestinal tract of animals. In this study, 12 LAB strains were isolated from milk of indigenous Indian goat breeds, and were characterized and evaluated for probiotic properties. Evaluation of LAB from milk of Indian goats has not been previously reported to our knowledge, although milk and dairy products from goats are quite prevalent in the Indian subcontinent.
LAB were isolated by enrichment approach using MRS medium as mentioned in earlier studies.22,23. A total of 12 Gram-positive bacteria were isolated after anaerobic culture. Morphological features of these isolates were consistent of morphological features of LAB. Thereafter 12 isolated obtained from goat milk were evaluated in characterization studies. Lactic acid production, a feature consistent with LABs, was observed in 9 of the 12 isolates pointing to their probiotic potential. Out of the 12 LAB isolates, four strains were identified by 16S rRNA sequencing and three isolates were identified to be L. plantarum while the other one isolate was identified as E. durans. All the three isolates proven to be probiotic further subjected for 16SrRNA sequencing for identification suggested previously.24
Antimicrobial activity is one of the most important characteristics of any viable probiotic strain. The LAB strains investigated in this investigation have varying degrees of antagonistic activity against the indicator pathogens. The antimicrobial effect of isolated LAB strains was tested against known pathogens such as S. aureus, K. pneumoniae, E. coli and S. typhi by well diffusion method, which is an established method to study antimicrobial activity in LAB.22 Highest zone of inhibition was observed against S. aureus by LAB strains from goat milk, whereas moderate activity was recorded against K. pneumoniae, E. coli and S. typhi. The LAB strains studied have a broad spectrum of antagonistic activity against both gram-negative and gram-positive pathogens, which is consistent with previous findings.22,25-27 The inhibitory activity of goat milk LAB was also observed against Gram-negative bacterial strains which is contradictory to findings of Anas et al.,28 who reported a higher antagonistic activity against gram-positive pathogens than against gram-negative pathogens. Studies have shown that antimicrobial activity is attributed to antimicrobial molecules such as bacteriocins, organic acids (e.g., acetic, lactic, propionic, succinic acids), short-chain fatty acids, hydrogen peroxide, and other low-molecular-weight chemicals produced by LAB.25 The minimum inhibitory activity also correlated with results of well diffusion tests and concentrations as low 6.5 mg/ml was found to be effective against all pathogens in some of the LAB isolates in this study.
The initial phase in bacterial colonization is bacterial adherence to the host’s tissues, which has long been thought to be a compelling trait among probiotics.29 The gastrointestinal tract of animals is one of the most complex microbial ecosystems, and it can influence nutrient absorption and metabolism, as well as the host’s nutritional and defensive activities.30 Meanwhile, LABs can live, colonize, and cling to host tissues as regular members of the healthy gut microflora. Survival of a probiotic strain in the gut depends on its ability to tolerate the presence of bile salts and low intestinal pH. In this study, all the LAB strains were able to withstand acidic pH condition of 2.0 without any damage to the cells. Also, the strains were also able to survive in buffer supplemented with bile salts showing higher viable counts after 6 h of incubation. These findings corroborated those of Mathara et al.31 who found that LAB strains from milk were tolerant to bile salts at concentrations of 0.1% to 0.5%. It has previously been reported that LAB from various sources may withstand bile salt.32
Adherence of microbial strains to adhere to intestinal cells is regarded as an essential characteristic for choosing the strain as a probiotic.33 LAB isolates from goat milk exhibited good adherence scores on Caco-2 cell lines. The adherence was higher in the E. durans as compared to the L. plantarum strains. These findings are in accordance to the findings of Collado et al.33 and Setyawardani et al.,26 who reported ability of LAB isolates from goat milk samples to adhere to intestinal cells. Probiotics are taken orally and must be able to colonize and compete with pathogens in the gastrointestinal tract.34 Inhibition of growth of pathogens along with adherence is an important attribute of probiotic strains, which has been demonstrated in the LAB strains isolated from goat milk in this study. The adherence of LAB strains is attributed to the production of cell surface proteins that, among other functions, facilitates binding of the bacterial cells to epithelium of the gastrointestinal tract. It has been reported that adherence further enables immunoregulation by LAB, which is also relevant in the removal of pathogens.35 Based on the probiotic properties like acid and bile tolerance, adhesion, lactic acid production and anti-pathogenic activity LAB 9 (L. plantarum) was further continued for the screening of bacteriocin production.
Detection of bacteriocin from CFS was carried out by using LC-MS, which facilitated detection of bacteriocin along with other compounds in the CFS of LAB isolates. Pre-treatment of samples by precipitation techniques was useful in improving bacteriocin detection as impurities were removed from the samples analyzed by LC-MS. The detection of bacteriocins from culture supernatants36 and entire producer cells37 has been reported using MALDI-TOF MS. These approaches were also successful in detecting and identifying bacteriocins with minimal sample preparation. In this study, lacticin was identified in the CSF of L. plantarum strain isolated from goat milk. Furthermore, information on retention time provided by LC-MS can be used as a reference for the identification of bacteriocin from new isolates. There is however scope of further improvements38 in the LC-MS technique employed in this study with respect to decrease in retention time and increase in resolution, which will be useful in high throughput screening of novel bacteriocins.
The findings of this study establish the probiotic potential of LAB isolates obtained for milk samples of indigenous goat breeds. The strains can be further evaluated for development of a potent dairy-based probiotic product which is effective against resistant bacterial pathogens such as MRSA. This emerging class of natural products can be promising alternatives to currently used antibiotics without side effects.
The milk of goat breeds indigenous to India proves to be an excellent source of LAB strains with the probiotic characteristics. To our knowledge, this is the first study on LAB from goat milk. LAB strains isolated and characterized in this study show good inhibitory action against S. aureus and other Gram-negative pathogens. LAB isolates also exhibited resistance to acidic conditions and bile salts for 2-3 h, indicating the ability to survive conditions of GI tract. The cells had the capability to adhere to epithelial cell lines for an extended time period along. All these features highlight the potential of the strains from goat milk to be employed in development of a potent probiotic product. Productions of both lactic acid as well as bacteriocin are added advantages that these strains will confer to the probiotic formulation. This study also highlights the probiotic potential of non-bovine milk sources such as goat milk, which is easily available to all sections of the society. Further studies at genetic level to improve probiotic properties of resistance and cellular adhesion, along with the identification of genes expressing bacteriocins can be carried on these LAB isolates.
Additional file: Additional Figure.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript and supplementary files.
- Nunez M, de Renobales M. IDF international symposium on sheep, goat and other non-cow Milk. Int Dairy J. 2016;58:1.
Crossref - Ranadheera CS, Naumovski N, Ajlouni S. Non-bovine milk products as emerging probiotic carriers, recent developments and innovations. Curr Opin Food Sci. 2018;22:109-114.
Crossref - Slacanac V, Bozanic R, Hardi J, Szabo JR, Lucan M, Krstanovic V. Nutritional and therapeutic value of fermented caprine milk. Int J Dairy Technol. 2010;63(2):171-189.
Crossref - Asteri I, Kittaki N, Tsakalidou E. The effect of wild lactic acid bacteria on the production of goat’s milk soft cheese. Int J Dairy Technol. 2010;63(2):234-242.
Crossref - Psoni L, Kotzamanidis C, Yiangou M ,Tzanetakis N, Litopoulou-Tzanetaki E. Genotypic and phenotypic diversity of Lactococcus lactis isolates from Batzos, a Greek PDO raw goat milk cheese. Int JFood Microbiol. 2007;114(2):211-220.
Crossref - Colombo E, Franzetti L, Frusca M, Scarpellini M. Phenotypic and Genotypic Characterization of Lactic Acid Bacteria Isolated from Artisanal Italian Goat Cheese. J Food Prot. 2010;73(4):657-662.
Crossref - Vera Pingitore E, Todorov SD, Sesma F, Franco BDG de M. Application of bacteriocinogenic Enterococcus mundtii CRL35 and Enterococcus faecium ST88Ch in the control of Listeria monocytogenes in fresh Minas cheese. Food Microbiol. 2012;32(1):38-47.
Crossref - Nero LA, De Mattos MR, De Aguiar Ferreira Barros M, Ortolani MBT, Beloti V, De Melo Franco BDG. Listeria monocytogenes and Salmonella spp. in Raw Milk Produced in Brazil: Occurrence and Interference of Indigenous Microbiota in their Isolation and Development. Zoonoses Public Hlth. 2008;55(6):299-305.
Crossref - Ortolani MB, Moraes PM, Perin LM, et al. Molecular identification of naturally occurring bacteriocinogenic and bacteriocinogenic-like lactic acid bacteria in raw milk and soft cheese. J Dairy Sci. 2010;93(7):2880-2886.
Crossref - Munoz-Atienza E, Gomez-Sala B, Araujo C, et al. Antimicrobial activity, antibiotic susceptibility and virulence factors of Lactic Acid Bacteria of aquatic origin intended for use as probiotics in aquaculture. BMC Microbiol. 2013;13(1):15.
Crossref - Leroy F, Vuyst LD. Bacteriocins of lactic acid bacteria to combat undesirable bacteria in dairy products. Aust J Dairy Technol. 2010;65:143-149.
- Farkas-Himsley H, Musclow CE. Bacteriocins, effect on mammalian cells: mode of action analyzed by flow cytometry and cell sorting. Cell Mol Biol Incl Cyto Enzymol. 1980;26(6):597-603.
- Sahl H, Bierbaum G. Multiple Activities in Natural Antimicrobials. Microbe Magazine. 2008;3(10):467-473.
Crossref - Cotter PD, Ross RP, Hill C. Bacteriocins – a viable alternative to antibiotics? Nat Rev Microbiol. 2013;11(2):95-105.
Crossref - Dischinger J, Basi Chipalu S, Bierbaum G. Lantibiotics: promising candidates for future applications in health care. Int J Med Microbiol. 2014;304(1):51-62.
Crossref - Ortolani MBT, Yamazi AK, Moraes PM, Vicosa GN, Nero LA. Microbiological Quality and Safety of Raw Milk and Soft Cheese and Detection of Autochthonous Lactic Acid Bacteria with Antagonistic Activity Against Listeria monocytogenes, Salmonella Spp., and Staphylococcus aureus. Foodborne Pathog Dis. 2010;7(2):175-180.
Crossref - Quigley EM. Gut bacteria in health and disease. Gastroenterol Hepatol (N Y). 2013;9(9):560-569.
- Perin LM, Moraes PM, Vicosa GN, Silva Junior A, Nero LA. Identification of bacteriocinogenic Lactococcus isolates from raw milk and cheese capable of producing nisin A and nisin Z. Int Dairy J. 2012;25(1):46-51.
Crossref - Raoult D. Probiotics and obesity: a link? Nat Rev Microbiol. 2009;7(9):616.
Crossref - Kruger MF, Barbosa MS, Miranda A, et al. Isolation of bacteriocinogenic strain of Lactococcus lactis subsp. lactis from Rocket salad (Eruca sativa Mill.) and evidences of production of a variant of nisin with modification in the leader-peptide. Food Control. 2013;33(2):467-476.
Crossref - Karimi S, Azizi F, Nayeb-Aghaee M, Mahmoodnia L. The antimicrobial activity of probiotic bacteria Escherichia coli isolated from different natural sources against hemorrhagic E. coli O157:H7. Electron Physician. 2018;10(3):6548-6553.
Crossref - Reuben RC, Roy PC, Sarkar SL, Rubayet Ul Alam ASM, Jahid IK. Characterization and evaluation of lactic acid bacteria from indigenous raw milk for potential probiotic properties. J Dairy Sci. 2020;103(2):1223-1237.
Crossref - Yazdi MKS, Davoodabadi A, Zarin HRK, Ebrahimi MT, Dallal MMS. Characterisation and probiotic potential of lactic acid bacteria isolated from Iranian traditional yogurts. Food Qual Safety. 2017;16(2):185-188.
Crossref - Dallal MMS, Zamaniahari S, Davoodabadi A, Hosseini M, Rajabi Z. Identification and characterization of probiotic lactic acid bacteria isolated from traditional persian pickled vegetables. GMS Hyg Infect Control. 2017;12:Doc15.
Crossref - Grosu-Tudor S-S, Stancu M-M, Pelinescu D, Zamfir M. Characterization of some bacteriocins produced by lactic acid bacteria isolated from fermented foods. World J Microbiol Biotechnol. 2014;30(9):2459-2469.
Crossref - Setyawardani T, Rahayu WP, Maheswari RR, Palupi NS. Antimicrobial activity and adhesion ability of indigenous lactic acid bacteria isolated from goat milk. Int Food Res J. 2014;21:959-964.
- Jose NM, Bunt CR, Hussain MA. Comparison of Microbiological and Probiotic Characteristics of Lactobacilli Isolates from Dairy Food Products and Animal Rumen Contents. Microorganisms. 2015;3(2):198-212.
Crossref - Anas M, Eddine HJ, Mebrouk K. Antimicrobial activity of Lactobacillus species isolated from Algerian raw goat’s milk against Staphylococcus aureus. World J Dairy Food Sci. 2008;3(2):39-49.
- Piwat S, Sophatha B, Teanpaisan R. An assessment of adhesion, aggregation and surface charges of Lactobacillus strains derived from the human oral cavity. Lett Appl Microbiol. 2015;61(1):98-105.
Crossref - Coman MM, Verdenelli MC, Cecchini C, et al. Probiotic characterization of Lactobacillus isolates from canine faeces. J Appl Microbiol. 2019;126(4):1245-1256.
Crossref - Mathara JM, Schillinger U, Kutima PM, et al. Functional properties of Lactobacillus plantarum strains isolated from Maasai traditional fermented milk products in Kenya. Curr Microbiol. 2008;56(4):315-321.
Crossref - Tang H, Qian B, Xia B, et al. Screening of lactic acid bacteria isolated from fermented Cornus officinalis fruits for probiotic potential. J Food Safety. 2018;38(6):e12565.
Crossref - Collado MC, Jalonen L, Meriluoto J, Salminen S. Protection mechanism of probiotic combination against human pathogens: in vitro adhesion to human intestinal mucus. Asia Pac J Clin Nutr. 2006;15(4):570-575.
- Neal-McKinney JM, Lu X, Duong T, et al. Production of organic acids by probiotic lactobacilli can be used to reduce pathogen load in poultry. PLoS One. 2012;7(9):e43928.
Crossref - Cunningham-Rundles S, Ahrne S, Bengmark S, et al. Probiotics and immune response. Am J Gastroenterol. 2000;95(1):S22-S25.
Crossref - Zendo T, Nakayama J, Fujita K, Sonomoto K. Bacteriocin detection by liquid chromatography/mass spectrometry for rapid identification. J Appl Microbiol. 2008;104(2):499-507.
Crossref - Hindre T, Didelot S, Le Pennec J-P, Haras D, Dufour A, Vallee-Rehel K. Bacteriocin detection from whole bacteria by matrix-assisted laser desorption ionization-time of flight mass spectrometry. Appl Environ Microbiol. 2003;69(2):1051-1058.
Crossref - Alshammari E, Patel M, Sachidanandan M, Kumar P, Adnan M. Potential Evaluation and Health Fostering Intrinsic Traits of Novel Probiotic Strain Enterococcus durans F3 Isolated from the Gut of Fresh Water Fish Catla catla. Food Sci Anim Resour. 2019;39(5):844-861.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.