ISSN: 0973-7510
E-ISSN: 2581-690X
The vaginal microbial composition of wellbeing women were predominated by Lactobacillus group which play prime character in health-promoting activities including the prevention of the colonization of pathogenic microbe. The present study focused to characterize the probiotic and cytotoxicity potential of the Lactobacillus strains collected from the vaginal environment of the healthy women. Lactobacillus strains were identified via biochemical characterization and 16SrRNA gene sequencing. Strains were also evaluated for their probiotic potentials that includes antibiotics sensitivity; survivability to gastrointestinal conditions; antibacterial activities, biofilm production, auto aggregation and adhesion abilities to hydrocarbon. Cytotoxic activities were assessed through 3-[4,5-dimethylthiazole-2-yl]-2,5-diphenyltetrazolium bromide (MTT) assay using HeLa (cervical), AGS (gastric cancer) and A549 (lung cancer) cell lines. Results identified five bacterial strains viz., (Lactobacillus mucosae K76, L. fermentum K81, L. fermentum K85, L. reuteri K97 and L. reuteri K99) contains high probiotic potential and three strains with high cytotoxic activities against cancer cell lines can be further explodes for the human beneficial activities. Finding from this study also indicates that vaginal environment represents a novel source of probiotic candidates.
Microbial diversity, bacterial vaginosis, antimicrobial activity.
The microbial populations living in vaginal microenvironment are known as vaginal microbiota form a complex network of the diverse groups of microbes. In normal healthy females, lactobacilli group predominate vaginal microenvironment and resists the pathogens by maintaining acidic pH in the vagina1. On certain situations such as unhygienic lifestyle, improper vaginal douching, random sexual behavior etc.,2 the Lactobacillus population declines which triggers the colonization of pathogenic microbes triggering bacterial vaginosis (BV). The BV leads to several clinical conditions such as vulnerability to certain pathogens, the Herpes Simplex Virus (HSV-2 and HIV-1) and in severe cases it may lead to late fetal loss and pre-termed delivery3. Therefore, the vaginal microbiota, in particular, the Lactobacillus group, in recent years has become an important subject of extensive research in view of their probiotic potential in BV.
Lactic acid bacteria (LAB) have been utilized as potential probiotic microorganisms in several microenvironments of an individual body e.g. gastrointestinal tract, oral cavity, and urogenital tract etc.4. Apparently, probiotic properties of the LAB are strain specific5. In vaginal microenvironment, lactobacilli are the resident microbiota which inhibits invasion of pathogenic microorganisms6,7. However, the probiotic potential of the vaginal lactobacilli strains is poorly known. It is important to note here that biodiversity of vaginal microorganism reported from different countries8-10 open up new possibilities of testing probiotic potentials of such microbes and supplementing them to vaginal ecosystem for curing diseases like bacterial vaginosis (BV).
Probiotics also exert immunomodulatory, anti-inflammatory, and anti-carcinogenic activities11,12. Among the LAB, Lactobacillus received major attention as probiotics. Lactobacilli are the resident microbiota which inhibits invasion of pathogenic microorganisms6,7. The activity of these bacteria is strain specific and depend on the products secrete by them5. Lactobacillus secretes organic acids, bacteriocin, biosurfactants, co-aggregation molecules and hydrogen peroxide which can restrict the growth of pathogenic microorganisms. Adhesion of pathogens to epithelial membrane was restricted by biosurfactants produced by lactobacilli. Organic acids, bacteriocin and hydrogen peroxide restrict the growth of pathogens and dispersion of pathogens were inhibited by coaggregation molecules. Although so many works are concentrated on the co-relation between probiotics and anti-cancer activities, none of them have explained the mechanisms13. Nevertheless, recently a few mechanisms have been put forwarded e.g., modification of microbiome; breaking down of carcinogenic molecules; regulation of apoptosis and competition with pathogens14.
In the present study, probiotic potentials and anticancer activity of some microorganisms with high antimicrobial activity were assessed which were isolated from the vaginal swab of women from North East India.
Sample collection
Vaginal swab samples were collected from 83 healthy pregnant and non-pregnant women during the period from 23 July 2012 to 29 November 2015. All women were at reproductive age i.e. 18-47. Ethical clearance for the study has been obtained from the Institutional Ethical Committee, Karimganj College (KC/IEC/2012/M-1/10, dated: 23 Jun 2012). The samples were collected by a practicing gynecologist of Red Cross Society Hospital, Karimganj, Assam, India. Initially, the experimental procedure was explained to the volunteers and when they agreed to participate in the study, they were asked to sign a written consent which was prepared as per the recommendation of the Institutional Ethical Committee. The samples were collected by a sterile tube and inoculated in De Man, Rogosa and Sharpe (MRS) agar media by streak-plate method15. After 48 h incubation at 37°C microbial colonies were obtained.
Physiological and biochemical identification
The isolates were characterized by colony morphologically done by Gram staining, catalase test and formation of spores. Biochemical characterization was done for IMViC test, nitrate reduction test and citrate utilization test by following Bergey’s Manual of Determinative Bacteriology. Carbohydrate fermentation pattern was also determined.
Molecular characterization
The genomic DNA of bacteria was isolated as per the method described by Purkayastha et al., (2017)16 with some modifications. The isolates were identified 16S rRNA gene was amplified using universal primers, 27s Forward primer (AGAGTTTGATCMTGGCTCAG) and 1492R Reverse primer (5ˈ-GGTTACCTTGTTACGACTT-3ˈ)17.
Sequence analysis
After obtaining the sequencing data, these were analyzed and edited using Sequence scanner software (v1.0). Thereafter, the sequence was matched using BLASTn (NCBI database) to identify its closest relatives. The sequences were submitted in the NCBI database using the submission tool Bankit and the accession numbers were obtained 18,19.
Studies of some probiotic characters
Effect of PH
Effect of pH was tested according to Shokryazdan et al.20, with modifications. All the LAB isolates (with final conc. of 7 to 8 log CFU/mL Phosphate Buffer Saline) were inoculated in sterile PBS buffer (1% V/V) and were adjusted to pH 3, 4, 5, 6, 7, 8 with 1 M HCl and 1 M NaOH. A control was prepared by PBS with pH of 7.2 and was incubated at 37°C for 3 hours. After that 100µL of each sample was incubated in MRS broth at 37°C for overnight. After 24 hours of incubation OD was measured (600 nm) and pH of the medium tested.
Antagonestic assay
The bacterial colonies were incubated in MRS broth (37°C, 48 hrs). Bacterial cells were separated by centrifugation (4200g at 4°C for 10 mins). The supernatant was taken out and filtered through membrane filter with pore size 0.22µm. which is used for antimicrobial assay.
Antimicrobial assay was performed followed by the method described by Vinderola et al.,21. Mueller-Hinton agar plates were prepared and seeded with pathogens viz., Bacillus subtilis MTCC 736, Escherichia coli MTCC 9492, Proteus mirabilis MTCC 425, Proteus vulgeris MTCC 771, Pseudomonas aeruginosa MTCC 3541, Staphylococcus aureus MTCC 3160 (0.5 McFarland standards). Wells were prepared with 5mm diameter. 50µl of Lactobacillus samples prepared for antimicrobial assay was loaded in each well. After incubation for 24 hrs at 37°C the pates were taken out from incubator and inhibition zone were measured for each well.
Susceptibility against antibiotics
Antibiotics susceptibility assay performed on 90% ISO + 10% MRS agar plates. An overnight culture (37°C, 24 h) of Lactobacillus species were seeded with a sterile cotton swab and allowed to air dry. 12 antibiotics (HiMedia, Mumbai, India) were used for the assay using agar-disc diffusion method. The test carried out triplicate and after incubation of 24 hrs at 37°C the average inhibition zone was calculated. A sterile plate seeded with overnight Lactobacillus culture was considered as a control for the experiment 22.
Screening of biofilm production
Overnight cultures (200µl) in MRS broth of Lactobacillus isolates were transferred into pre-sterilized polystyrene microtiter plates (GeNei, Bangalore), and incubated overnight. After incubation, 50µl of 1% crystal violet was added. After 10 min of incubation at room temperature wells were washed with distilled water. 200µl of 95% ethanol was added and incubated for 10 min. The optical density (OD) was measured at 540nm in Bio Spectrometer (Eppendorf Pvt. Ltd, Germany). OD >0.500 recognized high biofilm producers, OD between 0.500 and 0.100 biofilm producers and lower than 0.100 are considered as poor biofilm producers 23.
Determination of auto aggregation ability
Overnight cultured Lactobacillus strains were harvested by centrifuging in 5000g for 15 min. cells were diluted to reach an OD of 0.5 at 600nm by washing with PBS buffer (pH 7.4). Now, 4ml of cell suspensions of lactobacilli isolates were mixed by using vortex mixture for 10s. The aliquots were incubated for 4 h at 37°C. The percentages of autoaggregation (%) were determined by 100 x [1-(ODA/ODB)]; where ODA is the absorbance after 4 h of incubation and ODB is the absorbance before incubation 24.
Bacterial adhesion to hydrocarbon
Adhesion to the hydrocarbon is also related with the surface hydrophobicity of the bacterial cell surface. An overnight culture of Lactobacillus strains was produced as narrated for the autoaggregation assay (OD at 600nm = 0.5). Each of 1ml of hydrocarbon xylene and chloroform was mixed with 3ml of cell suspension, and incubated for 10 min at room temperature. After that, the heterogeneous solution was vortexed for 2 min and again incubated for 20 min. The absorbance of the aqueous layer was measured at 600 nm. The percentage of cell surface hydrophobicity was determined was determined by 100 x [1-ODA/ODB]; where ODA is the absorbance after mixing hydrocarbon and ODB is the absorbance before mixing hydrocarbon solvents 24.
Study of cytotoxicity of intracellular total protein
Cell lines and culture medium
HeLa (cervical), AGS (gastric cancer) and A549 (lung cancer) cell lines were procured from the National Centre for Cell Sciences (NCCS), Pune, India. Stock cells were cultured in the DMEM supplemented with 10% inactivated Fetal Bovine Serum (FBS), penicillin (100 IU/ml), streptomycin (100µg/ml) and amphotericin B (5µg/ml) in an humidified atmosphere of 5% CO2 at 37°C until confluent. The cells were dissociated with trypsin solution (0.2% trypsin, 0.02% EDTA, 0.05% glucose in PBS). The stock cultures were grown in 25 cm2 culture flasks, and all experiments were carried out in 96 microtitre plates (Tarsons India Pvt. Ltd., India).
Sample Preparation
The total protein of K76, K81 and K85 cultures were estimated using Bradfords method and BSA was used for standard. The K76, K81 and K85 were harvested at their log phases. The cultures were centrifuged at 8600g for 10 minutes. The supernatant was discarded and pellet was homogenized using sonicator at 4°C. The total protein was estimated and 50µg/ml concentration of all strains was used for experiment. The standard Doxorubicin (DOX) at the concentration of 5µg/ml was used as positive control.
Cell cytotoxicity
The cytotoxic effect of the intracellular total protein of K76, K81 and K85 was investigated using the MTT (Sigma, USA) on HeLa, A549 and AGS cell lines. The cells were seeded in 96-well plates at a density of 2 x 104 cells per well. After incubation for 20–24 h, the cells with 80–85% confluency was treated with the 50µg mL-1 of intracellular protein and incubated for 24 h. Then, 20µL of MTT (5mg/mL) solution was added to cells per well, and the plate was moved to a cell incubator for another 4 h. The medium was removed, and 150 mL of DMSO was added to the cells. The plate was gently shaken for 15 min to dissolve the formazan crystals generated by proliferating cells, and the measurement was performed using a Spectramax M2 Microplate Reader (Molecular Diagnostic, Inc.) at a wavelength of 570 nm. Relative viability was calculated taking wells with non-treated cells as 100% control. The results are expressed as mean values (±SD) of three repeats 25.
All the data’s were statistically analyzed using single factor ANOVA to assess significant differences between and within the bacterial isolates.
Identification of bacteria
Altogether 154 isolates were characterized through 16S rRNA gene sequencing technique by homology search with database of NCBI using BLASTn approach and EZ-Taxon26. Out of these, five isolates (Lactobacillus mucosae K76, L. fermentum K81, L. fermentum K85, L. reuteri K97 and L. reuteri K99 with corresponding accession number viz., KR264988, KR264990, KR264981, KR264984 and KR264986) were selected for further studies based on their antimicrobial activity against pathogens considered in this work.
Studies of some probiotic characters
Effect of PH
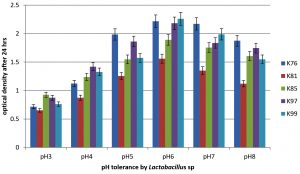
It was found that all the five Lactobacillus isolates can grow in different pH i.e., from 3 to 8 and at pH 6 the bacterial isolates showed highest optical density and it was also observed that after overnight incubation pH of all the individual culture falls or reaches to 4-4.5 (Table 1 and Fig. 1).
Table (1):
pH tolerance of Lactobacillus mucosae (K76), L. fermentum (K81), L. fermentum (K85), L. reuteri (K97) and L. reuteri K99) to different pH. Error bars indicate ± standard deviation.
| pH of the medium | pH of the medium after 24h of incubation | ||||
|---|---|---|---|---|---|
| K76 | K 81 | K 85 | K97 | K99 | |
| 3 | 3.54±0.6 | 3.53±0.34 | 3.76±0.34 | 3.55±0.25 | 3.62±0.26 |
| 4 | 4.43±0.5 | 4.27±0.15 | 3.86±0.15 | 3.95±0.35 | 3.89±0.4 |
| 5 | 4.4±0.3 | 4.18±0.3 | 4.02±0.25 | 4.19±0.28 | 4.02±0.38 |
| 6 | 4.23±0.21 | 4.06±0.25 | 4.16±0.21 | 4.2±0.35 | 4.18±0.26 |
| 7 | 4.18±0.12 | 4.29±0.18 | 4.2±0.3 | 4.25±0.41 | 4.28±0.35 |
| 8 | 4.28±0.25 | 4.37±0.22 | 4.36±0.16 | 4.37±0.38 | 4.32±0.45 |
Fig. 1. Tolerance of Lactobacillus mucosae (K76), L. fermentum (K81), L. fermentum (K85), L. reuteri (K97) and L. reuteri K99) to different pH. Error bars indicate ± standard deviation.
Table (2):
Percentage of auto-aggregation and cell-surface hydrophobicity of the Lactobacillus mucosae (K76), L. fermentum (K81), L. fermentum (K85), L. reuteri (K97) and L.reuteri K99). Error bars indicate ± standard deviation.
Selected Lactobacillus sp. |
Auto-aggregation (%) |
Hydrophobicity (%) |
|---|---|---|
K76 |
51.35±1.81 |
57.68±1.38 |
K81 |
50.02±1.65 |
48.35±1.62 |
K85 |
44.36±1.76 |
51.46±2.5 |
K97 |
42.58±1.92 |
58.23±1.74 |
K99 |
41.45±2.14 |
57.67±0.75 |
Antagonestic Assay
These vaginal lactobacilli showed very good antimicrobial activity against both Gram-positive (Staphylococcus aureus MTCC 3610, Bacillus subtilis MTCC 736) and Gram-negative bacteria ( Escherichia coli MTCC 9492, Proteus mirabilis, MTCC 425, Proteus Vulgaris MTCC771 and Pseudomonas aeruginosa MTCC 3541). Antagonistic activity is an important property for selecting probiotics and all five studied strains revealed strong antagonistic activity (Table 3).
Table (3):
Antagonistic activity of Lactobacillus mucosae (K76), L. fermentum (K81), L. fermentum (K85), L. reuteri (K97) and L. reuteri K99) and zone of inhibition (mm) against test pathogens. Error bars indicate ± standard deviation.
| Isolates | Test organisms (MTCC) | |||||
|---|---|---|---|---|---|---|
| Bacillus subtilisMTCC 736 | Escherichia coli MTCC 9492 | Proteus mirabilis MTCC 425 | Proteus vulgerisMTCC771 | Pseudomonas aeruginosa MTCC3541 | Staphylococcus aureus MTCC3160 | |
| K76 | 13.33 ± 1.33 | 11.66 ±0.33 | 14.33 ±0.87 | 10.66 ±0.66 | 17.66 ± 0.66 | 15.66 ± 0.62 |
| K81 | 10.66 ±0.33 | 17.66 ±0.66 | 15.66 ±0.62 | 13.33 ±1.33 | 14.33 ± 0.87 | 15.66 ± 0.62 |
| K85 | – | 11.66 ±0.33 | 10.66 ±0.33 | 11.66 ±0.33 | 11.66 ± 0.33 | 18.33 ± 0.57 |
| K97 | 16.33 ±0.57 | 19 ± 0.70 | 14.33 ±0.87 | 14.33 ±0.87 | 18.33 ± 0.57 | 15.66 ± 0.62 |
| K99 | 11.66 ±0.88 | 15.66 ±0.62 | 15.66 ±0.62 | 10.66 ±0.33 | 13.33 ± 1.33 | 15.66 ±0.62 |
Antibiotics susceptibility test
It was found that L. reuteri K99 was resistant to all antibiotics except for quinolone (norfloxacin). L. reuteri K97 was resistant against all the antibiotics except amoxyclav, ciprofloxacin, Methicillin and Norfloxacin. L. fermentum K85 showed sensitivity against all other test antibiotics except Norfloxacin, Nitrofurantoin, and Streptomycin. On the other hand, L. mucosae K76 and L. fermentum K81 were sensitive to all other test antibiotics. (Table 4).
Table (4):
Antibiotics susceptibility of Lactobacillus mucosae (K76), L. fermentum (K81), L. fermentum (K85), L. reuteri (K97) and L. reuteri K99). Error bars indicate ± standard deviation.
| Name of the Antibiotics | Quantity in the disc (mcg/disc) | Zone of inhibition (in mm) | ||||
|---|---|---|---|---|---|---|
| K76 | K81 | K85 | K97 | K99 | ||
| Amoxicillin (AMP) | 10 | 28.67±1.07 | 28±1.63 | 28.33±1.25 | R | R |
| Amoxyclav (AMC) | 30 | 26.33±1.01 | 26±0.82 | 27.33±0.47 | 24.67±1.25 | R |
| Chloramphenicol (C) | 30 | 24.33±1.01 | 25.33±1.7 | 25.67±1.25 | R | R |
| Ciprofloxacin (CIP) | 5 | 20.67±0.38 | 26.33±1.25 | 27.67±0.47 | 21.67±1.25 | R |
| Methicillin (MET) | 5 | 15 ± 1.0 | 14±1.63 | 15±0.82 | 15.67±0..47 | R |
| Norfloxacin (NX) | 10 | 30 ± 1.25 | 27.67±1.25 | R | 24.33±1.25 | 21.67±1.25 |
| Nitrofurantoin (NIT) | 300 | 26±1 | 21±0.82 | R | R | R |
| Streptomycin (HLS) | 300 | 26.67±1.5 | 25.67±1.25 | R | R | R |
| Tetracycline (TET) | 30 | 24±1 | 13.33±1.7 | 25.67±1.25 | R | R |
| Vancomycin (VA) | 30 | 32 ± 1.7 | 19.67±1.25 | 18.33±1.25 | R | R |
*R – resistant
Biofilm production
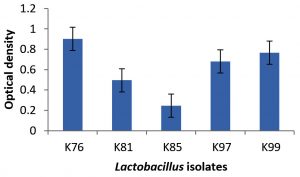
In the present experiment, L. mucosae K76 produced the highest percentage of biofilm. The property of biofilm production by the vaginal isolates is an important characteristic towards the adherence to the mucosal membrane (Fig. 2).
Fig. 2. Production of biofilm by Lactobacillus mucosae (K76), L. fermentum (K81), L. fermentum (K85), L. reuteri (K97) and L. reuteri K99). Error bars indicate ± standard deviation.
Cell cytotoxicity
The percentage cytotoxicity of intracellular protein of L. mucosae K76, L. fermentum K81, L. fermentum K85 and doxorubicin against HeLa were 71.55±0.24, 69.84±0.73, 63.47±2.90 and 74.46±6.68, respectively. Treatment against AGS cells shown cytotoxic effect of 56.96±1.89, 44.72±2.12, 47.85±6.41, 54.68±3.36, respectively and A549 cell treated with K76, K81, K85 and doxorubicin exhibited percent toxicity of 76.28±1.35, 77.45±2.14, 76.65±0.45, 79.23±4.23. The data expressed as Mean ± standard deviation.
According to of WHO27, many bacteria are proposed as probiotic. However, only LAB has been center of focus in the vaginal tract, even though only few of them were reported for their probiotic activities28-31. In the present experiment Lactobacillus mucosae K76, L. fermentum K 81, L. fermentum K 85, L. reuteri K97 and L. reuteri K99 were isolated from the vaginal swab of women of North East India and tested for their antimicrobial activity. Five isolates showed highest antimicrobial activity and were further studied for their probiotic properties. Antimicrobial activity against potential pathogens is an essential and novel property for selection of probiotics. It was observed that all the five Lactobacillus can inhibit the growth of both Gram positive and Gram negative bacteria. Probiotic strains secrete inhibitory products such as bacteriocins, organic acids, hydrogen peroxide etc which restrict the growth of other pathogens by competing for nutrients and colonization to the epithelial membrane 24,28,31.
The probiotic natures of these organisms were studied in respect of biofilm production, antibiotic susceptibility, auto-aggregation, adhesion to hydrocarbons, pH tolerance. The pH tolerance by the lactobacilli under study was tested for pH range from 3 to 8. The results of the study indicated that at low pH (pH 3) (F= 37602, F crit = 3.47805, P= 7.79 E-21) highest growth was exhibited by L. fementum K85 followed by L. reuteri K97, L. reuteri K99, L. fermentum K81 and L. mucosae K76. Similarly, at high pH (pH 8) (F=245214, F crit = 3.47805, P=6.61 E-25), the best growth was exhibited by Lactobacillus mucosae K76 followed by L. fementum K85, reuteri K97, L. reuteri K99, and L. fermentum K81. However, all the bacteria showed maximum growth at pH 6 (F= 52.307, F crit = 2.32, P=2.27 E24), the highest being exhibited by L. reuteri K99. The survival of Lactobacillus strains at low pH less than 3 has also been reported by many other workers32-34. It is interesting to note here that after overnight growth the pH of all culture medium set at pH 3 to 8 stabilized in the range of pH 3.53 to 4.43. Tolerance against low pH is an important property for selecting bacterial isolates as potential probiotics candidates as the bacteria have to grow in low vaginal pH32.
It was found that L. reuteri K99 was resistant to all antibiotics except for quinolone (norfloxacin). L. reuteri K97 was resistant against all the antibiotics except amoxyclav, ciprofloxacin, Methicillin and Norfloxacin. L. fermentum K85 showed sensitivity against all other test antibiotics except Norfloxacin, Nitrofurantoin, and Streptomycin. On the other hand, L. mucosae K76 and L. fermentum K81 were sensitive to all other test antibiotics. This might be due to impermeability, lack of hydrogenase activity or resistance gene coding for b-lactamases, ribosomal protection, or antibiotic-efflux proteins present in these bacteria23,24,35.
In respect of biofilm production, highest performance was exhibited by L. mucosae K76 followed by L. reuteri K99, L. reuteri K97, L. fermentum K81 and L. fermentum K85. Martin et al.,22 also observed that many strains of Lactobacillus have the ability to produce biofilm but three species of L. jensenii were found to produce the highest amount of biofilm.
The surface properties e.g., auto-aggregation and surface hydrophobicity are parameters for studying the colonization ability of Lactobacillus species in the vaginal or intestinal wall. Workers reported the significant difference in hydrophobicity value within the same species. In the present study, it was also found that there is a difference in hydrophobicity value within the species. Workers also reported that there was a correlation between aggregation and the property of surface hydrophobicity36 and studied has also reported that there was no correlation between the aggregation and hydrophobicity37. In our study, it has been observed that the strain with lowest auto-aggregation property showed height surface hydrophobicity value. From the above results, it is evident that all these species of lactobacilli have the potential for use as probiotic organisms.
For cell cytotoxicity, it was observed that all the isolates have cytotoxic activity on cancer cell line, among the three cancer line highest activity showed against A549 (lung cancer) followed by HeLa (cervical) and AGS (gastric cancer). Similar result was also reported Maqsood et al., 201838. However, from the viewpoint of antimicrobial activity and antibiotic sensitivity, L. mucosae K76 and L. fermentum K81 seems more effective.
In the present work five Lactobacillus strains were studied in vitro for different probiotic properties and it was found that all the isolates can be selected for potential probiotic candidate. But L. mucosae K76 was found to be safer depending on the antibiotic resistance profile and also showed best cytotoxic property against all the cancer cell lines. Further investigation may be carried out on the animal model to study the in vivo activity of these strains.
ACKNOWLEDGMENTS
The authors are very grateful to all the volunteer women involved in this study, who dedicated their time, effort and samples for this research. Authors acknowledge with thanks financial assistance received from Department of Biotechnology (DBT), Govt. of India (GOI) under NER Twinning Project (vide DBT Sanction Order No. BT/220/NE/TBP/2011 dated 30th May, 2012) and for providing infrastructural facility DBT, GOI sponsored Advanced Institutional Biotech Hub, Karimganj College (BCIL/NER-BPMC/2010; dated 30th November, 2010) and Advanced State Biotech Hub, Mizoram (No. BT/04/NE/2009; dated 22/ 12/ 2010).
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
SDP, MKB, HKP conceived this study. SDP collected samples. SDP, MKB, HKP designed wet-lab experiments. SDP, SDM & VM carried out wet-lab experiments. SDP, MJB, SDM & VM designed and performed computational analysis, prepared tables and figures and interpreted results. NSK, HKP, MJB offered technical consultation and professional suggestions. SDP wrote the manuscript.
FUNDING
The study was supported by the following projects:
DBT sponsored Twinning project (vide DBT Sanction Order No. BT/220/NE/TBP/2011 dated 30th May, 2012).
DBT sponsored Advanced Institutional Biotech Hub, Karimganj College (BCIL/NER-BPMC/2010; dated 30th November, 2010).
DBT, New Delhi sponsored State Biotech Hub, Mizoram (No. BT/04/NE/2009; dated 22/ 12/ 2010).
ETHICS STATEMENT
Ethical clearance for the study has been obtained from the Institutional Ethical Committee, Karimganj College (KC/IEC/2012/M-1/10, dated: 23 Jun 2012)
AVAILABILITY OF DATA
The data’s (sequences) may be accessed by using their Accession number in the public database NCBI.
- Linhares IM, Summers PR, Larsen B, Giraldo PC & Witkin SS. Contemporary perspectives on vaginal pH and lactobacilli. American journal of obstetrics and gynecology, 2011; 204: 120 e121-125,
Crossref - Verstraelen H et al. Longitudinal analysis of the vaginal microflora in pregnancy suggests that L. crispatus promotes the stability of the normal vaginal microflora and that L. gasseri and/or L. iners are more conducive to the occurrence of abnormal vaginal microflora. BMC Microbiology, 2009; 9: 116,
Crossref - Goldenberg RL, Culhane JF, Iams JD & Romero R. Epidemiology and causes of preterm birth. Lancet, 2008; 371: 75-84.
Crossref - Martinez RM, Hulten KG, Bui U & Clarridge JE, 3rd. Molecular analysis and clinical significance of Lactobacillus spp. recovered from clinical specimens presumptively associated with disease. Journal of Clinical Microbiology, 2014; 52: 30-36.
Crossref - Aureli P et al. Probiotics and health: an evidence-based review. Pharmacological Research, 2011; 63: 366-376
Crossref - Mercenier A, Pavan S & Pot B. Probiotics as biotherapeutic agents: present knowledge and future prospects. Current Pharmaceutical Design, 2003; 9: 175-191.
Crossref - Sartor RB. Probiotic therapy of intestinal inflammation and infections. Current opinion in gastroenterology, 2005; 21: 44-50.
- Anukam KC, Osazuwa EO, Ahonkhai I & Reid G. Lactobacillus vaginal microbiota of women attending a reproductive health care service in Benin city, Nigeria. Sexually Transmitted Diseases, 2006; 33: 59-62,
Crossref - Pavlova SI et al. Genetic diversity of vaginal lactobacilli from women in different countries based on 16S rRNA gene sequences. Journal of Applied Microbiology, 2002; 92: 451-459,
Crossref - Pendharkar S et al. Identification and characterisation of vaginal lactobacilli from South African women. BMC Infectious Diseases, 2013; 13: 43.
Crossref - Zhu Y, Michelle Luo T, Jobin C & Young HA. Gut microbiota and probiotics in colon tumorigenesis. Cancer Letters, 2011; 309: 119-127.
Crossref - Zhong L, Zhang X & Covasa M. Emerging roles of lactic acid bacteria in protection against colorectal cancer. World Journal of Gastroenterology, 2014; 20: 7878-7886.
Crossref - Hassan M, Watari H, AbuAlmaaty A, Ohba Y & Sakuragi N. Apoptosis and molecular targeting therapy in cancer. BioMed Research International, 2014; 2014: 150845.
Crossref - Awaisheh S et al. In vitro cytotoxic activity of probiotic bacterial cell extracts against Caco-2 and HRT-18 colorectal cancer cells. Milchwissenschaft, 2016; 69: 27-31.
- Man J, Rogosa MA & Sharpe M. A Medium for the Cultivation of Lactobacilli. Journal of Applied Microbiology, 2008; 23: 130-135,
Crossref - Antibacterial Activity of Weissella confusa Isolated From Vaginal Swab of Indian Women. International Journal of Advances in Chemical Engineering and Biological Sciences, 2017; 4,
Crossref - Turner S, Pryer KM, Miao VPW & Palmer JD. Investigating Deep Phylogenetic Relationships among Cyanobacteria and Plastids by Small Subunit rRNA Sequence Analysis1. Journal of Eukaryotic Microbiology, 1999; 46: 327-338.
Crossref - Thompson JD, Gibson TJ & Higgins DG. Multiple sequence alignment using ClustalW and ClustalX. Current Protocols in bioinformatics, 2002; Chapter 2, Unit 2 3,
Crossref - Tamura K, Nei M & Kumar S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proceedings of the National Academy of Sciences of the United States of America, 2004; 101: 11030-11035,
Crossref - Shokryazdan P et al. Probiotic potential of Lactobacillus strains with antimicrobial activity against some human pathogenic strains. BioMed Research International, 2014; 2014, 927268,
Crossref - Vinderola G et al. Usefuless of a set of simple in vitro tests for the screening and identification of probiotic candidate strains for dairy use. LWT – Food Science and Technology, 2008; 41: 1678-1688.
Crossref - Jalilsood T et al. Inhibition of pathogenic and spoilage bacteria by a novel biofilm-forming Lactobacillus isolate: a potential host for the expression of heterologous proteins. Microbial Cell Factories, 2015; 14: 96,
Crossref - Martin R et al. Characterization of indigenous vaginal lactobacilli from healthy women as probiotic candidates. International microbiology: The Official Journal of the Spanish Society for Microbiology, 2008; 11: 261-266.
- Chew SY, Cheah YK, Seow HF, Sandai D & Than LT. Probiotic Lactobacillus rhamnosus GR-1 and Lactobacillus reuteri RC-14 exhibit strong antifungal effects against vulvovaginal candidiasis-causing Candida glabrata isolates. Journal of Applied Microbiology, 2015; 118: 1180-1190
Crossref - Mosmann T. Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. Journal of Immunological Methods, 1983; 65: 55-63.
Crossref - Das Purkayastha S et al. Contrasting diversity of vaginal lactobacilli among the females of Northeast India. BMC Microbiology, 2019; 19: 198.
Crossref - Hotel A. Health and Nutritional Properties of Probiotics in Food Including Powder Milk with Live Lactic Acid Bacteria – Joint FAO/WHO Expert Consultation, 2001 2014.
- Dunne C et al. Probiotics: from myth to reality. Demonstration of functionality in animal models of disease and in human clinical trials. Antonie van Leeuwenhoek, 1999; 76: 279-292.
Crossref - Saavedra JM, Bauman NA, Oung I, Perman JA & Yolken RH. Feeding of Bifidobacterium bifidum and Streptococcus thermophilus to infants in hospital for prevention of diarrhoea and shedding of rotavirus. Lancet, 1994; 344: 1046-1049.
Crossref - Montrose DC & Floch MH. Probiotics used in human studies. Journal of Clinical Gastroenterology, 2005; 39: 469-484.
Crossref - Joshi V, Sharma S & Ran N. Production, Purification, Stability and Efficacy of Bacteriocin from Isolates of Natural Lactic Acid Fermentation of Vegetables. Food Technology and Biotechnology, 2006; 44.
- Tokatli M, Gulgor G, Bagder Elmaci S, Arslankoz Isleyen N & Ozcelik F. In Vitro Properties of Potential Probiotic Indigenous Lactic Acid Bacteria Originating from Traditional Pickles. BioMed Research International, 2015; 2015, 315819.
Crossref - Argyri AA et al. Selection of potential probiotic lactic acid bacteria from fermented olives by in vitro tests. Food Microbiology, 2013; 33: 282-291.
Crossref - Bevilacqua A, Corbo MR & Sinigaglia M. In vitro evaluation of the antimicrobial activity of eugenol, limonene, and citrus extract against bacteria and yeasts, representative of the spoiling microflora of fruit juices. Journal of Food Protection, 2010; 73: 888-894.
Crossref - Church DL, Bryant RD, Sim V & Laishley EJ. Metronidazole Susceptibility and the Presence of Hydrogenase in Pathogenic Bacteria. Anaerobe, 1996; 2: 147-153.
Crossref - Bautista-Gallego J et al. Screening of lactic acid bacteria isolated from fermented table olives with probiotic potential. Food Research International, 2013
Crossref - Ramos CL, Thorsen L, Schwan RF & Jespersen L. Strain-specific probiotics properties of Lactobacillus fermentum, Lactobacillus plantarum and Lactobacillus brevis isolates from Brazilian food products. Food Microbiology, 2013; 36: 22-29.
Crossref - Maqsood M et al. In vitro anticancer activities of Withania coagulans against HeLa, MCF-7, RD, RG2, and INS-1 cancer cells and phytochemical analysis. Integrative Medicine Research, 2018; 7: 184-191.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.