ISSN: 0973-7510
E-ISSN: 2581-690X
Salmonella is the leading cause of foodborne disease and a public health concern. In this study, food samples were collected from fresh markets within northeastern Thailand. Antibiotic-resistant Salmonella were detected in fresh vegetables and fermented meat products. Of 172 collected food samples, 25 (14.53%) were contaminated with Salmonella resistant to at least 1 of 9 antibiotics. Eight strains of 7 serovars, including S. Derby, S. Paratyphi B, S. Stanley, S. Dabou, S. Rissen, S. Hvittingfoss, and S. Brenderup, were resistant to S, TE, CN, SXT, and NA, accounting for 20%, 16%, 12%, 12%, and 12% of all antibiotics, respectively. To determine the anti-Salmonella activity of lactic acid bacteria, Pediococcus pentosaceus PS34 isolated from fermented fish was selected from 5 strains of food-derived lactic acid bacteria according to the antibacterial and acid-resistance properties. When added to 3 log CFU/ml cultures of Salmonella, the PS34 cell-free supernatant (CFS) reduced the numbers of Salmonella after 1, 15, 30, and 60 minutes of incubation; the reduction reached 99.9% within 30 minutes. The minimal arbitrary inhibition concentration of freeze-dried PS34 CFS was a 1:4 dilution. With the exception of neutralization to pH 6.5, treatment with proteolytic enzymes and heating at 100°C and 120°C did not reduce the freeze-dried PS34 CFS anti-Salmonella activity, indicating that the extracellular PS34 bioactive substances were heat stable and highly effective against Salmonella in vitro. Thus, bioactive substances produced by P. pentosaceus PS34 have the potential to reduce the risk of antibiotic-resistant Salmonella from contaminated food reaching consumers.
Antibiotic-resistant Salmonella, Pediococcus pentosaceus, bioactive substance, fresh vegetable, fermented food.
Salmonella is a leading cause of foodborne disease and a public health concern worldwide. In humans and animals, Salmonella infection causes high morbidity and mortality rates as well as large economic losses1. The source of infection is mainly cross-contamination from animal faces and affects a variety of foods, such as poultry, pork, beef, fish, seafood, eggs, milk, cheese, fresh products, and fruit2-4. Salmonella is becoming increasingly resistant to various antibiotics via resistance genes against 3rd– and 4th-generation beta-lactams5, which is becoming a serious problem in Thailand and many countries6. The inappropriate uses of antibiotics in animal production is the main reason for antimicrobial resistance in Salmonella, increasing the virulence and invasiveness of strains, making the treatment of the associated infections increasingly difficult and leading to prolonged illnesses7,8. In Thailand, Salmonella is present in food products at high levels due to improper hygiene practices during handling and food processing9. However, there is still limited information on the incidence of antibiotic-resistant Salmonella in food products10.
In recent years, people have begun to demand natural foods without chemical preservatives11 due to health concerns regarding chemical food additives12. There is increasing interest in the use of natural antimicrobials for food protection and safety that can control both spoilage and pathogenic microorganisms12,13. Thus, the use of natural antimicrobials derived from probiotics has become appealing14. Many probiotics are lactic acid bacteria (LAB) mainly comprised of Bifidobacterium and Lactobacillus15. Other species, such as Pediococcus acidilactici and Pediococcus pentosaceus16, also have probiotic properties. In addition to organic acids, probiotics produce a variety of antimicrobial compounds, such as hydrogen peroxide, carbon dioxide, diacetyl, acetaldehyde, D-isomer amino acids, biosurfactants, bacteriocin, and reuterin13.
In a previous study, several LAB strains were isolated from various traditional Thai fermented foods and Mekong freshwater fish. Among these, P. pentosaceus PS34 showed the greatest potential probiotic properties, producing many antimicrobial peptides, proteolytic and lipolytic enzymes and several health-benefit-associated substances. These strains also inhibited the growth of the standard S. Typhimurium DMST 562 (ATCC13311) strain17. Thus, the analysis of the ability of potential LAB strains to inhibit the spread of antibiotic-resistant Salmonella serovars without causing toxicity to food and the environment has gained our attention. The aims of this study were to determine the prevalence of antibiotic-resistant Salmonella in fermented foods and vegetables and to evaluate the antimicrobial activity of the selected probiotic, P. pentosaceus PS34, against antibiotic-resistant Salmonella isolated from food.
Bacterial strains
Salmonella Typhimurium DMST562 is a standard strain. The antibiotic-resistant Salmonella isolates used in this study originated from fermented foods and vegetables. Five LAB strains of Pediococcus pentosaceus PS34, NP6, BD33, CF32 and SW5, were isolated from fermented fish, fermented pork, Mae Kong river fish18.
Collection of food samples
A total of 172 food samples, including 100 vegetables and 72 Thai fermented foods (pla-som and nham), were purchased from fresh markets in Khon Kaen, Kalasin, Roiet and Mahasarakham provinces, Northeastern Thailand. After collection, the samples were tightly sealed and kept in plastic bags and chilled at 5-8°C in a delivery cooler box and analyzed in the laboratory within 24 h.
Isolation and identification of Salmonella
The isolation and identification of Salmonella was performed according to ISO 6579:2002. Briefly, 25 g of each food sample was added to 225 ml of buffered peptone water (BPW; Oxoid, Basingstoke, Hampshire, UK) and mixed using a stomacher for 2 minutes. The samples were pre-enriched at 37°C for 18 ± 2 h. The cultures obtained from the pre-enrichment were subcultured in two selective enrichment broths: Rappaport-Vassiliadis medium with soy and a Müllere-Kauffmann tetrathionate broth (Oxoid), and incubated for 24 ± 3 h at 41.5 ± 0.5°C and 37 ± 1°C, respectively. The enriched cultures were streaked on the XLD and Hextoen (Oxoid) selective plating media and incubated at 37°C for 24 h. Salmonella colonies were confirmed by biochemical tests using triple iron agar (TSI), motile indole lysine agar (MIL), urea agar and indole tests. The presence of the invasion-associated gene iagA in presumptive colonies displaying typical Salmonella biochemical properties was confirmed by an iQ-Check Salmonella II real-time PCR detection kit (Bio-Rad) according to the manufacturer’s instructions. All Salmonella isolates were stored at -80°C in 20% glycerol until further examination. Presumptive Salmonella colonies were further confirmed by serological tests. The serological somatic O and H antigen tests for Salmonella were determined by the Kauffmann-White method19 using commercial antisera (S&A, Thailand).
Screening of antibiotic-resistant Salmonella by disc diffusion
The antibiotic susceptibility was determined on Muller-Hinton agar using the standard disc diffusion method, according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI; Clinical and Laboratory Standards Institute, 2012). A total of 172 Salmonella isolates were screened with nine antibiotic discs (Oxoid, UK), including gentamycin 10 µg (CN), tetracycline 30 µg (TE), ciprofloxacin 5 µg (CIP), sulfamethoxazole-trimethoprim 25 µg (SXT), nalidixic acid 30 µg (NA), sulfonamides 300 µg (S), cephalothin 30 µg (KF), neomycin 30 µg (N), and nitrofurantoin 300 µg (F). The plates were incubated overnight at 37°C and 30°C. The diameters of the inhibition zones were measured and compared with CLSI standards (CLSI, 2017). Quality control tests were performed using E. coli ATCC 25922 and S. aureus ATCC 259223 as standard strains. The experiment was carried out in duplicate.
In vitro antimicrobial activity of P. pentosaceus BD33, CF32, NP6, PS34 and SW5 LAB crude cell-free supernatant (CFS) against antibiotic-resistant Salmonella
Preparation of crude CFS
One loop full of fresh culture of each of the 5 LAB strains was cultured in 10 ml of MRS agar at 35°C for 24 h and subcultured with 1% of LAB culture in 100 ml of MRS at 35°C for 24 h. After incubation, the CFS filtrate was obtained by centrifugation at 12,000 x g for 15 minutes. The supernatant pH was measured, and the supernatant was sterilized by filtration through a 0.2 µm pore size filter. The antimicrobial activity of the untreated CFS (pH 4.0) and CFS (pH 6.5) neutralized with 1N NaOH was tested.
Antimicrobial activity determination by agar well diffusion
The antimicrobial activity of CFS derived from P. pentosaceus BD33, CF32, NP6, PS34 and SW5 was tested against 8 Salmonella isolates representing 7 antibiotic-resistant serovars, including S. Paratyphi B, S. Hvittingfoss, S. Stanley, S. Brenderup, S. Rissen, S. Dabou, and S. Derby. A 24 h culture grown in LB at 37°C was adjusted to 0.5 McFarland standards, swabbed on LB agar plates with a sterile cotton swab, allowed to dry, and a hole was punched in the agar with a sterile 6 mm cork-borer. Each well was filled with 60 µl of CFS obtained from the P. pentosaceus isolates. After incubation at 37°C for 24, 36 and 48 h, the inhibition zones around the well were measured. A 0.8% lactic acid solution and MRS broth were used as positive and negative controls, respectively. The results were considered positive if the diameter of the inhibition zone was greater than 10 mm. The experiment was carried out in triplicate.
Antimicrobial activity determination by broth method
Eight strains of antibiotic-resistant Salmonella were grown in LB broth at 37°C for 24 h. Serial dilutions were performed in LB broth to obtain approximate cell concentrations at 3 Log CFU/ml, and 5 ml of CFS was added. After incubation for 1, 15, 30, and 60 minutes, samples were taken, and Salmonella survival was assessed by plating the samples on LB agar plates and counting the colonies after incubation at 37°C for 24 h. S. Typhimurium DMST562 and sterilized distilled water were used as positive and negative controls, respectively. The number of surviving Salmonella was calculated as follows:
% Reduction = (1-10-L) x 100, L= Log reduction
% Survival = 100 – (1-10-L) x 100
In vitro antimicrobial activity of lyophilized and concentrated PS34 CFS against antibiotic-resistant Salmonella
One loop full of fresh culture of the 8 LAB strains was cultured in 10 ml of MRS at 35°C for 24 h and subcultured with 1% (v/v) of LAB culture in 100 ml MRS at 35°C for 24 h. The crude P34 CFS was concentrated and preserved by the freeze-drying method. The freeze-dried CFS was resuspended in 5 ml of sterile distilled water and filtered through a 0.2 µm cellulose acetate filter. The optimal inhibitory activity of the CFS against Salmonella was assessed. The CFS was serially diluted to 1:1, 1:2, 1:4, 1:8, 1:16, 1:32 and 1:64 ratios in MRS broth, and 10 µl of each was dropped on an LB agar. The inhibition zone was measured. The experiment was carried out in triplicate.
Effects of proteolytic enzymes, pH and temperature on the antimicrobial activity of the CFS
The lyophilized CFS was treated with different conditions as follows: heat by boiling at 100°C for 30 minutes and autoclaving at 121°C for 15 minutes; treated with trypsin, pepsin, proteinase K at a final concentration of 1 mg/ml, and the pH was adjusted to 6.5 using 1 M NaOH. MRS broth was used as the negative control. Then, the antimicrobial activity of the treated CFS was tested against Salmonella by drop diffusion as described above.
Prevalence and serotyping of antibiotic-resistant Salmonella isolated from vegetables and fermented foods
Salmonella were detected in 25 of all 172 samples (14.53%). The isolates were derived from 16 of 100 vegetables (16%) and 9 of 72 fermented meat samples (12.5%). All Salmonella isolates expressed the invasion-associated gene iagA, as assessed by an iQ-Check Salmonella II real-time PCR detection kit. In this study, 25 Salmonella isolates were classified into 6 serogroups and 15 serovars. The type and numbers of the serogroups were as follows: B-3, C-8, E-1, I-1, X-1 and Y-1, in which serogroup C (32%) was the most abundant. Antibiotic-resistant Salmonella were detected in 64%, 20%, and 16% of vegetable, fermented fish, and fermented pork samples, respectively (Table 1).
Table (1):
Distribution of Salmonella serovars in vegetables and fermented pork and fish.
| Serogroup (Total) | Serovar | Vegetable | Fermented pork | Fermented fish | Total serotypes |
|---|---|---|---|---|---|
| B (n=3) | S. Derby | 1 | 0 | 0 | 1 |
| S. Paratyphi B | 1 | 0 | 0 | 1 | |
| S. Stanley | 0 | 2 | 1 | 3 | |
| C (n=8) | S. Brenderup | 0 | 1 | 0 | 1 |
| S. Brunei | 3 | 0 | 1 | 4 | |
| S. Choleraesuis | 1 | 0 | 0 | 1 | |
| S. Dabou | 1 | 0 | 0 | 1 | |
| S. Newport | 1 | 0 | 0 | 1 | |
| S. Rissen | 1 | 1 | 0 | 2 | |
| S. Tennessee | 0 | 0 | 2 | 2 | |
| S. Virchow | 1 | 0 | 0 | 1 | |
| E (n=1) | S. Weltevreden | 1 | 0 | 1 | 2 |
| I (n=1) | S. Hvittingfoss | 3 | 0 | 0 | 3 |
| X (n=1) | S. Bergen | 1 | 0 | 0 | 1 |
| Y (n=1) | S. Djakarta | 1 | 0 | 0 | 1 |
| Total positive samples | 16 | 4 | 5 | 25 | |
| Percentage | 64 | 16 | 20 | 100 | |
Antimicrobial susceptibility of antibiotic-resistant Salmonella isolates
In total, the antimicrobial susceptibility of 25 isolates covering 15 serovars was tested against 9 antimicrobial agents. Resistant Salmonella isolates were categorized into 8 serovars (S. Brenderup S. Dabou, S. Derby, S. Hvittingfoss, S. Paratyphi B, S. Rissen, S. Stanley and S. Tennessee). These resistant Salmonella isolates were resistant to at least one antibiotic. The percentages of strains resistant to sulfonamides (300 µg), tetracycline (30 µg), gentamicin (30 µg), nalidixic acid (30 µg), and sulfamethoxazole (25 µg) were 20%, 16%, 12%, 12%, and 12%, respectively. None of the isolates were found to be resistant to ciprofloxacin (50 µg), cephalothin (30 µg), neomycin (30 µg) or nitrofurantoin (300 µg), as shown in Tables 2 & 3.
Table (2):
Antibiotic resistance profiles of Salmonella isolated from vegetables and fermented meats.
Salmonella |
Food type |
Food origin |
Antibiotic-resistance profiles |
|---|---|---|---|
S. Paratyphi B |
Vegetable |
Lettuce |
CN |
S. Dabou |
Vegetable |
Parsley |
NA |
S. Hvittingfoss |
Vegetable |
Water cress |
NA, CN |
S. Rissen |
Vegetable |
Lettuce |
TE, S, SXT |
S. Derby |
Vegetable |
Coriander |
NA, TE, S, SXT |
S. Brenderup |
Fermented pork |
Nham |
S |
S. Stanley (P) |
Fermented pork |
Nham |
TE, CN |
S. Stanley (F) |
Fermented fish |
Pla-som |
TE, S, SXT |
Table (3):
Antibiotic-resistance percentages of Salmonella isolated from vegetables and fermented meats.
| Antibiotics (µg) | No. of Salmonella isolates (%) (n=25) | |||
|---|---|---|---|---|
| S | I | R | ||
| Cephalothin (30) | KF | 25 (100) | 0 (0) | 0 (0) |
| Ciprofloxacin (5) | CIP | 25 (100) | 0 (0) | 0 (0) |
| Gentamycin (10) | CN | 22 (88) | 0 (0) | 3 (12) |
| Nalidixic acid (30) | NA | 20 (80) | 2 (0) | 3 (12) |
| Neomycin (30) | N | 0 (0) | 25 (100) | 0 (0) |
| Nitrofurantoin (300) | F | 25 (100) | 0 (0) | 0 (0) |
| Sulfonamides (300) | S | 17 (68) | 3 (12) | 5 (20) |
| Sulfamethoxazole-Trimethoprim (25) | SXT | 22 (88) | 0 (0) | 3 (12) |
| Tetracycline (30) | TE | 20 (80) | 1 (4) | 4 (16) |
S: Susceptible; I: Intermediate; R: Resistant
Antimicrobial activity of LAB strains against antibiotic-resistant Salmonella
Due to their potential probiotic properties, including susceptibility to many antibiotics, inhibition of Gram positive and negative bacterial pathogens, and tolerance to low pH in the presence of inulin (a prebiotic)17,18, the antimicrobial activity of crude P. pentosaceus BD33, CF32, NP6, PS34 and SW5 CFS was investigated. The CFS of all P. pentosaceus strains was prepared as untreated (pH 4) and neutralized (pH 6.5) and tested against Salmonella using the agar well diffusion method. The untreated CFS of all strains showed an inhibition zone of approximately 12.3 mm against S. Typhimurium after 24 h of incubation (Fig. 1). However, no inhibition zone was detected for neutralized CFS (pH 6.5). PS34 exhibited the greatest tolerance in the presence of inulin to pH 2 after 24 h18, therefore PS34 CFS samples were tested against 8 isolates of antibiotic-resistant Salmonella and S. Typhimurium DMST562. The results showed that after prolonged incubation for 36 and 48 h, the inhibition zone increased slightly (Table 4). The 0.8% lactic acid control showed an inhibition zone of 11.6 mm, and the zone size was not altered after prolonged incubation. Generally, LAB produce the greatest amount of lactic acid at 24 h incubation. Thus, the increase in the inhibition zone after 24 h was derived possibly from other factors, such as bioactive substances.
Table (4):
The growth inhibition of 8 isolates of antibiotic-resistant Salmonella and S. Typhimurium DMST562 by crude PS34 CFS (pH 4) with 0.8% lactic acid and MRS broth used as positive and negative controls.
| Strains | Inhibition zone (mm) | ||||
|---|---|---|---|---|---|
| 24 h | 36 h | 48 h | 0.8% lactic acid | MRS | |
| Antibiotic-resistant Salmonella S. Typhimurium DMST562 |
12.5 | 12.7 | 13.5 | 11.6 | – |
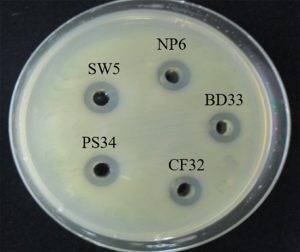
Fig. 1. Inhibitory effect of untreated CFS from overnight cultures of 5 strains of P. pentosaceus in MRS broth at pH 4.0 on S. Typhimurium DMST562.
Antimicrobial activity of crude PS34 CFS when mixed with Salmonella cultures for 1, 15, 30, and 60 minutes
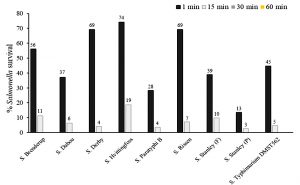
PS34 CFS reduced the number of Salmonella cells during the first one minute incubation by as much as 87% (survived 13%) for S. Stanley (P) (Fig. 2). The numbers of all tested Salmonella serovars decreased significantly with the survival at only 3-11% after 15 minutes. Incubation with the CFS reduced the numbers of Salmonella by approximately 99.9% within 30 minutes. Finally, the PS34 CFS eliminated all tested Salmonella at 60 minutes, in which the cell numbers were reduced by 99.98%. After 24 h, Salmonella cell survival was not detected, suggesting that Salmonella were completely eliminated.
Fig. 2. Survival of antibiotic-resistant Salmonella serovars after contact with crude P. pentosaceus PS34 CFS for 1, 15, 30 and 60 minutes.
Determination of the antimicrobial activity of freeze-dried P. pentosaceus PS34 CFS by drop diffusion
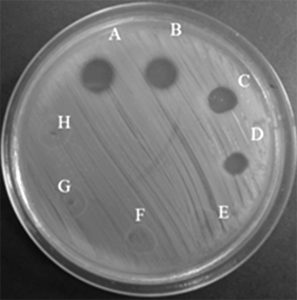
The 2-fold diluted solution of freeze-dried PS34 CFS was mixed with the cultures of the 8 antibiotic-resistant Salmonella isolates, and the inhibition zone was measured. The minimal arbitrary inhibition concentration of all samples was found to be a 1:4 dilution, and the growth inhibition effect was lost beyond a 1:8 dilution (Fig. 3).
Fig. 3.Minimal arbitrary inhibition concentration of freeze-dried PS34 CFS against S. Typhimurium DMST562 (A undiluted, B 1:1, C 1:2, D 1:4, E 1:8, F 1:16, G 1:32, and H 1:64).
Effect of heat, proteolysis, and pH on the anti-Salmonella activity of the freeze-dried PS34 CFS
When tested with all 8 isolates of Salmonella, the anti-Salmonella activity of the PS34 CFS remained but decreased slightly after heating (100°C for 30 minutes and 121°C for 15 minutes) and digestion with proteolytic enzymes (trypsin, pepsin, and proteinase K; Table 5). Only a pH of 7 negatively affected the inhibitory activity. It has been reported that lactic acid from LAB fermentation broth is sensitive to heat at approximately 111°C20. Thus, the inhibitory activity of PS34-CFS at 121°C was likely derived from antimicrobial agents and/or acid.
The resistance of Salmonella to many antibiotics has become a serious health problem worldwide14. In this study, the prevalence of Salmonella in vegetables and traditional Thai fermented pork (nham-mooh) and fermented fish (pla-som fak) sold in the local fresh markets and supermarkets in the northeastern region was 64% in vegetables and 36% in meat/fish. In this study, S. Typhimurium and S. Enteritidis, major infectious serovars in humans, were not detected. Of the Salmonella isolates, 20%, 16%, 12%, 12%, and 12%, were resistant to sulfonamides, tetracycline, gentamicin, nalidixic, and sulfamethoxazole, respectively. Among the antibiotic-resistant isolates, 12.5% and 16% were isolated from fresh vegetables and fermented meats, respectively. In agreement with previous studies, Salmonella from the meat products in retail markets in Thailand were resistant to ampicillin, sulfonamides and tetracycline21.
These results are similar to those in a report from Tunisia (20%)22 but lower than those detected previously in Morocco (75.43%)23, Italy (84.2%)24 and China (74%)25. In this study, 14.7% of Salmonella isolates (5/34) were resistant to two types of antibiotics. These rates are lower than those found in Botswana (20.3%)26 and Egypt (100%)27. The percentage of isolates resistant to tetracycline was 16.0%. Resistance to these antimicrobials is common in Salmonella isolates and has been observed previously in Morocco23, Tunisia22, Malaysia28 and China29. The prevalence of sulfonamide resistance is signi cant (20%), and might be a consequence of the widespread use of these antimicrobials in livestock. This percentage is consistent with that detected in Morocco (20.5%)30 but higher than that found in Tunisia (1.2%)22.
The Salmonella observed in this present study were determined by the detection of invasion-associated (iagA) genes. All positive isolates expressed the iagA gene, indicating that the consumption of raw nham-mooh, pla-som fak and fresh vegetables might lead to an increased risk of antibiotic-resistant Salmonella infection due to the transfer of resistance genes to other bacteria. However, further epidemiological studies and more data regarding the link between contaminated food consumption, human illness and the presence of virulence genes may be necessary to address food safety concerns.
The presence of several Salmonella serovars in foods points to natural cross-contamination in the environment. The prevalence of the predominant Salmonella serotypes in this study might be different from that in other geographical regions due to several factors, such as the environment and temperature30. The cross-contamination of the raw material usually occurs at its point of origin, such as the cross-contamination in fish, which usually occurs via exposure of the fish to various pathogens in aquatic environments31. The cross-contamination of vegetables usually occurs by fecal contamination of the water used for irrigation and food processing31. The low hygienic quality of food processing facilities mainly leads to cross-contamination due to a lack of good handler hygiene practices, such as hand washing during raw material handling, delivery and food processing steps32,33. In addition, techniques used for the isolation of pathogenic bacteria might be different30.
In Thailand, the small-scale production of many traditional fermented Thai meats such as nham-moo and pla-som-fak are processed by indigenous microorganisms, which could be contaminated with pathogens. Traditional Thai fermented food products are commonly produced at home or by small-scale industrial production and sold at small retail markets in upper northeastern Thailand. Hence, contamination with widespread foodborne pathogens, such Salmonella, could easily occur, and approaches to prevent this occurrence should be investigated.
In this study, we sought to determine the potential of natural antimicrobials to control pathogenic microorganism growth for food protection and safety as an alternative choice to current food preservatives12,13. The CFS from potential probiotic bacteria P. pentosaceus PS34 showed strong inhibitory activity against antibiotic-resistant Salmonella strains isolated from fresh vegetables and fermented foods. Our results are consistent with those of a previous study reporting that probiotic LAB inhibit the growth of various Gram-positive and Gram-negative pathogenic bacteria. The use of CFS from L. rhamnosus and L. casei combined with amikacin and gentamicin strongly inhibited the development of a Pseudomonas aeruginosa infection34. Qureshi et al. (2019)35 reported that the use of probiotics combined with a conventional antibiotic regimen appeared has great potential for eliminating Helicobacter pylori infections. However, the use of probiotics with antibiotic-resistant microorganisms could have both beneficial effects and risks. Therefore, the regulation of the use of probiotics in both humans and animals should be appropriately established8.
P. pentosaceus PS34 was isolated from fermented fish and is a potential probiotic strain due to its susceptibility to many antibiotics18. The CFS from P. pentosaceus PS34 may have inhibited the growth of Salmonella through the production of various antimicrobial compounds. LAB normally produce weak organic acids in the medium. Weak organic acids, such as lactic acid, can penetrate through the cell membranes of Gram-negative bacteria, causing intracellular acidification after the release of hydrogen ions36. This acidification leads to the loss of ATP from the cell. The intracellular hydrogen ions diffuse passively out of the bacterial cells across the membrane, resulting in cell damage and cell death37. Bacteriocin is a ribosomally synthesized antimicrobial peptide commonly produced by LAB. Bacteriocin works mainly against Gram-positive bacteria38. Many bacteriocins are heat stable, and the widely studied bacteriocin from LAB has gained increased attention as an alternative means to inhibit pathogens growth in foods39. In this study, the antimicrobial effect of CFS against Salmonella might also involve bacteriocin activity since the inhibition remained after the CFS was heated at high temperatures. Out of the total samples, Salmonella were isolated from 36% of fermented meats, which normally have a low pH, indicating that Salmonella are naturally able to tolerate acid. The inhibitory effect observed in this study was possibly derived from bioactive substances. However, the loss of CFS activity at neutral pH indicates that the PS34-derived bioactive substance favors pH 4.
The high prevalence of antibiotic-resistant Salmonella (at 64%) in fresh vegetables observed in this study is a public safety concern. To minimize the risk of human infection, it is important to continue surveillance and take measures to prevent food and environmental contamination by antimicrobial-resistant bacteria as well as to educate the public that avoiding raw food consumption is important to prevent antimicrobial-resistant bacterial infections. Thus, public health authorities should monitor and provide information and knowledge to customers for safe food consumption. New decontamination methods are needed to guarantee the safety of various foods40-43. Therefore, the regulation of the use of probiotics in both humans and animals should be appropriately established8.
In conclusion, the contamination of vegetables and fermented meats with antibiotic-resistant Salmonella is widespread, indicating the inappropriate use of antimicrobial antibiotics in humans and livestock. Salmonella cross-contaminate the environment, and thus the transference of antibiotic-resistance genes to other bacteria is possible. This study revealed the antimicrobial properties of CFS from P. pentosaceus PS34 originally derived from fermented fish, which was effective against a variety of antibiotic-resistant Salmonella serovars isolated from foods. Thus, PS34 has the potential to inhibit the growth of antimicrobial-resistant Salmonella, indicating that it may be useful in food or in the environment to improve the quality of food products by inhibiting the growth of contaminating antibiotic-resistant Salmonella.
ACKNOWLEDGEMENTS
We would like to express our grateful thanks to Ms. Siriyanapat Tasaku for her assistance with the LAB experiments.
CONFLICTS OF INTEREST
The authors declare that there is no conflict of interest.
AUTHOR CONTRIBUTION
WS and NR designed the experiments. NR performed the experiments. WS and NR analyzed the data and wrote the manuscript. WS, NR and SS read and approved the manuscript.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Scallan, E, Hoekstra RM, Angulo FJ, Tauxe RV, Widdowson MA, Roy SL, Jones RL, Griffin PM. Foodborne illness acquired in the United States-major pathogens. Emerging infectious diseases, 2011; 17(1): 7-15.
Crossref - Bangtrakulnonth A, Pornreongwong S, Pulsrikarn C, Sawanpanyalert P, Hendriksen R S, Lo Fo Wong DMA, Aarestrup FM. Salmonella serovars from humans and other sources in Thailand, 1993–2002. Emerging infectious diseases, 2004; 10(1): 131-136.
Crossref - Gorski L, Parker CT, Liang A, Cooley MB, Jay-Russell MT, Gordus AG, Mandrell R E. Prevalence, distribution, and diversity of Salmonella enterica in a major produce region of California. Appl. Environ. Microbiol., 2011; 77(8): 2734-2748.
Crossref - Andino A, Hanning I. Salmonella enterica: survival, colonization, and virulence differences among serovars. The Sci. World J., 2015; 520179.
Crossref - Maka L, Popowska M. Antimicrobial resistance of Salmonella isolated from food. Roczniki Panstwowego Zakladu Higieny, 2016; 67(4): 343-358.
- Utrarachkij F, Nakajima C, Siripanichgon K, Changkaew K, Thongpanich Y, Pornraungwong S, Suzuki Y. Genetic diversity and antimicrobial resistance pattern of Salmonella enterica serovar Enteritidis clinical isolates in Thailand. J. Infection Chemotherapy, 2016; 22(4): 209-215.
Crossref - Murgia M, Bouchrif B, Timinouni M, Al-Qahtani A, Al-Ahdal MN, Cappuccinelli P, Paglietti B. Antibiotic resistance determinants and genetic analysis of Salmonella enterica isolated from food in Morocco. Int. J. Food Microbiol., 2015; 215: 31-39.
Crossref - Imperial IC, Ibana JA. Addressing the antibiotic resistance problem with probiotics: reducing the risk of its double-edged sword effect. Frontiers in Microbiology, 2016; 7: 1983.
Crossref - Trongjit S, Angkititrakul S, Tuttle RE, Poungseree J, Padungtod P, Chuanchuen R. Prevalence and antimicrobial resistance in Salmonella enterica isolated from broiler chickens, pigs and meat products in Thailand–Cambodia border provinces. Microbiology and Immunology, 2017; 61(1): 23-33.
Crossref - Whistler T, Sapchookul P, McCormick DW, Sangwichian O, Jorakate P, Makprasert S, Supcharassaeng S. Epidemiology and antimicrobial resistance of invasive non-typhoidal Salmonellosis in rural Thailand from 2006-2014. PLoS Neglected Trop. Diseases, 2018; 12(8): e0006718.
Crossref - da Silva Sabo S, Vitolo M, Gonzalez JMD, de Souza Oliveira RP. Overview of Lactobacillus plantarum as a promising bacteriocin producer among lactic acid bacteria. Food Research Int., 2014; 64: 527-536.
Crossref - Saraoui T, Cornet J, Guillouet E, Pilet MF, Chevalier F, Joffraud JJ, Leroi F. Improving simultaneously the quality and safety of cooked and peeled shrimp using a cocktail of bioprotective lactic acid bacteria. Int. J. Food Microbiol, 2017; 241: 69-77.
Crossref - Hartmann HA, Wilke T, Erdmann R. Efficacy of bacteriocin-containing cell-free culture supernatants from lactic acid bacteria to control Listeria monocytogenes in food. Inter. J. Food Microbiol, 2011; 146(2): 192-199.
Crossref - Nami Y, Haghshenas B, Abdullah N, Barzegari A, Radiah D, Rosli R, Khosroushahi AY. Probiotics or antibiotics: future challenges in medicine. J. Medical Microbiology, 2015; 64(2): 137-146.
Crossref - Nami Y, Haghshenas B, Khosroushahi AY. Molecular identification and probiotic potential characterization of lactic acid bacteria isolated from human vaginal microbiota. Adv. Pharm. Bulletin, 2018; 8(4): 683.
Crossref - Martinez MP, Gonzalez Pereyra ML, Pena GA, Poloni V, Fernandez Juri G, Cavaglieri LR. Pediococcus acidilactici and Pediococcus pentosaceus isolated from a rainbow trout ecosystem have probiotic and ABF1 adsorbing/degrading abilities in vitro. Food Additives & Contaminants: Part A, 2017; 34(12), 2118-2130.
Crossref - Tasaku S, Siripornadulsil S, Siripornadulsil W. Inhibitory activity of food-originated P. pentosaceus NP6 against S. enterica serovar Typhimurium in Nile Tilapia by-products. Chiang Mai J. Sci, 2017; 44, 383-393.
- Tasaku, S. Isolation and characterization of lactic acid bacteria for potential use as probiotics in fermented Tilapia belly flap meat. Master of Science Thesis, Khon Kaen University, Khon Kaen, Thailand, 2014.
- Issenhuth-Jeanjean S, Roggentin P, Mikoleit M, Guibourdenche M, de Pinna E, Nair S, Fields PI, Weill FX. Supplement 2008-2010 (no. 48) to the White-Kauffmann-Le Minor scheme. Research Microbiol, 2014; 165: 526-530.
Crossref - Chen L, Zeng A, Dong H, Li Q, Niu C, A novel process for recovery and refining of L-lactic acid from fermentation broth. Bioresource Technology, 2012;112: 280–284.
Crossref - Wannaprasat W, Padungtod P, Chuanchuen R. (2011). Class 1 integrons and virulence genes in Salmonella enterica isolates from pork and humans. Int. J. Antimicrob. Agents, 2011;37: 457–61.
Crossref - Abbassi-Ghozzi I. Jaouani A, Hammami S, Martinez-Urtaza J, Boudabous M, Gtari A. Molecular analysis and antimicrobial resistance of Salmonella isolates recovered from raw meat marketed in the area of “Grand Tunis” Tunisia. Pathol. Biol. 2012; 60: e49-54.
Crossref - Chaiba A, Rhazi Filali F, Chahlaoui A, Soulayman Benchikh S, Zerhouni M. Prevalence and anti-microbial susceptibility of Salmonella isolates from chicken carcasses and giblets in Meknes, Morocco. Afr. J. Microbiol. Res. 2009; 3(5): 215-219.
- Proroga YTR, Capuano F, Carullo MR, La Tela I, Capparelli R, Barco L, Pasquale V. Occurrence and antimicrobial resistance of Salmonella strains from food of animal origin in southern Italy. Folia Microbiol, 2015; 61: 21-27.
Crossref - Yang X, Huang J, Wu Q, Zhang J, Liu S, Guo W, Yu S. Prevalence, antimicrobial resistance and genetic diversity of Salmonella isolated from retail ready-to-eat foods in China. Food Control, 2016; 60: 50-56.
Crossref - Mrema N, Mpuchane S, Gashe BA. Prevalence of Salmonella in raw minced meat, raw fresh sausages and raw burger patties from retail outlets in Gaborone, Botswana, Food Control, 2006; 17(3): 207-212.
Crossref - Sallam KI, Mohammed MA, Hassan MA, Tamura T. Prevalence, molecular identi cation and antimicrobial resistance pro le of Salmonella serovars isolated from retail beef products in Mansoura, Egypt. Food Control, 2014; 38: 209-214.
Crossref - Adzitey F, Rusul G, Huda N. Prevalence and antibiotic resistance of Salmonella serovars in ducks, duck rearing and processing environments in Penang, Malaysia. Food Res. Int. 2012; 45: 947-952.
Crossref - Li Y, Cai Y, Tao J, Kang X, Jiao Y, Guo R, Wang G, Pan Z, Jiao X. Salmonella isolated from the slaughterhouses and correlation with pork contamination in free market, Food Control, 2016;59: 591-600.
Crossref - Ed-dra A, Filali FR, Karraouan B, El Allaoui A, Aboulkacem A, Bouchrif B. Prevalence, molecular and antimicrobial resistance of Salmonella isolated from sausages in Meknes, Morocco. Microbial Pathogenesis, 2017; 105: 340-345.
Crossref - Benhamed S, Guardiola FA, Mars M, Esteban Mֱ. Pathogen bacteria adhesion to skin mucus of fishes. Vet. Microbiol, 2014; 171(1-2): 1-12.
Crossref - Guevremont E, Lamoureux L, Genereux M, Cote C. Irrigation water sources and time intervals as variables on the presence of Campylobacter spp. and Listeria monocytogenes on Romaine lettuce grown in muck Soil. J. Food Protection, 2017; 80(7): 1182-1187.
Crossref - Falardeau J, Johnson RP, Pagotto F, Wang S. Occurrence, characterization, and potential predictors of verotoxigenic Escherichia coli, Listeria monocytogenes, and Salmonella in surface water used for produce irrigation in the Lower Mainland of British Columbia, Canada. PLOS ONE, 2017; 12(9): e0185437.
Crossref - Aminnezhad S, Kermanshahi RK, Ranjbar R. Evaluation of synergistic interactions between cell-free supernatant of Lactobacillus strains and amikacin and genetamicin against Pseudomonas aeruginosa. Jundishapur J. Microbiology, 2015; 8(4): e16592.
Crossref - Qureshi N, Li P, Gu Q. Probiotic therapy in Helicobacter pylori infection: a potential strategy against a serious pathogen? Appl. Microbiol. Biotechnol., 2019; 103(4): 1573-1588.
Crossref - Mariam SH, Zegeye N, Aseffa A, Howe R. Diffusible substances from lactic acid bacterial cultures exert strong inhibitory effects on Listeria monocytogenes and Salmonella enterica serovar Enteritidis in a co-culture model. BMC Microbiology, 2017; 17(1): 35.
Crossref - Kavitake D, Kandasamy S, Devi PB, Shetty PH. Recent developments on encapsulation of lactic acid bacteria as potential starter culture in fermented foods–A review. Food Bioscience, 2018; 21: 34-44.
Crossref - Cotter PD, Hill C, Ross RP. Food microbiology: bacteriocins: developing innate immunity for food. Nature Rev. Microbiology, 2005; 3(10): 777-788.
Crossref - Iglesias MB, Lopez ML, Echeverroa G, Vinas I, Zudaire L, Abadias M. Evaluation of biocontrol capacity of Pseudomonas graminis CPA-7 against foodborne pathogens on fresh-cut pear and its effect on fruit volatile compounds. Food Microbiology, 2018; 76: 226-236.
Crossref - Xu W, Wu C. The impact of pulsed light on decontami-nation, quality, and bacterial attachment of fresh raspberries. Food Microbiology, 2016; 57: 135-143.
Crossref - Gurtler, J. B. Pathogen decontamination of food crop soil: a review. J. food protection, 2017; 80(9): 1461-1470.
Crossref - Pignata C, D’angelo D, Fea E, Gilli G. A review on microbiological decontamination of fresh produce with nonthermal plasma. J. Applied Microbiology, 2017; 122(6): 1438-1455.
Crossref - Tastan O, Pataro G, Donsi F, Ferrari G, Baysal T. Decontamination of fresh-cut cucumber slices by a combination of a modified chitosan coating containing carvacrol nanoemulsions and pulsed light. Int. J. Food Microbiol, 2017; 260: 75-80.
Crossref
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.