ISSN: 0973-7510
E-ISSN: 2581-690X
Hepatitis C virus (HCV) is an important contributor to chronic morbidity and mortality in developing countries. The study’s objective was to determine the genotype distribution and risk factors associated with the transmission of HCV infections in pediatric patients. Rapid screening and confirmation by the enzyme-linked immunosorbent assay (ELISA) were used to analyze 585 pediatric blood specimens hospitalized and visited the outpatient department of the largest tertiary care hospital in Pakistan. Detection and genotyping of HCV RNA were performed using a real-time polymerase chain reaction (RT-PCR). Demographic data and a history of risk factors were gathered through a survey questionnaire. HCV RNA was detected in 323 (72.4%) cases which showed viral load ranging from Log10 IU/mL < 3 to > 6 IU/mL. HCV genotype 3a was detected in 256 (79.3%) cases while type 3b and 1a was observed in 36 (11.1%) and 31 (9.6%) patients, respectively. HCV positivity was significantly associated with the cases from rural areas [p = 0.005; odds ratio (OR) 1.65; 95% CI 1.16-2.23] and also significantly associated with low-income group [p < 0.001; OR 5.75; 95% CI 3.90-8.40]. The primary risk factors associated with HCV transmission in children were family history (p = 0.002), blood transfusion (p = 0.03), surgical procedures (p = 0.02), and history of injections (p = 0.05). HCV genotype 3a is the most common genotype in children. The main risk factors for HCV transmission in children are blood transfusion, surgical procedures, and injection practices by informal health care providers.
Hepatitis C virus, Genotype 3, Risk factors, HCV transmission, Children
Viral hepatitis is a global health issue that recently has acquired considerable recognition and significance, given the enactment of the World Health Organization (WHO) elimination strategies.1 Hepatitis C virus (HCV) infection has emerged as a leading cause of chronic hepatitis, liver cirrhosis, and hepatocellular carcinoma. HCV was categorized as a distinct entity in 1989 and considered discrete hepatitis that gradually develops cirrhosis.2 The virus is not only responsible for liver problems but also lymphadenopathies.3 HCV is a blood-borne infection transmitted through reused needles, contaminated equipment, and blood transfusion, which chronically affects nearly 71 million humans worldwide.1 Until 2010, 40% of blood transfusions had no adequate screening facility for infectious diseases in Pakistan.4 The incidence of HCV infections is comparatively low in economically developed countries.5 Inadequate health care facilities, unsuitable screening facilities, low literacy rate, low socioeconomic status, lack of awareness on prevention and control measures contribute to the higher incidence of HCV in developing countries.6
The prevalence of HCV in pediatric patients has been reported as 3.26 million. Pakistani, Chinese, Indian, and Nigerian children account for more than 50% of the world’s HCV pediatric cases.7 HCV infections are predominantly transmitted to children horizontally; however, the vertical transmission route cannot be excluded.8 Unhygienic syringes, intravenous drug abuse, barbers, sharing of razors, skin piercing needles, and toothbrushes are the primary horizontal transmission routes in Pakistani adults.9 Pediatric transmission is mostly associated with the infected mother and children with congenital disorders who receive blood transfusions.10 The rate of HCV infections is tremendously higher in children suffering from thalassemia-major.11 Breastfeeding has no significant role in the transmission of HCV infection unless the nipples of the infected mother bleed and the blood mix up directly to the infant’s blood.12
HCV infections are prevalent worldwide, highest in Egypt and the WHO Eastern Mediterranean Region.1,13 Developed countries such as Australia, America, and Europe are comparatively less affected due to better facilities and public awareness of transmission sources.14 Nevertheless, in many parts of the world, HCV testing rates and care are still low.15 Pakistan is considered to be among the countries with a higher HCV infection rate. Notably, Northern areas have been reported with the highest HCV infection rate in the tested population.6 At least six major genotypes have been identified worldwide; a remarkable HCV feature in diagnosis, disease complications, treatment plan, and vaccine manufacturing.3
There is no effective HCV vaccine, and only preventive measures could decrease disease incidence.16 Evaluation of HCV infection involves diagnosing HCV in patients by using serological and molecular testing; furthermore, it also aids in evaluating the severity of hepatic diseases. Nowadays, real-time polymerase chain reaction (PCR) is the most widely used diagnostic marker for hepatitis infection with excellent sensitivity.17,18 Limited data is available on the prevalence of HCV infections, genotypic distribution, and associated risk factors in pediatric patients emphasized to carry out the current study.
Specimen collection
The study was conducted on 585 pediatric patients admitted or visited the outpatient department of the largest tertiary care public sector hospital of Lahore, Pakistan, following ethics approval (Ref. No. 1040/1041). Well-trained nurses and phlebotomists collected 3-4 mL of the whole blood in a silica gel vacutainer and transported it to the laboratory for serum separation. Samples were collected consecutively from children under 15 years old.
Data collection
The demographic information was collected as a part of confidential questionnaires following informed patients’ consent. The socioeconomic data was scaled as high, middle, and low.19 History of the potential HCV transmission risk factors recorded from the parents includes family history, invasive medical and dental procedures, blood transfusions, toothbrushes, circumcision, and ear piercing.
Serological diagnosis
A rapid immunochromatography (ICT) technique was used to detect HCV antibodies’ presence using the single-stage HCV test device (AccuQuik™ Test Kit, Cheyenne USA) using the patient’s serum. Screening positive samples were confirmed by the Enzyme-Linked Immunosorbent Assay (ELISA)20 tests (Southern Biotech ELISA Kit, Birmingham USA). Only positive HCV antibody specimens were quantified using the ELISA technique included in the study.
Real-time Polymerase Chain Reaction (RT-PCR)
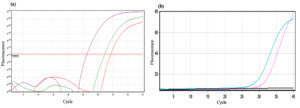
QIAamp DSP Virus Spin Kit (QIAGEN) has been used to extract the genome from HCV ELISA positive samples from silica-membrane technology. HCV RNA amplified and quantitated by the artus® HCV RG RT-PCR Kit (QIAGEN) on Rotor-Gene Q (QIAGEN). All the QIAGEN kits and instruments were purchased from Hilden, Germany. The quantification of HCV IU/mL RNA was evaluated using four standards, and fluorescence signals greater than the cut-off value were found to be positive (Fig. 1a). Internal control was amplified through testing, and each lot included positive and negative HCV RNA control.
Genotyping
RT-PCR positive cases were processed to detect HCV genotype from the extract RNA using AmpliSens® HCV-genotype-FRT (Moscow, Russia). Internal control was used to amplify each sample, and positive and negative controls were included with each tested batch (Fig. 1b).
Fig 1. (a) Negative control and patients’ samples (below threshold). Positive control and HCV RNA positive cases (above threshold) are shown in a higher degree of fluorescence. (b) Fluorescence peak of a positive HCV genotype 1a in a patient sample.
Statistical Analysis
The data was analyzed (SPSS ver. 26.0) using binary logistical regression analysis, and odds ratios (OR) with 95% confidence intervals (CI) were calculated. Variables with a p-value of < 0.10 were further analyzed by multivariate logistic regression analysis to predict independent variables associated with HCV infection. A p-value < 0.05 was considered statistically significant.
There were 585 HCV antibody-positive cases screened using ICT, of which antibodies were detected from 446 (76.2%) cases based on ELISA. Anti HCV ELISA positive cases were treated for final confirmation using RT-PCR, which detected HCV RNA in 323 (72.4%) cases.
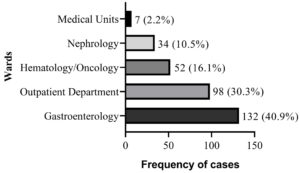
The HCV RT-PCR positive patients’ demographic information showed that 175 (54.1%) males and 148 (45.8%) females were infected. However, no significant association of HCV incidence with the gender of the patients was observed (p = 0.22). Majority of the cases (205, 63.5%) were detected in children > 10 years of age which was statistically insignificant (p = 0.07; OR=1.33 and 95% CI 0.97-1.83). A statistically significant association (p = 0.005) was observed in rural patients compared to urban areas (58.8% vs. 41.1%) with OR 1.65 and 95% CI 1.16-2.23. HCV positive cases were also significantly associated (p < 0.001) with the low-income group with OR 5.75 and 95% CI 3.90-8.40. Results of multivariate analysis also showed significant association of low-income group (p = 0.008; OR=1.41 and 95% CI 1.15-2.72) and rural area (p = 0.02; OR=1.78 and 95% CI 1.09-2.93) with HCV infection (Table 1). The highest numbers of HCV RT-PCR positive cases were 132 (40.9%), isolated from the gastroenterology, and 98 (30.0%) from the outpatient department. Other HCV cases belonged to hematology/ oncology (52; 16.1%) and nephrology (34; 10.5%), while few cases (7; 2.2%) were detected from medical units (Fig. 2).
Table (1):
Demographic characteristics of HCV patients enrolled in the study (n = 585).
| Binary logistic analysis | Multivariate logistic analysis | |||||
|---|---|---|---|---|---|---|
| Demographics | HCV Positive (n = 323) |
HCV Negative (n = 262) |
p-value | OR (95% CI) | p-value | OR (95% CI) |
| Sex | ||||||
| Male | 175 (54.1%) | 156 (59.5%) | 0.22 | 1.24 (0.90-1.3) | ||
| Female | 148 (45.8%) | 106 (40.4%) | ||||
| Age groups (Years) | ||||||
| <10 years | 118 (36.5%) | 109 (41.60%) | 0.07 | 1.33 (0.97-1.83) | 0.57 | 0.97 (0.60-1.34) |
| >10 years | 205 (63.4%) | 153 (57.2%) | ||||
| Residence | ||||||
| Rural area | 190 (58.8%) | 184 (70.2%) | 0.005 | 1.65 (1.16-2.23) | 0.008 | 1.41 (1.15-2.72) |
| Urban area | 133 (41.1%) | 78 (29.7%) | ||||
| Socioeconomic status | ||||||
| Low | 186 (57.5%) | 50 (19.0%) | < 0.001 | 5.75 (3.90-8.40) | 0.02 | 1.78 (1.09-2.93) |
| Middle | 95 (29.4%) | 155 (59.2%) | ||||
| High | 42 (13%) | 57 (21.8%) | ||||
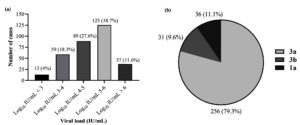
HCV viral load Log10 IU/mL > 5-6 was observed in 125 (38.7%) cases followed by Log10 IU/mL 4-5 in 89 (27.6%), Log10 IU/mL 3-4 in 59 (18.3%) and Log10 IU/mL < 3 in 13 (4%) patients (Fig. 3a). There were 37 (11.5%) patients who had Log10 IU/mL > 6 serum viral load. HCV genotype 3a was detected in 256 (79.3%) cases, 3b in 36 (11.1%) patients while 31 (9.6%) cases had genotype 1a (Fig. 3b).
Fig 3. (a) HCV viral load (IU/mL) among the pediatric HCV confirmed cases (n = 323). (b) Prevalence of HCV genotypes among positive cases.
The analysis of various risk factors showed a significant association of HCV transmission with the family history (p = 0.002), blood transfusion (p = 0.03), surgical procedures (0.02), and history of injections (0.05). The other risk factors in our study include catheterization (p = 0.77), male circumcision (p = 0.1) and female ear piercing (p = 0.12) had no significant association. Multivariate analysis revealed a significant association of family history of HCV infection with the disease (p = 0.01). All other risk factors exhibited a non-significant association (Table 2).
Table (2):
Risk factor associated with the transmission of HCV infection.
| Binary logistic analysis | Multivariate logistic analysis | |||||
|---|---|---|---|---|---|---|
| Risk Factors | HCV Positive (n = 323) |
HCV Negative (n = 262) |
p-value | OR (95% CI) | p-value | OR (95% CI) |
| History of HCV infection in Family | ||||||
| Yes | 25 (7.7%) | 5 (1.9%) | 0.002 | 4.31 (1.6-11.42) | 0.01 | 3.75 (1.25-11.25) |
| No | 298 (92.2%) | 257 (98.9%) | ||||
| Blood transfusion | ||||||
| Yes | 57 (17.6%) | 29 (11.0%) | 0.03 | 1.24 (1.06-2.78) | 0.93 | 1.02 (0.55-1.89) |
| No | 266 (82.3%) | 233 (88.9%) | ||||
| Catheterization | ||||||
| Yes | 30 (9.2%) | 27 (10.3%) | 0.77 | 0.91 (0.51-1.54) | ||
| No | 293 (90.7%) | 235 (89.6%) | ||||
| Surgical procedures | ||||||
| Yes | 217 (64.0%) | 151 (57.6%) | 0.02 | 1.50 (1.02-1.32) | 0.15 | 1.39 (0.88-2.21) |
| No | 106 (35.9%) | 111 (42.3%) | ||||
| Male circumcision | ||||||
| Yes | 133 (41.1%) | 90 (34.3%) | 0.1 | 1.01 (0.95-1.80) | ||
| No | 190 (58.8%) | 172 (65.6%) | ||||
| Female ear piercing | ||||||
| Yes | 35 (10.8%) | 41 (15.6%) | 0.12 | 0.65 (0.40-1.06) | ||
| No | 288 (89.1%) | 221 (84.3%) | ||||
| Injection’s history | ||||||
| Both formal and informal healthcare provider | 153 (44.2%) | 103 (39.3%) | 0.05 | 1.38 (1.01-1.45) | 0.91 | 1.02 (0.60-1.57) |
| Formal health care provider | 170 (55.7%) | 159 (60.6%) | ||||
HCV is a major global health concern, and its paradigm varies from region to region based on its historical and present risk factors. Chronic hepatitis C virus (HCV) infection affects more than 71 million people worldwide, mostly due to its predisposition to hepatitis C fibrosis, cirrhosis, and liver cancer. HCV prevalence is highest in Mongolian children (1.74%), followed by 1.02% HCV affected Pakistani children.7 Pediatric HCV prevalence in Pakistani Punjab has been reported as 1.88%.21 This study represents the prevalence of HCV infection among children, different genotypes, and associated risk factors. Among 323 HCV-positive children, the male gender has a higher which is in (54.1%) in comparison to females (45.9%), which is in accordance with some previously reported studies.6,22 We found more number (63.4%) of HCV-positive pediatric cases in the age group of >10 years that is similar to a work performed in Brazil.23 Age has been argued to be a major factor in HCV studies, with infection more predominant in older people.24
Our study reports 79.3% prevalence of genotype 3a, 11.1% 3b and 9.6% type 1a. HCV type 3a has been reported to be the most prevalent genotype.25,26 Genotype 3 is also frequently reported in adult patients in Pakistan.27 These findings are also supported by two other studies in which genotype 3 was reported as 72.9% and 89%.22,28 However, HCV genotype 1b was reported in Mankidia and Juanga tribes in India.29 We observed that most of the HCV-positive cases belong to rural areas. The contributing factors in HCV transmission were lack of awareness, low literacy rate, socioeconomic barriers, and improper screening facilities. The Pakistani community’s limited knowledge about the sources responsible for HCV transmission is an important reported risk factor.
The primary contributing factor in HCV infection transmission among pediatric patients is blood products’ transfusion.11 Blood transfusion practices remain a challenge in developing countries due to the absence of standardized infrastructures, a permanent energy source, well trained and equipped workers. The crucial factor in implanting a healthy transfusion is to ensure the blood donor’s accurate screening.9 The most significant risk factors among HCV-positive children observed in the current study were surgical intervention and blood transfusion.30,31 This higher rate of HCV transmission is due to the lack of public awareness of the virus’s spread and the use of no-sterilized syringes.9 The use of invasive procedures and blood transfusion remained a substantial HCV transmission source in Cambodia.32
The logistic regression analysis showed a significant association among children with blood transfusion history who had HCV infection compared to the anti-HCV negative children (p = 0.03). Two previous studies from Pakistan agree with the current findings, according to which blood transfusion, unsafe dental practices, and non-sterile procedures are the main factors of HCV acquisition.9,33 More research is needed with more powered sample sizes, diverse ethnicities, and a high number of individuals infected with different HCV genotypes.34 Our study highlights the dire need for intervention programs; awareness campaigns should be appropriately focused. By ensuring safe blood supplies and healthcare-related procedures should be the highest priority. The study’s limitation includes considering cases admitted only in a tertiary care hospital, whereas chronic hepatitis patients are also brought to the district and sub-divisional hospitals. Moreover, this is a single-center study probably with biased selection and data confined to Pakistan and cannot be extrapolated to other countries.
The study’s findings could play a vital role in strengthening public health initiatives. HCV genotype 3a is predominant among pediatric HCV-infected patients. HCV viral load varies from Log10 < 3 to > 6 IU/mL among the infected children. Information on transmission, risk factors, and mortality from HCV infection is of particular importance for rural communities and populations with low socioeconomic status. The assessment of different risk factors, including blood transfusion, surgical procedures, and injection practice from informal healthcare providers, is the key contributing factor in disseminating HCV infections. It is a dire need to implement infection control preventive strategies and treat pediatric patients with child-specific regimens to minimize HCV cases.
ACKNOWLEDGMENTS
The author extends gratitude to all the colleagues who helped in the study.
CONFLICT OF INTEREST
The author declares that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors have made a substantial, direct, and intellectual contribution to the work and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
The study was approved by the Ethics Committee of SAHS, The Children’s Hospital, Lahore, Pakistan (Ref. No. 1040/1041).
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- WHO hepatitis C fact sheets – Hepatitis C. Geneva, Switzerland World Health Organization (WHO). 2020. https://www.who.int/news-room/fact-sheets/detail/hepatitis-c#:~:text=Globally%2C%20an%20estimated% 2071%20million,chronic%20hepatitis%20C%20virus%20infection. Accessed 27 July 2021.
- Wuytack F, Lutje V, Jakobsen JC, et al. Sexual transmission of Hepatitis C Virus infection in a heterosexual population: A systematic review. HRB Open Res. 2018;1:1-17.
Crossref - Attaullah S, Khan S, Ali I. Hepatitis C virus genotypes in Pakistan: a systemic review. Virol J. 2011;8:433.
Crossref - Khan AA, Khan A. The HIV epidemic in Pakistan. J Pak Med Assoc. 2010; 60(4):300-307.
- Lee MH, Yang HI, Yuan Y, L’Italien G, Chen CJ. Epidemiology and natural history of hepatitis C virus infection. World J Gastroenterol. 2014;20(28):9270-9280.
Crossref - Kumar T, Ahmad N, Hayat MK, et al. Prevalence and genotypic distribution of hepatitis C Virus in Peshawar KPK, Pakistan. HAYATI J Biosci. 2017;24(1):22-25.
Crossref - Schmelzer J, Dugan E, Blach S, et al. Global prevalence of hepatitis C virus in children in 2018: a modelling study. Lancet Gastroenterol Hepatol. 2020;5(4):374-392.
Crossref - Indolfi G, Thorne C, El Sayed MH, Giaquinto C, Gonzalez-Peralta RP. The Challenge of Treating Children With Hepatitis C Virus Infection. J Pediatr Gastroenterol Nutr. 2017;64(6):851-854.
Crossref - Waheed Y, Shafi T, Safi SZ, Qadri I. Hepatitis C virus in Pakistan: a systematic review of prevalence, genotypes and risk factors. World J Gastroenterol. 2009;15(45):5647-5653.
Crossref - Umer M, Iqbal M. Hepatitis C virus prevalence and genotype distribution in Pakistan: Comprehensive review of recent data. World J Gastroenterol. 2016;22(4):1684-1700.
Crossref - Schwimmer JB, Balistreri WF. Transmission, natural history, and treatment of hepatitis C virus infection in the pediatric population. Semin Liver Dis. 2000;20(1):37-46.
Crossref - Tovo PA, Calitri C, Scolfaro C, Gabiano C, Garazzino S. Vertically acquired hepatitis C virus infection: Correlates of transmission and disease progression. World J Gastroenterol. 2016;22(4):1382-1392.
Crossref - Lauer GM, Walker BD. Hepatitis C virus infection. N Engl J Med. 2001;345(1):41-52.
Crossref - Puchades Renau L, Berenguer M. Introduction to hepatitis C virus infection: Overview and history of hepatitis C virus therapies. Hemodial Int. 2018;22(S1):S8-S21.
Crossref - Bajis S, Dore GJ, Hajarizadeh B, Cunningham EB, Maher L, Grebely J. Interventions to enhance testing, linkage to care and treatment uptake for hepatitis C virus infection among people who inject drugs: A systematic review. Int J Drug Policy. 2017;47:34-46.
Crossref - Rodrigue JR, Balistreri W, Haber B, et al. Impact of hepatitis C virus infection on children and their caregivers: quality of life, cognitive, and emotional outcomes. J Pediatr Gastroenterol Nutr. 2009;48(3):341-347.
Crossref - Wilkins T, Akhtar M, Gititu E, Jalluri C, Ramirez J. Diagnosis and Management of Hepatitis C. Am Fam Physician. 2015;91(12):835-842.
- Junaid K, Rasool H, Ul Mustafa A, et al. Association of IL28 B and IL10 Polymorphism with HCV Infection and Direct Antiviral Treatment. Ann Clin Lab Sci. 2021;51(4):512-520.
- Sawsan I, Fahmy AF, El Sherbini. Determining simple parameters for social classifications for health research. Bull High Inst Public Health. 1983;13:95-107.
- Olaniyan MF, Azeez MM. Frequency of Anti-HCV, HBsAg, HIV1-p24 Antigen, Acid Fast Bacilli and Plasmodium spp. Herbal Home And Hospital Patients With Elevated TNFα (≥ 50 pg/ml) in Saki, West Nigeria, J Pure Appl Microbiol. 2019;13(2):841-846.
Crossref - Seerat I, Mushtaq H, Rafiq M, Nadir A. Frequency and Associated Risk Factors of Hepatitis B Virus and Hepatitis C Virus Infections in Children at a Hepatitis Prevention and Treatment Clinic in Lahore, Pakistan. Cureus. 2020;12(5):e7926.
Crossref - Baig S, Siddiqi NM, Shaikh IA. Distribution frequency of hepatitis C virus genotypes in patients attending Liaquat University Hospital Jamshoro/Hyderabad. J Postgrad Med Inst. 2014;28(4):367-371.
- Casanova YS, Boeira Tda R, Sisti E, et al. A complete molecular biology assay for hepatitis C virus detection, quantification and genotyping. Rev Soc Bras Med Trop. 2014;47(3):287-294.
Crossref - Mengal MA, Abbas F, Mengal MA, et al. Passive surveillance of anti-hepatitis C virus antibodies in human subjects of four medical units of Balochistan, Pakistan. Int J Agric Biol. 2012;14(4):585-589.
- Nazir N, Jan MR, Ali A, et al. Prevalence of hepatitis-C virus genotypes and potential transmission risks in Malakand Khyber Pakhtunkhwa, Pakistan. Virol J. 2017;14(1):160.
Crossref - Wu N, Rao HY, Yang WB, et al. Impact of hepatitis C virus genotype 3 on liver disease progression in a Chinese national cohort. Chin Med J. 2020;133(3):253-261.
Crossref - Saleha S, Kamal A, Ullah F, Khan N, Mahmood A, Khan S. Prevalence of hepatitis C virus genotypes in district bannu, khyber pakhtunkhwa, pakistan. Hepat Res Treat. 2014;2014:165826.
Crossref - Aziz S, Rajper J, Noorulain W. Treatment outcome of HCV infected paediatric patients and young adults at Karachi, Pakistan. J Ayub Med Coll Abbottabad. 2012;24(3-4):56-58.
- Kar SK, Sabat J, Ho LM, Arora R, Dwibedi B. High Prevalence of Hepatitis C Virus Infection in Primitive Tribes of Eastern India and Associated Sociobehavioral Risks for Transmission: A Retrospective Analysis. Health Equity. 2019;3(1):567-572.
Crossref - Esmat G, Hashem M, El-Raziky M, et al. Risk factors for hepatitis C virus acquisition and predictors of persistence among Egyptian children. Liver Int. 2012;32(3):449-456.
Crossref - Kumar K, Singh RE, Chandrappa N, Hanumanthappa A, Vijayanath V, Anitha M. Seroprevalence of Hepatitis C Virus in Diagnostic Patients and Blood Donors in and Around Davangere. J Pure Appl Microbiol. 2011;5(1):275-279.
- Nouhin J, Iwamoto M, Prak S, et al. Molecular epidemiology of hepatitis C virus in Cambodia during 2016-2017. Sci Rep. 2019;9(1):7314.
Crossref - Jafri W, Jafri N, Yakoob J, et al. Hepatitis B and C: prevalence and risk factors associated with seropositivity among children in Karachi, Pakistan. BMC Infect Dis. 2006;6:101.
Crossref - Younas S, Mukhtar H, Gohar UF, et al. Diagnostic approach to elucidate the efficacy and side effects of direct-acting antivirals in HCV infected patients. J Infect Dev Ctries. 2021;15(10):1489-1496.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.