ISSN: 0973-7510
E-ISSN: 2581-690X
The present study was carried out to detect the association of miR-146a haplotypes polymorphisms with Behcet’s Disease in Iraqi patients, PCR-SSCP technique used in present study, blood was used to DNA extraction, the results show that there was strong association between miR-146a and Behçet’s Disease there were two patterns (A and B), polymorphisms show significant differences (p>0.05) between patients and control where haplotype A was appeared in control with (4% ) while in patients (43%) and haplotype B was appeared in control with (96% ) while in patients (57%). The present study concluded that there was association between miR-146a polymorphisms with Behçet’s Disease, our finding need more investigation to use this polymorphism as early indication of Behçet’s Disease incidence.
miR-146a, PCR-SSCP technique, Haplotypes, Polymorphisms
Behcet’s Disease (BD) is a inflammatory disease of unclear etiology characterized by three associated symptoms including attacks of ocular lesions, oral aphthous ulcers, and genital sores, (Sakane et al., 1999). The complications of disease result from the variation the intensity of relapsing episodes and self-limiting in time. Behcet’s Disease is associated with morbidity and mortality, particularly in males with early age onset (Mendes et al., 2009). Several studies are assumed that the disease may occur in individuals who are genetically susceptible in association with environmental agents. (de Menthon et al., 2009)
miRNAs, are families of non-coding RNAs which is the regulation of gene expression their main functions where they participate in the regulation of immune response pathways and several gene. (Chatzikyriakidou et al., 2012). miRNAs, can mediated the immune disorders and inflammation pathways by regulating their cellular and molecular targets (Chen et al., 2013). One of miRNAs that are related with apoptosis and inflammation processes is miR-146a which have important role in autoimmine diseases such as systemic lupus erythematosus, psoriasis and rheumatoid arthritis. (Zhang et al., 2014, Mohsen, 2018). So the present study was aimed to detect the role of miR-146a in the pathogenesis of Behcet’s Disease (BD).
1- Sample and data collection; about 2 ml of whole blood was collected from patients of Behcet’s Disease in Marjan hospital. All subjects in this study were taken written consent before participation in this study according to ethical approval of Iraq ministry of health, while control collected from healthy.
2- DNA extraction; DNA was extracted from whole blood using Favor gene extraction kit and concentration and purity were detected using nanodrope (Al-Terehi et al., 2016).
3- miR-146a (rs2910164 C-G) primer was (F:5′-GGGTCTTTGCACCATCTCTG-3’’ for the upstream primer and R: 5′-TCCAGTCTTCCAAGCTCTTCA-3′ for downstream.( Vinci,et al.,2013).
4- PCR conditions and size products miR-146a denaturation for 5min at 94°C, then 35 cycles (30 s at 94°C, 30 s at 57.8°C, 30 sat 72°C, and finally 10 min at 72°C). PCR products were determined by electrophoresis pattern in agarose gel (1.5% agarose, 70 V, 20 mA for 45 min) with ethidium bromide staining, the PCR size product were (195) bp for miR-146a.Statics, the results were statically analysis using odd ratio at CI 95% and p value <0.05).
5- SSCP technique, PCR products were denaturation using SSCP dye (EDTA, formamid and bromophynol blue) 1/1V:V in water bath for 5 min at 95°C then its child in ice for 2min.
6- SSCP electrophoresis, the products were electrophoresis as a following About 10 µl of the samples (sample+ dye) were loaded into wells of 8% acrylamide/bis gel containing 7%glycerol, and 1X TBE buffer. In more details; for recipe a 20× 20 0.1 cm gel format. 8 ml of 40% acrylamide/bis (stoke solution 37.5:1) mixed with 8 ml of 5X TBE, 2.8 ml,100%glycerol, then 40 µl TEMED and 400 µl of 10% ammonium per sulfate were added with 20.8 ml of dH2O After gel was casting sample were loaded and Run under the following conditions. Buffer 5.5 X TBE, Buffer temperature 10°C, Runtime 1.5 h and 100V. Then gel was staining using ethedium bromide for 15 min.
7- Haplotype frequency were determination by variety of bands between patients and control.
8- The statics analysis implemented using Qi square and odd ratio at p value <0.05.
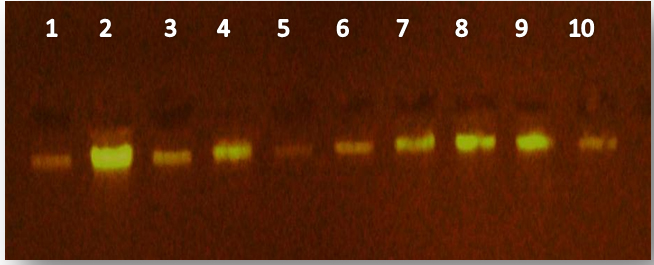
The results of present study show that the DNA has (50-200)ng and purity (1.7-2.2) as show in figure (1).
Fig. 1. Electrophoresis pattern of gnomic DNA in study groups, lane 1-5 DNA from patients lane 6-10 DNA from control
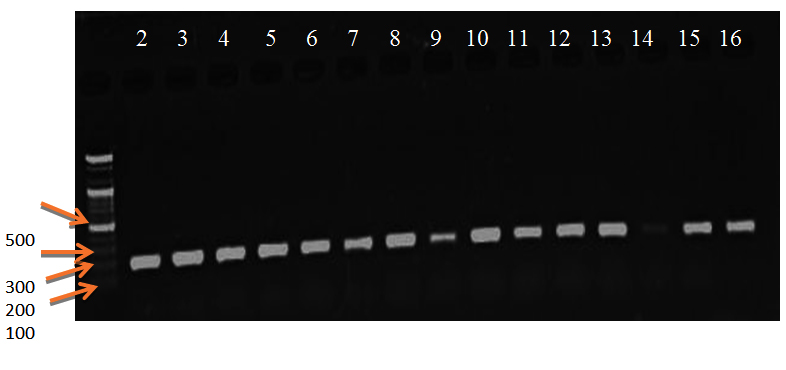
Fig. 2. Electrophoresis pattern of PCR product of miR-146a gene, ane 2-10 for patients, lane 11-16 for control, DNA marker (195 bp)
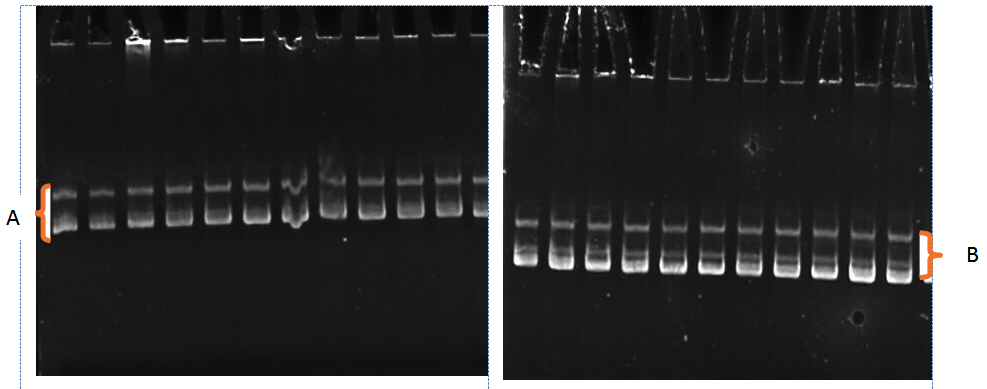
Fig. 3. The electrophoresis pattern of PCR-SSCP technique for MIR-146a gene,
( Pattern A:2 bands) and (Pattern B:3 bands)for both patients and control
Table (1):
The genotype distribution and odd ratio of MIR-146a gene polymorphism for patients and control.
P-value |
CI 95% |
OD ratio |
Control% |
Patients% |
Pattern name |
|---|---|---|---|---|---|
6.1749 – 53.0860 |
18.1053 |
4% |
43% |
A |
|
0.0188 – 0.1619 |
0.0552 |
96% |
57% |
B |
The results of miR-146a (rs2910164) of gene polymorphism which show in Table 1 and Figure 3 clarified the variation of haplotypes in patients and control, there were two patterns (A and B), polymorphisms show significant differences(p>0.05) between patients and control where haplotype A was appeared in control with (4% ) while in patients (43%) and haplotype B was appeared in control with (96% ) while in patients (57%). Behcet’s Disease is classified as an inflammatory disease which is included within autoimmune disease, and there were opinion that microRNAs have role in the regulation autoimmune diseases and participated in their pathogenesis. Several studies indicate that microRNAs can regulate immune response in negative or positive ways where they interplay with inflammation pathways and inhibit it( Boldin and Baltimore, 2012). MIR-146a role is interplay with the activity of nuclear factor kappa B (NF-5ØßB)which is type of proteins found in all cells have roles in controlling DNA transcription, survival of cells and production of cytokines where the NF-5ØßB activation lead to miR-146a gene is activated transcriptionally and thus lead to inhibition of TRAF6 and IRAK1 by miR-146a and cause the expression dampens of NF-5ØßB.( Hajishengallis and Chavakis, 2013) The polymorphisms in miR-146a gene lead to activation of NF-5ØßB as a result of downregulation of miR-146a and this lead to expansion of Th1 clonal and this state correlate with the occurrence of inflammation in autoimmune disease and uveitis that represent essential symptom in behchet disease (Zhou et al., 2014). In a study showed that the frequency of homozygous rs2910164 GG genotype increase in patients with Behcet’s Disease more than the CC cases and GC cases and this associated strongly with increase in the interleukin-17 and tumer necrosis factors which are represents the inflammatory markers in autoimmune disease (Zhou et al.,2014). The study of (Park et al.,2016) found the presence of an association between the functional SNP (rs2910164) of miR-146a with susceptibility to Behcet’s Disease.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
- Sakane T, Takeno M, Suzuki N. Behcet’s disease. N Engl JMed. 1999; 341(17):1284–1291.
- Mendes D, Correia M, Barbedo M, Vaio T, Mota M, Goncalves O, Valente J. Behcet’s disease—a contemporary review. J Autoimmun. 2009; 32(3–4): 178–188.
- de Menthon M1, Lavalley MP, Maldini C, Guillevin L, Mahr A. HLA-B51/B5 and the risk of Behcet’s disease: a systematic review and meta-analysis of case-control genetic association studies. Arthritis Rheum. 2009; 61(10):1287-96.
- Mohsen ,I.H.(2018). Study Polymorphisms of IL-4 Gene Using PCR-SSCP Technique in Iraqi Systemic lupus erthymatous Patients. J. Pharm. Sci. & Res., 2018; 10(3): 613-614.
- Al-Terehi, M; Hasan, AH; Muhammed, HJ; Mohsen , IH ; Al-Saadi, AH; Zaidan H.K and Al-Kiam ,ZH. Polymorphisms of Glutathione-S-Transferase M1 and T1 Genes in Breast Cancer Tissue in Iraqi patients. JCPS, 2016; 9(4), 2911-2914.
- Chatzikyriakidou, A., Voulgari, P. V., Georgiou, I. & Drosos, A. A. miRNAs and related polymorphisms in rheumatoid arthritis susceptibility. Autoimmunity reviews, 2012; 11: 636–641.
- Chen, H. F. et al. Association between miR-146a rs2910164 polymorphism and autoimmune diseases susceptibility: a meta-analysis. Gene, 2013; 521, 259–264.
- Zhang, W. et al. A single-nucleotide polymorphism of miR-146a and psoriasis: an association and functional study. Journal of cellular and molecular medicine 2014; 18: 2225–34.
- Vinci, S.; Gelmini, S.; Mancini, I. ; Malentacchi, F.; Pazzagli, M.; Beltrami, C.; Pinzani, P.; Orlando, C. Genetic and epigenetic factors in regulation of microRNA in colorectal cancers,. Methods 2013; 59: 138–146
- Park R, Lee WJ, Ji JD. Association between the three functional miR-146a single-nucleotide polymorphisms, rs2910164, rs57095329, and rs2431697, and autoimmune disease susceptibility: A meta-analysis. Autoimmunity. 2016; 49(7):451-458.
- Zhou Q, Hou S, Liang L, Li X, Tan X, Wei L, Lei B, Kijlstra A, Yang P. MicroRNA-146a and Ets-1 gene polymorphisms in ocular Behcet’s disease and Vogt-Koyanagi-Harada syndrome. Ann Rheum Dis. 2014; 73(1):170-6.
- Hajishengallis G, Chavakis T. ”Endogenous modulators of inflammatory cell recruitment”. Trends in Immunology. 2013; 34(1): 1–6.
- M. P. Boldin and D. Baltimore, “MicroRNAs, new effectors and regulators of NF-kappaB,” Immunological Reviews, 2012; 246(1): pp. 205–220.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.