ISSN: 0973-7510
E-ISSN: 2581-690X
The aim of this study was to identify and compare the plasmid replicons of clinical uropathogenic Escherichia coli (UPEC) isolates, involving extended spectrum β-lactamase (ESBL)-positive and ESBL-negative, E. coli ST131 and non-ST131 and various ST131 subclones. Plasmid replicon typing on 24 clinical UPEC isolates was carried out using polymerase chain reaction-based replicon typing. A statistical analysis was performed to assess the associations between plasmid replicon types and ESBL carriage, and to evaluate the link between ST131 isolates and high replicon carriage. Eight replicons, I1α, N2, Iγ, X1, FIIS, K, FIA, and FII were detected. The FII was the most common replicon identified here. ESBL-positive E. coli isolates were highly associated with I1α, N2, Iγ, X1, and FIIS replicons, while FIA was present only in ESBL-negative group. ST131 isolates were highly associated with I1α and N2 replicons compared to non-ST131. No link was found between replicon carriage and the number or type of ESBLs in E. coli isolates. The diversity observed in replicon patterns of our clinical E. coli isolates indicates that they might be originated from different sources. The presence of replicons reported previously in animal sources suggests a possible transfer of antimicrobial resistance between animal and human bacterial isolates.
Plasmid, replicon typing, E. coli, ESBL, ST131, H30
Urinary tract infections (UTIs) are commonly caused by Escherichia coli (E. coli),1 and it is shown that uropathogenic E. coli (UPEC) subset are responsible for approximately 80% of UTIs.2 Over the past two decades, the levels of antibiotic resistance and extended spectrum β-lactamases (ESBLs) carriage of UPEC have increased markedly.3,4
ESBLs are enzymes capable of degrading β-lactams antibiotics, rendering these agents inactive.5 ESBL genes have evolved into several hundreds of variants, complicating the fight against ESBL-producing bacterial isolates causing UTIs.6 For instance, blaCTX-M gene has nearly 170 gene variants so far.7 Among CTX-M gene variants, blaCTX-M-15 is the most common and it belongs to the CTX-M-1 phylogroup.8 Plasmids carry the vast majority of ESBL genes, and they are major players in the transfer and dissemination of ESBL-mediated resistance among clinical bacterial strains.9 There has been a link between particular plasmid types and ESBL-encoding genes. For instance, IncFII plasmids frequently carry blaCTX-M-15, while IncK plasmids commonly harbor blaCTX-M-14.10
The worldwide dissemination of the multidrug resistant (MDR) E. coli sequence type 131 (E. coli ST131) clone represents a major challenge to public health.11 E. coli ST131 is often fluoroquinolone (FQ) insusceptible and commonly carry CTX-M-15 ESBL on IncFII plasmids.12,13 ST131 isolates are subdivided into different subclones, and H30 is among the most common ST131 subclone. H30 comprises two subsets, H30R and H30Rx.11
We previously determined the antimicrobial sensitivity, ESBL production, ST131 clonal status, virulence associated gene carriage and metabolic potential of a panel of clinical UPEC isolates.14-16 The identification of plasmid replicon types is important in understanding the epidemiological perspective of prevalence and transmission of ESBL encoding genes in E. coli, and this can help in tracking their origin by comparing them to environmental or animal isolates and in the diagnosis of clinically important ESBL-producing E. coli.17 Given that information on plasmid replicon diversity of clinical E. coli isolates is very scarce in Saudi Arabia, this study sought to describe and compare the replicon diversity of clinical E. coli urine isolates, including ST131 and non-ST131, ESBL-positive and ESBL-negative, and different ST131 subclones. The link between plasmid replicon and ESBL carriage was also evaluated.
Bacterial isolates
This study involved a total of 24 E. coli isolates that obtained from urine specimens of hospitalized in-patients at a tertiary hospital in Riyadh, Saudi Arabia. Details on these E. coli isolates are shown in Table 1.
Table (1):
Information on the E. coli isolates used in this study.
| Isolate ID | MDRa | ESBL | ESBL type (s) | ST131 | ST131 subclone | Reference |
|---|---|---|---|---|---|---|
| U9 | MDR | + | CTX-M-15 | + | H30-nonRx |
14,15 |
| U10 | MDR | + | CTX-M-15 | + | H30-nonRx | |
| U12 | MDR | + | CTX-M-15 | + | H30-nonRx | |
| U16 | MDR | + | CTX-M-15 | – | NAb | |
| U24 | MDR | + | CTX-M-15 | + | H30-Rx | |
| U27 | MDR | + | CTX-M-15 | + | H30-Rx | |
| U46 | MDR | + | CTX-M-15 & OXA | + | H30-nonRx | |
| U57 | MDR | + | CTX-M-15 | + | H30-Rx | |
| U68 | MDR | + | CTX-M-15 | – | NA | |
| U71 | MDR | + | CTX-M-15 | + | Non-H30 | |
| U78 | MDR | + | CTX-M-15 | + | H30-nonRx | |
| U82 | MDR | + | CTX-M-15, OXA & TEM | + | H30-Rx | |
| U3 | Non- MDR | – | NA | – | NA | |
| U6 | Non- MDR | – | NA | – | NA | |
| U19 | Non- MDR | – | NA | + | H30-Rx | |
| U25 | MDR | – | NA | – | NA | |
| U30 | Non- MDR | – | NA | – | NA | |
| U32 | Non- MDR | – | NA | – | NA | |
| U34 | MDR | – | NA | – | NA | |
| U35 | Non- MDR | – | NA | – | NA | |
| U37 | Non- MDR | – | NA | – | NA | |
| U45 | MDR | – | NA | – | NA | |
| U79 | MDR | – | NA | – | NA | |
| U99 | MDR | – | NA | + | Non-H30 |
a MDR phenotype refers to displaying resistance to at least 1 antibiotic in ≥3 antibiotic groups 43.
b NA: Not applicable.
Plasmid replicon typing assays
Plasmid replicon typing was performed by polymerase chain reaction (PCR)-based assay using 2.0 kit (Diatheva, Italy) following the manufacturer’s instructions. Replicon typing was done through eight multiplex PCR reactions that can detect up to 30 replicon types (Table 2). Each reaction mix has 25µl containing 1U DNA polymerase. The setup of the PCR reactions included: 1 denaturation cycle for 10 minutes at 95°C; 25 denaturation cycles for 60 sec at 95°C, annealing for 30 sec at 60°C, and extension for 60 sec at 72°C; and 1 cycle of final extension for 5 minutes at 72°C. PCR products were visualized after being run on 2.5% agarose gel containing ethidium bromide. The assays were performed in triplicate showing fully concordant results.
Table (2):
Information on replicon types provided by PBRT kit.
Multiplex |
Target |
Amplicon size (bp) |
|---|---|---|
M1 |
HI1 |
534 |
HI2 |
298–308 |
|
I1-α |
159 |
|
M2 |
M |
741 |
N |
514 |
|
I2 |
316 |
|
B/O |
159 |
|
M3 |
FIB |
683 |
FIA |
462 |
|
P1 |
345 |
|
W |
242 |
|
M4 |
L |
854 |
X3 |
284 |
|
I1-γ |
161 |
|
M5 |
T |
750 |
A/C |
418 |
|
FIIS |
259–260 |
|
N2 |
177 |
|
M6 |
U |
843 |
X1 |
370 |
|
R |
248 |
|
FIIK |
142–148 |
|
M7 |
FIB KN |
631 |
X2 |
376 |
|
FIB KQ |
258 |
|
K |
190 |
|
M8 |
HIB-M |
570 |
FIB-M |
440 |
|
FII |
288–292 |
|
X4 |
172 |
Statistical analysis
Prism GraphPad (version 9.3.0) was used for statistical analysis. Fisher’s exact test was used for comparisons of various isolate groups, while Mann-Whitney U test was used to determine the mean replicon scores. The threshold for statistical significance was a P-value ≤ 0.05.
Plasmid replicon carriage of all isolates
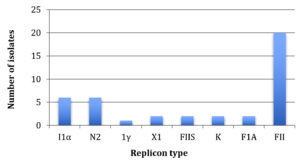
Of the 30 replicons tested in this study, a total of 8 (26.6%) replicons, I1α, N2, 1γ, X1, FIIS, K, FIA and FII, were detected for the 24 E. coli isolates (Table 3). Some isolates, such as U24, U46 and U3, carried only one replicon type, while others, such as U71, U9 and U68, concomitantly harbored 2, 3 and 4 plasmid replicons, respectively. The FII was the most common replicon that was found either solely or concomitantly in 20 (83.3%) of the 24 isolates, however, 1γ was the least common replicon and it was present in a single isolate (4.2%) (Fig. 1). The remaining 6 replicons were variably distributed between isolates (Table 3).
Table (3):
Plasmid replicon types detected for all tested E. coli isolates.
Isolate ID |
Number of replicons |
Replicon type(s) |
|---|---|---|
U9 |
3 |
I1α, N2 & FII |
U10 |
3 |
I1α, N2 & FII |
U12 |
3 |
I1α, N2 & FII |
U16 |
3 |
1γ, X1 & FII |
U24 |
1 |
FII |
U27 |
1 |
FII |
U46 |
1 |
FIIS |
U57 |
1 |
FII |
U68 |
4 |
N2, FIIS, X1 & K |
U71 |
2 |
N2 & FII |
U78 |
3 |
I1α, N2 & FII |
U82 |
1 |
FII |
U3 |
1 |
FII |
U6 |
1 |
FII |
U19 |
2 |
FIA & FII |
U25 |
1 |
I1α |
U30 |
1 |
FII |
U32 |
2 |
K & FII |
U34 |
2 |
FIA & FII |
U35 |
1 |
FII |
U37 |
1 |
FII |
U45 |
1 |
FII |
U79 |
1 |
FII |
U99 |
1 |
I1α |
Comparison of plasmid replicon carriage between ESBL-positive and ESBL-negative isolates
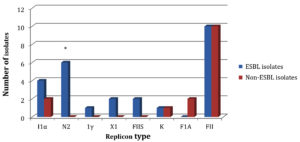
Our plasmid replicon typing results demonstrated a similarity between these isolate groups in their capability of harboring two replicon types: K and FII (Fig. 2). However, some differences in their replicon carriage were identified. Four replicons: N2, 1γ, X1 and FIIS were only carried by ESBL-positive isolates, while ESBL-negative isolates were exclusively carried the FIA replicon (Fig. 2). The ability of ESBL-positive isolates to harbor I1α replicon was more than that of ESBL-negative E. coli isolates. The difference between ESBL-positive and ESBL-negative isolates in carrying the N2 replicon was significant (P = 0.01) (Fig. 2).
Fig. 2. The plasmid replicon types detected for ESBL-producing and non-ESBL-producing isolates. The asterisk refers to presence of significant difference between the two isolate groups for N2 replicon carriage (P = 0.01).
Comparison of plasmid replicon carriage of ST131 and non-ST131 isolates
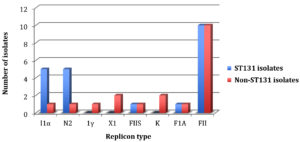
Our data demonstrated a similarity between both isolate groups to carry three replicon types: FIA, FIIS and FII (Fig. 3). Nonetheless, few insignificant differences in their replicon carriage were observed. Non-ST131 isolates exclusively harbored 1γ, X1 and K replicons, with at least one isolate showing capability of carrying these replicons. However, ST131 isolates were more able to carry I1α and N2 compared to non-ST131. No statistical difference in the median replicon carriage score between these two isolate groups was detected (P = 0.60) (Fig. 3).
Comparison of plasmid replicon carriage of E. coli ST131 subclones
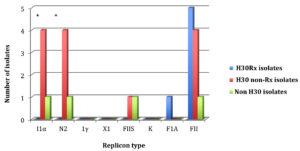
All ST131 isolates were similar in their incapability of carrying three replicons: 1γ, X1 and K, however there was a slight variability between ST131 subclones in carrying five replicons: I1a, FIA, N2, FIIS and FII. In general, the of replicon carriage of H30 isolates were nearly comparable to non-H30 isolates, and there was no specific replicon profile of a particular ST131 subclone (Fig. 4). The replicon carriage of H30 non-Rx subclone was higher than H30Rx, and a significant difference between these subclades in their carriage of I1α and N2 replicons was found (P = 0.05) (Fig. 4). Non-H30 isolates was not significantly associated with a specific replicon type compared to H30 isolates (Fig. 4).
Fig. 4. The plasmid replicon types detected for isolates belonging to ST131 subclones. The asterisks refer to the presence of significant difference between the H30Rx and H30 non-Rx isolate groups for I1a and N2 replicon carriage (P = 0.05).
Relating the plasmid replicon carriage to ESBL carriage
The results of relating the plasmid replicon carriage of the twenty-four E. coli isolates to their ESBL carriage are shown in Table 4. In general, there was no specific relationship between replicon carriage and the number or type of ESBLs carried by E. coli isolates. Our data showed that FII was found in 10 (83.3%) CTX-M-15-producing isolates, however this replicon type was not exclusively harbored by ESBL-positive isolates, as it was also detected in 10 (83.3%) ESBL-negative isolates. Additionally, isolates carrying more than one ESBL type, such as U46 and U82, did not carry more replicons in comparison to those carrying single ESBL type (Table 4).
Table (4):
Relating the plasmid replicon type(s) to ESBL carriage.
Isolate ID |
No. of replicons |
Replicon type(s) |
ESBL type(s) |
|---|---|---|---|
U9 |
3 |
I1α, N2 & FII |
CTX-M-15 |
U10 |
3 |
I1α, N2 & FII |
CTX-M-15 |
U12 |
3 |
I1α, N2 & FII |
CTX-M-15 |
U16 |
3 |
1γ, X1 & FII |
CTX-M-15 |
U24 |
1 |
FII |
CTX-M-15 |
U27 |
1 |
FII |
CTX-M-15 |
U46 |
1 |
FIIS |
CTX-M-15 & OXA |
U57 |
1 |
FII |
CTX-M-15 |
U68 |
4 |
N2, FIIS, X1 & K |
CTX-M-15 |
U71 |
2 |
N2 & FII |
CTX-M-15 |
U78 |
3 |
I1α, N2 & FII |
CTX-M-15 |
U82 |
1 |
FII |
CTX-M-15, OXA & TEM |
U3 |
1 |
FII |
NA |
U6 |
1 |
FII |
NA |
U19 |
2 |
FIA & FII |
NA |
U25 |
1 |
I1α |
NA |
U30 |
1 |
FII |
NA |
U32 |
2 |
K & FII |
NA |
U34 |
2 |
FIA & FII |
NA |
U35 |
1 |
FII |
NA |
U37 |
1 |
FII |
NA |
U45 |
1 |
FII |
NA |
U79 |
1 |
FII |
NA |
U99 |
1 |
I1α |
NA |
Characterizing bacterial plasmids in MDR bacteria, such as ExPEC, is essential in elucidating the role these plasmids play in the global spread of antimicrobial resistance. Several techniques have successfully been used to characterize plasmids with variable degrees of applicability. Among these techniques, PCR-based replicon typing is an easy useful tool with proven specificity and sensitivity.18,19 In Enterobacteriaceae, IncF, IncI, IncA/C, IncL, IncN and IncH are among the most frequently reported plasmids associated with carrying antibacterial resistance genes.20
In our analysis, eight replicons were detected in the 24 isolates, and there was a high degree of plasmid replicon diversity among these isolates. This is common for E. coli isolates and concurs with a previous report demonstrating high diversity in replicon types among UPEC isolates.21 The diversity observed in replicon patterns of our clinical E. coli isolates indicates that they might be originated from different sources. With respect to the distribution of replicons, our data found that IncF plasmids were the most encountered group. FII replicons were found alone in 11 isolates and combined with FIA in two isolates. This is concordant with several reports showing that IncF plasmids are limited to Enterobacteriaceae and mainly present in E. coli,22 and is also in agreement with many reports describing the epidemiology of IncF plasmids in E. coli.23-25 However, it is in contrast to a previous study, showing that FIB and B/O were the most common replicon types among a collection of UPEC isolates.21
We also found four isolates showing the replicon combination I1a, N2 & FII. The presence of such a combination is not uncommon and the cointegration of IncF plasmids with IncI1 and IncN in E. coli was previously reported.26,27
Our study compared the replicon types of ESBL-positive and negative isolates, and demonstrated that K and FII replicons were detected in both isolate groups in similar frequency, and that ESBL-positive isolates were more probable to carry FIIS, I1a, N2, 1γ and X1 isolates in comparison to ESBL-negative isolates. It has been shown that ESBL genes are mostly found on IncF plasmids, that also encode for carbapenemases, aminoglycoside-modifying enzymes, and quinolone resistance,10 which agrees with our findings. However, we showed that FII carriage was comparable between ESBL-positive and negative isolates, suggesting that this replicon is not specific to ESBL-producing E. coli. Given the little focus towards characterizing plasmid replicon carriage of non-ESBL producing E. coli strains in the literature, it is crucial to perform comparative large-scale studies on the epidemiology of IncF plasmid carriage of ESBL-positive and ESBL-negative E. coli isolates.
Our analysis also showed the ability of some isolates to carry replicons that are harbored by E. coli isolates from animal sources. For example, IncIγ, the replicon detected in one isolate in this study, was also reported to carry blaCMY-2 gene in E. coli isolate from animal source.28 Additionally, we detected IncK plasmid in two isolates, and several previous studies reported the presence of IncK plasmids in E. coli isolated from animal sources, mainly carrying blaCMY-2 and blaCTX-M-14 genes.29-32 In this study, N2 replicon was found in six isolates, all of them also have FII replicons. This is in accordance with a previous study reported the colocalization of IncN with IncF plasmids in E. coli.33 Several ESBL genes were carried on IncN plasmids in E. coli isolates including blaCTX-M-1,34 blaCTX-M-14 and blaCTX-M-27.35 IncN2 plasmids were reported in E. coli from human hosts and carried blaNDM-1 gene in Thailand,36 Australia,37 and China.38 IncX1 plasmid was present in two isolates, both are ESBL-producing non-ST131 isolates. IncX1 encoding qnrS1 gene was identified in a previous study in a quinolone-resistant E. coli isolate from animal source.39
Currently, E. coli ST131 is considered as the main contributor for the spread of multidrug resistance and certain genes coding for CTX-M ESBLs on IncFII plasmids. Surprisingly, our data has not found higher plasmid replicon carriage of ST131 compared to non-ST131, although almost all our ST131 isolates were MDR and CTX-M-15 producing. Moreover, FII replicon was detected in similar frequencies in both ST131 and non-ST131 isolates. Despite our sample size is low, this finding is important and highlights the importance of performing comparative studies on different successful ExPEC clones to evaluate the difference in their plasmid replicon carriage.
Two fluoroquinolone-resistance clades are believed to lead the global expansion of ST131. The two clades are H30R (Clade 1), and H30Rx (Clade 2), both have fimH30 allele.40,41 A recent plasmidome meta-analysis-based evidence has indicated that CTX-M-encoding ST131 plasmids are clade-specific, meaning that clonal expansion is the cause of the global expansion of ST131 rather than horizontal gene transfer. The analysis found that IncF plasmids coding for CTX-M-27 are found only in clade 1, whereas IncF plasmids coding for CTX-M-15 are found only in clade 2.42 Our analysis showed that, among the 12 ST131 isolates included in the study, five are H30R, five are H30Rx and two are non-H30. All H30R isolates have identical replicons (I1α, N2 and FII) except one isolate that possessed only FIIS replicon. All H30Rx isolates posses FII replicon except one of them that had both FII and FIA. However, in contrast to the recent evidence, most of these subclones were having CTX-M-15 regardless of the subclone.
With respect to the limitations of this study; it examined a small number of UPEC isolates, therefore the associations observed in the current analysis might not be fully definitive. It also provided a description and a comparison of the replicon diversity of E. coli urine isolates collected from Riyadh city, which does not necessarily reflect the plasmid replicon diversity of E. coli isolates collected from other geographical parts in Saudi Arabia.
This study is the first to identify and compare the plasmid replicon diversity of different UPEC isolate groups from Saudi Arabia. This diversity observed in replicon patterns of our isolates indicates that they might be originated from different sources. The presence of replicons reported previously in animal sources suggests a possible transfer of antibiotic resistance between animal and human bacterial isolates. The lack of specific replicon carriage of the globally disseminated MDR ExPEC clone, ST131, or its subclones was also demonstrated here. In future, studying the plasmid replicon diversity of major MDR E. coli clones is essential to define the role of these plasmids in making MDR ExPEC clones such successful pathogens.
ACKNOWLEDGMENTS
The authors extend their appreciation to the College of Applied Medical Sciences Research Centre at King Saud University for their support of this research.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
This study was approved by the Research Ethics Committee at College of Applied Medical Sciences, King Saud University, Saudi Arabia. (CAMS 042-3839).
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Poolman JT, Wacker M. Extraintestinal pathogenic Escherichia coli, a common human pathogen: challenges for vaccine development and progress in the field. J Infect Dis. 2016;213(1):6-13.
Crossref - Kaper JB, Nataro JP, Mobley HL. Pathogenic Escherichia coli. Nat Rev Microbiol. 2004;2(2):123-140.
Crossref - Foxman B. The epidemiology of urinary tract infection. Nat Rev Urol. 2010;7(12):653-660.
Crossref - Pitout JD, Nordmann P, Laupland KB, Poirel L. Emergence of Enterobacteriaceae producing extended-spectrum β-lactamases (ESBLs) in the community. J Antimicrob Chemother. 2005;56(1):52-59.
Crossref - Perez F, Endimiani A, Hujer KM, Bonomo RA. The continuing challenge of ESBLs. Curr Opin Pharmacol. 2007;7(5):459-469.
Crossref - Brolund A, Sandegren L. Characterization of ESBL disseminating plasmids. Infect Dis. 2016;48(1):18-25.
Crossref - Canton R, Coque TM. The CTX-M β-lactamase pandemic. Curr Opin Microbiol. 2006;9(5):466-475.
Crossref - D’Andrea MM, Arena F, Pallecchi L, Rossolini GM. CTX-M-type β-lactamases: a successful story of antibiotic resistance. Int J Med Microbiol. 2013;303(6-7):305-317.
Crossref - Upadhyay S, Joshi S. TEM mediated extended spectrum cephalosporin resistance in clinical & environmental isolates of Gram negative bacilli: A report from northeast India. Indian J Med Res. 2015;142(5):614-617.
Crossref - Rozwandowicz M, Brouwer M, Fischer J, et al. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J Antimicrob Chemother. 2018;73(5):1121-1137.
Crossref - Nicolas-Chanoine M-H, Bertrand X, Madec J-Y. Escherichia coli ST131, an intriguing clonal group. Clin Microbiol Rev. 2014;27(3):543-574.
Crossref - Peirano G, Pitout JDD. Molecular epidemiology of Escherichia coli producing CTX-M [beta]-lactamases: the worldwide emergence of clone ST131 O25: H4. Int J Antimicrob Agents. 2010;35(4):316-321.
Crossref - Nicolas-Chanoine MH, Blanco J, Leflon-Guibout V, et al. Intercontinental emergence of Escherichia coli clone O25: H4-ST131 producing CTX-M-15. J Antimicrob Chemother. 2008;61(2):273-281.
Crossref - Alqasim A, Abu Jaffal A, Alyousef AA. Prevalence of Multidrug Resistance and Extended-Spectrum β-Lactamase Carriage of Clinical Uropathogenic Escherichia coli Isolates in Riyadh, Saudi Arabia. Int J Microbiol. 2018;2018:3026851.
Crossref - Alqasim A, Jaffal AA, Alyousef AA. Prevalence and molecular characteristics of sequence type 131 clone among clinical uropathogenic Escherichia coli isolates in Riyadh, Saudi Arabia. Saudi J Biol Sci. 2020;27(1):296-302.
Crossref - Alqasim A, Jaffal AA, Almutairi N, Alyousef AA. Comparative phenotypic characterization identifies few differences in the metabolic capacity between Escherichia coli ST131 subclones. Saudi J Biol Sci. 2021;28(1):762-769.
Crossref - Brolund A. Overview of ESBL-producing Enterobacteriaceae from a Nordic perspective. Infect Ecol Epidemiol. 2014;4(1):24555.
Crossref - Carattoli A, Bertini A, Villa L, Falbo V, Hopkins KL, Threlfall EJ. Identification of plasmids by PCR-based replicon typing. J Microbiol Methods. 2005;63(3):219-228.
Crossref - Carloni E, Andreoni F, Omiccioli E, Villa L, Magnani M, Carattoli A. Comparative analysis of the standard PCR-Based Replicon Typing (PBRT) with the commercial PBRT-KIT. Plasmid. 2017;90:10-14.
Crossref - Carattoli A. Resistance plasmid families in Enterobacteriaceae. Antimicrob Agents Chemother. 2009;53(6):2227-2238.
Crossref - Johnson TJ, Wannemuehler YM, Johnson SJ, et al. Plasmid replicon typing of commensal and pathogenic Escherichia coli isolates. Appl Environ Microbiol. 2007;73(6):1976-1983.
Crossref - Awosile BB, Michael A. Genetic Environments of Plasmid-Mediated blaCTXM-15 beta-lactamase gene in Enterobacteriaceae from Africa. Microbiol Res. 2021;12(2):383-394.
Crossref - Carattoli A, García-Fernández A, Varesi P, et al. Molecular epidemiology of Escherichia coli producing extended-spectrum β-lactamases isolated in Rome, Italy. J Clin Microbiol. 2008;46(1):103-108.
Crossref - Hopkins KL, Liebana E, Villa L, Batchelor M, Threlfall EJ, Carattoli A. Replicon typing of plasmids carrying CTX-M or CMY β-lactamases circulating among Salmonella and Escherichia coli isolates. Antimicrob Agents Chemother. 2006;50(9):3203-3206.
Crossref - Coque TM, Novais A, Carattoli A, et al. Dissemination of clonally related Escherichia coli strains expressing extended-spectrum beta-lactamase CTX-M-15. Emerg Infect Dis. 2008;14(2):195-200.
Crossref - Osborn AM, da Silva Tatley FM, Steyn LM, Pickup RW, Saunders JR. Mosaic plasmids and mosaic replicons: evolutionary lessons from the analysis of genetic diversity in IncFII-related repliconsThe EMBL accession numbers for the sequences reported in this paper are AJ009980 (pGSH500 alpha replicon) and AJ009981 (pLV1402 alpha replicon). Microbiology. 2000;146(9):2267-2275.
Crossref - Froehlich B, Parkhill J, Sanders M, Quail MA, Scott JR. The pCoo plasmid of enterotoxigenic Escherichia coli is a mosaic cointegrate. J Bacteriol. 2005;187(18):6509-6516.
Crossref - Hiki M, Usui M, Kojima A, Ozawa M, Ishii Y, Asai T. Diversity of plasmid replicons encoding the blaCMY-2 gene in broad-spectrum cephalosporin-resistant Escherichia coli from livestock animals in Japan. Foodborne Pathog Dis. 2013;10(3):243-249.
Crossref - Valverde A, Canton R, Garcillan-Barcia MP, et al. Spread of blaCTX-M-14 is driven mainly by IncK plasmids disseminated among Escherichia coli phylogroups A, B1, and D in Spain. Antimicrob Agents Chemother. 2009;53(12):5204-5212.
Crossref - Hordijk J, Wagenaar JA, Kant A, et al. Cross-sectional study on prevalence and molecular characteristics of plasmid mediated ESBL/AmpC-producing Escherichia coli isolated from veal calves at slaughter. PLoS One. 2013;8(5):e65681.
Crossref - Stokes MO, Cottell JL, Piddock LJV, et al. Detection and characterization of pCT-like plasmid vectors for blaCTX-M-14 in Escherichia coli isolates from humans, turkeys and cattle in England and Wales. J Antimicrob Chemother. 2012;67(7):1639-1644.
Crossref - Guo Y-F, Zhang W-H, Ren S-Q, et al. IncA/C plasmid-mediated spread of CMY-2 in multidrug-resistant Escherichia coli from food animals in China. PLoS One. 2014;9(5):e96738.
Crossref - Szmolka A, Fortini D, Villa L, Carattoli A, Anjum MF, Nagy B. First report on IncN plasmid-mediated quinolone resistance gene qnrS1 in porcine Escherichia coli in Europe. Microb Drug Resist. 2011;17(4):567-573.
Crossref - Dolejska M, Villa L, Hasman H, Hansen L, Carattoli A. Characterization of IncN plasmids carrying blaCTX-M-1 and qnr genes in Escherichia coli and Salmonella from animals, the environment and humans. J Antimicrob Chemother. 2013;68(2):333-339.
Crossref - Ma J, Liu J-H, Lv L, et al. Characterization of extended-spectrum β-lactamase genes found among Escherichia coli isolates from duck and environmental samples obtained on a duck farm. Appl Environ Microbiol. 2012;78(10):3668-3673.
Crossref - Netikul T, Sidjabat HE, Paterson DL, et al. Characterization of an IncN2-type blaNDM-1-carrying plasmid in Escherichia coli ST131 and Klebsiella pneumoniae ST11 and ST15 isolates in Thailand. J Antimicrob Chemother. 2014;69(11):3161-3163.
Crossref - Poirel L, Lagrutta E, Taylor P, Pham J, Nordmann P. Emergence of metallo-β-lactamase NDM-1-producing multidrug-resistant Escherichia coli in Australia. Antimicrob Agents Chemother. 2010;54(11):4914-4916.
Crossref - Hao Y, Shao C, Geng X, Bai Y, Jin Y, Lu Z. Genotypic and phenotypic characterization of clinical Escherichia coli sequence type 405 carrying IncN2 plasmid harboring blaNDM-1. Front Microbiol. 2019;10:788.
Crossref - Norman A, Hansen LH, She Q, Sorensen SJ. Nucleotide sequence of pOLA52: a conjugative IncX1 plasmid from Escherichia coli which enables biofilm formation and multidrug efflux. Plasmid. 2008;60(1):59-74.
Crossref - Johnson TJ, Danzeisen JL, Youmans B, et al. Separate F-type plasmids have shaped the evolution of the H30 subclone of Escherichia coli sequence type 131. MSphere. 2016;1(4):1-15.
Crossref - Pitout JD, DeVinney R. Escherichia coli ST131: a multidrug-resistant clone primed for global domination. F1000Research. 2017;6:195.
Crossref - Kondratyeva K, Salmon-Divon M, Navon-Venezia S. Meta-analysis of pandemic Escherichia coli ST131 plasmidome proves restricted plasmid-clade associations. Sci Rep. 2020;10(1):36.
Crossref - Magiorakos A-P, Srinivasan A, Carey R, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268-281.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.