ISSN: 0973-7510
E-ISSN: 2581-690X
The lactic acid bacteria (LAB) involved in production of L’ben, an Algerian traditional fermented cow milk product, were isolated and identified. 10 samples of this product, collected from different regions of western Algeria, were analyzed. A total of 30 LAB strains were isolated and identified by phenotypic and biochemical analyses and API 20 Strep identification system. The results obtained showed that the isolates belong to the species: Enterococcus faecium (36,6%), Enterococcus faecalis (30%), Enterococcus avium (13,3%), Leuconostoc mesenteroides (13,3%), Lactococcus lactis ssp cremoris (3,3%) and Lactococcus lactis ssp lactis (3,3%). Thereafter, growth kinetic and technological properties of these bacteria including: the acidification ability and the proteolytic activity were studied in a purpose of a possible industrial use. The obtained results were very heterogeneous; this led us to use the Principal Components Analysis “PCA” for strains grouping and comparison of their properties facilitated by graphics presenting all properties of each strain simultaneously to elect among all strains those with the most interesting technological profiles. Only four strains form a distinct group in the positive sense of the vectors of the variables formed and could become leavens for fermenting different kinds of milks, especially that of cow, which represents the ecosystem from which they originate.
Acidification, Identification, Lactic acid bacteria, Proteolysis, PCA
Various types of traditionally fermented dairy products exist throughout the world , Their nature depends on the type of milk used, the pretreatment, the conditions of fermentation and subsequent processing (Tamime, 1997; Stanely, 1998). These products have a very important role in the daily diet of people. Indeed, they have been the subject of several studies in: Algeria ( Harrati ,1974) , Morocco (Ouadghiri et al., 2009), Romania (Zamfir et al., 2006), Lebanon (Chammas et al., 2006), Egypt (El-Baradei et al., 2008), Burkina Faso (Savadogo et al., 2004), Kenya (Mathara et al., 2004), South Africa (Beukes et al., 2001) etc.
In Algeria, the fermentation of milk gives the “L’ben”, a popular traditional fermented milk product, produced mainly in the rural areas. Transformation of cow milk into traditional Algerian dairy products, such as L’ben, achieved through spontaneous fermentation without the addition of any selected starter. Milk fermentation process has been relied on the activity of LAB, which play a crucial role in converting milk as raw material to fermented milk products. Such products generally present irregular sensorial qualities. The dairy industry has developed considerably, thanks to the use of selected lactic acid bacteria, the choice being based on their production of lactic acid, aromatic compounds, bacteriocins, and CO2 and their resistance to phages (De Roissart, 1986; Herrero et al., 1996; AL’benzio et al., 2001).
The main objective of this study was to isolate firstly and identify the predominant LAB responsible for the fermentation of the traditional fermented milk L’ben and secondly to study in these strains, some properties with technological potential as acidifying activity and proteolysis. This new isolates may attract the attention of the dairy industry for development of starter cultures because the value of these strains lies in their ability to produce specific metabolites, especially those responsible for flavor then use them in an industrial process for other fermented products with better hygienic quality and thorough therapeutic character.
Samples collection
A total, of 10 samples of L’ben traditionally maked were collected from cow farms in western of Algeria. The samples were collected aseptically in sterile bottles , transported immediately to the laboratory in a cool isotherm box and stored at 4°C before analysis.
Isolation of lactic acid bacteria
All samples were serial ten-fold diluted on NaCl solution (0,9%) and 1 ml of these dilutions was pour-plated at surface in the media of lactic acid bacteria, M17 (Terzaghi et Sandine, 1975) and MRS (De Man et al., 1960) adjusted to pH 5.5. After incubation at 30°C for 24 to 48 h, the colonies were randomly picked from plates having a colony number between 30 and 300 and several representative strains displaying the general characteristics of lactic acid bacteria were chosen for more purification by successive streaks on M17 and MRS plates (Harrigan and Mc Cance,1976). The screening of the purified isolates was carried out according to the Gram staining and catalase test. The strains of lactic acid bacteria were stored at –20°C in M17 and MRS broth with glycerol (80%).
Identification of lactic acid bacteria
Gram-positive and Catalase negative strains were presumptively identified as LAB. Microscopic appearances of cultures were judged using Gram stained preparations (Gerhardt et al., 1981) .Further identification was performed by using the following biochimical properties : growth at different temperatures (10, 15, 20 25, 30, 37 and 45°C) and at pH 9.6 as well as the ability to grow in the different concentrations of NaCl (2, 3, 4, 4.5, 5, 6.5 and 10%,); survival after heating of 60°C for 30 min (Samelis et al., 1994);gas production from glucose, determined in M17 and MRS broth containing inverted Durham ; hydrolysis of arginine, tested on M17 and MRS with bromocresol purple (Thomas, 1973); and production of acetoin from glucose, determined by using the Voges-Prokauer test (Zourari et al., 1991).
The fermentation of carbohydrates was determined in MRS and M17 broth without glucose and meat extract, but containing bromocresol purple (0.02 g/l) used as a pH indicator, and supplemented with 1% of the following carbohydrates: glucose, fructose, lactose, saccharose, xylose, arabinose, maltose, mannose, cellobiose, raffinose and Mannitol. To ensure anaerobic conditions, each tube was topped up with drop of sterile liquid paraffin after inoculation (Samelis et al., 1994).
The isolates were selected for identification to species level using the API 20 Strep galleries and API 20 Strep medium (BioMerieux S.A., Marcy-l’Etoile, France) according to the manufacturer’s instructions. The galleries were incubated at 30°C and reactions were observed after 24 and 48 h. The API Web database (API Strepto V7.0 2007/02 BioMerieux S.A.) was used to interpret the obtained results. All tests for phenotypic characterization were conducted twice for each strain.
Technological characterization
Measurements of Acidity and growth kinetics parameters
Sterilized skimmed-milk was inoculated with 1% of each precultured strain and incubated at 30°C for 18 h. At a regular interval time (1 hour), samples were aseptically collected and the strains growth was determined by measuring the optical density at ë = 600 nm as described by Boutrou et al. (1998). The determination of kinetic growth parameters (Nmax; ìmax and T-lag) , which are respectively, Maximum cell density, growth rate and latency, was obtained from each strain growth curve using specialized software: MicroFit © version 1.0 of the “Institute of Food Research (IFR), UK” designed by Dr. Peter DG Wilson as the Baranyi growth model (Baranyi and Roberts, 1994). The Dornic acidity (1°D = 0.1 g lactic acid/liter) was performed according to the method described by Accolas et al. (1977) by titration of 10 ml of sample cultures with NaOH (1/9N) in the presence of phenolphthalein indicator (1% in alcohol) at a regular interval time (1 hour).
Measurements of Proteolytic Activity
All the thirty bacterial cultures were evaluated for their ability to the proteolysis in skim milk agar plate (skim milk powder 4 % and agar 1.5%). In this test to detect protein hydrolysis, all selected bacteria (LAB), were seeded at the surface of skim milk agar plates and were incubated at 30°C for 48 h, followed by cooling in a refrigerator (4°C) for 3 days as described by (Fransen et al, 1997; Pailin et al., 2001). Protein hydrolysis was observed by the production of clear halos surrounding isolated colonies and the diameter of each clear zone and it’s corresponding colony was measured and recorded in mm. Duplicate trials were conducted and all results were averaged. The proteolytic activity (PA) was expressed as the ratio of the diameter of the clear zone to the diameter of the colony.
Statistical analysis (Principal Component Analysis)
Principal component analysis (PCA) is probably the most popular multivariate statistical technique and it is used by almost all scientific disciplines. It is also likely to be the oldest multivariate technique. This technique is a widely used multivariate analytical statistical procedure that can be applied to data to reduce the set of dependent variables to a smaller set of underlying variables (called factors) based on patterns of correlation among the original variables (Lawless and Heymann, 1998).
For the evaluation and visualization of results of the technological characterization of isolates, the multivariate statistic method Principal Component Analyses (PCA) was carried out to describe the relations between the proteolytic activity (PA), acidity and growth kinetics parameters (Nmax; ìmax and T-lag) used as variables for this statistical analysis. The results were arranged into data matrix and analyzed by statistic program (STATISTICA software for Windows), version 7.0.
Isolation of lactic acid bacteria
Several authors have recorded the predominance of LAB in traditional fermented cow milk (Hamama et Bayi, 1991; Samolada et al., 1998; Lin et al., 1999; Mathara, 1999; Abdelgadir et al., 2001; Beukes et al., 2001). These investigators found the main LAB genera to comprise lactobacilli, lactococci and leuconostocs. The presence of enterococci and pyogenic streptococci has also been reported (Samolada et al., 1998; Mathara, 1999; Beukes et al., 2001).
A total of 30 isolates obtained from L’ben samples, catalase negative and Gram positive, were considered as belonging to the group of lactic acid bacteria (LAB), and were correctly identified by morphological, physiological and biochemical analysis and completed by using the API 20 Strep system as described by the manufacture instructions. From the 30 isolates, 24 isolates were identified as belonged to the genus Enterococcus, 4 isolates to Leuconostoc, 2 isolates to Lactococcus. Figure 1 show the percentage distribution of the 30 strains isolated from L’ben samples and their characteristics are shown in Table 1.
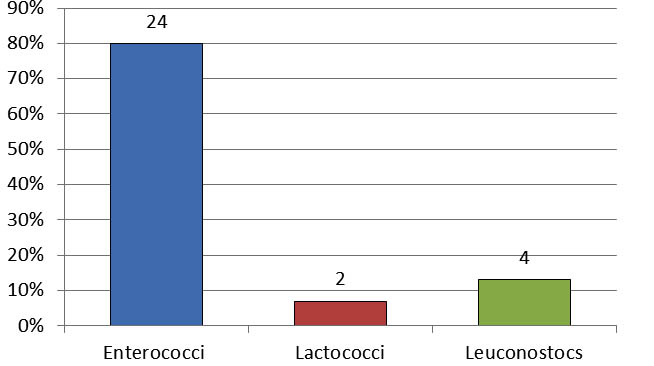
Fig. 1. The percentage distribution of the 30 strains isolated from Algerian L’ben samples
Table (1):
Biochimical and physiological characteristics of LAB strains isolated from Algerian L’ben samples
| Tests | 1 | 2 | 3 | 4 | 5 | 6 | |
|---|---|---|---|---|---|---|---|
| Gram | + | + | + | + | + | + | |
| Catalase | – | – | – | – | – | – | |
| Growth at | 15°C | + | + | + | + | + | + |
| 20°C | + | + | + | + | + | + | |
| 25°C | + | + | + | + | + | + | |
| 30°C | + | + | + | + | + | + | |
| 37°C | + | + | + | + | + | + | |
| 45°C | + | + | + | – | – | – | |
| pH = 9.6 | + | + | + | – | – | – | |
| 2% NaCl | + | + | + | + | + | + | |
| 3% NaCl | + | + | + | + | + | + | |
| 4% NaCl | + | + | + | + | + | – | |
| 4.5% NaCl | + | + | + | + | – | – | |
| 5% NaCl | + | + | + | + | – | – | |
| 6.5% NaCl | + | + | + | – | – | – | |
| Survival after heating at 60°C for 30mn | + | + | + | – | – | – | |
| Production of CO2 from glucose | – | – | – | + | – | – | |
| Hydrolysis of Arginine | + | + | – | – | + | – | |
| Fermentation of | glucose | + | + | + | + | + | + |
| fructose | + | + | + | + | + | + | |
| lactose | + | + | + | + | + | + | |
| saccharose | + | + | + | + | + | + | |
| xylose | – | – | – | + | – | – | |
| arabinose | + | – | – | – | – | – | |
| maltose | + | + | + | + | + | – | |
| mannose | + | + | + | + | + | + | |
| cellobiose | + | + | + | + | + | + | |
| raffinose | – | – | – | – | – | – | |
| Mannitol | – | – | – | + | – | – | |
| Number of species | 11 | 9 | 4 | 4 | 1 | 1 | |
Key:
Species: 1. Enterococcus faecium; 2. Enterococcus faecalis; 3. Enterococcus avium; 4. Leuconostoc mesenteroides; 5. Lactococcus lactis ssp lactis; 6. Lactococcus lactis ssp cremoris; +, positive; -, negative
The 24 Enterococci isolated were characterized by their ability to grow from 15 to 45°C in 6.5% NaCl and pH 9.6 broth. They also survive at 60°C for 30 min and form NH3 from arginine, but not CO2 from glucose. At this level, the two classical species Enterococcus faecalis and Enterococcus faecium have been identified, the latter having the same profile but differing by the hydrolysis of L-Arabinose where Enterococcus faecium metabolizes this sugar unlike Enterococcus faecalis, as reported by Boubekri et Ohta (1996).
The complementary identification up to the level of species through the use of API 20 Strep galleries have revealed that, 11 strains of 24 were identified at Enterococcus faecium (LBO6, LBO7, LBO10, LBO12, LBO14, LBO18, LBO19, LBO21, LBS1, LBS2 and LBS3), 9 strains, to Enterococcus faecalis (LBO1, LBO2, LBO4, LBO5, LBO15, LBO16, LBO17, LBO25 and LBR1) and 4 to Enterococcus avium (LBO8, LBO9, LBO20 and LBO23).
The four isolates Gram-positive ovoid cocci form (LBO3, LBO11, LBO22 and LBO26), assigned to the genus of Leuconostoc, are heterofermentative characterized by the production of gas from glucose (which is an important characteristic for distinguishing them), mesophilic (grow from 15 to 37°C, but not at 45°C), thermosensitive and were unable to grow at pH 9.6 also at 6.5% NaCl. All of these isolates were ADH negative and fermented maltose and galactose but not arabinose and raffinose. According to the scheme of Villani et al. (1997), they were presumptively identified as Leuconostoc sp. The results of the complementary identification of these four strains with the API 20 Strep system belonged them to Leuconostoc mesenteroides.
The two isolates (LBO13 and LBO24) belonged to Lactococcus lactis, are homofermentative, mesophilic and thermosensitive and unable to grow neither at pH 9.6 nor at 6.5% NaCl. By reference to the API 20Strep identification system and the ADH test they were subdivided into two subspecies; Lactococcus lactis subsp lactis (ADH positive) ,coded LBO24, and Lactococcus lactis subsp cremoris (ADH negative) coded LBO13, moreover, this latter can’t grow at 4% of NaCl and can only use very few carbohydrates.
This study has revealed that L’ben samples are dominated majorly by the genus of Enterococcus, followed shyly by Leuconostoc and Lactococcus, with a total absence of lactobacilli. These same observations already reported in earlier work by Harrati, (1974), where he also noted the absence of these bacteria in the Algerian L’ben and made the following assumptions: Fermentation of milk by lactococci would not be sufficiently advanced allow the lactobacilli to grow, the presence of inhibitory factors for lactobacilli in Algerian milk, Their development is favored when the milk is coagulated and when the L’ben reaches a high acidity.
According to Ouadghiri et al.(2009), the low presence of lactobacilli in the L’ben or even their total absence can be explained by the delayed growth of the latter under mesophilic conditions. Another study by Abo-Elnaga et al.(1977), carried out on iraquien L’ben, showed that the flora of this product is dominated by lactobacilli because it has a titratable acidity of 131° D in comparison with the Algerian L’ben, in which it is lower, around 78° D.
The predominance of enterococci in our L’ben samples may be related to a lack of bacteriocin-producing strains (Benkerroum et al., 2000), although this has not been verified in this study. The two classic species Enterococcus faecalis and Enterococcus faecium have been isolated in significant quantities by several authors; from Algerian Klila (Boubekri and Ohta, 1996), from the Moroccan Jben (Ouadghiri et al., 2005) and from the Algerian artisanal cheese “Bouhezza” (Aissaoui et al., 2012). Similarly, Mechai et al. (2014) identified Enterococcus faecium in the Algerian samples of L’ben, Jben and Rayeb. Ouadghiri et al. (2009) described the prevalence of Enterococcus faecium species in other enterococci in morrocan L’ben samples.
According to Švec and Franz (2014), enterococci are important bacteria in the diet as well as in public health, its association with food seems harmful because they cause deterioration, or on the other hand, they can play an important role in the maturation and development of aromas of some traditional fermented products such as cheese especially, those that are manufactured in Mediterranean regions. Entrococci are also used as probiotics and are useful in the treatment of diarrheal diseases caused by food-related pathogens.
As for the Leuconostocs, their presence may be linked to post-preparation contamination, since Mathara et al. (2004) reported that strains of Leuconostoc have requirements nutrients and are uncompetitive during milk fermentation.
Holzapfel (2002) notes that leuconostocs frequently dominates the early stages of most spontaneous fermentations. All members of the genus Leuconostoc are heterofermentative, fermenting glucose via the hexose-monophosphate pathway to produce equimolar amounts of lactic acid, ethanol and CO2 (Garvie, 1984; Jay, 1992). Additionally, they are able to convert citrate to aroma compounds such as acetoin and diacetyl, a characteristic that would be of functional significance towards aroma development in L’ben samples.
These bacteria are generally associated with different habitats, and have an indisputable importance in the fermentation of food, but they can also cause deterioration (Björkroth et al., 2014). Despite this, these bacteria pose no health risk. Leuconostocs are considered to be safe microorganisms (GRAS) (Ogier et al., 2008).
For lactococci, an identical result by Harrati (1974) has been described in Algerian fermented milk “L’ben”,where the two strains : Lactococcus lactis subsp. cremoris and Lactococcus lactis subsp. lactis, obtained in our case, were identified. These bacteria are known for their use as starter cultures of fermented dairy products. They metabolize milk lactose, and their main role included in texture development by the production of exopolysaccharides (EPS) and flavor by the production of aromatic compounds (alcohols and aldehydes) and / or through the metabolism of citrate, amino acids and lipids (Kim , 2014).
Technological Characterization of Lactic Acid Bacteria
All the results relating to growth parameters, acidity and proteolysis are given in table 2.
Table (2):
The thirty strains responses relating to growth parameters, acidity and proteolysis
Strains |
Nmax |
µmax |
T-lag |
Acidity(°D) after 19h |
Proteolsis H/C |
|---|---|---|---|---|---|
LBO 1 |
1.17 |
0.17 |
2.29 |
91 |
4.6 |
LBO 2 |
1.38 |
0.21 |
3.11 |
92 |
5 |
LBO 3 |
2.23 |
0.55 |
8.53 |
117 |
2.8 |
LBO 4 |
1.12 |
0.17 |
3.05 |
91 |
3.75 |
LBO 5 |
2.20 |
0.11 |
7.43 |
82 |
3.6 |
LBO 6 |
1.54 |
0.25 |
3.46 |
68 |
2.1 |
LBO 7 |
1.95 |
0.52 |
2.83 |
99 |
2.3 |
LBO 8 |
1.46 |
0.23 |
3.88 |
85 |
5.25 |
LBO 9 |
1.18 |
0.20 |
2.48 |
82 |
3.5 |
LBO 10 |
1.41 |
0.57 |
5.84 |
74 |
2.4 |
LBO 11 |
2.21 |
0.55 |
3.99 |
112 |
8 |
LBO 12 |
2.02 |
0.36 |
4.68 |
88 |
3 |
LBO 13 |
2.21 |
0.33 |
3.55 |
86 |
2.8 |
LBO 14 |
1.84 |
0.32 |
3.70 |
84 |
3.2 |
LBO 15 |
1.78 |
0.34 |
5.95 |
91 |
4 |
LBO 16 |
2.21 |
0.45 |
6.70 |
79 |
3.4 |
LBO 17 |
2.24 |
0.47 |
5.91 |
75 |
3.6 |
LBO 18 |
1.97 |
0.32 |
2.55 |
78 |
4.75 |
LBO 19 |
2.14 |
0.33 |
2.73 |
79 |
3.6 |
LBO 20 |
1.93 |
0.31 |
3.08 |
70 |
3.6 |
LBO 21 |
1.88 |
0.31 |
3.48 |
70 |
2.3 |
LBO 22 |
1.51 |
0.47 |
3.41 |
71 |
3.4 |
LBO 23 |
1.53 |
0.60 |
4.57 |
81 |
4.2 |
LBO 24 |
1.69 |
0.35 |
7.28 |
77 |
2.7 |
LBO 25 |
1.93 |
0.44 |
7.97 |
69 |
3.3 |
LBO 26 |
1.52 |
0.36 |
2.97 |
77 |
3.7 |
LBS 1 |
2.28 |
0.56 |
4.60 |
85 |
2.4 |
LBS 2 |
1.17 |
0.51 |
4.81 |
82 |
2.6 |
LBS 3 |
1.34 |
0.19 |
2.69 |
83 |
3.6 |
LBR 1 |
1.23 |
0.17 |
1.14 |
83 |
3.8 |
Kinetics of growth and dornic acidity evolution
An accurate evaluation of the isolates growth in skim milk is important. To be done, an examination of the growth profiles of the thirty strains must be realized. The variation of growth was monitored for all isolates, as shown in Figure 2.
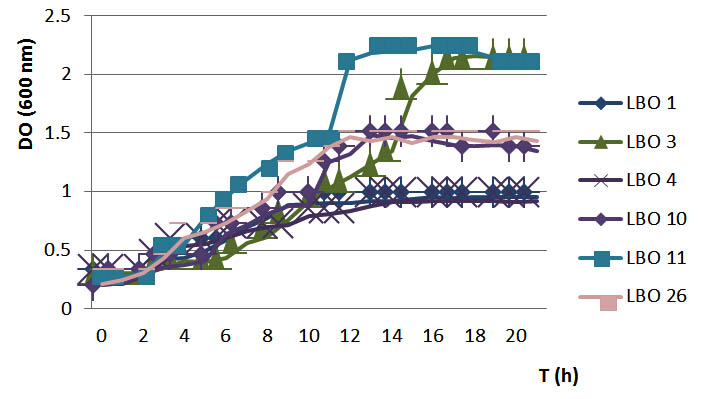
Fig. 2. Example of the growth kinetics of six strains
Estimation of growth parameters
Growth parameters, estimated by using the Baranyi model (Baranyi et Roberts, 1994), are: Nmax [maximum cell concentration log (cfu.ml-1)]; µmax [maximum specific growth rate (h-1)] and T-lag [duration time of lag phase (h)]
Values of the growth parameters obtained varied between strains. It depends to their ability to growth in milk. The highest biomass and growth rates, together with shorter lag phases, were obtained in the strains LBO11 “Leuconostoc mesenteroides” and LBO13 “Lactococcus lactis subsp. cremoris”. In fact, after only four hours of latency, this strains and especially LBO11 grew faster than the other strains and register biomass increase until 19 h before their growth leveled off at around 6.63. The low biomass produced by the other strains correlated well with the lower amount of lactic acid produced resulting in a higher pH can be justified by the accumulation of lactic acid in the medium. Therefore these strains were found to be unable to tolerate high lactic acid concentrations. This can be also explained by the complex growth requirements (Gomes and Malcata, 1999) of this species, exhibiting poor growth in media without the addition of large amounts of supplements, such as yeast extract and peptone (Taillandier et al., 1996 ; Elli et al., 1999).
Lactic acid production
The accurate estimation of the lactic acid production is of paramount importance. In fact, the diminution of pH of the milk is due to the production of lactic acids from lactose fermentation (Thomson et Gentry, 1994). The amount of lactic acid varies according to the strains and their capacity and the rate of degradation of the lactose (figure 3), indeed the amount of lactic acid produced after 19 h of incubation is between (70 and 100 °D) in the majority of strains with the exception of two Leuconostoc strains (LBO3 and LBO11) which stand out from the set of strains since they produce a higher amount of lactic acid between 112 and 117°D.
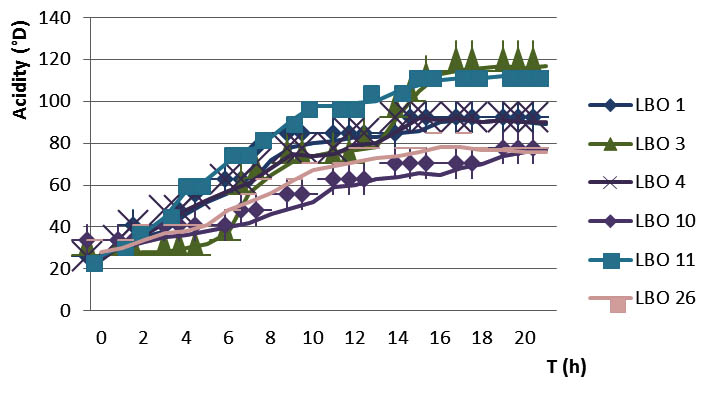
Fig. 3. Example of Dornic acidity kinetics of six strains
Thus, according to their ability of acidification, the strains were divided as follows: highly acidifying isolates (include LBO3 and LBO11) that coagulate milk before 10 h of incubation, low acidifying isolates (LBO6, LBO20, LBO21, LBO22 and LBO25) that coagulate milk after 19 h of incubation, the remaining isolates are recognized as moderately acidifying isolates, they coagulate milk after 10 to 19 h of incubation.
The varying acidities observed between different isolates were explained by De Roissart (1986). Indeed, the acidifying aptitude characterizes each isolate because it is related to the particular aptitudes which have it to degrade the compounds of the medium, to make them more assimilable and to transport the nutritive elements in the cytoplasm. In addition, the differences are due to a deficiency in the transport system of the nutritive substances of fermentable sugars (AL’benzino et al., 2001).
The fast acidifying strains are therefore good candidates for dairy fermentation process as primary starter culture while poor acidification strains can be used as adjunct cultures depending on other properties (Ayad et al., 2004).
Proteolytic activity
The appearance of the clear zones of the proteolytic activity in skim milk agar medium was easily detected (Figure 4).

Fig. 4. Example of Proteolytic activity of some strains
The proteolytic activity of mesophilic strains do not differ greatly between the thirty strains studied with the exception of a single strain of Leuconostoc (LBO11 ) which emerges from the set of strains because of their largest zone . In fact, an examination of the ratio (halo / colony) (figure 5), shows that this strain presents a maximum activity in contrary of the others that have a low proteolytic activity. This is an indication that the LAB strains have very low proteolytic activities in Milk, in line with the findings of other investigators on LAB from other sources (Ayan et al., 2005; Thapa et al., 2006; Hassaine et al., 2007). The proteolytic activity of dairy LAB is essential for the growth of the organisms in milk and it is involved in the development of organoleptic properties of different fermented products (Christensen et al., 1999; Hassaine et al., 2007).
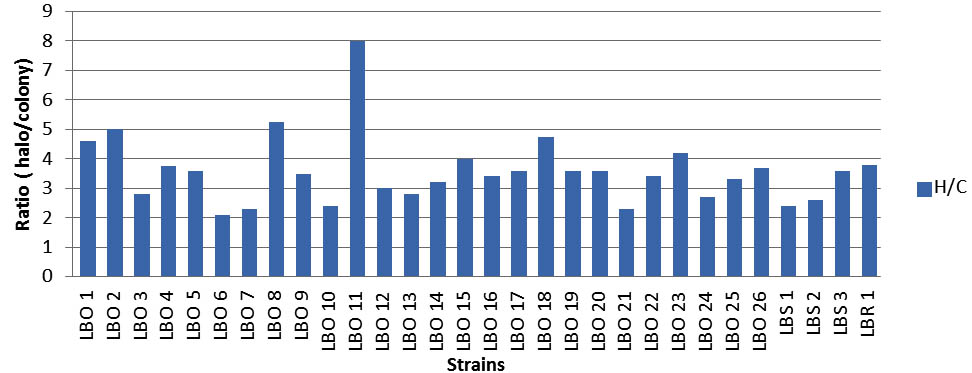
Fig. 5. Histogram representation of ratio (H/C) of the thirty strains
Data analysis
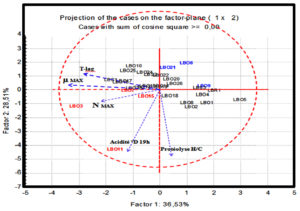
An analysis in principal components “PCA” was applied in order to classify the strains simultaneously for all the characters studied: growth parameters (Nmax ,µmax ,Tlag), dornic acidity and proteolysis. The purpose of use of PCA was to join, into clusters, the strains showing a very heterogeneous technological interests profiles. The different strains are divided along two axes described by the five variables studied as shown in figure 6.
The first two axes (F1, F2) together account for 65, 04 % of the variance. The LBO3, LBO7, LBO11 and LBO15 strains (shown in red) form a distinct group in the positive direction of the variables vectors, those with a higher level of growth, acidifying and proteolytic activity. Another group is formed opposite to the latter (shown in blue), composed of strains LBO6, LBO9, LBO21, which are characterized by slow growth, low acidity and proteolytic activity and between these two groups, the majority of the remaining strains form a third intermediate group (presented in black), which show a great heterogeneity compared to the variables studied.
This analysis also shows that the LBO11 strain stands out slightly from its group, formed by the strains demonstrating an interesting technological profile, thanks to the high values recorded for this strain in terms of growth and both activities studied (acidification and proteolysis). The same goes for the LBO3 strain, which can be also distinguished from its group, but mainly because of its growth and acidification, which are important.
Based on these results, it seems obvious that these two strains prove to present the most interesting technological profiles for possible small or large scale applications.
This method offers us a powerful discriminatory analysis enough to sort and elect the most successful strains among a set of strains showing a very heterogeneous profiles, it’s a powerful tool, which should prove to be useful in the field of strains selection, for industry as for research (Zourari et al., 1991).
The results arising from this work showed that several LAB species and genera can be isolated from the Algerian fermented cow milk “L’ben”. A dominance of enterococci “80%” against 13% and 7% respectively for leuconostoc and lactococci was detected. These isolates showed interesting technological properties notably for the two strains of Leuconostoc mesenteroides LBO3 and LBO11, both strains were selected by the discriminatory analysis and could be used as starters in the manufacture of dairy products with sensorial qualities liked by tasters. Optimal exploitation of them requires specific conditions of use and further, more detailed studies of their technological properties to evaluate the selected strains on a pilot scale individually and in mixed cultures.
- Tamime, A.Y., Marshall, V.M.E. Microbiology and technology of fermented milks, In: B.A. Law (Ed). Microbiology and Biochemistry of Cheese and Fermented Milk, Blackie Academic and Professional, 1997; pp 57-152.
- Stanley, G. Cheeses, In: B.J.B. Wood (Ed). Microbiology of Fermented Foods, second ed., 1, Blackie Academic and Professional, 1998; pp263-307.
- Harrati, E. Recherches sur le L’ben et le klila algériens. Ph. D. thesis, U.E.R. Sciences de la Vie, Université de Caen (France). 1974.
- Ouadghiri, M., Vancanneyt ,M., Vandamme, P., Naser ,S., Gevers ,G., Lefebvre, K., Swings, J. Identification of lactic acid bacteria in Moroccan raw milk and traditionally fermented skimmed milk ‘L’ben‘. J Appl Microbiol., 2009; 106: 486-495.
- Zamfir, M., Vancanneyt, M., Makras, L., Vaningelgem, F., Lefebvre, K., Pot, B., Swings, J., De Vuyst, L. Biodiversity of lactic acid bacteria in Romanian dairy products. Syst Appl Microbiol., 2006; 29: 487-495.
- Chammas, G.I., Saliba, R., Béal, C. Characterization of the fermented milk “Laban” with sensory analysis and instrumental measurements. J Food Sci., 2006; 71: 156-162.
- El-Baradei ,G., Delacroix-Buchet, A., Ogier, J.C. Bacterial biodiversity of traditional Zabady fermented milk. Int J Food Microbiol., 2008; 121: 295–301.
- Savadogo, A., Ouattara, C.A.T., Bassole, I.H.N., Traore , A.S. Antimicrobial activities of lactic acid bacteria strains isolated from Burkina Faso fermeted milk. Pakistan J Nutr., 2004; 3(3):174-179.
- Mathara, J.M., Schillinger, U., Kutima, P.M., Mbugua, S.K., Holzapfel, W.H. Isolation, identification and characterisation of the dominant microorganisms of Kule naoto: the Maasai traditional fermented milk in Kenya. Int J Food Microbiol, 2004; 94: 269-278.
- Beukes, E.M., Bester, B.H., Mostert, J.F. The microbiology of South African traditional fermented milks. Int J Food Microbiol., 2001; 63(3): 189-197.
- De Roissart, H.B. Bactéries lactiques. In Laits et produits ! laitiers vache, brebis et chevre. Tome 3. Paris, Technique et documentation. Lavoisier, 1986; pp 343-408.
- Herrero, M., Mayo, B., Gonzalez, B., Suarez, J.E. Evaluation of technologically important traits in lactic acid bacteria isolated from spontaneous fermentation. Journal of Applied Bacteriology., 1996; 81: 565-570.
- AL’benzio, M., Corb, M.R., Rehman, S.U., Fox, P.F., De Angelis, M., Corsetti, A., Sevi, A., Gobetti, M. Microbiological and biochemical characteristics of canestrato pugliese cheese made from raw milk, pasteurized milk or by heating the curd in hot whey. International Journal of Food Microbiology., 2001; 67: 35-48.
- Terzaghi, B.E., Sandine, W.E. Improved medium for lactic streptococci and their bacteriophages. Applied Microbiology., 1975; 29: 807-813.
- De Man, J.C., Rogosa, M., Sharpe, M.E. A medium for the cultivation of lactobacilli. J Appl Bacteriol; 1960., 23: 130-135.
- Harrigan, W.F., McCance, M.E. Laboratory Methods in Food and Dairy Microbiology. Academic Press, London. 1976.
- Gerhardt, P., Murray, R.G.E., Costilow, R.N., Nester, E.W., Wood, W.A., Krieg, N.R., Phillips, G.B. Manual of Methods For General Bacteriology. American Society for Microbiology, Washington, DC. 1981.
- Samelis, J., Maurogenakis, F., Metaxopoulos, J. Characterization of lactic acid bacteria isolated from naturally fermented Greek dry salami. International Journal of Food Microbiology., 1994 ; 23: 179-196.
- Thomas,T.D. Agar medium for differentiation of Streptococcus cremoris from the other bacteria. Dairy Sci Technol., 1973; 8: 70-71.
- Zourari, A., Roger, S., Chabanet, C., Desmazeaud, M.J. Caracterisation des bacteries lactiques thermophiles isolees de yaourts artisanaux grecs. I. Souches de Streptococcus salivarius subsp thermophilus. Le Lait ; 1991 ;71(4) : 445-461.
- Boutrou, R., Sepulchre ,A., Gripon, J.C., Monnet, V. Simple tests for predicting the lytic behaviour and proteolytic activity of lactococcal strains in cheese. Journal of Dairy Science., 1998 ; 81: 2321-2328.
- Baranyi, J., Roberts,T.A. A dynamic approach to predicting bacterial growth in food. Food Microbiol., 1994; 23: 277-294.
- Accolas, J.P., Bloquel, R., Regnier , J. Acidifying properties of the thermopiles lactic bacteria in relation to the manufacture of yoghourt. Milk., 1977; 67: 1-23.
- Fransen, N.G., O’Connell, M.B., Arendt, E.K. A modified agar medium for screening of proteolytic activity of starter cultures of meat fermentation purpose. International Journal of Food Microbiology., 1997; 36: 235-239.
- Pailin, T ., Kang, D.H., Schmidt, K., Fung, D.Y.C. Detection of extracellular bound proteinases In EPS-producing lactic acid bacteria cultures on skim milk agar. Letters in Applied Microbiology., 2001; 33: 45-49.
- Lawless, H.T., Heymann, H. Sensory evaluation of food: Principles and practices. New York : Kluwer Academic/Plenum Publishers, 1998; pp 362-378.
- Hamama, A., Bayi, M. Composition and microbiological profile of two Moroccan traditional dairy products: raib and jben. J Soc Dairy Technol .,1991; 44(4): 118-120.
- Samolada, M., Litopoulou-Tzanetaki, E., Xanthopoulos, V., Tzanetakis, N. Changes in microbial flora during manufacture of a traditional fermented milk from ewe’s milk. Food Microbiology .,1998; 15(1): 43-50.
- Lin, C.W., Chen, H.L., Liu, J.R. Identification and characterization of lactic acid bacteria and yeasts isolated from kefir grains in Taiwan. Aust J Dairy Technol., 1999; 54:14-18.
- Mathara, J.M. Studies on lactic acid producing microflora in mursik and kule naoto, traditional fermented milks from Nandi and Maasai communities in Kenya. Ph.D. thesis, University of Nairobi. 1999.
- Abdelgadir, W.S., Hamad, S.H., Moller, P.L., Jakobsen, M. Characterisation of the dominant microbiota of Sudanese fermented milk rob. International Dairy Journal., 2001; 11(1-2): 63-70.
- Boubekri, K., Yoshiyuki, O. Identification of lactic acid bacteria from Algerian traditional cheese, El-klila. J Sci Food Agr., 1996; 70(4):501-505.
- Villani, F., Moschetti, G., Blaiotta, G., Coppola, S. Characterization of strains of Leuconostoc mesenteroides by analysis of soluble whole-cell protein pattern. DNA finger printing and restriction of ribosomal DNA. Journal of Applied Bacteriology., 1997; 82: 578-588.
- Abo-elnaga, I.G ., El-aswad, M ., Moqui, M. Some chemical and microbiological characteristics of Leben. Milchwissenschait., 1977; 32: 521-524.
- Benkerroum, N., Oubel, H., Zahar, M., Dlia, S ., Filali-Maltouf, A. Isolation of a bacteriocin-producing Lactococcus lactis subsp. lactis and application to control Listeria monocytogenes in Moroccan jben. Journal of Applied Microbiology., 2000; 89: 960-968.
- Ouadghiri, M., Vancanneyt, M., Amar, M., Swings, J. Biodiversity of lactic acid bacteria in Moroccan soft white cheese (jben). Fems Microbiology Letters ., 2005; 251: 267-271.
- Aissaoui, O., Pediliggieri, C., Benatallah, L., Lortal, S., Licitra, G., Zidoune, M.N., Carpino , S. Bouhezza, a traditional Algerian raw milk cheese, made and ripened in goat skin bags. Journal of Food Agriculture and Environment., 2012 ; 10(2): 289- 295.
- Mechai, A., Debabza, M., Kirane, D. Screening of technological and probiotic properties of lactic acid bacteria isolated from Algerian traditional fermented milk products. Int Food Res J ., 2014; 21(6): 2451-2457.
- Švec, P., Franz, M.A.P.C. The genus Enterococcus, In: Lactic Acid Bacteria, Biodiversity and Taxonomy. UK: John Wiley et Sons, 2014 ; pp 175-211.
- Holzapfel ,W.H. Appropriate starter culture technologies for small-scale fermentation in developing countries. Int J Food Microbiol., 2002; 75(3):197-212.
- Garvie, E.I. Taxonomy and identification of dairy bacteria. In F. L. Davies, & B. A. Law (Ed), Advances in the microbiology and biochemistry of cheese and fermented milk. London: Elsevier Applied Science Publishers,1984; pp 35–66.
- Jay , J.M. Modern food microbiology. New York: Chapman &Hall. 1992.
- Björkroth, J., Dicks, M.T., Endo, A., Holzapfel, W.H. The genus Leuconostoc, Lactic Acid Bacteria, Biodiversity and Taxonomy. UK: John Wiley et Sons LTD, 2014; pp 391-404.
- Ogier, J.C., Casalta, E., Farrokh, C., Saïhi, A. Safety assessment of dairy microorganisms: the Leuconostoc genus. International Journal of Food Microbiology., 2008; 126 : 286-290.
- Kim, W. The genus Lactococcus, Lactic Acid Bacteria, Biodiversity and Taxonomy. UK: John Wiley et Sons, 2014; pp 430-443.
- Gomes, A.M.P ., Malcata, F.X. Bifidobacterium spp. & Lactobacillus acidophilus: biological, biochemical, technological and therapeutical properties relevant for use as probiotics. Trends Food Sci Tech., 1999; 10: 139-157.
- Taillandier, P., Gilis, F., Portugal, F.R., Laforce, P ., Strehaiano, P. Influence of medium composition, pH and temperature on the growth and viability of Lactobacillus acidophilus. Biotechnology Letters., 1996; 18: 775-780.
- Elli, M., Zink, R., Reniero, R., Morelli, L. Growth requirements of Lactobacillus johnsonii in skim and UHT milk. International Dairy Journal., 1999 ; 9: 507-513.
- Thomson, J., Gentry, C.H and Weeks. Métabolisme des sucres par les bactéries lactiques. In Derroissart. H. Luquet. FM. (Ed) Bactéries lactiques aspects fondamentaux et technologiques. Uriage. Lorica, 1994; pp239-290.
- Ayad, E.H.E., Nashat, S., El-Sedek, N., Metwaly, H., El-Soda, M. Selection of wild lactic acid bacteria isolated from traditional Egyptian dairy products according to production and technological criteria. Food Microbiology, 2004; 21: 15-725.
- Ayan, K., Durlu-Ozkaya, F., Tunail, N. Commercially important characteristics of Turkish origin domestic strains of Streptococcus thermophilus and Lactobacillus delbrueckii ssp bulgaricus. International Journal of Dairy Technology ., 2005 ; 58(3): 150-157.
- Thapa, N., Pal, J., Tamang, J.P. Phenotypic identification and technological properties of lactic acid bacteria isolated from traditionally processed fish products of the Himalayas. International Journal of Food Microbiology., 2006; 107: 33-38.
- Hassaine, O., Zadi-Karam, H., Karam, N. Technological important properties of lactic acid bacteria isolated from raw milk of three breed of Algerian dromedary (Camelus dromedarius). Afr J Biotechnol., 2007; 6(14):1720-1727.
- Christensen, J.E., Dudley, E.G., Pederson, J.A., Steele, L.J. Peptidases and amino acid catabolism in lactic acid bacteria. Antonie Van Leeuwenhoek., 1999; 76: 217-246.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.