ISSN: 0973-7510
E-ISSN: 2581-690X
Thirty eight isolates nodulating acacia were isolated from different Moroccan areas to determine their ability to survive under various stress conditions, in order to select resistant ones. The response of these isolates to different environmental stress was evaluated to show their preliminary variability based on phenotypical characteristics. Tolerance to increasing levels of NaCl (0.5-10% w/v), pH (4.8-11), temperature (4-55°C) and resistance response to seven heavy metals and five antibiotics were studied as phenotypic markers. The majority of isolates from different Acacia trees were fast growing. Numerical analysis of these phenotypic characteristics showed that the isolates were divided into three distinct clusters presenting intra and inter-cluster diversity. The symbiotic effectiveness of different isolates was studied and showed significant differences with respect to nodule number, nitrogen fixation and improving plants growth (shoot and root dry weight) compared to the uninoculated and non-fertilized control under greenhouse conditions. On the basis of phenotypic and symbiotic studies, six isolates (I1, I3, I5, I6, I16 and I28) were selected for genotypic characterization. The selection of such rhizobial isolates able to adapt to various environmental stress conditions and to improve plant’s growth is important for their future valorization to develop inoculants for Acacia especially in arid areas.
Rhizobium, Acacia, phenotypic characterization, symbiotic effectiveness, Morocco.
Nitrogen, an essential nutrient for plant’s growth and development, is sometimes limiting because it is generally present as unusable form even though the atmosphere contains 78% of N2 by its volume. Hence, N2 fixing associations are very important to convert the most stable and abundant form of N2 to a useful form (NH3). The Rhizobium-legumes symbiosis is of particular agricultural importance, producing about 50% of 175 million tons of total nitrogen biologically fixed and providing nearly half of the nitrogen used in agriculture per year1. In addition, the increasing use of organic / bio-fertilizers would reduce the need for chemical fertilizers and their adverse effects such as environmental pollution and nature degradation2. Previous studies have been performed to investigate and to identify rhizobia associated with legumes3-4. These nitrogen fixing bacteria are classified into 13 genera containing about 98 species5.
Acacia is a legume tree qualified as anti-desert plant which plays a crucial role in the dynamic and structure of plant communities, provides several services by agro-ecosystems and contributes to the restoration of degraded soils6-7 due to its ability to form a nitrogen-fixing symbiosis with rhizobia. Acacia plantations cover an area of 1,128,000 ha of natural forests in Morocco and are among the fast growing, N2-fixing multipurpose woody legumes which are prominent among the exotic species planted in arid and semiarid lands8. This tree covers different areas such as Atlantic Plain (Mediterranean climate), the Middle Atlas (relatively humid) and Souss, presahara and Sahara arid regions.
In this study, N2 fixing bacteria were isolated from Acacia root nodules and the effects of some environmental stresses including temperature, alkalinity/acidity, heavy metals, antibiotics and salt on their growth were investigated to provide a phenotypic characterization.
Moreover, their symbiotic effectiveness was studied and a genotypic characterization was conducted for the most performant ones.
Collection of nodules
Nodules were collected according to the method recommended by Vincent9 and Beck et al.10 from two Acacia species (A. saligna and A. horrida) growing in three Moroccan regions (Oriental, center and North-West) (table 1). A dug of 15 – 25 cm approximately was carried around the plant to a depth of at least 20 – 50 cm to extract a part of its root system. Then, roots were rinsed delicately with running water and nodules were detached to 1-2 cm of their attachment point.
Table (1):
Geographic origin and geodesic coordinates of the sampling sites..
| Strain name | Acacia species | Geographic origin | Coordinates °N, °W | ||
|---|---|---|---|---|---|
| Latitude | Longitude | altitude | |||
| I1 | Acacia. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
| I2 | A. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
| I3 | A. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
| I4 | A. saligna | Cap de l’eau (Nador) | 35° 08’10’’N | 2° 25’ 30’’W | 1 m |
| I5 | A. saligna | Cap de l’eau (Nador) | 35° 08’10’’N | 2° 25’30’’W | 1 m |
| I6 | A. saligna | Saidia | 35° 05’ 15’’ N | 2° 17’ 16’’ W | 20 m |
| I7 | A. saligna | Berkane | 34°55’12’’ N | 2° 19’ 11’’ W | 173 m |
| I8 | A.saligna | Meknes | 33° 53’ 36’’ N | 5° 32’ 50’’ W | 548m |
| I9 | A. saligna | Saidia | 35° 05’ 15’’ N | 2° 17’ 16’’ W | 20 m |
| I10 | A. saligna | Agadir | 30° 25’ 12’’ N | 9° 33’ 53’’ W | 16 m |
| I11 | A. saligna | Berkane | 34°55’12’’ N | 2° 19’ 11’’ W | 173 m |
| I12 | A. saligna | Berkane | 34°55’12’’ N | 2° 19’ 11’’ W | 173 m |
| I13 | A. saligna | Fez | 34° 02’ 13’’ N | 4° 59’ 59’’ W | 104 m |
| I14 | A. saligna | Fez | 34° 02’ 13’’ N | 4° 59’ 59’’ W | 104 m |
| I15 | A. saligna | Fez | 34° 02’ 13’’ N | 4° 59’ 59’’ W | 104 m |
| I16 | A. saligna | Berkane | 30° 25’ 12’’ N | 9° 33’ 53’’ W | 16 m |
| I17 | A. saligna | Casablanca | 33° 35’ 17’’ N | 7° 36’ 40’’ W | 27 m |
| I18 | A. horrida | Casablanca | 33° 35’ 17’’ N | 7° 36’ 40’’ W | 27 m |
| I19 | A. horrida | Casablanca | 33° 35’ 17’’ N | 7° 36’ 40’’ W | 27 m |
| I20 | A. horrida | Rabat | 34°01’ 31’’ N | 6°50’10’’ W | 79m |
| I21 | A. horrida | Rabat | 34°01’ 31’’ N | 6°50’10’’ W | 79m |
| I22 | A. horrida | Rabat | 34°01’ 31’’ N | 6°50’10’’ W | 79m |
| I23 | A. horrida | Rabat | 34°01’ 31’’ N | 6°50’10’’ W | 79m |
| I24 | A. saligna | Fez | 34° 02’ 13’’ N | 4° 59’ 59’’ W | 104 m |
| I25 | A. saligna | Fez | 34° 02’ 13’’ N | 4° 59’ 59’’ W | 104 m |
| I26 | A. saligna | Fez | 34° 02’ 13’’ N | 4° 59’ 59’’ W | 104 m |
| I27 | A. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
| I28 | A. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
| I29 | A. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
| I30 | A. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
| I31 | A. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
| I32 | A. saligna | Agadir | 30° 25’ 12’’ N | 9° 33’ 53’’ W | 16 m |
| I33 | A.saligna | Meknes | 33° 53’ 36’’ N | 5° 32’ 50’’ W | 548m |
| I34 | A.saligna | Meknes | 33° 53’ 36’’ N | 5° 32’ 50’’ W | 548m |
| I35 | A. horrida | Fez | 34° 02’ 13’’ N | 4° 59’ 59’’ W | 104 m |
| I36 | A. horrida | Fez | 34° 02’ 13’’ N | 4° 59’ 59’’ W | 104 m |
| I37 | A. saligna | Saidia | 35° 05’ 15’’ N | 2° 17’ 16’’ W | 20 m |
| I38 | A. saligna | Oujda | 34°41’12’’ N | 1° 54′ 41’’ W | 450 m |
Isolation of rhizobia from nodules and phenotypic characterization of isolates
The nodules, surface sterilized by ethanol 95% for 10 sec and mercuric chloride solution (HgCl2) for 4 min according to Vincent9, were aseptically crushed with a sterile glass rod in NaCl (9 ‰) drops10. Rhizobia were isolated on Yeast-Mannitol-Agar medium (YMA) supplemented with 0.0025 % (w/v) Congo red using a standard procedure9. The single colonies were selected and checked for purity by repeated streaking on YMA11. Individual colonies morphology was characterized based on size, color, mucosity, borders, transparency, elevation and Gram stain reaction.
Isolates authentification
Acacia cyanophylla’s seeds were surface sterilized by rinsing in ethanol 95% (v/v), soaking for 4 min in 0.2% HgCl2 (w/v), and washed three times in sterile distilled water. They were subsequently scarified with H2SO4 95% and germinated on 0.9% agar for 2 to 3 days in the dark12. Well germinated seeds were aseptically transferred into pots filled with sterile sandy-loam soil at rate of three seeds per pot. The pots were then kept in a greenhouse. Each seedling was inoculated with 1 ml of a freshly prepared bacterial suspension (108 UFC/ml). Three pots were used for each isolate. Non-inoculated plants were used as nitrogen-free control (C0). Nitrogen control (NC) was considered as non-inoculated plants receiving weekly 0.5% KNO3 (w / v) as nitrogen source. After six months, plants were harvested, nodules separated from roots then rhizobia reisolated from nodules.
Bromothymol blue test
The ability of isolates to acidify or basify the YMA medium was evaluated by adding bromothymol blue (BBT) at 0.0025% (w/v) as pH indicator (yellow for acid reaction and dark blue for basic reaction)9. The inoculated plates were incubated at 28°C for 3 to 10 days.
Effect of extrinsic factors
Determination of Salt tolerance
Salinity tolerance of the isolates was examined on YMA containing different NaCl concentrations (0.5%, 1.5%, 2%, 5%, 7% to 10% (w/v)), after an incubation period of 7 days at 28°C.
Determination of pH tolerance
The ability of rhizobial isolates to grow in basic or acidic media was evaluated by inoculating them on YMB medium which pH values were adjusted to 4.8, 5.8, 6.8, 8.8 and 11 with NaOH or HCl13. The results were obtained after 7 days of incubation at 28°C.
Determination of temperature tolerance
Isolates were incubated at different temperatures (4 °C, 14 °C, 28 °C, 37 °C, 44 °C, 54 °C) to estimate the maximal and optimal growth temperatures of each isolate, and to evaluate their tolerance to different temperatures14. The bacterial growth was evaluated after 7 days of incubation.
Effect of intrinsic factors
Determination of heavy metals resistance
Resistance of tested rhizobial isolates was assessed against seven heavy metals using different concentration levels on YMA medium. Heavy metals stock solutions were filter sterilized (0.22 µm) and added to sterile agar medium to obtain concentrations which vary from 0.025 to 3 mmol l-1 of: AsNa2HPO4, CuSO4, NiCl2, K2Cr2O7, ZnSO4, 7H2O, Cd(No3)2, 7 H2O, and Pb(No3)2. The plates were inoculated with about 108cell/ml (10µl) and the bacterial growth was evaluated after 7 days of incubation at 28°C15. Isolates were considered resistant when growth occurred and sensitive when no growth was detected.
Determination of antibiotic resistance
Antibiotic resistance was tested for different concentrations (10-100 µg)16 of five antibiotics (Ampicillin, Kanamycin, Streptomycin, Tetracycline and Chloramphenicol) which were added to agar media as filter-sterilized aqueous stock solutions. 10µl (~108cell/ml) of each tested isolate were seeded, incubated at 28°C then their growth (Presence/absence) was scored after 7days for resistance/sensitivity17.
Evaluation of the symbiotic effectiveness of different rhizobial isolates
Isolates were tested for their symbiotic effectiveness by inoculating them separately to young plantlets of A. cyanophylla and A. horrida. Those plantlets were grown in plastic pots maintained in the greenhouse under the same conditions described above for the test of isolates authentication. Three replications were carried out for inoculated and non-inoculated plants. Plants were harvested six months after inoculation. The root nodules formed on each plant were counted; plant’s shoot and root dry matters were measured after drying at 70 ° C for 48 h. The average plant’s dry weight from three replicates inoculated with the same isolate was used to evaluate the relative efficiency (RE) expressed as the percentage of gain in aerial parts dry matter of inoculated plants (PDPi) compared to non inoculated control plants receiving KNO3 (PDCn):
RE = PDPi / PDCn x 100
Then aerial part’s nitrogen percentage and total nitrogen content were determined by using Kjedahl method18.
PCR amplification and sequencing of 16S rRNA gene
Total genomic DNA from bacterial isolates was extracted by a quick cell lysis in boiling water bath. The nucleotide sequences of the 16S rRNA gene (rDNA) were determined by direct sequencing of appropriate PCR products from each strain with the universal primers 8F (5’- AGA GTT TGA TCC TGG CTC AG -3’) and 1540R (5’- AAG GAG GTG ATC CAG CC-3’). Each 50 µl reaction contained 1 µl of the cell lysate (approx. 20ng DNA), 1,25 U of GoTaqR G2 DNA polymerase (Promega), 1X reaction buffer, 0,2 mMdNTPs, 0,15 µM of each primer, as well as 5% DMSO. The PCR protocol was set to: 4 min 95°C initial denaturation, 35 cycles of amplification: 1 min 95°C for denaturation, 1 min 55°C for annealing, 2 min 72°C for elongation; the final elongation step was set to 7 min. The nucleotide sequence of the PCR products was determined for both strands by Sanger sequencing. DNA sequences were compared to the GenBank database by basic local alignment search tool (BLAST) requests using the blast-n algorithm and optimization for highly similar sequences (megablast).
Numerical Analysis
The numerical analysis includes comparison of phenotypic characteristics between studied isolates taken in pairs. This method of analysis was used as a primary method of characterization and grouping of unidentified isolates19. A dendrogram performed with Ward method and Euclidian distance on the numerical analysis of phenotypic traits was obtained with XLSTAT software.
Statistical analysis
Means were compared using analysis of variance at P = 0.05. The used Software was Statgraphics Centurion XVI.
Morphologic characterization
All isolates tested were gram negative bacilli and most of them were fast-growing bacteria (63%) as indicated by the observation of a color variation from blue to yellow in YEM agar plates containing BBT, whereas, 21% were intermediate growing and 16% were slow growing.
Effect of extrinsic factors
Results of isolate’s tolerance to some environmental stress factors are presented in Table 2.
Table (2):
Effect of NaCl concentration, pH and temperature on rhizobial isolates.
| Isolates | NaCl % | pH | Temperature (C°) | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0.5 | 1 | 2 | 5 | 7 | 10 | 4.8 | 5.8 | 6.8 | 8.8 | 11 | 4 | 14 | 28 | 37 | 44 | 55 | |
| I1 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I2 | + | + | + | + | – | – | + | + | + | + | + | + | + | + | + | + | – |
| I3 | + | + | + | + | – | – | + | + | + | + | + | + | + | + | + | + | – |
| I4 | + | + | + | + | – | – | + | + | + | + | + | + | + | + | + | + | – |
| I5 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – | – | – |
| I6 | + | + | + | + | – | – | + | + | + | + | + | + | + | + | + | + | – |
| I7 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – | – |
| I8 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I9 | + | + | + | + | + | – | + | + | + | + | + | + | + | + | + | + | – |
| I10 | + | + | + | + | + | – | + | + | + | + | + | + | + | + | + | + | – |
| I11 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I12 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I13 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I14 | + | + | + | + | + | – | + | + | + | + | + | + | + | + | + | + | – |
| I15 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I16 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I17 | + | + | + | + | + | – | + | + | + | + | + | + | + | + | + | + | – |
| I18 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I19 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I20 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I21 | + | + | + | + | + | – | + | + | + | + | + | + | + | + | + | + | – |
| I22 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – | – |
| I23 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I24 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – | – |
| I25 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – | – |
| I26 | + | + | + | + | + | + | + | + | + | – | – | + | + | + | – | – | – |
| I27 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I28 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I29 | + | + | + | + | – | – | + | + | + | + | + | + | + | + | + | + | – |
| I30 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I31 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I32 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I33 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I34 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I35 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – | – |
| I36 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – |
| I37 | + | + | + | + | + | + | + | + | + | + | + | + | + | + | + | – | – |
| I38 | + | + | + | + | + | – | + | + | + | + | + | + | + | + | + | + | – |
All tested isolates grew in 5% NaCl, 87% were tolerant to 7% NaCl and 71% could persist to high salinity of 10% NaCl. Furthermore, the fast growing isolates showed higher salt tolerance than slow-growers.
All isolates tested were well adapted to high or low pH values: 100% were able to grow in variable pH values from 4.8 to 6.8, 97% survived at pH of 8.8 and 11.
It was also found that all isolates were able to survive at temperature ranging between 4 and 28°C. 95% of them presented a tolerance to a temperature of 37°C and 79% to 44°C. However, all isolates couldn’t tolerate high temperature (55°C).
Intrinsic resistance to heavy metals
Results of rhizobial isolate’s tolerance to different heavy metals concentrations are reported in Table 3. The majority of isolates were able to tolerate low concentrations of the tested heavy metals, but at higher concentrations bacterial growth was negatively affected. Indeed, the highest resistance was recorded for As with a growth rate of 100% at 3mmol l-1, followed by Cr, Zn and Pb (68%, 31% and 29% at 3mmol l-1). The resistance of isolates to different tested concentrations of cadmium and copper seems to be similar. These two metals are considered as the most growth inhibitors which induced a lethal effect on all isolates at concentrations of 2mmol l-1. Morever, the isolates growth is inhibited for nickel at a concentration of 3mmol l-1.
Table (3):
Metal’s concentraion (mmol l-1) |
AS |
Cr |
Cu |
Cd |
Zn |
Ni |
Pb |
|---|---|---|---|---|---|---|---|
0.250 |
100% |
100% |
65% |
63% |
100% |
84% |
100% |
0.500 |
100% |
100% |
60% |
60% |
100% |
71% |
100% |
1.0 |
100% |
94% |
48% |
23% |
94% |
55% |
100% |
2.0 |
100% |
94% |
0% |
0% |
92% |
29% |
95% |
3.0 |
100% |
68% |
0% |
0% |
31% |
0% |
29% |
Intrinsic resistance to antibiotics
Strong resistance was recorded for ampicillin, chloramphenicol, kanamycin and streptomycin, while the lowest resistance was observed for tetracycline. At a concentration of 10 µg / ml, the majority of the isolates showed good resistance with a growth rate of 71% for ampicillin and kanamycin and 73% for chloramphenicol. While at the same concentration of tetracycline and streptomycin, only 44 to 50% of isolates were able to grow in the presence of these antibiotics. At high concentrations (50 and 100 µg / ml), the inhibitory effect was highly marked especially in the case of three antibiotics, tetracycline, kanamycin and streptomycin. Resistance percentages of 50% and 39%, respectively, were noted for the other two antibiotics (ampicillin and chloramphenicol) (Figure 1).
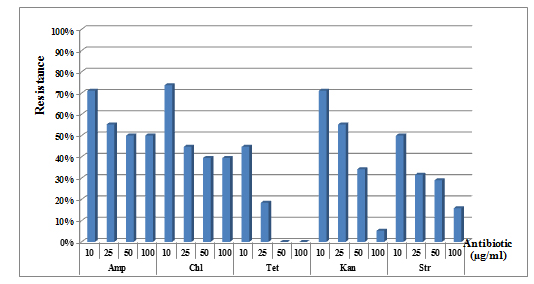
Fig. 1. Resistance of isolates to different concentrations of the tested antibiotics
Numerical analysis
For the numerical Analysis, the most discriminating characters of this analysis were grouped and presented in table 4. All phenotypic characteristics were treated by XLSTAT software to build the dendrogram represented in Figure 2. At 45% of similarity degree, the isolates were divided into two delimited clusters with an independent lineage represented by the strain I26, which presented a great capacity to tolerate the extrinsic stress factors (temperatures from 4-44 °C, 10% NaCl and pH 4.8 – 6.8) but was less resistant to intrinsic factors: sensitivity towards high concentrations of some heavy metals and antibiotics (Table 4).
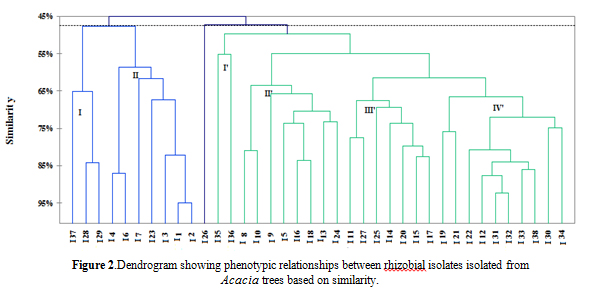
Fig. 2. Dendrogram showing phenotypic relationships between rhizobial isolates isolated from Acacia trees based on similarity
Table (4):
Results of phenotypic tests differentiating between isolates nodulating Acacia of different clusters formed by numerical analysis.
| Cluster 1 | Cluster 2 | I 26 | |
|---|---|---|---|
| Number of isolates | 10 | 27 | 1 |
| Host plants | A. saligna A. horrida |
A. saligna A. horrida |
A. saligna |
| Heavy metals resistance (mmol l-1) | |||
| As 0.25 | 10 | 27 | 1 |
| As 0.5 | 10 | 27 | 1 |
| As 1.0 | 10 | 27 | 1 |
| As 2.0 | 10 | 27 | 1 |
| As 3.0 | 10 | 27 | 1 |
| Cr 0.25 | 10 | 27 | 1 |
| Cr 0.5 | 9 | 26 | 1 |
| Cr 1.0 | 9 | 26 | 1 |
| Cr 2.0 | 9 | 17 | 1 |
| Cr 3.0 | 4 | 6 | 1 |
| Cu 0.25 | 7 | 17 | 0 |
| Cu 0.5 | 7 | 15 | 0 |
| Cu 1.0 | 6 | 10 | 0 |
| Cu 2.0 | 0 | 0 | 0 |
| Cu 3.0 | 0 | 0 | 0 |
| Cd 0.25 | 7 | 16 | 1 |
| Cd 0.5 | 7 | 15 | 1 |
| Cd 1.0 | 4 | 5 | 0 |
| Cd 2.0 | 0 | 0 | 0 |
| Cd 3.0 | 0 | 0 | 0 |
| Zn 0.25 | 10 | 27 | 1 |
| Zn 0.5 | 10 | 27 | 1 |
| Zn 1.0 | 10 | 25 | 1 |
| Zn 2.0 | 10 | 24 | 1 |
| Zn 3.0 | 5 | 7 | 0 |
| Ni 0.25 | 9 | 21 | 1 |
| Ni 0.5 | 8 | 18 | 0 |
| Ni 1.0 | 7 | 13 | 0 |
| Ni 2.0 | 4 | 5 | 0 |
| Ni 3.0 | 0 | 0 | 0 |
| Pb 0.25 | 10 | 27 | 1 |
| Pb 0.5 | 10 | 27 | 1 |
| Pb 1.0 | 10 | 27 | 1 |
| Pb 2.0 | 10 | 26 | 0 |
| Pb 3.0 | 1 | 9 | 0 |
| Antibiotic resistance (µg ml-1) | |||
| Amp 10 | 10 | 15 | 1 |
| Amp 25 | 10 | 10 | 1 |
| Amp 50 | 10 | 10 | 1 |
| Amp 100 | 10 | 9 | 1 |
| Chl 10 | 10 | 16 | 1 |
| Chl 25 | 8 | 7 | 1 |
| Chl 50 | 8 | 5 | 1 |
| Chl 100 | 8 | 5 | 1 |
| Tet 10 | 5 | 12 | 0 |
| Tet 25 | 1 | 5 | 0 |
| Tet 50 | 0 | 0 | 0 |
| Tet 100 | 0 | 0 | 0 |
| Kan 10 | 10 | 14 | 1 |
| Kan 25 | 10 | 9 | 0 |
| Kan 50 | 8 | 3 | 0 |
| Kan 100 | 2 | 0 | 0 |
| Str 10 | 9 | 9 | 1 |
| Str 25 | 9 | 4 | 0 |
| Str 50 | 9 | 2 | 0 |
| Str 100 | 5 | 1 | 0 |
| pH tolerance | |||
| 4.8 | 10 | 27 | 1 |
| 5.8 | 10 | 27 | 1 |
| 6.8 | 10 | 27 | 1 |
| 8.8 | 10 | 27 | 0 |
| 11 | 10 | 27 | 0 |
| Tolerance to temperature (°C) | |||
| 4 | 10 | 27 | 1 |
| 14 | 10 | 27 | 1 |
| 28 | 10 | 27 | 1 |
| 37 | 10 | 25 | 1 |
| 44 | 9 | 20 | 1 |
| 55 | 0 | 0 | 0 |
| Tolerance to NaCl (%) | |||
| 0.5 | 10 | 27 | 1 |
| 1 | 10 | 27 | 1 |
| 2 | 10 | 27 | 1 |
| 5 | 10 | 26 | 1 |
| 7 | 6 | 26 | 1 |
| 10 | 4 | 13 | 1 |
The first cluster contains ten isolates and the second one twenty seven isolates
The first cluster is formed by N2 fixing bacteria isolated from A. saligna and A. horrida and is divided at 47% of similarity degree in two subclusters. The subcluster’s I isolates (65% of similarity) tolerate different stress factors (salinity, pH and temperature), resist to high antibiotic concentrations except Tetracycline, but are less resistant to heavy metals especially cadmium and copper. The subcluster’s II isolates present a similarity level of 58% and show tolerance / resistance to both extrinsic and intrinsic stress factors excluding I4 and I6 which are sensitive for low concentrations of chloramphenicol.
The 2nd cluster which is more heterogeneous is divided in four subclusters at 48% of similarity. The subcluster I’contains two isolates with 55% of similarity, presenting sensitivity to high concentrations of antibiotics and heavy metals. These two isolates are tolerant to different abiotic stresses with the exception of the high concentration of NaCl (10%).
The subcluster II’ contains eight isolates with 63% of similarity, presenting sensitivity to high concentrations of antibiotics and moderate resistance to heavy metals except I18, and having a good tolerance to abiotic stress. The subcluster III’ consists of seven isolates (67% of similarity) tolerant to high salinity percentage (7-10%) and temperatures (37- 44°C), to acid/alkaline medium. The isolates of this subcluster present high resistance to heavy metals but are more sensitive to the majority of tested antibiotics.
The subcluster IV’ contains ten isolates, at 63% of similarity, generally tolerant to extrinsic stress factors, sensitive to low concentrations of cadmium, nickel and copper, and highly sensitive to the tested antibiotics.
Evaluation of the rhizobial isolate’s symbiotic effectiveness
The symbiotic effectiveness of the strains was evaluated based on shoot dry mass, nodule’s number and dry mass, and total nitrogen of the plant. The obtained results showed that acacia could be nodulated by all isolates tested. A large variability of the infective capacity of the isolates was demonstrated (Table 5). Indeed, even if the plants inoculums concentration of different isolates was the same (108 UFC/ml), the average nodules number formed by plant ranged from 31 to 81. The most infective isolate was I1 with 81±3.53 nodules formed by plant, while the less infective isolate (I23) was able to induce the formation of 31 nodules.
Table (5):
Plant growth parameters: (SDW: Shoot dry weight, RDW: root dry weight), nodule number (Nod N) and nodule dry weight (NDW) values of Acacia trees evaluated six months after plants inoculation.
isolate |
SDW g/plant |
RDW g/plant |
Nod N /plant |
NDW g/plant |
|---|---|---|---|---|
I1 |
15,32±0,55 |
9,97±0.74 |
81,3.53 |
0,365±0,04 |
I2 |
7.32±0.89 |
7.32±0.57 |
61±8.48 |
0,212±0.02 |
I3 |
7,25±1.15 |
4,84±1.71 |
37±3.53 |
0,132±0.01 |
I5 |
5,44±0,77 |
3,12±0.14 |
57±3.53 |
0,148±0.03 |
I6 |
8,72±0,33 |
6,01±1.22 |
56±5.65 |
0,143±0.01 |
I14 |
8,45±0.08 |
4,37±0.19 |
52±4.24 |
0,143±0.01 |
I16 |
5,54±1,15 |
2,89±0.20 |
56±5.65 |
0,129±0.004 |
I20 |
6,68±1.35 |
3,24±0.55 |
53±4.94 |
0,103±0.01 |
I23 |
3,44±0.26 |
1,05±0.09 |
31±4.94 |
0,124±0.02 |
I25 |
8,30±1.44 |
6,91±1.03 |
42±9.89 |
0,147±0.004 |
I28 |
13,25±0,55 |
8,75±0.28 |
67±4.24 |
0,141±0.02 |
NC |
12.02±1.13 |
5.82±0.24 |
0.00±0.00 |
0.00±0.00 |
C0 |
1.26±0.06 |
0.37±0.14 |
0.00±0.00 |
0.00±0.00 |
P-value |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
F |
38.23 |
30.4 |
41.82 |
36.51 |
Plant response to inoculation reveals different relative efficiencies (RE) (Figure 3). The results showed that the shoot dry weight (SDW) of plants inoculated with the isolates I1 and I28 exceeded the SDW of plants fertilized with mineral nitrogen (Table 5). In fact, shoot biomass, used as an indicator of relative effectiveness, indicated that isolate I1 was the most effective with an 127% dry biomass of the NC control followed by I28 with 110% of RE. The least effective isolate was I23 with only 29% of the dry biomass of the C0 control. The dry biomass of C0 control was 10% of the NC control.
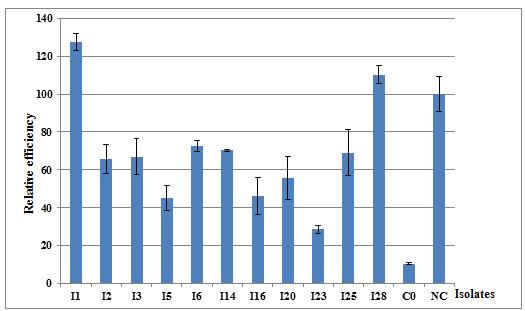
Fig. 3. Relative efficiency of isolates nodulating acacia evaluated six months after plants inoculation
Regarding the performance in root dry weight (RDW), the results showed a significant difference between inoculated isolates, the highest RDW was detected in I1 while the lowest RDW was showed by I23 with values of 9.97 and 1.05 g/plant respectively (Table 5).
The percentage of total nitrogen derived from atmospheric N2 and fixed by acacia was evaluated in the rhizobium inoculation field trial. In fact, it varied from 2.1% for isolate I20 to 4.9% for the isolate I23, with significant difference between the different isolates (P= 0.000) (Figure 4).
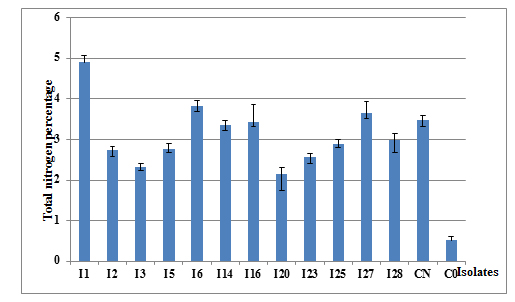
Fig. 4. Total nitrogen percentage in the aerial parts of acacia inoculated with different isolates of Rhizobium, evaluated six months after inoculation. (C0 = nitrogen-free control, NC nitrogen control fertilized with KNO3)
The shoot’s total nitrogen content showed also significant differences between the studied isolates (P = 0.000) (Figure 5). Plants inoculated with the isolate I1 accumulated more nitrogen in their aerial parts (0.75 g/plant) followed by the ones inoculated with the isolates I6and I28which accumulated 0.34 and 0.38 g/plant respectively.
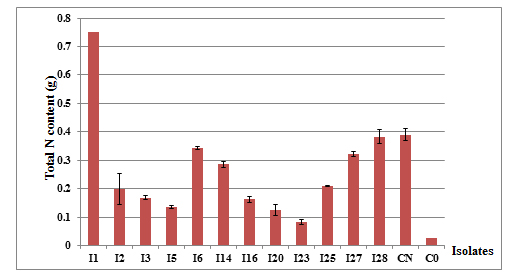
Fig. 5. Total nitrogen content in the aerial parts of acacia inoculated with different isolates of Rhizobium, evaluated six months after inoculation
These results showed not only a differential effect of the studied isolates on acacia’s growth but also showed a significant biodiversity of Rhizobium bacteria isolated from Moroccan soils.
Molecular classification of the plant-growth promoting isolates
The sequencing of 16S rRNA genes of the tested isolates was repeatedly reported as the preferred method in order to characterize microbial communities, found in the plant rhizosphere20-21. According to the sequences similarity, local rhizobial isolates are identified as members Rhizobium and Phyllobacterium genera (Table 6).
Table (6):
Sequence analysis of 16S rDNA for six rhizobial strains nodulating Acacia sp..
Isolate |
Sequence (bp) |
Accession (no) |
Homology to the reference strains |
Identity (%) |
|---|---|---|---|---|
I1 |
471 |
Rhizobium pusenseMB17a |
98 |
|
I3 |
537 |
Rhizobium sp.Moz90 |
99 |
|
I5 |
603 |
Rhizobium sp. LAR-14 |
99 |
|
I6 |
650 |
Rhizobium sp. SAB12b |
99 |
|
I16 |
549 |
Rhizobium sp. DG22 |
99 |
|
I28 |
491 |
LC133652.1 |
Phyllobacterium sp. JCM 28305 |
98 |
This study provides phenotypic characterization of rhizobial isolates from different moroccan Acacia tree species. The majority of isolates were fast growing. The response of the various isolates against different environmental stress was evaluated in order to show their preliminary variability based on phenotypical characteristics.
In the present study, all isolates grew in medium salt concentrations (5.0%) and the majority of them (71%) could persist to high salinity reaching 10%. This NaCl range tolerance agree with previous reports showing that Rhizobium isolates from arid and saline area’s various collections are highly salt-tolerant and withstand at high NaCl levels up to 5-10 %3. Furthermore, the fast growing isolates showed higher salt tolerance than slow-growers, which is consistent with previous studies13; 22. However, other studies revealed that salt tolerance is not correlated with growth rate23 but is correlated with other physiological and biochemical mechanisms24. Furthermore, salinity is one of the major factors restricting the symbiotic nitrogen fixation25 and nodulation in legumes. Therefore, these isolates may be performant candidates for inoculation in saline soils frequently observed in some irrigated areas in Morocco. Especially because salinity is among the main factors inducing desertification and lands degradation26.
The results showed that most of the isolates in this study could grow within a wide temperature range (4°C to 44°C) which is concordant with Maâtallah et al27 finding that Rhizobium could grow within a wide temperature range (4°C to 44°C). The growth inhibition observed for all the tested isolates at high temperature (55°C) agree with Zahran et al28 and Fentahun et al29 findings. Furthermore, Rhizobia are known to be mesophilic and have optimum culture growth temperatures in the range of 28-31 °C30. Maximum temperature degrees (Tmax) for free-living rhizobia ranged between 35-45 °C 19; 28. Even if the rhizobia grow at high temperatures it does not mean that it is an efficient N2-fixer29. In fact, the exposure to high temperatures can lead to loss of symbiotic plasmid and consequently the loss of the bacterial infective capacity31. So, high soil temperature can generate a major problem for biological nitrogen fixation in leguminous because the high temperature may control the bacterial infection in host plant.
According to our results, the isolates studied are generally slightly tolerant to acidity at pH 4.8-5.8, and mostly tolerant to alkalinity. This finding is concomitant with results of previous studies which indicated that Rhizobia were able to grow at a wide pH range in soil conditions32-34 and even at pH 12 in agar culture35. The isolates which can survive on a wide pH range are candidates for further strain improvement to highly acidic or alkaline conditions. Several authors have reported the sensitivity of slow-growing rhizobial strains to higher acid pH than fast growing rhizobia36-37. Other studies showed that fast-growing rhizobia are generally more sensitive to acidity than bradyrhizobia11. Similarly, Marsudi et al.38 reported that the slow growing Bradyrhizobium strains isolated from Acacia saligna were found to be sensitive to alkaline pH and tolerant to acid pH.
However, in our study, no obvious relationship between the isolates growth rate and their tolerance to different pH range was found. Mohamed et al.39 also deducted in their results that there is no correlation between pH tolerance of strains and their growth’s rate. Regarding the intrinsic resistance to heavy metals, several studies showed that they might have been toxic to rhizobia when present in soil at moderate to high concentrations.
In our study, strong resistance has been recorded, on one hand, for arsenate, chrome, plumb and zinc. On the other hand, low resistance has been recorded for nickel, copper and cadmium. This result is consistent with the literature that showed that Rhizobium could withstand to high concentrations of arsenate, zinc and plumb40 and was less resistant to high concentrations of cadmium and cooper41. The selection of isolates resistant to heavy metals presents actually great practical interest for bio-remediation investigations. Abd-Alla et al.42 have shown that an excess of Cd, Zn, and Cu in soil can have an adverse effect for symbiotic microorganisms as well as for the symbiosis establishment. Similarly, Sepehri et al.43 revealed that Cd can affect the symbiotic properties of Sinorhizobium meliloti strains and therefore S. meliloti-alfalfa symbiosis.
As with Cd, the tested isolates were found to be more sensitive to Cu than other metals, which is consistent with previous studies showing that high Cu concentrations may be toxic to soil microorganisms by affecting their structural diversity and their tolerance to metals44-45.
Antibiotic resistance of bacterial strains isolated from acacia was tested against different antibiotics. In general, the isolates showed strong resistance to Ampicillin, Chloramphenicol and Kanamycin. However, the majority of the tested isolates were sensitive to high concentrations of Streptomycin, Kanamycin and Tetracycline. This is in agreement with the results of Gauri et al.46 and Bakhoum et al.16. Similar results were also found by Hilali et al. (2006)31 in a rhizobacterial population nodulating lupine in Morocco in which tetracycline, streptomycin and kanamycin were considered as inhibitors of the plasma membrane and protein synthesis, and which had the most radical effect on the isolates growth.
Graham et al.30 reported that the inhibitory effect of an antibiotic varies with its nature, its concentration in the medium, and the degree of inhibition varies from one species to another and from one strain to another. In similar studies, Kanamycin has been reported to be growth’s inhibitor of some strains of Bradyrhizobium nodulating lupine47 and some strains nodulating different Acacia’s species23. However, in our study, tetracycline was recorded as the most detrimental for the isolates growth. The selection of isolates with multiple resistance to different antibiotics is very important because this property can be used as a great marker for strain’s identification and study of their diversity48.
Previous studies mentioned that the strains associated with Acacia spp. in Africa can belong to Rhizobium, Agrobacterium, Ensifer, Mesorhizobium and Bradyrhizobium49-52. In this study, the identified strains isolated from Acacia tree was found to belong to Rhizobium and Phyllobactrium genus. In fact, it has been reported that Phyllobacterium genus was able to nodulate some Acacia species53-54.
The six strains were isolated from the same species (Acacia saligna), and were found to belong to different rhizobial species. This is consistent with previous studies which reported that changes in the nature of rhizobia associated with the same legume can be affected by the different geographic and meteorological characteristics of each sampling site55-56.
Also, it was found that I1 and I28 isolated from the same geographical region are classified in two different genuses. However, a previous study showed that legume plants in the same geographic region had one rhizobial species in common57.
All tested isolates were able to infect their host plant and to fix atmospheric nitrogen leading to plant shoot production above the noninoculated and nitrogen-free controls. In fact, the results of the experiments showed that acacia’s inoculation by different isolates have significant effects on the level of infection of A. saligna (measured by the nodules number formed).
The results of isolate’s symbiotic effectiveness showed not only a differential effect of the studied isolates on Acacia seedlings growth but also a significant diversity of Rhizobial isolates in Moroccan soils. Indeed, I1, I2, I3 and I28 were all isolated from the same geographical region (Oujda), but induced significantly different plant dry matter yields.
The results of this study also showed differences in N2– fixation among the tested Acacia plants both for the %Ndfa and the total N fixed.
Several previous works also showed that selection and inoculation of effective rhizobium increased nodulation, shoot dry weight, height and total shoot nitrogen content58-59.
These results are extremely encouraging for the use of acacias in land’s rehabilitation and/or restoration projects. Rhizobial isolates identified as effective in the greenhouse trial can be useful in field inoculation trials to improve soils fertility and transfer nitrogen to associated crops by symbiotic interaction Acacia/Rhizobia.
This study provides phenotypic and symbiotic characterization of rhizobial isolates from different Moroccan Acacia tree species and showed their large physiological and biochemical diversity. Most of them were fast growing and their response to different environmental stress showed a variable resistance against stress factors, namely, temperature, pH, salinity and also to antibiotics and heavy metals, which allowed us to make a preliminary classification and characterization of these isolates based on their phenotypic similarities. Moreover, this analysis enabled to bring out a phenotypic groups containing intra and inter- cluster diversity. This phenotypic diversity and the high symbiotic efficiency of the tested Rhizobial strains can help us to precisely select the right candidates for any inoculation test according to requirements of different environmental conditions considered in order to improve and to enhance plant’s crops and nitrogen fixation in semi-arid and arid areas.
- Ögütçü, H., Algur, Ö.F., Elkoca, E., Kantar, F. The Determination of Symbiotic Effectiveness of Rhizobium Strains Isolated from Wild Chickpeas Collected from High Altitudes in Erzurum. Turk. J. Agric. For., 2008: 32: 241–248.
- Farissi, M., Bouizgaren, A., Faissal, A., Mustapha, M., Ghoulam, C. Isolation and screening of rhizobial strains nodulating alfalfa for their tolerance to some environmental stresses. Pacesetter. J. Agric. Sci. Res., 2014; 2: 9–19.
- Gal, S.W., Choi, Y.J. Isolation and Characterization of Salt Tolerance Rhizobia from Acacia Root Nodules. Agric. Chem. Biotechnol., 2003; 46: 58–62.
- Berrada, H., Fikri-Benbrahim, K. Taxonomy of the Rhizobia: Current Perspectives. Br. Microbiol. Res. J., 2014; 4: 616–639.
- Weir, B. The Current Taxonomy of Rhizobia. 2008.
- Diabate, M., Munive, A., De Faria, S.M., Ba, A., Dreyfus, B., Galiana, A. Occurrence of nodulation in unexplored leguminous trees native to the West African tropical rainforest and inoculation response of native species useful in reforestation. New. Phytol., 2005; 166: 231–239. doi: 10.1111/j.1469-8137.2005.01318.x
- Teixeira, H., Rodríguez-Echeverría S. Identification of symbiotic nitrogen-fixing bacteria from three African leguminous trees in Gorongosa National Park. Syst. Appl. Microbiol., 2016; 39: 350–358. doi: 10.1016/j.syapm.2016.05.004
- FikriBenbrahim, K., Ismaili, M. Interactions in the Symbiosis of Acacia saligna with Glomusmosseae and Rhizobium in a Fumigated and Unfumigated Soil .Arid. Land. Res. Manag., 2002; 16: 365–376.
- Vincent, J.M. (ed): A manual for practical study of root nodule bacteria, International Biological Programme: Blackwell Scientific, 1970; pp 164.
- Beck, D.P., Meteron, L.A., Afandi, F.. Practical Rhizobium Legume technology manual, Technical Manual: International Center for Agricultural Research in the Dry Areas 1993; 19: pp 389.
- Jordan, D. Family III Rhizobiaceae Conn. 1938-254. Bergey’sMannual of Systematic Bacteriology. The Williams and Wilkins Co. (ed) Baltimor: Krieg N.R., Holt, J.G., 1984; 1: 235–244.
- Fikri-Benbrahim, K., Chraibi, M., Lebrazi, S., Moumni, M., Ismaili, M. Phenotypic and Genotypic Diversity and Symbiotic Effectiveness of Rhizobia Isolated from Acaciasp. Grown in Morocco. J. Agric. Sci. Technol., 2017; 19: 201–216.
- Küçük, Ç.,Kivanç, M., Kinaci, E. Characterization of Rhizobium sp. Isolated from Bean. Turk J. Biol., 2006; 30: 127–132.
- Hung, M.H., Bhagwath, A.A., Shen, F.T., Devasya, R.P., Young, C.C.. Indigenous rhizobia associated with native shrubby legumes in Taiwan. Pedobiologia. (Jena). 2005; 49: 577–584.doi: 10.1016/j.pedobi.2005.06.002
- Cevheri, C., Küçük, Ç.,Çetin, E.. Fungicide, antibiotic, heavy metal resistance and salt tolerance of root nodule isolates from Vicia palaestina. Afr. J. Biotechnol., 2011; 10: 2423–2429. doi: 10.5897/AJB10.1291
- Bakhoum, N., Le Roux, C., Diouf, D., Kane, A., Ndoye, F., Fall, D., Duponnois, R., Noba, K., Sylla, S.N., Galiana, A. Distribution and Diversity of Rhizobial Populations Associated with Acacia senegal (L.) Willd. Provenances in Senegalese Arid and Semiarid Regions. Open. J. For., 2014; 4: 136–143. doi: 10.4236/ojf.2014.42019
- Kenenil, A., Assefa, F., Prabu, P.C. Characterization of Acid and Salt Tolerant Rhizobial Strains Isolated from Faba Bean Fields of Wollo, Northern Ethiopia. J. Agric. Sci. Technol., 2010; 12: 365–376.
- Nelson, D.W., Sommers, L. Determination of total nitrogen in material. Agron. J., 1973; 65: 109–112.
- Zhang, X., Harper, T., Karsisto, M., Lindstrom, K. Diversity of Rhizobium Bacteria Isolated from the Root Nodules of Leguminous Trees. Int. J. Syst. Bacteriol., 1991; 41: 104–113.
- Majeed, A., Abbasi, M.K., Hameed, S., Imran, A., Rahim N. Isolation and characterization of plant growth-promoting rhizobacteria from wheat rhizosphere and their effect on plant growth promotion. Front. Microbiol., 2015; 6:198. 10.3389/fmicb.2015.00198
- Verma, P., Yadav, A.N., Khannam, K.S., Kumar, S., Saxena, A.K., Suman, A.. Molecular diversity and multifarious plantgrowth promoting attributes of Bacilli associated with wheat (Triticum aestivum L.) rhizosphere from six diverse agro-ecological zones of India. J. Basic Microbiol., 2016; 56:44–58.
- Berrada, H., Nouioui, I., Houssaini, M.I., El Ghachtouli, N., Gtari, M., FikriBenbrahim, K. Phenotypic and genotypic characterizations of rhizobia isolated from root nodules of multiple legume species native of Fez, Morocco. Afr. J. Microbiol. Res., 2012; 6: 5314–5324. doi: 10.5897/AJMR11.1505
- Zerhari, K., Aurag, J., Khbaya, B., Kharchaf, D., Filali-Maltouf, A. Phenotypic characteristics of rhizobia isolates nodulating Acacia species in the arid and Saharan regions of Morocco. Lett. Appl. Microbiol., 2000; 30: 351–357.
- Gouffi, K., Pica, N., Pichereau, V., Blanco, C. Disaccharides as a New Class of Non accumulated Osmoprotectants for Sinorhizobium meliloti. Appl. Environ. Microbiol., 1999; 65: 1491–1500.
- Shetta, N.D., Al-Shaharani, T.S., Abdel-Aal, M. Identification and Characterization of Rhizobium Associated with Woody Legume Trees Grown under Saudi Arabia Condition. Am. J. Environ. Sci. 2011; 10: 410–418.
- Fikri-Benbrahim, K., Ismaili, M., Fikri-Benbrahim, S., Tribak, A. Synthèse Problèmes de dégradation de l’environnement par la désertification et la déforestation/ : impact du phénomène au Maroc. Science et changement splanétaires/Sécheresse., 2004; 15: 307–320.
- Maâtallah, J., Sanjuan, J., Lluch, C. Phenotypic characterization of rhizobia isolated from chickpea (Cicerarietinum) growing in Moroccan soils. Agron., 2002; 22: 321–329. doi: 10.1051/agro:2002013
- Zahran, H., Abdel-Fattah, M., Yasser, M., Mahmoud, A., Bedmar, E. Diversity and environmental stress responses of rhizobial bacteria from Egyptian grain legumes. Aust. J. Basic Appl. Sci., 2012; 6: 571–583.
- Fentahun, M., Akhtar, M.S., Muleta, D., Lemessa, F. Isolation and characterization of nitrogen deficit Rhizobium isolates and their effect on growth of haricot bean. Afr. J. Agric, Res., 2013; 8: 5942–5952. doi: 10.5897/AJAR2012.6690
- Graham, P., Stress tolerance in Rhizobium and Bradyrhizobium, and nodulation under adverse soil conditions. Can. J. Microbiol.1992; 38: 475–484.
- El Hilali, I. La symbiose Rhizobium-lupin: biodiversité des microsymbiontes et mise en evidence d’une multiinfection nodulaire chez Lupinus luteus. Dissertation, Rabat University Mohammed V, 2006; pp 231.
- Küçük, Ç., K1vanç, M. Preliminary characterization of Rhizobium strains isolated from chickpea nodules. Afr. J. Biotechnol.; 2008; 7: 772–775.
- Youseif, S.H., Abd El-Megeed, F.H., Ageez, A., Mohamed, Z.K., Shamseldin, A., Saleh, S.A. Phenotypic characteristics and genetic diversity of rhizobia nodulating soybean in Egyptian soils. Eur. J. Soil. Biol., 2014; 60: 34–43. doi: 10.1016/j.ejsobi.2013.10.008
- Hannane, F.Z., Kacem M., Kaid-harche, M. Preliminary characterization of slow growing rhizobial strains isolated from Retama monosperma (L.) Boiss.root nodules from Northwest coast of Algeria. Afr. J. Biotechnol., 2016; 15: 854–867. doi: 10.5897/AJB2016.15226
- Surange, S., Wollum, A., Kumar, N., Nautiyal, C. Characterization of Rhizobium from root nodules of leguminous trees growing in alkaline soils. Can. J. Microbiol., 1997; 43: 891–894.
- Zablotowicz, R.M., Focht, D.D. Physiological characteristics of cowpearhizobia: evaluation of symbiotic efficiency in Vignaun guiculata. Appl. Environ. Microbiol., 1981; 41(3): 679-685.
- Temesgen, D., Assefa, F. Nodulation pattern and biodiversity of rhizobia of some important leguminous trees and shrubs of Ethiopia. Ethiop. J. Biol. Sci., 2010; 9(2): 153-171.
- Marsudi, N.D.S., Glenn, A.R., Dilworth, M.J. Identification and characterization of fast and slow growing root nodule bacteria from South Western Australian soils able to nodulate Acacia saligna. Soil. Biol. Biochem., 1999; 31: 1229-1238.
- Mohamed, S.H., Smouni, A., Neyra, M., Kharchaf, D., Filali-Matouf, A. Phenotypic characteristics of root-nodulating bacteria isolated from Acacia spp. grown in Libya. Plant. Soil. 2000; 224: 171-183.
- Carrasco, J.A., Armario, P., Pajuelo, E., Burgos, A., Caviedes, M.A., López, R., Chamber, M.A., Palomares, A.J. Isolation and characterisation of symbiotically effective Rhizobium resistant to arsenic and heavy metals after the toxic spill at the Aznalco´llar pyrite mine. Soil Biol. Biochem., 2005; 37: 1131–1140. doi: 10.1016/j.soilbio.2004.11.015
- Pajuelo, E., Dary, M., Palomares, A., Rodriguez-Llorente, I.D., Carrasco, J., Chamber, M. Biorhizoremediation of Heavy Metals Toxicity Using Rhizobium-Legume Symbioses. Biological Nitrogen Fixation: Towards Poverty Alleviation through Sustainable Agriculture 2008; 101–104.
- Abd-Alla, M.H., El-Enany, A.W.E., Nafady, N.A., Khalaf, D.M., Morsy, F.M. Synergistic interaction of Rhizobium leguminosarum bv. viciae and arbuscular mycorrhizal fungi as a plant growth promoting biofertilizers for faba bean (Viciafaba L.) in alkaline soil. Microbiol. Res., 2014; 169(1): 49-58.
- Sepehri, M., Rastin, N.S., Rahmani, H.A., Alikhani, H. Effects of soil pollution by cadmium on nodulation and nitrogen fixation ability of native strains of Sinorhizobium meliloti. JWSS-IsfahanUniversity of Technology, 2006; 10(1): 153-163.
- Dell’Amico, E., Mazzocchi, M., Cavalca, L., Allievi, L., Andreoni, V. Assessment of bacterial community structure in a long-term copper-polluted ex-vineyard soil. Microbiol. Res., 2008; 163(6): 671-683.
- Abdel-lateif, K.S. Isolation and characterization of heavy metals resistant Rhizobium isolates from different governorates in Egypt. Afr. J. Biotechnol.,2017; 16(13): 643-647.
- Gauri, A.S., Bhatt, R., Pant, S., Bedi, M., Naglot, A. Characterization of Rhizobium isolated from root nodules of Trifolium alexandrinum. J. Agric. Technol., 2011; 7: 1705–1723.
- Madrzak, C.J., Golinska, B., Kroliczak, J., Pudelko, K., Lazewska, D., Lampka, B., Sadowsky, M.J. Diversity among Field Populations of Bradyrhizobium japonicum in Poland. Appl. Environ. Microbiol., 1995; 61(4): 1194-1200.
- Sawada, Y., Miyashita, K., Yokoyama, T. Diversity within serogroups of Japanese isolates of Bradyrhizobium japonicum as indicated by intrinsic antibiotic resistance. Soil. Sci. Plant. Nut., 1990; 36: 501–504. doi: 10.1080/00380768.1990.10416919
- De Lajudie P., Willems, A., Pot, B., Dewettinck, D., Maestrojuan, G., Neyra, M., Collins, M., Dreyfus, B., Kersters, K., Gillis, M. Polyphasic Taxonomy of Rhizobia: Emendation of the Genus Sinorhizobium and Description of Sinorhizobium meliloti comb. nov., Sinorhizobium saheli sp. nov., and Sinorhizobium teranga sp. nov. Int. J. Syst. Bacteriol., 1994; 44: 715–733.
- Khbaya, B., Neyra, M., Normand, P., Zerhari, K., Filali-Maltouf, A. Genetic Diversity and Phylogeny of Rhizobia That Nodulate Acacia spp. in Morocco Assessed by Analysis of rRNA Genes. Appl. Environ. Microbiol., 1998; 64: 4912–4917.
- Odee, D.W., Haukka, K., Mcinroy, S.G., Sprent, J.I., Sutherland, J.M., Young, J.P.W. Genetic and symbiotic characterization of rhizobia isolated from tree and herbaceous legumes grown in soils from ecologically diverse sites in Kenya. Soil. Biol. Biochem., 2002; 34: 801–811.
- Amrani, S., Noureddine, N.E., Bhatnagar, T., Argandoñ, M., Nieto, J.J., Vargas, C. Phenotypic and genotypic characterization of rhizobia associated with Acacia saligna (Labill.) Wendl. in nurseries from Algeria. Syst. Appl. Microbiol., 2010; 33: 44–51. doi: 10.1016/j.syapm.2009.09.003
- Hoque M.S., Broadhurst, L.M., Thrall, P.H. Genetic characterization of root-nodule bacteria associated with Acacia salicina and A. stenophylla (Mimosaceae) across south-eastern Australia. Int. J. Syst. Evol. Microbiol., 2011; 61: 299–309. doi10.1099/ijs.0.021014-0
- Crisóstomo, J.A., Rodríguez-Echeverría, S., Freitas, H. Co-introduction of exotic rhizobia to the rhizosphere of the invasive legume Acacia saligna, an intercontinental study. Appl. Soil. Ecol., 2013; 64: 118–126.
- Gu, C., Wang, E., Sui, X., Chen, W.F., Chen, W.X. Diversity and geographical distribution of rhizobia associated with Lespedeza spp. in temperate and subtropical regions of China. Arch. Microbiol., 2007; 188: 355–365.
- Lafay, B., Burdon, J.J. Molecular Diversity of Legume Root-Nodule Bacteria in Kakadu National Park, Northern Territory, Australia. PLoS One., 2007; 2(3): e277. doi: 10.1371/journal.pone.0000277
- Russell, J., Moreau, C., Goldman-Huertas, B., Fujiwara, M., Lohman, D., Pierce, N. Bacterial gut symbionts are tightly linked with the evolution of herbivory in ants. Proc. Natl. Acad. Sci., 2009; 106: 21236–21241.
- Burdon, J., Gibson, A., Searle, S., Woods, M., Brockwell, J. Variation in the effectiveness of symbiotic associations between native rhizobia and temperate Australian Acacia: within species interactions. J. Appl. Ecol. 1999; 36: 398–408.
- Yohannes, Y., Assefa, F. Phenotypic characteristics of root nodule bacteria and arbuscular mycorrhizal fungi infecting Acacia polyacantha growing in Ghibe wooded grasslands. Ethiop. J. Nat. Sci., 2007; 9(1): 123-139.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


