ISSN: 0973-7510
E-ISSN: 2581-690X
The coronavirus disease (COVID-19) pandemic has expanded the use of chlorhexidine digluconate, a biocide frequently used in hospitals, to inhibit the spread of infection. Genes responsible for resistance against the quaternary ammonium compound qac in Staphylococcus aureus isolates have been shown to confer tolerance to a number of biocidal chemicals, including chlorhexidine. The aim of this study was to determine the occurrence of antiseptic resistance genes (qacA/B and qacC) in clinical isolates of methicillin-susceptible (MSSA) and methicillin-resistant Staphylococcus aureus (MRSA). The study also aimed to investigate the association between the presence of the mecA, qacA/B, and qacC genes in MRSA isolates and the susceptibility of the isolates to chlorhexidine to evaluate its future use in the Theodor Bilharz Research Institute (TBRI) hospital, following the Centers for Disease Control and Prevention recommendations for patients with MRSA. S. aureus isolates (n = 100) were collected from inpatients and outpatients at TBRI. A minimal inhibitory concentration of chlorhexidine was also detected. Polymerase chain reaction was used to detect the mecA, qacA/B, and qacC genes. The results revealed that 84% of S. aureus isolates were MRSA. MRSA (61.9%) and MSSA (68.8%) isolates were susceptible to chlorhexidine. The qacA/B gene was more dominant, being detected in 34%, while qacC was detected in only 5% of S. aureus isolates. All S. aureus isolates with reduced susceptibility to chlorhexidine harbored either the qacA/B or qacC genes. The clinical use of chlorhexidine may continue to increase, emphasizing the significance of continuous caution underlining the emergence of new clones with reduced susceptibility and avoiding antiseptic misuse.
Staphylococcus aureus, MRSA, MSSA, Chlorhexidine, qacA/B ,qacC genes, Antiseptic Resistance
Methicillin-resistant Staphylococcus aureus (MRSA) is a nosocomial infection-causing bacterium that can spread extensively among hospital patients and healthcare providers. Infection control strategies used to prevent MRSA infections in hospitals include hand hygiene, along with disinfection of contaminated areas, instruments, and colonized patients. Many antiseptics are used to control MRSA, although abuse and sublethal doses of antiseptics have resulted in the occurrence of MRSA that is less susceptible to antiseptics.1 An overall increase in the use of antiseptics that occurred after the coronavirus disease 2019 (COVID-19) pandemic has aggravated this problem of the decreased susceptibility of MRSA to antiseptics.2
Chlorhexidine is a cationic biguanide that is used as the main antiseptic for infection control in hospitals. This compound is used to decolonize surfaces, bathe patients in the intensive care unit, piercing sites of central venous catheters, preoperative preparation, and hand asepsis.3 Chlorhexidine kills bacteria by damaging the cytoplasmic membrane and outer cell layers, allowing intracellular contents to escape.4
Clinical isolates of Staphylococcus with mechanisms resistant to chlorhexidine have been reported.3 The qac genes encode for these mechanisms, which are responsible for resistance to organic cations via proton motive force-dependent multidrug efflux. In staphylococci, six plasmid-encoded qac efflux pumps have been identified that belong to two main protein families: the small multidrug resistance family encoded by qacC, qacG, qacH, and qacJ, and the major facilitator superfamily, including qacA and qacB genes.5
Resistance to monovalent and divalent organic cations is determined by qacA, whereas resistance to monovalent organic cations is governed by a closely related qacB determinants.5 The qacB is virtually identical to qacA, except for seven to nine bases that are difficult to distinguish using simple Polymerase chain reaction (PCR).1,3 qacC confers resistance to chlorhexidine and codes for a much smaller membrane protein than the qacA/B gene family.3 MRSA transmission in the intensive care unit can be prevented with a chlorhexidine-based surface antiseptic program; however, strains bearing qacA/B genes may be unaffected or spread more promptly.6
This study aimed to detect the presence of resistance genes (qacA/B and qacC) in patients with methicillin-susceptible S. aureus (MSSA) and MRSA clinical isolates. We also studied the correlation between existence of mecA gene and different antiseptic resistance genes (qacA/B and qacC) in MRSA isolates and determined the association between the occurrence of qacA/B and qacC genes and susceptibility to chlorhexidine to evaluate its future use at Theodor Bilharz Research Institute (TBRI) hospital to follow Centers for Disease Control and Prevention (CDC) recommendations for hand hygiene and body wash for patients with MRSA.
Clinical Isolates
This study was conducted with 100 clinical S. aureus isolates obtained from various clinical samples. Specimens were processed at the microbiology laboratory of TBRI between January 2017 and November 2020. S. aureus isolates were isolated from 48 urine (48%), 28 (28%) from blood, 16 (16%) from pus, and 8 (8%) from sputum specimens. All specimens included in the study were archived and codes were used, instead of the patient names. The patients’ informed consent was waived as all patient data were anonymized. The study protocol was reviewed and approved by the TBRI institutional review board under Federal Wide Assurance (FWA00010609) and the work was carried out in accordance with the Code of Ethics of the World Medical Association (Declaration of Helsinki) for Experiments in Humans and its later amendments (GCP guidelines) or comparable ethical standards. The total number of specimens received at the microbiology lab during this period was 5172; No organisms were detected in 1188 specimens. Out of remaining 3984 specimens, Gram negative was detected in 2880 & Gram positive in 1104, among them, 900 were staphylococci, 100 S. aureus and 800 Methicillin-resistant coagulase negative Staphylococci (MRConS). S. aureus was identified using traditional methods, at the species level (morphology of the colony, gram staining, and coagulase and catalase tests). Mannitol fermentation was examined using mannitol salt agar (Bio-Rad). Confirmation of the identified S. aureus isolates was carried out using the Vitek 2 compact automated system that uses fluorogenic methodology for organism identification and a turbidimetric method for susceptibility testing using a 64-well card that provides results in 5–8 h (BioMerieux, France). Of the 100 S. aureus isolates, 84 were MRSA and 16 were MSSA. All S. aureus isolates were kept in glycerol broth at –70°C and then subcultured onto blood agar for 48 h, prior to further investigations, where only 84 and 16 isolates of MRSA and MSSA, respectively, could be elicited with a total of 100 isolates. The MRSA ATCC 43300 standard strain was used as the control.
Detection of mecA-mediated Oxacillin Resistance
Disc diffusion tests were carried out on Mueller–Hinton agar (MAST Diagnostics, UK), and cefoxitin (30 μg) disc were applied. After 16–18 h of incubation at 33–35°C, the zone of inhibition was measured. Clinical Laboratory Standards Institute (CLSI) criteria were used to make the interpretations: mecA-positive was ≤ 21 mm and mecA-negative was ≥ 22 mm.7 S. aureus isolates that were mecA-positive were reported to be oxacillin-resistant.
Susceptibility of MRSA Strains to Antimicrobial Agents
Antimicrobial susceptibility testing (AST) for different antibiotics was performed using the Vitek 2 compact system. The minimum inhibitory concentrations (MICs) were detected using the AST cards of Vitek 2 compact system for oxacillin (OXA), cefoxitin (FOX), ciprofloxacin (CIP), moxifloxacin (MOX), vancomycin (VA), trimethoprim/sulfamethoxazole (SXT), gentamicin (CN), clindamycin (DA), erythromycin (E) and linezolid (LZD). The steps were completed following the manufacturer’s instructions, and the results were read after 10–20 h (BioMerieux, France). MIC results were obtained and CLSI guidelines were used to interpret the findings.7
Broth Microdilution Test for Chlorhexidine
The CLSI and Manual of Clinical Microbiology recommend a broth microdilution procedure using 96-well microtiter plates with two-fold serial dilutions to determine the MIC of chlorhexidine. The MIC is defined as the lowest concentration of antimicrobial agent that prevents observable growth following a 20-h incubation period at 37°C in Mueller–Hinton broth.7
The isolates were considered resistant to chlorhexidine when the MIC was ≥ 4 μg/mL,3,8 and the susceptibility breakpoint for chlorhexidine applied in this study was derived from the MIC epidemiological cut-off, a standard approach employed when validated breakpoints are not available, allowing our results to be compared to those in published reports.3,8,9 An increase in MIC values to chlorhexidine does not mean “resistance,” because this agent can be used at high concentrations without causing toxicity. Thus the terms “reduced susceptibility” or “increased tolerance” are better used for pathogens exhibiting an elevated MIC to chlorhexidine.10,11
When the MIC for chlorhexidine was less than 4 μg/mL, the isolates were deemed susceptible.3,8 Because chlorhexidine can be used in high quantities without causing toxicity, an increase in MIC values does not imply “resistance.” Thus, microorganisms with an increased MIC to chlorhexidine are better described as having “reduced susceptibility” or “enhanced tolerance”.10,11
Detection of mecA, qacA/B, and qacC Genes using PCR
Following the manufacturer’s instructions, genomic DNA was extracted from pure colonies of the isolated S. aureus using a DNA Mini Purification Kit (Qiagen, USA). The mecA, qacA/B, and qacC genes (Table 1) were detected using a conventional PCR test in a volume of 20 µL with approximately 2 mg of extracted DNA, using 10 µM primers, ready-made PCR master mix, and solution nuclease-free water (Thermo Scientific, USA).1
Table (1):
Primers used for the detection of mecA, qacA/B and qacC genes and expected product size.
Primer |
Sequence (5′-3′) |
Amplicon size (bp) |
Reference |
|---|---|---|---|
mecA forward mecA reverse |
F: GTA GAA ATG ACT GAA CGT CCG ATA A R: CCA ATT CCA CAT TGT TTC GGT CTA A |
310 |
[1] |
qacA/B: forward qacA/B reverse |
F: CTA TGG CAA TAG GAG ATA TGG TGT R: CCA CTA CAG ATT CTT CAG CTA CAT G |
321 |
[1] |
qacC: forward qacC reverse |
F: GGC TTT TCA AAA TTT ATA CCA TCC T R: ATG CGA TGT TCC GAA AAT GT |
249 |
[1] |
Initial denaturation at 95°C for 5 min was followed by 35 cycles of denaturation at 95°C for 30 s, annealing (59°C for mecA, 58°C for qacA/B, and 56°C for qacC) for 30 s, and extension at 72°C for 30 s. This was followed by a 10-minute final extension at 72°C. The predicted amplicons for mecA, qacA/B, and qacC were 310, 321, and 249 bp in length, respectively.
Statistical Analysis
The statistical tool SPSS version 21 was used to code and enter data. Descriptive statistics were used to summarize the data: number and percentage for qualitative variables, and mean and standard deviation for normally distributed quantitative variables. For qualitative variables, the chi-square test was used to assess statistical differences across groups. Statistical significance was set at P ≤ 0.05.
A total of 100 S. aureus clinical isolates were collected from different clinical specimens representing 1.9% of the total specimens received (100/ 5172), 2.5% of all clinical isolates (100/ 3984), 9% of the total Gram positive isolates (100/1104). In the present study, S. aureus represented 11.1% of the total isolated staphylococci (100/ 900). This study was performed on only 84 and 16 isolates of MRSA & MSSA respectively that could be elicited with the total number of 100 S. aureus isolates.
The specimens collected from the outpatient clinic were representing 16% (16/100) while specimens collected from inpatients were 84% (84/100). Urine were the major specimens collected from inpatients were (39.2%) followed by blood (32.1%) and then pus (19.04%), while in outpatients it was mainly urine (93.7%). Most of the inpatients specimens were recovered from the intensive care unit representing 28.5% (24/84) of the total specimens followed by urology department in 22.6% (19/84).
MRSA isolates, 84/100 S. aureus (84%), were phenotypically identified using Kirby–Bauer disc diffusion method via cefoxitin disc and using the Vitek 2 system, which determined oxacillin and cefoxitin MICs. All the isolates were sensitive to vancomycin and linezolid. Conversely, all the MSSA isolates were susceptible to linezolid, vancomycin, and trimethoprim/sulfamethoxazole (Table 2).
Table (2):
Antibiotic susceptibility of MRSA and MSSA isolates by Vitek2 system.
| Antibiotic(s) | MRSA | MSSA | ||
|---|---|---|---|---|
| Sensitive No (%) |
Resistant No (%) |
Sensitive No (%) |
Resistant No (%) |
|
| Vancomycin | 84(100%) | 0 (0%) | 16 (100%) | 0 (0%) |
| Linezolid | 84 (100%) | 0 (0%) | 16 (100%) | 0 (0%) |
| Ciprofloxacin | 59 (70.23%) | 25 (29.7%) | 14 (87.5%) | 2 (12.5%) |
| Moxifloxacin | 59 (70.23%) | 25 (29.7%) | 14 (87.5%) | 2 (12.5%) |
| Clindamycin | 57(67.8%) | 27 (32.1%) | 14 (87.5%) | 2 (12.5%) |
| Erythromycin | 54 (64.2%) | 30 (35.7%) | 14 (87.5%) | 2 (12.5%) |
| Gentamicin | 40 (47.6%) | 44 (52.3%) | 13 (81.25%) | 3 (18.75%) |
| Trimethoprim/ Sulphamethoxazole |
36 (42.8%) | 48 (57.1%) | 16 (100%) | 0 (0%) |
Data are expressed as number (%).
Broth Microdilution Test for Chlorhexidine
The minimum inhibitory concentration (MIC) of chlorhexidine was then determined. The final chlorhexidine concentrations ranged from 0.5–256 µg/mL, and the MIC results ranged from 0.5–128 µg/mL (Table 3). The MIC inhibiting 90% of isolates (MIC90) for chlorhexidine against MRSA were 16 µg/mL and in MSSA was 8 µg/mL. Out of 84 MRSA isolates, 61.9% (52/84) were sensitive to chlorhexidine, whereas 38.1% (32/84) isolates were resistant. On the other hand, 68.8% (11/16) MSSA isolates were sensitive to chlorhexidine and 31.2% (5/16) were resistant (Table 3). No statistically significant difference was found in the chlorhexidine MIC results among the MRSA and MSSA isolates (P = 0.603).
Table (3):
Chlorhexidine minimum inhibitory concentration values for MRSA and MSSA isolates.
| Chlorhexidine | Number (%) of isolates with related MIC,µg/ml | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0.5 | 1 | 2 | 4 | 8 | 16 | 32 | 64 | 128 | MIC50 µg/ml | MIC90 µg/ml | |
| MRSA (no= 84) |
15 (17.8%) | 19 (22%) | 18 (21.4%) | 8 (9.5%) | 13 (15.4%) | 2 (2.3%) | 2 (2.3%) | 6 (7.1%) | 1 (1.1%) | 2 | 16 |
| MSSA (no = 16) |
2 (12.5%) | 3 (18.7%) | 6 (37.5%) | 1 (6.2%) | 2 (12.5%) | 0 (0%) | 0 (0%) | 2 (12.5%) | 0 (0%) | 2 | 8 |
Molecular Detection of mecA, qacA, qacB, and qacC Genes using PCR
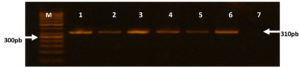
The presence of mecA was detected using PCR (Figure 1). Out of 100 S. aureus strains, 84 (84%) were found to contain mecA, verifying them as MRSA; 82.1% (69/84) were from inpatients, and 17.9% (15/84) outpatients. mecA gene was absent in the remaining 16 (16%) strains (MSSA).
Figure 1. Agarose gel electrophoresis of representative PCR products of S. aureus mecA positive gene (310bp) M: molecular weight marker (ladder 50 bp), Lane 1: positive control, Lane( 2-6): positive DNA samples mecA PCR amplification product, Lane 7: negative control
The presence of the quaternary ammonium resistance genes qacA/B and qacC was examined in 100 S. aureus strains. The qacA/B was the most common biocide resistance gene, being detected in 34% (34/100) of S. aureus isolates (Figure 2,3), of which 94.1% (32/34) were from inpatients, 5.9% (2/34) outpatients, while qacC was the least prevalent, only found in 5% (5/100) of S. aureus isolates, of which 80% (4/5) were from inpatients, and 20% (1/5) outpatients (Figure 3).
 Figure 2. Agarose gel electrophoresis of representative PCR products of S. aureus qacA/B positive gene (321bp) Lane M: molecular weight marker (ladder 50 bp), Lane 1: Negative control, Lane 2: positive control, Lane 3-4: positive DNA samples qacA/B PCR amplification product
Figure 2. Agarose gel electrophoresis of representative PCR products of S. aureus qacA/B positive gene (321bp) Lane M: molecular weight marker (ladder 50 bp), Lane 1: Negative control, Lane 2: positive control, Lane 3-4: positive DNA samples qacA/B PCR amplification product
Figure 3. Agarose gel electrophoresis of representative PCR products of S. aureus qacA/B positive gene (321bp) and qacC positive gene 249bp. Lane M: molecular weight marker (ladder 50 bp), Lane 1: negative control, Lane2: positive control, Lane (3-7): positive DNA samples qacA/B PCR amplification product, Lane 8: negative control, Lane 9: positive control for qacC gene, Lane (10-12): positive DNA samples qacC PCR amplification product
Compared to MSSA, genes for resistance to quaternary ammonium compounds were found to be more prevalent in MRSA. The qacA/B genes were detected in 34.5% (29/84) of MRSA isolates and in 31.2% (5/16) of MSSA isolates, while qacC was found out in 5.95% (5/84) of MRSA isolates, and it was not detected in any MSSA isolate. None of the S. aureus isolates harbored either of these genes. However, no statistically significant difference was found regarding qacA/B and qacC abundance, among MRSA and MSSA isolates (P = 0.8 and 0.317, respectively).
The presence of antiseptic resistance genes in S. aureus isolates was compared with the MIC results for chlorhexidine. The qacA/B gene was found in 34 isolates, 32 (94.1%) of which were resistant to chlorhexidine. This gene, instead, was found in 2 (5.9%) chlorhexidine-susceptible isolates. The qacC gene was found in five isolates that were resistant to chlorhexidine, whereas this gene was not detected in any of the chlorhexidine-susceptible isolates. In MRSA and MSSA isolates, there was a statistically significant association between qacA/B, qacC, and chlorhexidine resistance (P = 0.001 and 0.006, respectively; Table 4).
Table (4):
Comparison between MICs of chlorhexidine and presence of methicillin resistance genes and antiseptic resistance genes in 100 S. aureus isolates.
| Antiseptic | Methicillin resistance gene | Quaternary ammonium resistance genes | |||||
|---|---|---|---|---|---|---|---|
| mecA (+) (n = 84) |
mecA (-) (n = 6) |
qacA/B (+) (n= 34) |
qacA/B (-) (n= 66) |
qacC (+) (n= 5) | qacC (-) (n= 95) | ||
| Chlorhexidine | <4 μg/ml (Susceptible) (n= 63) | 52 (61.9%) | 11 (68.8%) | 2 (5.9%) | 61 (92.4%) | 0 (0%) | 63 (66.3%) |
| ≥ 4 μg/ml (Resistant) (n= 37) | 32 (38.1%) | 5 (31.2%) | 32 (94.1%) | 5 (7.6%) | 5 )100%) | 32 (33.7%) | |
| Chi-square test | 0.270 | 72.100 | 8.962 | ||||
| p-value | 0.603 | 0.001 | 0.006 | ||||
Recently, evidence for the development of resistance to antiseptics has been increasing.12 The decreased sensitivity to chlorhexidine is caused by qacA and qacB, which are placed in plasmids and are highly closely linked. The qacA and qacB genes encode a proton-dependent efflux pump (QacA/QacB), a member of the major facilitator superfamily of transport proteins, in the form of 14α-helical transmembrane segments able to pump chlorhexidine out of bacteria.1
Although chlorhexidine is not used in TBRI hospital, this study was performed to evaluate its future use to follow the CDC recommendations for hand hygiene and body washing of patients with MRSA (CDC, 2017). Instead, a panel of antiseptics and disinfectants are used in TBRI hospital, including alcohol hand rub gel (2-propanol 45+1-propanal 30) for hands and QAC foam of didecyldimethylammonium chloride+biguanide. For surfaces, QAC: benzyl-C12-18-alkyldimethyl–ammoniumchlorides N-(3 aminopropyl)-n-dodexalpropane-1,3-diamine, chlorine: sodium dichloroisocyanurate tablets and sodium hypochlorite, and sodium hydroxide are used. Effervescent granules of sodium dichloroisocyanurate are used for blood and body fluid spills. For surfaces, skin, and air, hydrogen peroxide: 0.5–1% (surfaces and fabrics), 3% (skin, minor cuts), 6% (airborne isolation), 12% (transplantation). While reprocessing of endoscopes and preoperative skin preparation, activated glutaraldehyde solution and povidone–iodine 10% W/V, respectively, are used.
This study aimed to determine the occurrence of antiseptic resistance genes (qacA/B and qacC) in clinical isolates of MSSA and MRSA. We also studied the correlation between mecA gene existence and different antiseptic resistance genes (qacA/B and qacC) in MRSA isolates and determined the association between the occurrence of qacA/B and qacC genes and susceptibility to chlorhexidine to evaluate its future use in TBRI hospital.
A total of 100 S. aureus clinical isolates were obtained from different clinical samples, representing 1.9% of the total specimens received (100/5172). A higher rate of S. aureus was isolated from clinical specimens (30/300, 10%) in a study from Rwanda.13 Additionally, a higher rate was detected in a study from Ehtiopia (14.3%).14 In our study, S. aureus was identified in 2.5% of all clinical isolates (100/3984). A previous study revealed that the proportion of S. aureus identified, in comparison with the total clinical isolates, was 19.96%.15 In the current study, S. aureus represented 10.9% of the total Gram positive isolates (120/1104). A higher prevalence of S. aureus was detected in Gram positive isolates in a previous study (48.8%).16 In the present study, S. aureus represented 11.1% of all staphylococci species isolated (100/900). A previous study reported a S. aureus prevalence of 54.3% of all Staphylococcus strains isolated.17
In the current study, it was found that (84/100) 84% of isolates were confirmed to be MRSA. Our result is comparable to a recent study by Elsherif et al. (2020), who reported a MRSA prevalence of 92.3% of all S. aureus isolates.18 According to many studies, the frequency of MRSA among hospitalized patients in Egypt ranges from 50 to 82 percent.19 However, lower rates were reported in two different studies conducted Abd El-Baky et al.20 and Ahmed et al.,21 who found that methicillin resistance represented 25.4% and 15% of S. aureus isolates, respectively.20,21 Additionally, Elshabrawy et al.,22 reported a prevalence of 40% in Mansoura University hospital.
Similar to our results, studies originating from the Kingdom of Saudi Arabia reported that MRSA accounted for 77.5 to 90% of all S. aureus isolates.23,24 In addition, studies from Nepal have reported MRSA prevalence rates ranging from 69.1 to 75.5%.25-27 Another study in Iran reported that methicillin resistance represented 85% of S. aureus isolates.28 However, in Europe, different lower prevalence rates of MRSA were recorded in various regions: Germany (1%), Belgium (2–43%), United Kingdom (5–23%), Spain (16%), Romania (34%), Italy (37%), France (38%), Greece (40%), and Portugal (49%).29 Furthermore, Diekema et al.,30 revealed lower prevalance in hospitals in the United States of America (USA), stating that MRSA accounts for 30–50% of all nosocomial S. aureus isolates. In USA, the incidence of hospital- and community-acquired MRSA infections decreased by 74% and 40%, respectively, from 2005 to 2016. The decline in MRSA infections could be attributed to a variety of infection control initiatives, including improvements in preventing device- and procedure-related infections, as well as efforts that aided in disrupting MRSA transmission in the hospitals.31
MRSA infection prevalence was a noticeable difference in the prevalence of MRSA infection. The difference in MRSA prevalence rates could be probably attributed to differences in the studied populations and discrepancies in the infection control procedures applied, along with unreasonable antibiotic use. Some researchers used active surveillance cultures, whereas others used a culture of clinical specimens.27 Furthermore, many hospital-acquired infections may be caused by S. aureus.22,32
Therefore, the high rates stated in this study could be due to the long duration of the study and injudicious use of antibiotics, as the percentage of systemic antibiotics that can potentially elicit methicillin-resistant strains, traded at drug stores without a prescription is high in our community, resulting in an elevated resistance to antimicrobial agents and allowing the transfer of resistance between hospitals and the community.
In our study, the MRSA isolates were more resistant towards the tested antibiotics than the MSSA isolates. However, no isolates showed decreased sensitivity to vancomycin and linezolid. Interestingly, 57.1% of MRSA strains were resistant to trimethoprim/sulfamethoxazole, whereas none of the MSSA strains were resistant to it. MRSA infections, including endocarditis, bone and joint infections, and meningitis, are routinely treated with antibiotics. This treatment strategy could be worrying sign and lead to a rise in resistance toward the antibiotics.29
Some proportion of MRSA isolates (52.3% and 35.7%) exhibited elevated resistance to gentamicin and erythromycin, respectively. These two antibiotics are commonly and randomly used to treat a variety of illnesses, including generalized and pyogenic infections.27 In contrast, only 18.7% and 12.5% of MSSA isolates exhibited increased resistance to gentamicin and erythromycin, respectively. The trend of multiclass antibiotic resistance reported in this study is in concordance with an Egyptian and an Iranian study, both of which reported high resistance to trimethoprim/sulfamethoxazole in MRSA isolates and increased susceptibility towards vancomycin and linozelid.21,28 More than 25% of MRSA isolates were resistant to gentamicin, trimethoprim/sulfamethoxazole, and erythromycin, consistent with Gurung et al.’s results27; however; none of the MRSA isolates were resistant to vancomycin. Additionally, these antibiotics showed less than 50% resistance towards the isolated MSSA. This could be explained by the fact that the mecA gene in MRSA is found on the staphylococcal cassette chromosome mec, which also encodes genes for various antibiotics, including aminoglycosides.1,33
In the present study, 38.1% and 31.2% of MRSA and MSSA isolates had decreased sensitivity to chlorhexidine, respectively, in line with Askar et al.1 study which stated that 42% of MRSA had reduced susceptibility to chlorhexidine, whereas all MSSA isolates were susceptible. Ignak et al.10 reported that 50% of MRSA and 15% of MSSA had reduced susceptibility to chlorhexidine. Moreover, Dittmann et al.11 reported that 65% of MRSA and 5% of MSSA had decreased sensitivity to chlorhexidine.
In the current study, all S. aureus isolates were tested for the presence of the quaternary ammonium resistance genes qacA/B and qacC using PCR. qacA/B was more dominant, being found in 34% of S. aureus isolates (MRSA 34.5% and MSSA 31.2%), whereas qacC was less common, only detected in 5% of S. aureus isolates (MRSA 5.95% and MSSA 0%). This result is comparable to the Egyptian study by Askar et al.,1 where qacA/B was found in 47.4 % of MRSA isolates, whereas qacC was found in 28.9 % of these isolates. Moreover, John et al.,34 and Htun et al.35 revealed that the prevalence of qacA/B gene was 35.08% and 46.6%, while that of qacC was 4.7% and 13.6%, respectively.34,35 Our results are in partial agreement with a study from Japan, which reported qacA/B genes to be more abundant in MRSA (32.6%) and MSSA (7.5 %) isolates than qacC genes, which were found to be 3.3% in MRSA and 5.9% in MSSA.36 Meyer et al.,37 discovered that qacA/B genes were more common in 42% of S. aureus (63% MRSA and 12% MSSA) than the qacC gene, which was found in 5.8% of S. aureus (6.4% MRSA and 5% MSSA). A Malaysian study detected qacA/B gene at a much higher rate (83.3%) than qacC (1.9%) among their MRSA isolates.38 According to Duran et al.,39 the prevalence of qacA/B genes in MRSA and MSSA isolates was 43.8% and 3.3%, respectively, whereas the prevalence of the qacC gene was 5% and 2.5%, respectively.
In Japan, however, qacA/B genes were found in 14% of clinical MRSA isolates, while qacC genes in 28%.40 Longtin et al.41 found that in 334 MRSA isolates obtained from two Canadian intensive care units, qacC genes were more common (7%) than qacA/B genes (2%), and that no isolate carried both genes. Additionally, a study in Turkey reported that qacC (13.8%) was more common than qacA/B (10.3%) among S. aureus isolates.10 ‘Qac’ symbolizes two major families, separating QacA and QacB (MFS superfamily members) from the qacC proteins that belong to the SMR superfamily, which could explain the differential predominance of qac genes (qacA/B and qacC).28 Except for some shared substrates and similar efflux functions, these two large protein families have little in common. The qacA and qacB genes were found on the chromosomes, and the plasmids pSK1 and pSK23. The genes are mostly prevalent in S. aureus and S. epidermidis, and show modest sequence variation. The qacC gene has been found on both conjugative and non-conjugative (rolling-circle) plasmids, and it has been identified and characterized in both S. aureus and coagulase-negative Staphylococcus species.5
Despite the fact that MRSA had more qacA/B and qacC genes than MSSA, there was no statistically significant difference in the presence of these genes between MRSA and MSSA in this study. In contrast to our findings, Zhang et al.42 found that 50% of MRSA and 16% of MSSA were positive for the qacA/B gene, a statistically significant difference. Furthermore, several researchers found the frequency of qacA/B and qacC to be statistically significant among MRSA and MSSA.28,39 The lack of statistical significance in our study could be attributed to the relatively low number of MSSA isolates. The higher prevalence of qac genes in MRSA than in MSSA can be attributed to the fact that the resistance of MRSA isolates to disinfectants and antiseptic compounds is mediated mainly by plasmid-encoded qacA/B and qacC determinants. The occurrence of such determinants is occupied by MDR resistance patterns in MRSA isolates toward multiple targets, leading to cross-resistance between antibiotic and antiseptics in S. aureus isolates.43 In addition, the mecA gene on a mobile genomic island staphylococcal cassette chromosome is a carrier for virulence and other drug-resistant genes, as well as a vehicle for genetic exchange between staphylococci.28 More research is needed to understand gene transfer selection that leads to the dissemination of resistance genes for designing long-term strategies to cure microbial diseases.43
In the present study, all S. aureus isolates exhibited with decreased sensitivity to chlorhexidine (MIC ≥4 mg/L) and harbored either qacA/B or qacC, in agreement with other studies reporting that all isolates carrying qacA/B or qacC had a chlorhexidine MIC ≥ 4 µg/mL.1,42 Despite all previous studies revealing a clear association between qacA/B and reduced susceptibility to chlorhexidine, the present study unravels that the distribution of the qacA/B genes is not confined to strains showing chlorhexidine resistance, whereas two isolates harboring qacA/B genes were susceptible to chlorhexidine, in agreement with other reports.3,35 This finding could be explained by the fact that gene expression is not guaranteed by the presence of the qacA/B genes. Transcription regulators, intensity of exposure, and past contact with cationic chemicals influence their expression.3 In a meta-analysis of the effectiveness of chlorhexidine gluconate bathing in minimizing infections among individuals in the intensive care unit (ICU), evidence indicated the benefit of daily chlorhexidine digluconate bathing in reducing MRSA infections. However, effectiveness may be influenced by the underlying baseline risk of these events in the ICU population. As a result, chlorhexidine digluconate bathing appears to be most beneficial when infection rates in the ICU population are high.44
In this study, a statistically significant association between the existence of qacA/B and qacC and lower susceptibility to chlorhexidine was observed, consistent with research from Iran and Singapore, which found an association between the presence of qac genes and a chlorhexidine MIC ≥ 4 µg/mL.28,37 The existing chlorhexidine MICs for staphylococci harboring the qac gene may permit existence at in-use concentrations, even when relatively augmented resistance may allow continuance when leftover disinfectants are present.28 It must be noted that the overuse of chlorhexidine as a decontaminant may increase the number of MRSA isolates with qacA/B resistance genes.37
In our study, inpatients had a higher rate of S. aureus infection (84%) than outpatients (16%), consistent with Rajbhandari et al.,45 and Gurung et al.,27 who found a greater prevalence of S. aureus in the inpatient setting (62.3% and 55.7%), compared to in outpatients (37.7% and 44.3%), respectively.45,27 Likewise, the distribution of mecA, qacA/B, and qacC genes in the patients was found to be 82.1%, 94.1, 80%, respectively, in inpatients and 17.9%, 5.9%, and 20%, respectively, in outpatients.
Our result is in accordance with a study that found that mecA, qacA/B, and qacC were more dominant in inpatients than in outpatients.27 These findings suggest that nosocomial infections are more common in hospitals. Longer hospital stays, antibiotic treatment, underlying immunocompromised conditions, hospital environment, overuse of antiseptics and disinfectants, instrumentation, and use of other invasive devices could be the risk factors that predispose patients to MRSA acquisition.
This study revealed a high percentage of S. aureus isolates harboring the antiseptic resistance genes qacA/B or qacC, which were more common in MRSA than in MSSA isolates. The presence of qacA/B or qacC was linked to decreased susceptibility to chlorhexidine. However, the presence of qacA/B did not always imply a reduction in chlorhexidine susceptibility. The clinical use of chlorhexidine may continue to increase, implying the development of new clones with reduced susceptibility.
ACKNOWLEDGMENTS
The authors would like to thank Prof. Abdel Fattah Hassan Attia, Professor of Medical Microbiology and Immunology, Faculty of Medicine, Cairo University, Egypt for assigning this work, and supplied all necessary facilities for its success.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
HB conceptualized the research. AR analyzed, and interpreted the data. MS performed the microbiological identification and interpreted the results. AF collected samples, analyzed and interpreted the data. AS performed the molecular detection. NY performed the MIC and molecular detection. MS and AF wrote the manuscript. HB, AR, AF, RM, NY revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
This study was funded by Theodor Bilharz Research Institute under Internal Project (No. 110).
DATA AVAILABILITY
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
ETHICS STATEMENT
The protocol of the study was approved by TBRI Institutional Review Board under Federal Wide Assurance (FWA00010609) and the work has been carried out in accordance with the Code of Ethics of the World Medical Association (Declaration of Helsinki) for Experiments in Humans and its later amendments (GCP guidelines) or comparable ethical standards.
- Askar H, Badaway W, Hammad E. Aminoglycoside and chlorhexidine resistance genes in Staphylococcus aureus isolated from surgical wound infections. Int Arab J Antimicrob Agents. 2016;16(3):571-578.
Crossref - Dindarloo K, Aghamolaei T, Ghanbarnejad A, et al. Pattern of Disinfectants Use and Their Adverse Effects on the Consumers after COVID-19 Outbreak. J Environ Health Sci Eng. 2020;18(2):1301-1310.
Crossref - Do Vale B, Acacia G, Thiago A, Lopes M, De Melo M. Decreased Susceptibility to Chlorhexidine and Distribution of QacA/B Genes among Coagulase-Negative Staphylococcus Clinical Samples. BMC Infect Dis. 2019;19(1):1-5.
Crossref - Prag G, Karin F, Jacobsson S, Bengt H, Magnus U, Soderquist B. Decreased Susceptibility to Chlorhexidine and Prevalence of Disinfectant Resistance Genes among Clinical Isolates of Staphylococcus Epidermidis. APMIS. 2014;122(10):961-967.
Crossref - Trudy W, Ussery D, Nielsen L, Hanne Ingmer. Review and Phylogenetic Analysis of Qac Genes that Reduce Susceptibility to Quaternary Ammonium Compounds in Staphylococcus Species. Eur J Microbiol Immunol. 2015;5(1):44-61.
Crossref - Batra R, Cooper BS, Whiteley C, Patel AK, Wyncoll D, Edgeworth JD. Efficacy and Limitation of a Chlorhexidine-Based Decolonization Strategy in Preventing Transmission of Methicillin-Resistant Staphylococcus aureus in an Intensive Care Unit. Clin Infect Dis. 2010;50(2):210-217.
Crossref - Clinical and Laboratory Standards Institute. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. Approved standard M7-A7. Clinical and Laboratory Standards Institute, Wayne, PA. 2019.
- Sheng W, Wang J, Lauderdale T, Weng C-M, Chen D, Chang S-C. Epidemiology and susceptibilities of methicillin-resistant Staphylococcus aureus in Taiwan: emphasis on chlorhexidine susceptibility. Diagn Microbiol Infect Dis. 2009;63(3):309-313.
Crossref - Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing, 30th ed CLSI supplement M100 Clinical and Laboratory Standards Institute, Wayne, PA. 2020.
- Ignak S, Nakipoglu Y, Gurler B. Frequency of antiseptic resistance genes in clinical staphycocci and enterococci isolates in Turkey. Antimicrob Resist Infect Control. 2017;6:88.
Crossref - Dittmann K, Schmidt T, Muller G, et al. Susceptibility of livestock-associated methicillin-resistant Staphylococcus aureus (LA-MRSA) to chlorhexidine digluconate, octenidine dihydrochloride, polyhexanide, PVP-iodine and triclosan in comparison to hospital-acquired MRSA (HA-MRSA) and community-aquired MRSA (CA-MRSA): a standardized comparison. Antimicrob Resist Infect Control. 2019;8:122.
Crossref - Guo Y, Song G, Sun M, Wang J, Wang Y. Prevalence and Therapies of Antibiotic-Resistance in Staphylococcus aureus. Front Cell Infect Microbiol. 2020;10:107.
Crossref - Noel G, Benigne B, Vincent M. Prevalence of Staphylococcus aureus among Clinical Isolates and their Responses to Selected Antibiotics at Centre Hospitalier Universitaire de Kigali (CHUK). J Microbiol Exp. 2017;5(4):00158.
Crossref - Dilnessa T, Bitew A. Prevalence and antimicrobial susceptibility pattern of methicillin resistant Staphylococcus aureus isolated from clinical samples at Yekatit 12 Hospital Medical College, Addis Ababa, Ethiopia. BMC Infect Dis. 2016;16:398.
Crossref - Sapkota J, Sharma M, Jha B Bhatt CP. Prevalence of Staphylococcus aureus Isolated from Clinical Samples in a Tertiary Care Hospital: A Descriptive Cross-sectional Study. JNMA J Nepal Med Assoc. 2019;57(220):398-402.
Crossref - Mamtora D, Saseedharan S, Bhalekar P, Katakdhond S. Microbiological profile and antibiotic susceptibility pattern of Gram-positive isolates at a tertiary care hospital. J Lab Physicians. 2019;11(2):144-148.
Crossref - Cuevas O, Cercenado E, Vindel A, et al. Evolution of the antimicrobial resistance of Staphylococcus spp. in Spain: five nationwide prevalence studies, 1986 to 2002. Antimicrob Agents Chemother. 2004;48(11):4240-4245.
Crossref - Elsherif M, Helal H, El-Ansary R, et al. Staphylococcal Enterotoxins and Toxic Shock Syndrome Toxin-1 and Their Association among Bacteremic and Infective Endocarditis Patients in Egypt. BioMed Research International. 2020;6981095:1-9.
Crossref - Abdel-Maksoud M, El-Shokry M, Ismail G,et al. Methicillin-resistant Staphylococcus aureus recovered from healthcare- and community-associated infections in Egypt. Int J Bacteriol. 2016;2016:5751785.
Crossref - Abd El-Baky R, Rady H, Gad G. Prevalence and Conjugal Transfer of Vancomycin Resistance among Clinical Isolates of Staphylococcus aureus. Advances in Research. 2014;2(1):12-23.
Crossref - Ahmed F, Gad F, Abdalla M, Hasaneen M, Abdelwahab F. Prevalence of methicillin resistant Staphylococcus aureus among Egyptian patients after surgical interventions. Surginfect. 2014;15(4):404-411.
Crossref - Elshabrawy WO, Zaki M E, Kamel MF. Genetic and phenotypic study of methicillin-resistant Staphylococcus aureus among patients and health care workers in Mansoura University Hospital, Egypt. Iran J Microbiol. 2017;9(2):82-88.
- Baddour MM, Abuelkheir MM, Fatani, AJ. Trends in antibiotic susceptibility patterns and epidemiology of MRSA isolates from several hospitals in Riyadh, Saudi Arabia. Ann Clin Microbiol Antimicrob. 2006;5:30.
- Alzolibani A, Al Robaee AA, Al Shobaili HA, Bilal JA, Ahmad MI. Documentation of Vancomycin-Resistant Staphylococcus Aureus (VRSA) among Children with Atopic Dermatitis in the Qassim Region, Saudi Arabia. Acta Dermatovenerol Alp Pannonica Adriat. 2012;21(3):51-53.
- Rijal K, Shrestha N, Pahari N, et al. Methicillin Resistant Staphylococcus Aureus in patients visiting Western Regional Hospital, Pokhara. Journal of Institute of Medicine Nepal. 2008;30(1):21-25.
- Tiwari HK, Sapkota D, Das AK, Sen M R. Assessment of different tests to detect methicillin resistant Staphylococcus aureus. Southeast Asian J Trop Med Public Health. 2009;40(4):801-806.
- Gurung RR, Maharjan P, Chhetri GG. Antibiotic resistance pattern of Staphylococcus aureus with reference to MRSA isolates from pediatric patients. Future Sci OA. 2020;6(4):FSO464.
Crossref - Damavandi MS, Dehkordi MS, Dehghan A, Heibati F, Taghaddosi R, Gholipoural A. Detection of Antiseptic Resistance Genes among Staphylococcus aureus Colonising Nurses and Coagulase-Negative Staphylococci Isolated from Clinical Specimens at Teaching Hospitals in Southwest of Iran. Jundishapur J Microbiol. 2017;10(1):e39285.
Crossref - Elshimy R, Khattab RA, Zedan H, Hosny AS, Elmorsy TH. Study on prevalence and genetic discrimination of methicillin-resistant Staphylococcus aureus (MRSA) in Egyptian hospitals. Afr J Microbiol Res. 2018;12(27):629-646.
Crossref - Diekema DJ, Pfaller MA, Schmitz FJ, et al. Survey of Infections Due to Staphylococcus Species: Frequency of Occurrence and Antimicrobial Susceptibility of Isolates Collected in the United States, Canada, Latin America, Europe, and the Western Pacific Region for the SENTRY Antimicrobial Surveillance Program, 1997-1999. Clin Infect Dis. 2001;32(Suppl 2):S114-32.
Crossref - Kourtis AP, Hatfield K, Baggs J, et al. Vital Signs: Epidemiology and Recent Trends in Methicillin-Resistant and in Methicillin-Susceptible Staphylococcus aureus Bloodstream Infections- United States. MMWR. Morbid Mortal Wkly Rep. 2019;68(9):214-219.
Crossref - Karakonstantis S, Kalemaki D. Antimicrobial overuse and misuse in the community in Greece and link to antimicrobial resistance using methicillin-resistant S. aureus as an example. J Infect Public Health. 2019;12(4):460-464.
Crossref - Fatholahzadeh B, Emaneini M, Gilbert G, et al. Staphylococcal cassette chromosome mec (SCCmec) analysis and antimicrobial susceptibility patterns of methicillin-resistant Staphylococcus aureus (MRSA) isolates in Tehran, Iran. Microb Drug Resist. 2008;14(3):217-220.
Crossref - John MA, Burden J, Stuart JI, et al. Comparison of three phenotypic techniques for detection of methicillin resistance in Staphylococcus spp. reveals a species-dependent performance. J Antimicrob Chemother. 2009;63(3):493-396.
Crossref - Htun HL, Hon PY, Holden M, Ang B, Chow A. Chlorhexidine and octenidine use, carriage of qac genes, and reduced antiseptic susceptibility in methicillin-resistant Staphylococcus aureus isolates from a healthcare network. Clin Microbiol Infect. 2019;25(9):1154.e1-1154.e7.
Crossref - Alam MM, Ishino M, Kobayashi N. Analysis of genomic diversity and evolution of the low-level antiseptic resistance gene smr in Staphylococcus Aureus. Microb Drug Resist. 2003;9(Suppl 1):S1-7.
Crossref - Meyer B, Cookson B. Does microbial resistance or adaptation to biocides create a hazard in infection prevention and control? J Hosp Infect. 2010;76(3):200-205.
Crossref - Shamsudin M, Alreshidi M, Hamat R, Alshrari A, Atshan S, Neela V. High prevalence of qacA/B carriage among clinical isolates of meticillin-resistant Staphylococcus aureus in Malaysia. J Hosp Infect. 2012;81(3):206-208.
Crossref - Temiz M, Duran N, Duran GG, Eryılmaz N, Jenedi K. Relationship between the resistance genes to quaternary ammonium compounds and antibiotic resistance in staphylococci isolated from surgical site infections. Med Sci Monit. 2014;20:544-550.
Crossref - Noguchi N, Suwa J, Narui K, et al. Susceptibilities to antiseptic agents and distribution of antiseptic-resistance genes qacA/B and smr of methicillin-resistant Staphylococcus Aureus isolated in Asia during 1998 and 1999. J Med Microbiol. 2005;54(6):557-565.
Crossref - Longtin J, Seah C, Siebert K, et al. Distribution of antiseptic resistance genes qacA, qacB, and smr in methicillin-resistant Staphylococcus Aureus isolated in Toronto, Canada, from 2005 to 2009. Antimicrob Agents Chemother. 2011;55(6):2999-3001.
Crossref - Zhang M, O’Donoghue M, Ito T, Hiramatsu K, Boost M. Prevalence of antiseptic-resistance genes in Staphylococcus aureus and coagulase-negative staphylococci colonising nurses and the general population in Hong Kong. J Hosp Infect. 2011;78(2):113-117.
Crossref - Saber N, Kandala NJ, Mohammed HA. Detection a New Antiseptic Resistant Variant of Qac Gene in Some Multi Drug Resistant Staphylococcus Aureus Isolated from Different Clinical Sources. Baghdad Sci J. 2019;16(3):571-579.
Crossref - Frost SA, Alogso M, Metcalfe L, et al. Chlorhexidine bathing and health care-associated infections among adult intensive care patients: a systematic review and meta-analysis. Crit Care. 2016;20(1):379.
Crossref - Rajbhandari R, Manandhar SP, Shrestha J. Comparative study of MRSA and its antibiotic susceptibility pattern in indoor and outdoor patients in Bir Hospital, Nepal.Nepalese J Microbiol. 2003; 1: 62-65.
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.