ISSN: 0973-7510
E-ISSN: 2581-690X
Tuberculosis (TB) is a serious disease that has been observed since ancient times. In the mid-to late-20th century, the main clinical approach to this disease involved focusing on its diagnosis, prevention, and treatment. However, in the 21st century, the focus has shifted toward the diagnosis and treatment of drug-resistant TB. With the use of the Xpert MTB/RIF assay at the frontlines in India, interpreting indeterminate results to treatment with rifampicin, an antitubercular drug, can be challenging. This is further exacerbated by a lack of knowledge regarding mutation frequency in antitubercular drug-resistant genes in this region. Among antitubercular drugs, rifampicin is the most potent and effective drug for the treatment of tuberculosis; hence, understanding the pattern of rifampicin resistance (rpoB) gene mutations will provide insights into the genetic basis of this resistance, which may help in the prevention and treatment of TB. This retrospective observational study presents sociodemographic details, sample types, Mycobacterium tuberculosis load, types of probe mutations detected, and rifampicin indeterminate results from the Xpert MTB/RIF assay. Of the 314 samples analyzed, 258 showed rifampicin resistance as detected by MTB, with 56 samples of MTB-detected rifampicin indeterminate results. Type E probe mutation (58.9%) was the most common type, while the least frequent mutation was Type C probe (1.5%). No missing probe was observed in approximately 8.9% of samples. Among the 56 rifampicin indeterminate results, the maximum Cycle threshold value did not cross 34.5 in six samples.
Xpert MTB/RIF, rpoB Mutation, Rifampicin Indeterminate
India is currently the leading country in tuberculosis (TB) cases and deaths worldwide, accounting for one-fifth of the global TB cases, and the second largest number of multi-drug resistant TB (MDR-TB) cases, after China.1 Most recently, with an increase in the utilization of the Xpert MTB/RIF assay, a rapid molecular test for the identification of TB and its resistance to rifampicin (RIF), a greater number of rifampicin-resistant TB (RRTB) cases have been reported.2 MDR-TB is resistant to isoniazid and rifampicin, two of the most potent antitubercular drugs. However, only rifampicin resistance (RR) can be detected by the Xpert MTB/RIF assay, due to the fact that over 90% of these cases are also resistant to isoniazid.3 rpoB mutations are observed in 96.1% of RRTB strains throughout the world, typically observed in between the 507th and 533th amino acid residuals (81 bp) in the rpoB gene, denoted as the rifampicin-resistance determining region (RRDR).4 The Xpert MTB/RIF assay uses five various overlapping probes (A, B, C, D and E) to identify the mutations in codons from 507-511, 511-518, 518-523, 523-529, and 529-533, respectively. These probes can be used to differentiate the wild type from the mutated type, the latter of which is related to rifampicin resistance. Since the effects of mutations in this gene on bacterial physiology and survival depend on the different types of mutations, understanding the types, patterns, and geographical tally of these mutations could benefit future interventions.5 The Xpert MTB/RIF assay is a rapid, automated molecular test that detects multi-drug-resistant tuberculosis using rifampicin as a predictor. However, it has limitations. For example, when the bacilli load was very low, indeterminate results for M. tuberculosis were observed, indicating that the test could not determine whether the bacteria were resistant to rifampicin. This represents a challenge for clinicians wishing to initiate antitubercular treatment (ATT) in their patients based on indeterminate rifampicin results. To address this, this study determined the type of rpoB gene mutation common in the geographical area of Kalaburagi, India, in relation to rifampicin indeterminate results.
Objectives
- To determine the most common type of rpoB gene mutation and its frequency
- To analyze rifampicin indeterminate results in the Xpert MTB/RIF assay
This retrospective study was conducted at a tertiary care center at the ESIC Hospital Kalaburagi, Karnataka, India. The district tuberculosis office was informed when the present study was conducted.
Inclusion criteria
All M. tuberculosis detected rifampicin resistance/rifampicin indeterminate TB cases diagnosed by the Xpert MTB/RIF, including all age groups and suspected, symptomatic, referred, pulmonary, and extrapulmonary cases of tuberculosis.
Exclusion criteria
Cases of M. tuberculosis detected with rifampicin sensitivity.
Data were collected retrospectively from the last five years for all types of specimens received and tested using the Xpert MTB/RIF assay at tertiary care hospitals in the CBNAAT Laboratory from March 2018 to February 2023. The collected data included sociodemographic details, sample types, MTB loads, types of probes where the mutation was detected, and indeterminate results for rifampicin.
Using hemi-nested real-time PCR, the Xpert MTB/RIF assay (Cepheid, CA, USA) four module version was used to simultaneously identify M. tuberculosis and rifampicin resistance in patient samples. Targeting the 81-bp rpoB gene, five partially overlapping fluorescent probes (A, B, C, D, and E) were used. Based on the cycle threshold (CT) value, the probes were semi-quantitatively graded as High, Medium, Low, or Very Low. In cases with M. tuberculosis that were sensitive to rifampicin, all five probes matched the amplified DNA with similar CT values. Samples resistant to rifampicin were indicative of a mutation that caused differences in the CT values among the probes. A difference ≥4 between the maximum CT value and minimum CT value among the five probes was considered to indicate rifampicin resistance.6
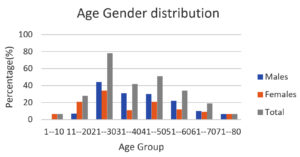
Of the 314 samples analyzed, 258 showed rifampicin resistance by MTB, with indeterminate rifampicin resistance detected in the other 56 samples. The incidence of MTB rifampicin resistance in this region was 2.36%, whereas that of indeterminate rifampicin resistance was 0.5%. Among the 258 rifampicin-resistant patients, 147 (57%) were males and 111 (43%) were females. The age group between 21 and 30 years (30.2%) was comprised of the largest number of cases, followed by the age groups 41-50 (19.7%), 31-40 (16.2%), 51-60 (13.1%), 11-20 (10.8%), 61-70 (7.3%), 71-80 (1.5%), and 1-10 (0.7%) (Table 1 and Figure 1).
Table (1):
Age-Gender distribution and MTB Bacillary Load vs Sample type
| Age group | Males (%) | Females (%) | Total | Bacillary Load | Pulmonary (%) | Extra Pulmonary (%) |
|---|---|---|---|---|---|---|
| 1-10 | 0.00 | 1.80 | 2 | High | 28.63 | 10.00 |
| 11-20 | 4.76 | 18.92 | 28 | Medium | 33.47 | 20.00 |
| 21-30 | 29.93 | 30.63 | 78 | Low | 20.56 | 30.00 |
| 31-40 | 21.09 | 9.91 | 42 | Very Low | 17.34 | 40.00 |
| 41-50 | 20.41 | 18.92 | 51 | Total | 100.00 | 100.00 |
| 51-60 | 14.97 | 10.81 | 34 | |||
| 61-70 | 6.80 | 8.11 | 19 | |||
| 71-80 | 2.04 | 0.90 | 4 | |||
| Total | 100.00 | 100.00 | 258 | |||
Maximum type of clinical sample was from Pulmonary 248 and extrapulmonary samples were 10.
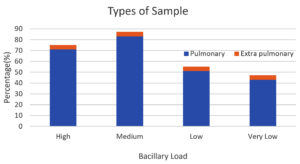
The grading of bacillary load according to CT value showed maximum cases of medium load followed by High, Low and very low. (Table 1, Figure 2).
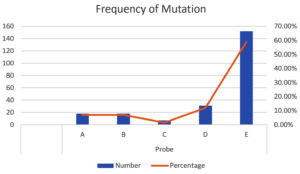
The most common probe that was positive for mutations was probe E, followed by probes D, A, B, and C (Figure 3). Mutation combinations were also observed, as follows: AB, DE, AD, AE, CD, and BD (Table 2). No missing probe was observed in 23 samples.
Table (2):
Mutation Frequency of different Probes of rpoB gene
Codon site |
Number |
Percentage |
|---|---|---|
Probe |
||
A |
18 |
6.98 |
B |
18 |
6.98 |
C |
4 |
1.55 |
D |
31 |
12.02 |
E |
152 |
58.91 |
A&B |
4 |
1.55 |
A&D |
1 |
0.39 |
A&E |
1 |
0.39 |
C&D |
1 |
0.39 |
B&D |
1 |
0.39 |
D&E |
4 |
1.55 |
No missing probe |
23 |
8.91 |
Total |
258 |
100.00 |
Among the 56 Rifampicin indeterminate results, 6 samples had shown that the maximum CT value did not cross 34.5 (Table 3).
Table (3):
Rifampicin Indeterminate Result as shown
Maximum CT |
Minimum CT |
∆CT |
Remarks |
|---|---|---|---|
38.5 |
32.6 |
5.9 |
|
36.5 |
32.7 |
3.8 |
|
35.8 |
33.2 |
2.6 |
|
36.5 |
32.7 |
3.8 |
|
33.2 |
28.1 |
4.1 |
No probe has crossed 34.5 |
36.5 |
34.1 |
2.4 |
|
36.8 |
0 |
36.8 |
|
37.8 |
33 |
4.8 |
|
39.1 |
32.7 |
6.4 |
|
40.5 |
0 |
40.5 |
|
40.3 |
0 |
40.3 |
|
35.8 |
0 |
35.8 |
|
38.8 |
32.2 |
6.4 |
|
40 |
35.3 |
4.7 |
|
36.8 |
33.6 |
3.2 |
|
38.3 |
32.7 |
5.6 |
|
38.1 |
32.8 |
5.3 |
|
38.8 |
34.9 |
3.9 |
|
40.6 |
0 |
40.6 |
|
34.6 |
0 |
34.6 |
|
36.2 |
33 |
3.2 |
|
38.1 |
35 |
3.1 |
|
36.2 |
33.4 |
2.8 |
|
36.2 |
32.7 |
3.5 |
|
40.3 |
0 |
40.3 |
|
38.5 |
33 |
5.5 |
|
38.3 |
34.4 |
3.9 |
|
40.8 |
35 |
5.8 |
|
40.2 |
33.8 |
6.4 |
|
36.2 |
33.1 |
3.1 |
|
36.5 |
32.9 |
3.6 |
|
36.5 |
32.3 |
4.2 |
|
36.1 |
33.4 |
2.7 |
|
37.4 |
32.6 |
4.8 |
|
38 |
35.3 |
3.3 |
|
37 |
32.8 |
4.2 |
|
36.9 |
32.7 |
4.2 |
|
40.8 |
0 |
40.8 |
|
36.7 |
32.8 |
3.9 |
|
36.8 |
33.4 |
3.4 |
|
35.2 |
0 |
35.2 |
|
33.3 |
0 |
33.3 |
Not Crossed 34.5 |
36.9 |
32.6 |
4.3 |
|
36.4 |
32.4 |
4 |
|
39.1 |
36.4 |
2.7 |
|
41 |
32.4 |
8.6 |
|
36.7 |
34 |
2.7 |
|
35.7 |
0 |
35.7 |
|
33.2 |
0 |
33.2 |
Not Crossed 34.5 |
37.3 |
0 |
37.3 |
|
34.8 |
0 |
34.8 |
|
35.1 |
0 |
35.1 |
|
33.2 |
0 |
33.2 |
Not Crossed 34.5 |
36.1 |
32.7 |
3.4 |
|
32.5 |
0 |
32.5 |
Not Crossed 34.5 |
32.9 |
0 |
32.9 |
Not Crossed 34.5 |
Knowing the frequency and type of mutations associated with drug resistance in MTB in different settings helps in the development and implementation of rapid, large-scale genetics-based assays for the diagnosis of drug resistance.7
In the present study, 258 MTB-detected rifampicin-resistant cases and 56 MTB-detected rifampicin indeterminate cases were analyzed. Among the 258 rifampicin-resistant cases, 147 (57%) were Male predominant, with 111 (43%) in females. Huge variation was observed in the total samples of different groups (only two samples in the 1-10 age group compared to 78 in the 21-30 age group). A study in Ethiopia found that males are at greater risk for MDR-TB.8 In the present study, the 21–30 age group was found to be more affected by rifampicin resistance, which is consistent with previous studies from Mizoram, China, and Ethiopia.3,9,10 This could be due to the fact that this age group is more active and exposed to tuberculosis. Furthermore, more pulmonary samples (248) than extrapulmonary samples (10) were observed.
A greater number of cases presented with a medium bacillary load, followed by high, low, and very low loads among 248 sputum samples, whereas extrapulmonary cases (10) presented with very low (4), low (3), medium (2), and high (1) loads.
Among the five probes evaluated, most common mutations were observed with probe E (58.91%), which corresponds to codon 529-533, followed by probe D (12.02%) at codon 523-529, probe A (6.98%) at codon 507-511, probe B (6.98%) at codon 511-518, and probe C (1.55%) at codon 518-523, consistent with a previous study by NIRT Chennai.2 These results highlight the fact that this pattern is prevalent in south India, particularly when compared to studies conducted in Mizoram, Madhya Pradesh, which showed a different pattern.1,3 Some degree of correlation was also observed with studies conducted in Andhra Pradesh, Mumbai and Punjab, wherein the predominance of Probe E mutation was reported (Table 2).11-13
With respect to other countries, similar studies in Pakistan (Munir et al., Alimgir et al.), Uganda (Mboowa et al.), and Ethiopia (Alemu et al.) found that the predominant mutation was observed using probe E, consistent with the results of the present study, indicating that probe E mutation is the most common type worldwide.4,5,10,14 The least frequent mutation was that of probe C, which is also consistent with the majority of the aforementioned literature. This may be due to the lower frequency for this mutation in this region. The frequency of combined mutations was 4.65%, which is comparable to that reported in Nigeria (Ochang et al.). This highlights the non-adherence of patients to antitubercular treatment (ATT), which may have given rise to more than one mutation.15 A lack of missing probes was consistent with the results reported by Munir et al., who found that 8.3% of samples were missing probes, compared to other studies in which a comparatively lower percentage was reported.14
In the Xpert MTB/RIF assay, the criteria used to denote a rifampicin indeterminate result is if the Ct of the first probe is >34.5 cycles and the Ct of the last probe is >38 cycles. However, in the present study, among 56 rifampicin indeterminate results, six sample Ct values crossed neither >34.5 nor >38. This implies that the programmed/configured software was not completely accurate in interpreting the rifampicin sensitivity reports. Furthermore, in all six samples, a mutation was observed in one of the probes, which could be interpreted as rifampicin resistance. These results study emphasize that, along with the report, the Ct value should be analyzed and interpreted for resistance according to the interpretive guidelines (missing probe or >4 cycle difference between the minimum and maximum Ct values, provided that neither of the Ct values is >34.5). The rifampicin indeterminate results can also be analyzed by using Shewart control charts, as described by Abaate et al. (preprint).16 To overcome this, the World Health Organization (WHO) has endorsed the Xpert MTB/RIF Ultra, which relies on melt curve analysis to detect rifampicin sensitivity. The Xpert MTB/RIF Ultra is more specific for the diagnosis of rifampicin resistance than the Xpert MTB/RIF assay.17,18
This study was conducted in a high-TB-burden region of India, in which the highest prevalence of TB has been reported (the state of Karnataka, Kalaburagi). Data on rifampicin resistance and its pattern within this locality from the last five years was evaluated. As a result, probe E mutation was found to be the most common type, while probe C mutation was the least common. Approximately 8.3% of probes were missing, which may indicate a partial mutation. These results provide insights that will help clinicians understand the genetic basis of antitubercular drug resistance and initiate the right treatment. In addition, this study also highlights the importance of reading CT values in order to interpret rifampicin indeterminate results in cases where the minimum and maximum CT values do not cross 34.5. The latter can be overcome using the Xpert MTB/RIF Ultra or Shewhart control charts. Xpert MTB/RIF Ultra is superior to Xpert MTB/RIF in diagnosing rifampicin resistance because its results indicate either rifampicin sensitivity or rifampicin resistance rather than rifampicin indeterminate result. This highlights how the principle of detecting rifampicin resistance using Xpert MTB/RIF Ultra differs from that of Xpert MTB/RIF.
Limitation of the study
No standard comparative tests were used. Compared with the Xpert MTB/RIF assay, Xpert MTB/RIF Ultra was warranted.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the Institutional Ethics Committee, ESIC Medical College and Hospital, Kalaburagi, India, with reference number ESICMC/GLB/IEC(2)/05/2023.
- Sharma P, Singh R. GeneXpert MTB/RIF Based Detection of Rifampicin Resistance and Common Mutations in rpoB Gene of Mycobacterium Tuberculosis in Tribal Population of District Anuppur, Madhya Pradesh, India. J Clin Diagn Res. 2020;14(9):LM01-LM03.
Crossref - Rajendran P, Kumar MP, Thiruvengadam K, et al Characterization of probes associated with rifampicin resistance in M.tuberculosis detected by GenXpert from a national reference laboratory at Chennai. Tuberculosis. 2022;133:102182.
Crossref - Sailo CV, Lalremruata R, Sanga Z, et al. Distribution and frequency of common mutations in rpoB gene of Mycobacterium tuberculosis detected by Xpert MTB/RIF and identification of residential areas of rifampicin resistant-TB cases: A first retrospective study from Mizoram, Northeast India. J Clin Tuberc Other Mycobact Dis. 2022;29:100342.
Crossref - Mboowa G, Namaganda C, Ssengooba W. Rifampicin resistance mutations in the 81 bp RRDR of rpoB gene in Mycobacterium tuberculosis clinical isolates using Xpert® MTB/RIF in Kampala, Uganda: a retrospective study. BMC Infect Dis. 2014;14:481
Crossref - Alamgir M, Sajjad M, Baig SM, Noori YM. Mutational Frequencies in Mycobacterial rpoB gene using GeneXpert/ MTB Rif Assay in Rifampicin Resistant patients at a tertiary care setting in Urban Sindh, Pakistan: Analysis from a Five-Year Period. Pak J Med Sci. 2021;37(4):1151-1154.
Crossref - Ocheretina O, Byrt E, Maboub M-M, et al. False-positive Rifampin Resistant Results with Xpert MTB/RIF Version 4 Assay in clinical samples with a low bacterial load. Diagn Microbiol Infect Dis. 2016;85(1):53-55.
Crossref - Gupta A, Prakash P, Singh SK , Anupurba S. Rapid Genotypic Detection of rpoB and katG Gene Mutations in Mycobacterium tuberculosis Clinical Isolates from Northern India as Determined by MAS-PCR. J Clin Lab Anal2013;27(1):31-37.
Crossref - Asgedom SW, Teweldemedhin M, Gebreyesus H. Prevalence of multidrug-resistant tuberculosis and associated factors in Ethiopia: a systematic review. J Pathog. 2018;2018:7104921.
Crossref - Zhu M, Han G, Takiff HE, et al. Times series analysis of age-specific tuberculosis at a rapid developing region in China, 2011-2016. Sci Rep. 2018;8(1):8727.
Crossref - Alemu A, Tadesse M, Seid G, et al. Does Xpert® MTB/RIF assay give rifampicin resistance results without identified mutation? Review of cases from Addis Ababa, Ethiopia. BMC Infect Dis. 2020;20(1):87.
Crossref - Reddy R, Alvarez-Uria G. Molecular epidemiology of rifampicin resistance in Mycobacterium tuberculosis using the GeneXpert MTB/RIF assay from a rural setting in India. J Pathog. 2017;2017:6738095.
Crossref - Kanade S, Nataraj G, Mehta P, Shah D. Pattern of missing probes in rifampicin resistant TB by Xpert MTB/RIF assay at a tertiary care centre in Mumbai. Indian J Tuberc. 2019;66(1):139-143.
Crossref - Kaur R, Jindal N, Arora S, Kataria S. Epidemiology of rifampicin resistant tuberculosis and common mutations in rpoB Gene of Mycobacterium tuberculosis: a retrospective study from six districts of Punjab (India) Using Xpert MTB/RIF Assay. J Lab Phys. 2016;8(2):96-100.
Crossref - Munir MK, Shamim S, Rehman S, Hanif A, Saeed MS. Comparison of Genexpert® Probe missing in hotspot RRDR of rpoB gene among Primary and Acquired drug resistant cases of Pulmonary Tuberculosis. Biol Clin Sci Res J. 2022;3(1):165.
Crossref - Ochang EA, Udoh UA, Emanghe UE, et al. Evaluation of rifampicin resistance and 81-bp rifampicin resistant determinant region of rpoB gene mutations of Mycobacterium tuberculosis detected with XpertMTB/Rif in Cross River State. Nigeria Int J Mycobacteriol. 2016;5(Supl 1):S145-146.
Crossref - Abaate TJ, Alali AA. Analysis of rifampicin indeterminate results using shewhart control charts: implications for patients and tuberculosis control programmes. medRxiv. 2023.
Crossref - Dorman SE, Schumacher SG, Alland D, et al. Xpert MTB/RIF Ultra for detection of Mycobacterium tuberculosis and rifampicin resistance: a prospective multicentre diagnostic accuracy study. 2018:18(1):76-84.
Crossref - Ajbani K, Naik S, Kazi M, Shetty A, Rodrigues C. Interpreting very low Mycobacterium tuberculosis detected on Xpert Mycobacterium tuberculosis/rifampicin. Lung India 2019; 36(6):555-557.
© The Author(s) 2024. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.