ISSN: 0973-7510
E-ISSN: 2581-690X
Microbial sources are regularly used as reliable biocatalysts sources which are often used in the process and production industry. Demands for such organisms with greater capacity of intended enzyme production are on the rise. Lipase is important enzyme used in the biotechnological process of hydrolysis of fats in almost all the relevant industries We have utilized the local oil-contaminated soil resources to search for efficacious bacterial strains that have excellent lipase activity. We were successful in identifying two such bacterial sources, namely, Bacillus subtilis strain RCPS3 and Bacillus fumarioli strain RCPS4, responsible for lipase production from oil effluent contaminated soil of Telangana. This is the first report of these two strains from this part of India that are involved in lipase production. The strains were isolated, optimized, and purified using standard microbiology protocols and were characterized at the molecular level using the biomarker 16s ribosomal RNA genes of the strains. The identified and isolated bacterial strains were confirmed as Bacillus subtilis strain RCPS3, and Bacillus fumarioli strain RCPS4 through molecular and computational characterization.
Lipase, Bacillus subtilis, Bacillus fumarioli, lipase production, molecular characterization, oil-contaminated soil
Industrial biotechnology has proven its mettle in accelerating and benefitting multiple production processes for long1. Search for novel microorganisms and efficient biocatalysts remained an important part of the progress2. This ongoing process and efforts from the global research community successfully aided in the tremendous progress and contributed towards the ever-increasing industrial demand.
Biocatalysts remain the best solution till now in mediating an industrial process through accelerating or decelerating a particular reaction. It is cost-effective, easily producible, and highly specific in reaction mediation. Therefore, a huge demand for biocatalysts is present in different industries such as textiles, paints, food processing, beverage, and biomedicine. Fulfilling such requirements and meeting the industry standard for novel enzymes is an arduous and convoluted task for global researchers.
A large number of industrial enzymes are on regular demand for various essential processes. Lipase is one such unique and industrially important enzyme that is regularly used for various industrial processes3. Essentially, a lipase catalyzes the hydrolysis of the lipids or fats. This esterase family member is highly active and acts specifically on the glycerol backbone of lipid molecules. Industrially, in the baking and detergent industry, lipase is used abundantly along with generous use in fuel conversion from vegetable oil. The major source of lipase in the industry is fulfilled from the animal sources, but ample use of microbial sources is also on-demand. Moreover, there is an ever-growing need for more supply of microbial lipase in the industry. Lipase is in great demand due to the gigantic requirement for dairy product hydrolysis, flavor improvement in bakery product, improvement in aroma in beverages, transesterification process, flavor development in meat and fishes, esterification in cosmetics, biodegradable stain removal, and reduction of oil and grease through hydrolysis3. Several literature sources have documented such industrial applications of lipase and its origin from microbial sources4-5.
According to the enzyme nomenclature, recommended lipases are known as triacylglycerol acyl hydrolase, bearing the Enzyme Commission number (EC number) of EC 3.1.1.35. Sustainability, green production, and application of lipases can induce the lipase enzyme production further from the microbial sources6. The global food and feed industry enzyme market has usage of almost 10% lipase for various purposes7 which is having almost 8% annual growth7. Thus, lipase sources, production are the present industry requirement8. Searching novel enzyme sources will always aid in the further development of the industry that extensively uses lipase as a biocatalyst8. Several other literature sources have extensively documented the industrial need, applications, and effectiveness of using lipase9-10.
Urban waste management has become another challenge for our societies. The ever-growing population in the urban area, poor sanitation, lack of planning, and limited resources are fueling the poor management of urban waste. In such conditions, utilization of waste resources, especially for industrial production could serve as a great solution. Therefore, effluents, specifically oil contaminated waste, and waste volatile substances, could be an excellent source that can be utilized11. Different waste sources and bacterial origins have been used for better production of lipases, such as agricultural industrial waste and Aspergillus niger12. Scaling-up of the lipase production to meet the industry requirements is another important aspect that should be addressed with different technical and feasible approaches. Proper media sources may serve as a solution13. Solid fermentation technology and fungal sources have been also utilized for better lipase enzyme production14. Therefore, attention should be given to the isolation process, purification, and characterization of the enzyme production15-16. Presently, scientific research and investigations in this direction have turned into interdisciplinary and better outcomes are being drawn from computational approaches, chemical engineering, application microbiology and biotechnology, genetic engineering, and other disciplines.
In the present study, we have isolated, optimized, and characterized lipase enzymes from thermophilic Bacillus subtillis and Bacillus fumarioli that were collected from oil-contaminated soils of Warangal district.
Sample collection
Different oil-contaminated regions around the Warangal district were surveyed and various soil samples were collected that were contaminated with oil effluents.
Isolation of lipase producing bacteria
Isolation of lipase producing bacteria was done by the spread plate method using the serial dilution process on the nutrient agar medium. Bacterial colonies were grown and isolated by streaking over the Tributyrin and Rhodamine-B agar medium. Halos formed around the hydrolytic zones and the bacterial colonies with positive results were identified and separated. Such colonies were subcultured again for future preservation and use. The positive colonies were subcultured in the TBA (Tributyrin agar medium) medium in test tubes by preparing slants. These slants were inoculated with positive colonies and incubated for 24 hours. After incubation these slants were preserved at 4°C for further experiements.
Screening of bacterial strains
Primary screening of the bacterial strains was done by the method suggested by Limpon et al. (2006)17. Following the screening of the bacterial strains, the lipolytic activity of the isolated strains was assessed.
Lipase assay
The lipase activity was measured by the titrimetric method18. The isolated strains that showed positive results for lipase production, were further isolated for the production process.
Inoculum medium was prepared using 10 gm peptone, and 10 gm sodium acetate (CH3COONa). Other component used were 3H2O,10 gm yeast extract, 20 gm glucose, 0.03 gm manganese sulfate (MnSO4), 0.09 gm magnesium sulfate (MgSO4), copper sulfate 1.5 gm (CuSO4), 0.5 gm Potassium chloride (KCl), 5 ml Olive oil, and 1000 ml of distilled water. The pH was set to 10.8. Following the stock preparation, 15 ml of prepared inoculum was used for the inoculation of the isolated culture. Later, the inoculated flasks were incubated in the rotary shaker overnight at 120 rpm and 37°C temperature.
The production medium was prepared using 0.24 gm MgSO4, 2.5 gm Sodium chloride (NaCl),1 gm Ammonium dihydrogen phosphate (NH4H2PO4), 2 gm Peptone,0.24 gm Calcium dichloride hydrate (CaCl2H2O), Olive oil 12 ml, and 2-3 drops of tween 20 for every 1000 ml of water.
Further, 1 ml of the inoculated culture was added to the Erlenmeyer flasks containing 100 ml of production medium and incubated for 24 hours on a rotary shaker at 120 rpm, and the temperature was maintained at 37°C (temperature for wild type)19.
For every 24 hours, the inspection was done for consecutive 5 days. Every day, 1 ml of culture from the production medium was centrifuged at 8000 rpm for 60 mins. The centrifuged supernatant of cell-free culture was used as the crude enzyme source of lipase. The lipolytic activity was determined by the titrimetric method mentioned earlier.
Lipase activity
The lipase activity was measured by the titrimetric method using olive oil as substrate. To 1 ml of culture supernatant, 1ml of olive oil and 2 ml of phosphate buffer was added and incubated for 60 mins at 37°C temperature. After completion of incubation, the reaction was halted by adding 1 ml of acetone-ethanol solution that was prepared in 1:1 proportion. Later, the amount of acid released was determined by titration with 0.05M sodium hydroxide (NaOH) at pH 7.0 and the temperature was maintained at 30°C.
Assessment of Carbon sources
Different types of sugars were added as carbon sources and were optimized. Sugars such as Lactose, Maltose, and Dextrose hindered the lipase production, while starch enhanced of the lipase production. The best carbon source was identified as starch for lipase production in the present study. In many cases, reduction in the production of lipase was noted in the presence of sugars that were used as carbon sources, this might have happened due to catabolite repression as the carbon sources were already available in the medium.
Nitrogen sources
The effect of various organic and inorganic nitrogen sources was examined for better lipase production. Organic sources that were used were tryptone, yeast extract, beef extract, protease, and peptone. Considered inorganic nitrogen sources were ammonium nitrate and calcium nitrate. These nitrogen sources were added to the final concentration of 1% (w/v) to the medium and incubated for 24 hours in a rotary shaker of 150 rpm at 36°C.
Effect of temperature, pH, and agitation on the lipase production
The effect of temperature, pH, and agitation was examined for lipase production by altering the pH of the medium between 05.0 and 10.0. Similarly, the temperature was also altered between 20°C and 50°C. Moreover, the effect of agitation was tested by incubating the medium with culture on the orbital shaker with different agitation speed adjustments, ranging from 110 rpm to 200 rpm, for 24 hours.
Screening
The oil spilled soils that were collected from different areas of Warangal oil mills were used as a source for bacterial samples. These soil samples were serially diluted and then spread on the TBA plates by spread plate method. A total of 100 colonies showed hydrolytic zones that were displaying clear hallows around the colonies. Among the 100 colonies, we found 20 colonies having lipase positive bacterial strains. Upon screening, these 20 isolates were identified with the highest lipolytic activity. Further, the molecular characterization was done for these isolates.
Molecular identification of the isolated bacterial strains
To characterize the isolated strains further, DNA was isolated from the culture and the quality of the isolated DNA was evaluated using 1.0% agarose gel, and a single band of high molecular weight DNA was observed. Later on, the fragment of the 16S rDNA gene was used as a biomarker, hence, amplified by 27F and 1492R primers. A single discrete PCR amplicon band of 1500 bp was generated. Next, the PCR amplicon was purified to remove contaminants. Forward and reverse DNA sequencing reaction of PCR amplicon was carried out with forward and reverse primers using the BDT v3.1 Cycle sequencing kit on ABI 3730xl Genetic Analyzer.
The consensus sequence of the 16S rDNA gene was generated from the forward and the reverse sequence data using Aligner software. In the next stage, bioinformatics analysis was conducted. The 16S rDNA gene sequence was used to carry out BLAST with the NCBI GenBank database. Based on the maximum identity score, the first ten sequences were selected and were aligned using multiple sequence alignment program ClustalW. The distance matrix was generated and the phylogenetic tree was constructed using MEGA7.
Survey and soil sample collection
A detailed survey of oil-contaminated regions was done before the sample collection. Possible collection spots were identified and soil samples were collected from the designated places. The collected samples were used as potential bacterial sources that may have better lipase activity. The samples were containing oil and effluent mixed soil which were dry in nature.
Isolation and screening of bacterial strains
As mentioned in the methodology section, the bacterial samples were isolated applying serial dilution and spread-plate technique. Nutrient agar was used initially, and the Tributyrin agar medium was used later. A total of 70 bacterial colonies were obtained, out of which 30 were positive. From these 30, further we have chosen 10 positive colonies. The strains were morphologically circular, white, smooth, gram negative, cocci, and motile in appearance.
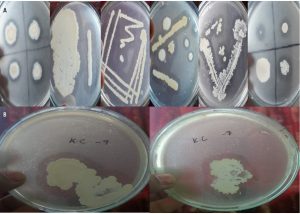
The obtained colonies are represented in Fig. 1. Different colony formation is presented in Fig. 1(A) and Fig. 1(B). The Figs. display the efficient lipase activity of some of the colonies.
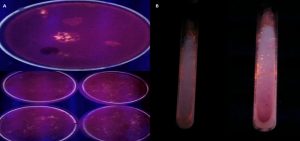
Positive colonies were detected for further use through the Rhodamine-B agar medium as mentioned in the prior section. Fig. 2 presents the results obtained from the assessment of the colonies using the Rhodamine-B medium. Detection of the colonies is shown in Fig. 2(A) and the formation of the halos is shown in Fig. 2(B).
Production medium for growing isolated bacterial strains
The increased production of the isolated strains from the colony was important to estimate the lipase activities of the strains. Hence, they were systematically grown in the production medium. A detailed description of the growth of the bacterial strains in the production medium is shown in Fig. 3. In Fig. 3, it is evident that the sample 1 and sample 2 have slight growth, whereas sample 3 and sample 4 are showing yellow color due to lipolysis. Besides, sample 5 is showing the color of light greenish-yellow due to the slow or reduced pace of lipolysis. Sample 6 presents no growth and was white as the fresh production medium. A total of 6 samples were inoculated in 6 flasks and incubated for lipase activity . The flasks 3, 4 and 5 showed lipolytic activity.
Fig.3. Enriched production medium and relevant outcomes for growth of extracellular lipase producing bacteria.
Assessment of Lipase activity by the bacterial strains
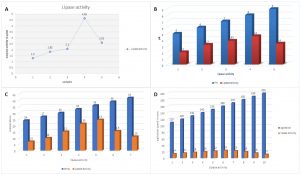
The lipase activity was assessed with extreme care for the selected bacterial strains. The observed results are presented in Fig. 4. Fig. 4 (A) presents the lipase activity of the sample where activities were measured in µl/ml. The maximum lipase activity value observed was 4.65 µl/ml whereas the lowest lipase activity recorded was 1.3 µl/ml.
Fig.4. Outcomes of the optimation done for the lipase production . (A) A presentation of lipase activity for the samples considered. (B) Effect of pH on lipase activity. (C) Effect of temperature on lipase activity. (D) Effect of agitation on lipase activity.
Optimization of Lipase activity
Along with lipase activity assessment, influencing factors were examined to optimize the lipase production capacity of the selected bacterial strains. Therefore, pH (Fig. 4B), temperature (Fig. 4C), and agitation (Fig. 4D) were tested extensively. The optimum pH was pH 8.0 with 4.6 µl/ml lipase activity. Similarly, the optimum temperature was observed at 36°C with 24.5 µl/ml lipase activity. Agitation was decided through the rpm as shown in (Fig. 4D), 160 rpm was found optimum with 24 µl/ml lipase activity.
Optimization of nutrient resources
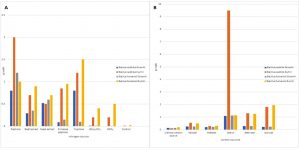
Nutrients resources play a vital role in bacterial growth. Therefore, optimization of the nutrient resources, specifically the carbon and nitrogen sources are important to optimize the lipase production and activity. Various nitrogen and carbon sources were considered and comparative growth of the selected strains was assessed. Fig. 5 presents the comparative estimation of growth in the presence of different nitrogen sources (Fig. 5A) and also in the presence of various carbon sources (Fig. 5B). Peptone was the optimum nitrogen source for both Bacillus subtilis and Bacillus fumarioli strains. Tryptone was a good source for Bacillus subtilis and yeast extract was effective for Bacillus fumarioli strains. Similarly, starch was found an excellent carbon source for both Bacillus subtilis and Bacillus fumarioli. Unfortunately, other carbon sources did not provide better outcomes.
Fig.5. Results of the varying carbon and nitrogen nutrition sources used. (A) Effect of nitrogen sources on the medium. (B) Effect of carbon sources on the medium.
Molecular characterization
Better results in the previous experimentations inspired us to further characterize and identify the isolated bacterial strains through molecular approaches.16SDNA based marker was used in this experimentation as mentioned in the method section. The primers used were, 27F and 1492R primers ((5→3´) for forward strand: “AGAGTTTGATCMTGGCTCAG”, and (3´→5´) for reverse strand: “AAGTCGTAACAAGGT”). The PCR amplicon band was developed in Agarose gel. Two different bacterial strain samples (Bacillus subtilis (sample SOSS_7_4; Accession Number: (MN173874), and Bacillus fumarioli (sample POSS_8_2; Accession Number: MT645092) that had maximum lipase activity were used for final molecular characterizations. Fig. 6 and Fig. 7 presents the results of the molecular characterizations.
Fig.6. Molecular identification of the samples isolated. (A) PCR amplification of isolated sample POSS_8_2. (B) DNA sequencing results from the amplified samples. (C) Development of a phylogenetic tree for sample characterization.
After DNA isolation, purification, and PCR amplification, the samples were assessed in the presence of the 100bp ladder marker (Promega, USA) in an agarose gel (Fig. 6A, Fig. 7A). Further, eluted DNA was sequenced for species identification. The obtained sequences are represented in Fig. 6B and Fig. 7B for both the samples.
Fig.7. (A) PCR amplification of isolated sample SOSS_7_4. (B) Sequencing result of the DNA of the sample. (C) Development of a phylogenetic tree for sample identification. The consensus sequence was used for species identification and BLAST analysis was done for both the sequences generated. The results of the top 10 BLAST hits are presented in Table 1 and Table 2 (sample POSS_8_2 and SOSS_7_4).
Table (1):
Results obtained after the BLAST analysis of sample POSS_8_2.
Sequences producing significant alignments: Description |
Max Score |
Total Score |
Query Cover |
E value |
Per. Ident |
Accession |
|---|---|---|---|---|---|---|
Bacillus niacini strain IFO15566 16S ribosomal RNA, partial sequence |
2254 |
2254 |
98% |
0 |
96.70% |
NR_024695.1 |
Bacillus drentensis strain IDA1967 16S ribosomal RNA, partial sequence |
2248 |
2248 |
98% |
0 |
96.63% |
NR_029002.1 |
Bacillus novalis strain IDA3307 16S ribosomal RNA, partial sequence |
2246 |
2246 |
98% |
0 |
96.56% |
NR_042168.1 |
Bacillus bataviensis strain IDA1115 16S ribosomal RNA gene, partial sequence |
2242 |
2242 |
98% |
0 |
96.56% |
NR_036766.1 |
Bacillus niacini strain NBRC 15566 16S ribosomal RNA, partial sequence |
2231 |
2231 |
97% |
0 |
96.81% |
NR_113777.1 |
Bacillus drentensis strain NBRC 102427 16S ribosomal RNA, partial sequence |
2230 |
2230 |
97% |
0 |
96.81% |
NR_114085.1 |
Bacillus fumariolis train NBRC 102428 16S ribosomal RNA, partial sequence |
2228 |
2228 |
97% |
0 |
96.73% |
NR_114086.1 |
Bacillus novalis strain NBRC 102450 16S ribosomal RNA, partial sequence |
2226 |
2226 |
97% |
0 |
96.73% |
NR_114094.1 |
Table (2):
Table 2: Results obtained after the BLAST analysis of sample SOSS_7_4.
Sequences producing significant alignments: Description |
Max Score |
Total Score |
Query Cover |
E value |
Per. Ident |
Accession |
|---|---|---|---|---|---|---|
Bacillus haynesii strain NRRL B-41327 16S ribosomal RNA, partial sequence |
1317 |
2548 |
100% |
0 |
98.78% |
NR_157609.1 |
Bacillus licheniformis strain DSM 13 16S ribosomal RNA, partial sequence |
1312 |
2543 |
100% |
0 |
98.65% |
NR_118996.1 |
Bacillus sonorensis strain NBRC 101234 16S ribosomal RNA, partial sequence |
1312 |
2530 |
99% |
0 |
98.65% |
NR_113993.1 |
Bacillus licheniformis strain BCRC 11702 16S ribosomal RNA, partial sequence |
1312 |
2526 |
99% |
0 |
98.65% |
NR_116023.1 |
Bacillus licheniformis strain ATCC 14580 16S ribosomal RNA, partial sequence |
1306 |
2532 |
100% |
0 |
98.51% |
NR_074923.1 |
Bacillus sonorensis strain NRRL B-23154 16S ribosomal RNA, partial sequence |
1306 |
2408 |
95% |
0 |
98.51% |
NR_025130.1 |
Bacillus licheniformis strain NBRC 12200 16S ribosomal RNA, partial sequence |
1304 |
2524 |
99% |
0 |
98.38% |
NR_113588.1 |
Bacillus subtilis strain IAM 12118 16S ribosomal RNA, complete sequence |
1277 |
2447 |
100% |
0 |
97.84% |
NR_112116.2 |
Bacillus mojavensis strain NBRC 15718 16S ribosomal RNA, partial sequence |
1277 |
2439 |
99% |
0 |
97.84% |
NR_112725.1 |
Our molecular and bioinformatics analysis confirmed that the isolated, separated, and characterized bacterial strains were Bacillus subtilis (sample POSS_8_2), and Bacillus fumarioli(sample SOSS_7_4). Further, it was confirmed that the strains were thermophilic and they were Bacillus subtilis strain RCPS3 and Bacillus fumarioli strain RCPS4 (Accession Number: MT645092).All these 10 sequences for both the query sequences were collected and multiple sequence alignment was developed for individual cases, respectively. Next, the phylogenetic tree was developed using the MEGA7 tool to understand their respective phylogenetic distances as presented in Table 3 and Table 4.
Table (3):
Phylogenetic distance estimated using MEGA7 for the sample POSS_8_2.
POSS_8_2 |
0.004 |
0.003 |
0.003 |
0.004 |
0.004 |
0.003 |
0.004 |
0.003 |
0.004 |
0.003 |
NR_024695.1 |
0.016 |
0.002 |
0.003 |
0.003 |
0.000 |
0.002 |
0.004 |
0.003 |
0.003 |
0.003 |
NR_029002.1 |
0.016 |
0.007 |
0.003 |
0.002 |
0.002 |
0.000 |
0.004 |
0.003 |
0.002 |
0.002 |
NR_042168.1 |
0.015 |
0.013 |
0.009 |
0.003 |
0.003 |
0.003 |
0.004 |
0.001 |
0.003 |
0.002 |
NR_036766.1 |
0.017 |
0.010 |
0.006 |
0.009 |
0.003 |
0.002 |
0.004 |
0.003 |
0.000 |
0.003 |
NR_113777.1 |
0.016 |
0.000 |
0.007 |
0.013 |
0.010 |
0.002 |
0.004 |
0.003 |
0.003 |
0.003 |
NR_114085.1 |
0.016 |
0.007 |
0.000 |
0.009 |
0.006 |
0.007 |
0.004 |
0.003 |
0.002 |
0.002 |
NR_114086.1 |
0.017 |
0.018 |
0.018 |
0.016 |
0.018 |
0.018 |
0.018 |
0.004 |
0.004 |
0.004 |
NR_114094.1 |
0.016 |
0.014 |
0.010 |
0.001 |
0.010 |
0.014 |
0.010 |
0.017 |
0.003 |
0.003 |
NR_114093.1 |
0.017 |
0.010 |
0.006 |
0.009 |
0.000 |
0.010 |
0.006 |
0.018 |
0.010 |
0.003 |
NR_148626.1 |
0.013 |
0.011 |
0.009 |
0.009 |
0.010 |
0.011 |
0.009 |
0.020 |
0.009 |
0.010 |
Table (4):
Phylogenetic distance estimated using MEGA7 for the sample SOSS_7_4.
SOSS_7_4 |
0.004 |
0.004 |
0.004 |
0.004 |
0.004 |
0.004 |
0.004 |
0.005 |
0.005 |
0.005 |
NR_118996.1 |
0.014 |
0.004 |
0.003 |
0.000 |
0.001 |
0.003 |
0.000 |
0.005 |
0.005 |
0.005 |
NR_113993.1 |
0.014 |
0.004 |
0.005 |
0.003 |
0.003 |
0.001 |
0.003 |
0.006 |
0.006 |
0.006 |
NR_116023.1 |
0.014 |
0.004 |
0.000 |
0.005 |
0.001 |
0.003 |
0.000 |
0.005 |
0.005 |
0.005 |
NR_074923.1 |
0.015 |
0.005 |
0.001 |
0.007 |
0.001 |
0.003 |
0.001 |
0.005 |
0.005 |
0.005 |
NR_025130.1 |
0.015 |
0.005 |
0.007 |
0.001 |
0.007 |
0.008 |
0.003 |
0.006 |
0.006 |
0.006 |
NR_113588.1 |
0.014 |
0.004 |
0.000 |
0.005 |
0.000 |
0.001 |
0.007 |
0.005 |
0.005 |
0.005 |
NR_112116.2 |
0.021 |
0.019 |
0.021 |
0.023 |
0.021 |
0.022 |
0.025 |
0.021 |
0.002 |
0.000 |
NR_112725.1 |
0.021 |
0.019 |
0.021 |
0.023 |
0.021 |
0.022 |
0.025 |
0.021 |
0.003 |
0.002 |
NR_113265.1 |
0.021 |
0.019 |
0.021 |
0.023 |
0.021 |
0.022 |
0.025 |
0.021 |
0.000 |
0.003 |
Many studies have been conducted on novel lipase producing bacteria identification where oil spilled and petrol spilled soils were used20-23. Successful isolation of bacterial strains requires further optimization of lipase production. Assessment of the enzyme production activity in the laboratory, followed by pilot-scale evaluation and industrial-scale quantification is necessary.
The amalgamation of different advanced culture techniques24-26, production27-29, and molecular techniques30-32, and advanced bioinformatics analysis33-36 can provide the intended results with maximum accuracy and help in the identification of exact strains with all necessary information. In the present study, we have isolated, assessed, and optimized novel species that are having excellent lipase production and activities from oil-contaminated soil sources in Warangal district of Telangana state, India. As a next step, pilot-scale and industrial-scale production can be attempted to use the identified and isolated organisms for the industry.
Identification of novel and cheap sources of efficient lipase producing bacterial strains is of pivotal importance considering the excessive industrial applications. Moreover, oil effluent sources can also help in managing environmental aspects. In the present study, we were successful in identifying, isolating, optimizing, and characterizing bacterial strains that are efficacious in lipase production and lipase activity from oil-contaminated soil samples in our region.
This work would help in further using the novel bacterial species that were found efficacious in lipase production for the industry. Necessary action should be taken for further optimization and advanced molecular techniques can be used to scale-up the activities of the strains.
ACKNOWLEDGMENTS
We are thankful to Head Department of Biotechnology,for providing necessary facilities and the financial assistance received from UGC-NFST is gratefully acknowledged.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest .
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This study was supported by grants from the UGC-NFST (2015-16/NFST-2015-17-ST-TEL-2636).
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Hatti-Kaul R, Tornvall U, Gustafsson L, Borjesson P. Industrial biotechnology for the production of bio-based chemicals–a cradle-to-grave perspective. Trends in Biotechnology. 2007;25(3):119-124.
Crossref - Cheetham PSJ. Screening for novel biocatalysts. Enzyme and Microbial Technology, 1987; 9(4):194-213.
Crossref - Verma N, Thakur S, Bhatt AK. Microbial lipases: industrial applications and properties (a review). International Research Journal of Biological Sciences. 2012;1(8):88-92.
- Andualema B, Gessesse A. Microbial lipases and their industrial applications: Review. Biotechnology. 2012;11(3):100-118.
Crossref - Houde A, Kademi A, Leblanc D. Lipases and their industrial applications: an overview. Appl Biochem Biotechnol. 2004;118(1-3):155-170.
Crossref - Agobo KU, Arazu VA, Uzo K, Igwe CN. Microbial Lipases: A Prospect for Biotechnological Industrial Catalysis for Green Products: A Review. Ferment Technol. 2017;6(144):2.
- Guerrand D. Lipases industrial applications: focus on food and agroindustries. 2017;24(4).
Crossref - Sarmah N, Revathi D, Sheelu G, et al. Recent advances on sources and industrial applications of lipases. Biotechnol Progress. 2018;34(1):5-28.
Crossref - Seitz EW. Industrial application of microbial lipases: a review. J Am Oil Chem Soc. 1974;51(2):12-16.
Crossref - Joshi R, Kuila A. Lipase and their different industrial applications: A review. Brazilian Journal of Biological Sciences. 2018;5(10):237-247.
Crossref - Muthumari GM, Thilagavathi S, Hariram N. Industrial Enzymes: Lipase Producing Microbes from Waste Volatile Substances. International Journal of Pharmaceutical Sciences and Research. 2016;7(5):2201-2108.
- Costa TM, Hermann KL, Garcia-Roman M, Valle RdCSC, Tavares LB. Lipase production by Aspergillus niger grown in different agro-industrial wastes by solid-state fermentation. Braz J Chem. 2017;34(2):419-427.
Crossref - Soleymani S, Alizadeh H, Mohammadian H, et al. Efficient media for high lipase production: One variable at a time approach. Avicenna J Med Biotechnol. 2017;9(2):82-86. PMID: 28496947
- Kumar DS, Ray S. Fungal lipase production by solid state fermentation-an overview. Journal of Analytical and Bioanalytical Techniques. 2014;6(230):1-10.
- Zouaoui B, Bouziane A. Isolation, Optimisation and purification of lipase production by Pseudomonas Aeruginosa. Journal Biotechnol Biomaterial. 2011;7(1):345-51.
Crossref - Sharma R, Chisti Y, Banerjee UC. Production, purification, characterization, and applications of lipases. Biotechnol Adv. 2001;19(8):627-662.
Crossref - Bora L, Kar A, Gogoi HK. Isolation and screening of bacterial strains for extracellular enzymatic potential from hot springs of Lohit district of Arunachal Pradesh, India. Asian J Microbiol Biotechnol Environ Sci. 2006; 8(2):267-270.
- Venkateshwarlu N, Reddy SM. Production of lipase by five thermophilic fungi. Indian J Microbiol. 1993;33(2):119-124.
- Kohno M, Enatsu M, Funatsu J, Yoshiizumi M, Kugimiya W. Improvement of the optimum temperature of lipase activity for Rhizopus niveus by random mutagenesis and its structural interpretation. Journal of Biotechnology. 2001;87(3):203-210.
Crossref - Bharathi D, Rajalakshmi G, Komathi S. Optimization and production of lipase enzyme from bacterial strains isolated from petrol spilled soil. J King Saud Univ Sci. 2018;31(4):898-901.
Crossref - Veerapagu M, Narayanan AS, Ponmurugan K, Jeya KR. Screening selection identification production and optimization of bacterial lipase from oil spilled soil. Asian J Pharm Clin Res. 2013;6(3):62-67
- Golani M, Hajela K, Pandey GP. Screening, identification, characterization and production of bacterial lipase from oil spilled soil. Int J Curr Microbiol Appl Sci. 2016;5(3):745-763.
Crossref - Mohan TS, Palavesam A, Immanvel G. Isolation and characterization of lipase-producing Bacillus strains from oil mill waste. African Journal of Biotechnology. 2008;7(15).
- Grooters AM, Whittington A, Lopez MK, Boroughs MN, Roy AF. Evaluation of microbial culture techniques for the isolation of Pythium insidiosum from equine tissues. J Vet Diagn Invest. 2002;14:288-294.
Crossref - Fujimoto K, Ogawa M, Higashi K, Miki N. Micro-tube mass production device for microbial culture. 2016 doi: 10.1109/EMBC.2016.7590744. In 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) (pp. 481-484). IEEE.
- Okafor N. Modern industrial microbiology and biotechnology. CRC Press. 2016.
Crossref - Srivastava N, Srivastava M, Ramteke PW, Mishra PK. Solid-State Fermentation Strategy for Microbial Metabolites Production: An Overview. In New and Future Developments in Microbial Biotechnology and Bioengineering, Elsevier. 2019:345-354.
Crossref - Carroll AL, Desai SH, Atsumi S. Microbial production of scent and flavor compounds. Curr Opin Biotechnol. 2016;37:8-15.
Crossref - Chubukov V, Mukhopadhyay A, Petzold CJ, Keasling JD, Martin HG. Synthetic and systems biology for microbial production of commodity chemicals. npj Syst Biol Appl. 2016;2:16009
Crossref - Torsvik V, Daae FL, Sandaa RA, Ovreas L. Novel techniques for analysing microbial diversity in natural and perturbed environments. J Biotechnol. 1998;64(1):53-62.
Crossref - Ranjard L, Poly F, Nazaret S. Monitoring complex bacterial communities using culture-independent molecular techniques: application to soil environment. Res Microbiol. 2000;151(3):167-177.
Crossref - Pereira RM, Silveira EL, Scaquitto DC, et al. Molecular characterization of bacterial populations of different soils. Braz J Microbiol. 2006;37:439-447
Crossref - Banerjee AK, Harikrishna N, Kumar JV, Murty US. Towards classifying organisms based on their protein physicochemical properties using comparative intelligent techniques. Applied Artificial Intelligence. 2011;25(5):426-439.
Crossref - Devulder G, Perriere G, Baty F, Flandrois JP. BIBI, a bioinformatics bacterial identification tool. J Clin Microbiol. 2003;41(4):1785-1787.
Crossref - Arora N, Banerjee AK, Mutyala S, Murty US. Comparative characterization of commercially important xylanase enzymes. Bioinformation. 2009;3(10):446-453.
Crossref - Livny J, Waldor MK. Identification of small RNAs in diverse bacterial species. Curr Opin Microbiol. 2007;10(2):96-101.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.