ISSN: 0973-7510
E-ISSN: 2581-690X
Acinetobacter baumannii became a serious endemic and widespread pathogen responsible for causing nosocomial infections due to restricted treatment options. This study was conducted to evaluate the role of both antibiotic resistance and genetic pattern in infection caused by MBLs resistance A. baumannii isolated in Baghdad hospitals. Finally, developing selective media in order to detect multidrug resistance A. baumannii isolates according to colony shape appearance changing. Collected 124 isolates from various clinical and environmental sources. Biofilm Formation was detected, Antibiotic sensitive for 21 antibiotic discs were determined, MDR index calculated, PCR was performed to investigate MBLs genes bla (IMP2, KPC, DIM, SPM, VIM, GIM, AIM, BIC, SIM, NDM, Pre-NDM-A, GES, Imp-1, and VIM). Increasing resistant prevalence of A. baumannii appear significantly in higher rates to seven antibiotics group and MDR index ranged between 0.29 to 1.0 from 6-21 antibiotics. It was clear in Biofilm formation evaluated 75.8% strong biofilm after 72 hours, indicated that there is a significant negative correlation between biofilm formation capacity and the inherent ability of bacteria to show multidrug resist (P< 0.001). Our findings indicated that non-MDR isolates tended to form more robust biofilm formation. Metallo b-Lactamase detection phenotype showed 113/124 isolates able to produced ESBL, a result confirmed by PCR assay to detect the resistance of MBL-genes, appeared that a high percentage of blaIMP 79% which left clinicians with limited treatment options. An increasing number of hospital MDR A. baumannii has been reported. Furthermore, the study will focus on the evolution and the emergence of multidrug resistance A. baumannii in Iraq. The common MBL gene is the blaIMP
Acinetobacter baumannii; Metallo-β-lactamase genes; Biofilm; Multidrug resistance.
Acinetobacter baumannii is one of the most important opportunistic pathogens that cause outbreaks in hospitals, health care and also associated with complications in hospitalized patients1.
Unstable genetic element, tolerate harsh environments, and Metallo ג-lactamases (MBLs) enzyme have led to the development of MDR resistance and epidemic A. baumannii2,4.
Different mechanisms of resistance and resistant to all commercial antibiotic caused alarming problems in a limited choice of antibiotic for the treatment of MDR. A. baumannii isolates3.
Increasing of MDR hospital isolates A. baumannii isolate has been stated from many countries around the world. Plus, inter-hospital environment dispersal of multidrug-resistant A. baumannii has been observed5. The ability of A. baumannii to rapid genetic change owing to their ability to acquire genes horizontally and vertically6.
This study came to highlight the current patterns of resistance to antibiotics as well as the prevailing genetic patterns of resistance in A. baumannii in order to achieve effective treatment in Baghdad hospitals for these bacteria with multiple resistance to antibiotics, which represents a threat to public health.
Sample collection
The study was carried out from December 2017 through March 2018 and involved 6 hospitals: Children’s Educational Hospital, Baghdad Teaching Hospital, Al-Hawraq Hospital, Educational Laboratories (Medical City), Imam Ali Hospital and Al-Kindi Hospital. After the patients were examined by the physician, samples were collected from various sources: wounds, burns, ulcers, spinal cord fluid-blood, blood, body fluid and swabs of the hospital environments and tested using conventional microbiological methods.
Laboratory Identification of Isolates
Morphological Examination:
The samples were diagnosed including Gram staining, traditional culture and using the selective media such as the chromo agar Acinetobacter medium and use modified Leeds agar base media by replacing fructose and sucrose and Manitol by xylose and glucose and add crystal violet dry powder dispense from selective media. Furthermore, the results of the identification of A. baumannii were confirmed by the API 20E system according to Forbes 7.
Temple preparation for PCR
Template DNA was prepared by the boiling method as described by8.
Molecular detection of A. baumannii
In order to confirm the detection, isolates were subjected to PCR against the 16s rRNA, oxa51 and RecA, PCR mixture and amplification were done as explained by9,10,11 respectively.
Antimicrobial susceptibility and ג-Lactamase detection
Isolates were plated on Mueller–Hinton agar and their susceptibilities to different antibiotics:Ampicillin/Sulbactam 20-10g (SAM30), Piperacillin/Tazobactam 100-10g (PTZ110),piperacillin (PRL100), Ticarcillin /Clavulanic acid 75-10ug (TIM85), Ceftazidime 30ug (CAZ30), Cefotaxime 30ug (CTX30), Ceftriaxone 30ug (CRO30), Imipenem 10ug (IMI10), Meropenem 10ug (MEM10), Colistin Sulphate 25 ug(CO25 ), Gentamicin 30 mg(GN 30), Tobramycin 10ug (TN10), Amikacin 30µg (AK30), Netilmicin 30ug (NET30), Doxycycline 30ug (DXT30), Tetracycline 30ug (T30), Lofloxacine 5ug (LEV5), Ciprofloxacin 5µg (CIP 5), Trimethoprim – sulfamethoxazole1.25/23.75µg (SXT1.25/23.75 ), were tested by disk diffusion method according to the Clinical and Laboratory Standard Institutes guidelines (2017).
Modified Hodge test (MHT)
Performed according to Bonnin12, for detection of carbapenemase production, Metallo ג-Lactamase and ESBLs production among isolates by using modified indirect three-dimensional methods according3.
Biofilm Formation
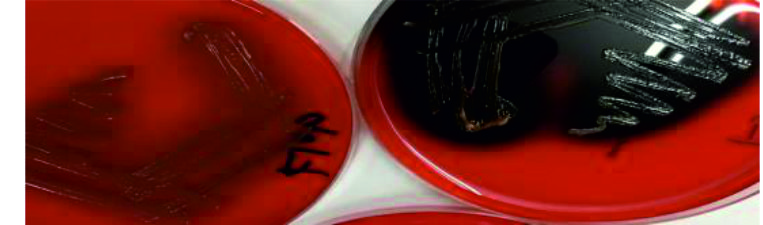
Congo red agar (CRA) method (qualitative Biofilm production assay): A simple qualitative assay for detection of biofilm was described by Mathur13.With modification as follows: (40) g\l blood agar base, (0.8) mg Congo red,(5)g\l d-glucose, (5) g/l xylose. These components were dissolved in 1L of distilled water and autoclaved. The agar then was dispensed into sterile Petri dishes and kept at 4°C until being used. Following inoculation, the agar plates were incubated at 37°C for 24–48 h. The appearance of black colonies with a dry crystalline consistency could be considered as strong evidence for the ability to form a biofilm. Each experiment was conducted in three repeats
Tissue culture plate method (quantitative biofilm production assay)
Generally considered to be the gold-standard technique for biofilm detection, Adopted the prescribed route by Lotfi14.
Motility Assay
Motility Assay Performed as per15.
PCR amplification of Metallo b –lactam genes:
PCR method was used for screening of the Metallo b-lactam genes: bla (IMP2, KPC, DIM, SPM, VIM, GIM, AIM, BIC, SIM, NDM, Pre-NDM-A, GES, Imp-1, and VIM). The primers and PCR programs used in this study were as previously described by (16; 17; 18 and 5 ). PCR product bands were analyzed after electrophoresis in a 1% agarose gel at 95 V for 45 min in 1X TBE containing Ethidium Bromide under UV radiation8.
Statistical analysis
One-way ANOVAs was applied exploitation SPSS 21 so as to match variations among Microtiter plate below standard and changed conditions. The experiments were performed in triplicate. P values of £ 0.05 were thought of as significant31
A total of 124A. baumanii isolates were collected from patients infected with significant and life-threatening diseases show in Fig. (1&2). The specimens were as follows: C.S.F 44/124 (35.5%), blood 32\124 (25.8%), sputum7/124 (5.65%), urine 5/124 (4.03%), burn and wounds and post-surgical swab 23/124 (18.54%), body fluid 3\124(2.42%), and swab from environment swab 10\124(8.06%). The diseases from which A. baumanii bacteria isolated were wound and urinary tract infection, septicemia and bacterima, pneumonia, and meningitis. Hence, the most prevalent disease related to A. baumanii was meningitis followed by septicemia. The age of patients ranged from 1 day to 70 years and male to female ratio was 1.375. And the child\adult ratio of 1.19.
 Fig. 1. Colony of Acinetobacter baumanii. A: On Chromo agar; B: OnMacConckey aga; C: On Blood agar r; D: On Modified Media
Fig. 1. Colony of Acinetobacter baumanii. A: On Chromo agar; B: OnMacConckey aga; C: On Blood agar r; D: On Modified Media Fig. 2.Agarose gel electrophoresis (1% agarose, 7 v/cm2 ) and Ethidium bromide staining to detect A: 16S rRNA gene size product (band 240bp); B: OXA51 gene size product (band 353bp) c- rec A gene .using template DNA prepared by boiling method. Right, Lane, molecular size DNA ladder (100 bp DNA Ladder); Left lanes DNAs isolated from A.baumannii sample showed Positive PCR.
Fig. 2.Agarose gel electrophoresis (1% agarose, 7 v/cm2 ) and Ethidium bromide staining to detect A: 16S rRNA gene size product (band 240bp); B: OXA51 gene size product (band 353bp) c- rec A gene .using template DNA prepared by boiling method. Right, Lane, molecular size DNA ladder (100 bp DNA Ladder); Left lanes DNAs isolated from A.baumannii sample showed Positive PCR.Antimicrobial susceptibility detection
The overall susceptibility patterns of A. baumannii isolates from various clinical sources is displayed in Table 1. Isolates showed high XDR 75%, MDR 23.4%, and PDR 1.6%, The MAR index for experimental isolates (6-21 from 21 antibiotics) ranged between 0.29 to 1.0.
Table 1:
Significance and Antibiotic groups distribution according to the number and percentage of resistance
| Antibiotics | Resp. | No. | % | P-value | Antibiotics | Resp. | No. | % | P-value |
|---|---|---|---|---|---|---|---|---|---|
| TE | S | 53 | 42.7 | 0.704
NS |
AK | S | 9 | 7.3 | 0.000 |
| I | 13 | 10.5 | I | 10 | 8.1 | ||||
| R | 58 | 46.8 | R | 105 | 84.7 | ||||
| Doxt | S | 69 | 55.6 | 0.099
NS |
NET | S | 25 | 20.2 | 0.000 |
| I | 5 | 4 | I | 8 | 6.5 | ||||
| R | 50 | 40.3 | R | 91 | 73.4 | ||||
| SXT | S | 13 | 10.5 | 0.000
HS |
PTZ | S | 7 | 5.6 | 0.000 |
| I | 8 | 6.5 | I | 8 | 6.5 | ||||
| R | 103 | 83.1 | R | 109 | 87.9 | ||||
| LEV | S | 37 | 29.8 | 0.002
HS |
TIM | S | 14 | 11.3 | 0.000 |
| I | 17 | 13.7 | I | 5 | 4 | ||||
| R | 70 | 56.5 | R | 105 | 84.7 | ||||
| CIP | S | 16 | 12.9 | 0.000
HS |
SAM | S | 13 | 10.5 | 0.000 |
| I | 6 | 4.8 | I | 16 | 12.9 | ||||
| R | 102 | 82.3 | R | 95 | 76.6 | ||||
| CTX | S | 4 | 3.2 | 0.000
HS |
IMP | S | 28 | 22.6 | 0.000
HS |
| I | 14 | 11.3 | I | 3 | 2.4 | ||||
| R | 106 | 85.5 | R | 93 | 75 | ||||
| CFM | S | 7 | 5.6 | 0.000 | MEM | S | 20 | 16.1 | 0.000
HS |
| I | 3 | 2.4 | I | 2 | 1.6 | ||||
| R | 114 | 91.9 | R | 102 | 82.3 | ||||
| CPM | S | 11 | 8.9 | 0.000
HS |
PIP | S | 3 | 2.4 | 0.000
HS |
| I | 10 | 8.1 | I | 3 | 2.4 | ||||
| R | 103 | 83.1 | R | 118 | 95.2 | ||||
| CRO | S | 2 | 1.6 | 0.000 | AUG | S | 4 | 3.2 | 0.000
HS |
| I | 7 | 5.6 | I | 7 | 5.6 | ||||
| R | 115 | 92.7 | R | 113 | 91.1 | ||||
| CN | S | 5 | 4 | 0.000
HS |
CAZ | S | 2 | 1.6 | 0.000
HS |
| I | 4 | 3.2 | I | 17 | 13.7 | ||||
| R | 115 | 92.7 | R | 105 | 84.7 | ||||
| TOB | S | 8 | 6.5 | 0.000 | S: Sensitive
R: Resistance I: Intermediate |
||||
| I | 9 | 7.3 | |||||||
| R | 107 | 86.3 | |||||||
β-Lactamase detection
Metallo β-Lactamase detection phenotype showed (n=113)91.1% isolates able to produced while only (n=1)0.8% overall the isolates produce ESBL.
Correlation between biofilm formation and multidrug-resistant isolates
The simple logistic regression test analysis indicated that there is a significant negative correlation between biofilm formation capacity and the inherent ability of bacteria to show multidrug -resistance index (P < 0.001). Our findings indicated that non-MDR isolates tended to form more robust biofilm formation Fig. (3). Among 124 strong biofilm producers, 94% (n = 91) were non-MDR and 6.2%% (n = 6) were MDR isolates. All of the negative biofilm isolates were MDR 100% (n=3) and weak biofilm producers were 50%MDR index isolates and 50% non-MDR.
Twitching motility pattern
The results showed that 6/124 isolates were non-motile (4.8%), while 118/125 (95.2%) isolates of A. baumannii had positive results, 95/124 (75%) of isolates had high mobility isolates with a large area of migration <25 mm after 24 h. of incubation period; while 23/124 (18.5%) isolates showed an average migration area of between 5 and 25 mm.
Beta-lactam genes distribution
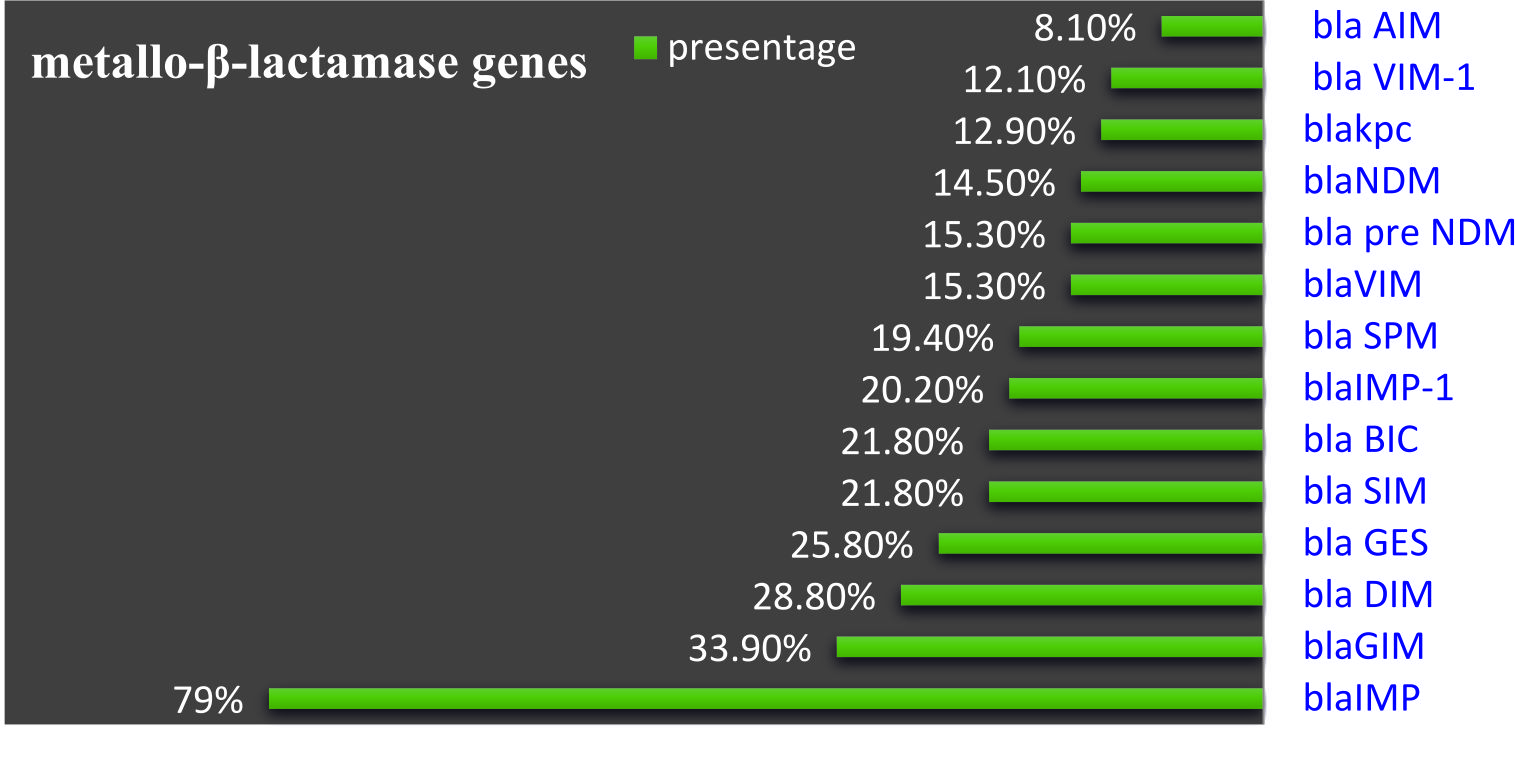
From all isolates among them, 0.8% (n=1) isolates were phenotypically confirmed ESBL producers while 99.2% (n=123) were phenotypically confirmed by non-ESBL producers. While Metallo beta-lactamase present in 91.2% (n=113), and8.9 % (n=11) non Metallo beta-lactamase, Out of the 14 beta-lactam (bla) genes studied, blaIMP was detected among (79%), followed by blaGIM in (33.9%) and variable detection of other genes range between 28.2% – 8.1%. (Fig. 4&5).
Association between bla genes, antibiotic sensitivity, and biofilm formation
The result can be divided to4 group description as shown in Table 2, Hence, the most prevalent group was D include strong bioflim production with highly MDR index and different patterns of gene resistant.
Table (2):
Grouping of A. baumannii according to antibiotic sensitivity, antibiotic genes patteren&bioflim formation
| Biofilm | N0. | Beta lactam antibiotic | Non Beta lactam antibiotic sensitive and intermediate | metallo-β-lactamase | Predominate pattern |
|---|---|---|---|---|---|
| Formation | and intermediate | sensitive genes | |||
| Negative | 3 | (IMP, MEM, TIM AUG-+) | DXT, TE( LEV, CN | IMP(DIM, SPM | |
| (SXT, NET_+) | SIM,VIM, GIM, | ||||
| BIC, NDM, PRE | Non | ||||
| NDM +_) | |||||
| Weak | 4 | CIP, CTX, CPM(SAM, IMP, CFMPTZ, TIM, MEM | TE,DXT,LEV, CN | IMP(VIM, BIC | 1-bla GIM, IMP ,BIC(50%) |
| , AUG +_) | KPC, GES, SIM, | 2- bla VIM, KPC (50%) | |||
| (SXT+-) | GIM, NDM_+) | 3- bla DIM (50%), | |||
| 4-blaGES, SIM,NDM,SPM (33.3%) | |||||
| Moderate | 23 | (CIP, CFM, MEM, | (TE, DXT, AK, | ( IMP, VIM, SIM, | 1. blaIMP, VIM (30.4%) |
| SAM, TIM, CTX | LEV, NET | GIM, GES, DIM, | 2-bla IMP GES (30.4%) | ||
| , CRO, PIP, IMP, | SXT, TOB | NDM, KPC, SIM, | 3-bla IMPalone (21.7%) | ||
| CPM, PTZ, | SXT, CN+_) | SPM, BIC, PRE | 4- bla VIM, SIM | ||
| CAZ+_) | NDM, VIM-1, | 5-blaDIM, GIM, DIM (17.3%) | |||
| IMP-1_+) | 6- bla IMP SIM ,GIM ,GES (8.7%) | ||||
| 7- bla GIM VIM-1(8.6%) | |||||
| 8 – TE, DXT, AK (1.6%) | |||||
| 9- TE, DXT, NET (4.8%) | |||||
| 10- (4.3%) for nine patterns | |||||
| Strong | 94 | (CIP, CFM, SAM, | AK, CN(TE,DXT,LEV, | IMP (VIM, SIM, | 1- bla IMP, AIM, IMP-1 (21.1%) |
| TIM, MEM, CPM, | SXT, NET, TOB_+) | GIM, GES, DIM, | 2-blaI MP, DIM (19.1%) | ||
| IMP, CRO, AUG, | NDM, KPC, SIM | 3-bla IMP, BIC, AND GES (17.02%) | |||
| PIP, CN, CTX, | ,SPM, BIC, PRE NDM | 4- bla IMP, GES (15.9%) | |||
| PTZ, CAZ_+) | , VIM-1, IMP- 1_+) | 5-bla IMP, SIM (14.9%) | |||
| 6-bla IMP, VIM-1 (14.06%) | |||||
| 7- bla IMP, IMP-1 (13.8%) | |||||
| 8-bla DIM, GIM (12.8% | |||||
| 9-bla IMP, SIM ,GIM(12.8%) | |||||
| 10-bla IMP, KPC (11.7%) | |||||
| 10- bla VIM, SIM (10.6%) | |||||
| 11-bla TE (61.6%) | |||||
| 12bla- DXT (53.2%) | |||||
| 13- blaLEV (47.9%) | |||||
| 14-, blaNET (44.7%) | |||||
| 15-bla TE, DXT (31.9%) | |||||
| 16 bla-DXT, NET (23.4%) | |||||
| 17- blaTE, LEV (20.2%) | |||||
| 18- bla TE,DXT,LEV(16.2) | |||||
| 19- bla DXT, LEV (17.02%) | |||||
| 20-blaTOB (15.96%) | |||||
| 21-, bla NET, DXT (11.7%) | |||||
| 22- blaTE, DXT, NET (10.6%) | |||||
| 23- blaTE,SXT,LEV( 8.5%) | |||||
| 24-blaDXT, SXT (7.4%) | |||||
| 25- blaTE, DXT, AK (6.4%) | |||||
| 26-bla -AK, NET (6.4%) | |||||
| 27- bla DXT, AK (6.4%) | |||||
| 28- bla LEV, TOB (5.3%) | |||||
| 29- bla TE, DXT, LEV, NET (3.2%) | |||||
| 30- bla LEV, CN (2.1%) | |||||
| 31 –blaTE,DXT,LEV(2.1%) | |||||
| 32- bla non predominate pattern show 1.06% | |||||
*Red color means constant
Allocation of specimens of A. baumannii may diverge with each hospital as each hospital aperient has a different environment associated with it. More than 91.94% of the A. baumannii isolates were obtained from CSF culture, Blood culture, burn swab, postoperative wound swab, sputum, urine, and body fluid Similar results had been obtained in different studies reported in Iraq by Hussein (19; 5) and (15). As well in different countries such as Doughari20,21,22.
This consideration reveals that a total 76/124 A. baumannii were isolated, d from systemic infection CSF and blood culture, This may be due to the fact that bacteria are opportunistic pathogens capable of penetrating the defenses of the body, although ability to adhere to the instruments or hands of workers, which makes it easier to transmitted between patients who are in hospitals, from above indicated horizontal transition of bacteria from the skin or infected organs to the blood causing Bacteremia.
The use of selective cultures in the diagnosis of A. baumannii in principle is one of the quickest methods and gives a good percentage of diagnosis. This study proved the modified media in the accurate diagnosis of A. baumannii compared to the routine culture media. On the other hand, the study clarified that the diagnosis of genus and species together by using the PCR reaction is quick, accurate and highly sensitive.
It is clear that the emergence of resistant A. baumannii isolates is increasing worldwide. In this research, the resistance of A. baumannii isolates to 15 out of 21tested antibiotics was above 80%.
The frequency of MbLs among isolates of A. baumannii in this study was 91.1%. In MגL-producing A. baumannii were multidrug resistant, the lowest resistance was observed against Tetracyclin and doxycycline which can be used as a treatment option. A significant increase in resistance to carbapenem antibiotics was observed, In this regard23 pointed out that the most important reasons for the emergence of resistance to carbapenem antibiotics are including the production of enzymes Carbapene-mases belonging to b-lactamase enzymes, and loss of protein in the outer membrane has been associated with anti-microbial resistance to Imipenem and Meropenem. It is worth mentioning that the resistance of Meropenem was higher than Imipenem. The high rate of resistance to most cephalosporins antibiotics tested indicates these bacteria possess multiple resistance mechanisms. In addition to their production enzymes b-lactamase and extend b-lactamase and have the ability to alter outer membrane proteins and efflux pump system that acts on efflux the antibiotic extracellular extrusion24. The resistance of A. baumannii to the quinolones and aminoglycoside was increased even to an active antibiotic such as Amikacin.
The study demonstrated the increase of antibiotic resistance to most classes of antibiotics, especially the antibiotics of choice for the treatment of A. baumannii infections include the fluoroquinolones, aminoglycosides, and carbapenems.
The MAR index values for all isolates showed high MDR, ranged between 6-21 from 21 antibiotics. The MAR index for experimental isolates ranged higher than 0.29 reach to 1.0. Suggesting the source of these isolates is from highly contaminated environments where it is excessive and randomly use of antimicrobial agents. Development of MAR pathogenic isolates of A. baumannii determined potential nosocomial infection in the hospital environment. Most isolates from CSF and respiratory system had MAR indices over (0.43), and over (0.76) show in UTI infection while the lowest MDR index (0.29) in body fluid confirming that there was a high antibiotic use and high selective pressure in these environments MDR strains become established in the hospital environment, these can persist for months.
Detection of biofilm formation
Microtiter plate assay
A. baumannii isolates were classified into strong, moderate, weak, and none biofilm producers, respectively at 550nm.Gentile25. demonstrated that there was a high rate of biofilm production between 60% of highly resistant A. baumannii isolates, our results agreed with Iraqi study15.
Twitching motility pattern
It was believed in A. baumannii strains that display a motility phenotype depending on autoinducer molecules related to phenomenon quorum sensing.McQueary26. Further revealed that the production of lipopolysaccharide (LPS) Pili has an important role in twitching motility and adhesion (inert or living surface) and contributes indicating biofilm formation. The results indicated that when the resistance to antimicrobial agent rate is relatively high, there is an increase in productivity of adhesion factors gradually, on the contrary of Ali and Khudhair27 study.
Genotype detection of the studied genes for MBLs
In our study, blaBICwere identified only in (21.8%) of A. baumannii isolates by PCR. In the other hand, the blaKPC gene was identified among 16 /124 isolate (12.9%) of A. baumannii. Also, the blaGES gene was detected among 32/124 isolate (25.8%) A. baumannii. Class A b-lactamases in A. baumannii may be considered a major problem in comparison to other carbapenemases and was usually not-carbapenem specific and often hydrolyze most compounds of the lower beta-lactam classes and take part in the high-level resistance to carbapenems28.
The high frequency of Class A carbapene-mases genes incidence may be because they are plasmid mediated so bacteria can gain this plasmid from another gram-negative bacteria presented in hospital environmental.
The most frequent Class B Metallo-b-lactamases genes detected in A.baumannii were blaIMP, IMP-1, VIM, VIM-1, GIM, SIM,pre-NDM and NDM. All these genes in this group chromosome encoded or carried by the plasmid. Among MBL genes, IMP is more important, especially in Iraq, because of its high presence 79% in A. baumannii isolates. The prevalence of b-lactamase-producing isolates is increasing at an alarming rate worldwide4.
The Metallo-beta lactamase (MBL)gene bla VIM and VIM-1 was the only carbapenemase gene detected among 15.3% and 12.1% A. baumannii isolate respectively, previous studies showed that 42.8 % of A. baumannii isolates from Baghdad/Iraq were VIM type positive3. Similar to this observation, AL-Jubori in 2016 reported 25% of A. baumannii isolates carried blaVIM11. Further bla VIM gene screening in Sulaimani/Iraq noticed only in 2.8% of A.baumannii isolates29.
The accurate identification of MDR-positive A. baumannii isolate represents an important step in infection control and prevention of the spread of multidrug-resistant isolates (30). First large series of clinical isolates from Asia and the UK was blaNDM have been reported by Yong31, since this publication, NDM-1 has an extensive in worldwide and now it is one of the most important carbapenemases in all Enterobacteriaceae and in A. baumannii6. While the NDM-1 reported in Iraq is found in 20%to 40% of A. baumannii1.
Carefully monitoring the presence of MDR A. baumannii among hospitalized patients is recommended to prevent wide dissemination of antibiotic resistance and limit the indiscriminate use of cephalosporins and carbapenems antibiotic in the hospital. Antibiotics Program, infection control, and monitor hospital environment should be applied to minimize the emergence of multiple drug-resistant bacteria.
The gaining and spread of unique ²-lactamase-mediated antibiotic resistance mechanism could finally assist the development of effective protection and control measures.
In conclusion, we have shown that metallo-beta-lactamase producing A. baumannii isolates is an emerging threat in Baghdad hospitals and Finally, in a proof of concept experiment the novel selective media were applied to determine the A. baumannii isolates as well as distinguish sensitive isolates from multiple antibiotic resistance.
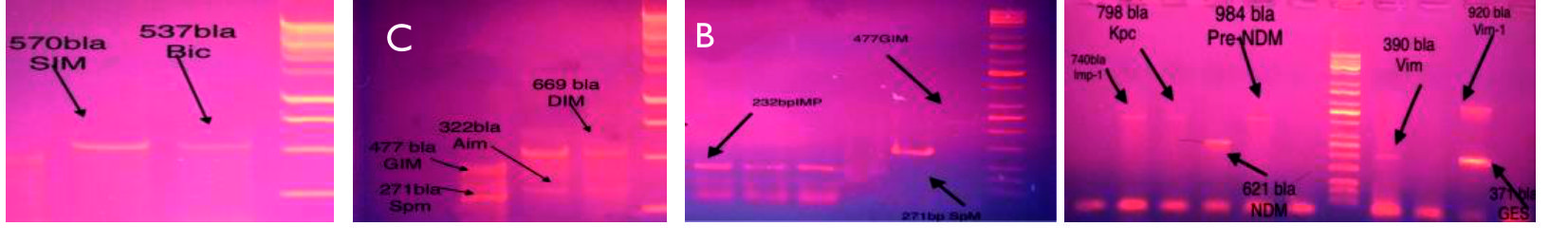 Fig. 5.Agarose gel electrophoresis (1% agarose, 7 v/cm2) and Ethidium bromide staining to detect A, B, C, D Metallo gene using template DNA prepared by boiling method. Right Lane, molecular size DNA ladder A and B (100 bp DNA Ladder); C and D (250 bp ladder) Left lanes DNAs isolated from A. baumannii sample showed Positive PCR
Fig. 5.Agarose gel electrophoresis (1% agarose, 7 v/cm2) and Ethidium bromide staining to detect A, B, C, D Metallo gene using template DNA prepared by boiling method. Right Lane, molecular size DNA ladder A and B (100 bp DNA Ladder); C and D (250 bp ladder) Left lanes DNAs isolated from A. baumannii sample showed Positive PCR Acknowledgments
We would like to express our thanks to Mustansiriyah University Baghdad, Iraq for their support.
Conflict Of Interest
The authors declares that there is no conflict of interest.
Authors’ Contribution
All authors have made substantial, direct and intellectual contribution to the work and approved it for publication.
Funding
None.
Data Availability
All datasets generated or analyzed during this study are included in the manuscript.
Ethics Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
- AL-Harmoosh R.A. And Jarallah E.M. First detection of the blaNDM-1 and blaNDM-2 genes in a clinical isolate of Acinetobacter baumannii in Hillah hospitals -Iraq. International Journal of Advanced Research, 2015; 3(10): 1407 – 1416.
- Aziz R.A. and Al-Jubori S.S. Characterization of Biofilm Production and Quorum Sensing Phenomenon among Antibiotic Resistant Acinetobacter baumannii isolated From Wound infections in Iraq. Journal of Global Pharma Technology, 2017; 05(9): 50-58
- Al Marjani M. and Khadam Z.F. b -Lactamases in Acinetobacter baumannii clinical isolates recovered from humans in Iraq. Advance Pharmaceutical Journal, 2016; 1(4): 81-89.
- Almasaudi S.B.. Acinetobacterspp. as nosocomial pathogens: Epidemiology and resistance features Saudi J. Biol. Sci. 2018; 25(3): 586–596.
- AL-Kadmy I.M., Ali, A.N.M., Salman I.M.A., & Khazaal S.S. Molecular characterization of Acinetobacter baumannii isolated from Iraqi hospital environment. New microbes and new infections, 2018; 21(1): 51-57.
- Fallah F., Taherpour A., Vola M.H. and Hashemi A. Global spread of New Delhi metallo-beta-lactamase-1(NDM-1). Iran. J. Clin. Infect. Dis., 2011; 6(4): 171-176.
- Forbes B.A., Sahm D.F. and, weissfeld A.S. Valley and Scotts diagnostic microbiology. 12th Ed. Mosby elsewhere. Texas, 2007 pp. 334- 339.
- Ali M.R., Al-Taai H.R.R., AL-Nuaeyme H.H. and Khudhair A.M. Molecular study of genetic diversity in Escherichia coli isolated from different clinical sources. Biochemical and Cellular Archives, 2018; 18(2): 2553-2560
- ROCHA. Carlos Andrי Narciso da. Persistence and dispersion of Acinetobacter spp. In the urban water cycle, 2010.
- Roberts S.A., Findlay R. and Lang S.D. Investigation of an outbreak of multi drug resistant Acinetobacter baumannii in an intensive care burns unit. Ghost Infect, 2001; 48:228–232.
- AL_Jubori S.S., AL_kadmy I.M.S. and AL_Ani Z.J. Emergence of Multidrugresistance (MDR) Acinetobacter baumannii Isolated from Iraqi hospitals. Advances in Environmental Biology, 2016; 10(5): 265-27.
- Bonnin R.A., Naas T., Poire Land Nordmann P. Phenotypes, Biochemical, and Molecular Techniques for Detection of Metallo-_-Lactamase NDM in Acinetobacter baumannii. Journal of Clinical Microbiology, 2012; 50(4): 1419–1421.
- Mathur T., Singhal S., Khan S., Upadhyay D.J., Fatma.T And Rattan A. Detection of biofilm formation among the clinical isolates of Staphylococci: an evaluation of three different screening methods. Indian Journal of Medical Microbiology,2006; 24 (1): 25-9.
- Lotfi G.H., Hafida H.A., Nihel K.L., Abdelmonaim K.H., Nadia A.I., Fatima N.A.S. and Walter Z.I. Detection of biofilm formation of a collection of fifty strains of Staphylococcus aureus isolated in Algeria at the University Hospital of Tlemcen. Afr. J. OfBacteriol. Res., 2014; 6(1): 1-6.
- Authman S.H., Ali F.S., and Al- Marjani M.F. Biofilm Formation in Imipenem Resistant Acinetobacter baumannii from the Intensive Care Unit.Journal of Global Pharma Technology, 2017; 10(9): 404-411.
- Lauretti L., Riccio M.L., and Mazzario l.A. Cloning and characterization of blaVIM, a new integration-borne metallo-b-lactamase gene from a Pseudomonas aeruginosa clinical isolates. Antimicrob Agents Chemothe, 1999; 43: 1584–1590.
- Tsakris A., Pournaras S., Woodford N., Palepou M.F.I., Babini G.S. and Douboyas J. Outbreak of infections caused by Pseudomonas aeruginosa producing VIM-1 carbapenemase in Greece. J. Clin. Microbiol., 2000; 38: 1290–2
- Mojtaba M. and Rahimzadeh M. Molecular detection of metallic-b-lactamase genes, blaIMP-1, blaVIM-2 and blaSPM-1 in imipenem resistant Pseudomonas aeruginosa isolated from clinical specimens in teaching hospitals of Ahvaz, Iran. Iran J. Microbiol., 2015; 7(1): 2–6PMCID: PMC4670463 PMID: 26644866
- Hussein N.H., Al-Mathkhury H.J.F. and Sabbah M.A. Identification of Imipenem-Resistant genes in Acinetobacter baumannii Isolated from Baghdad Hospitals. J. MED. Microb. Diagn., 2014; 3(170). Do: 10.4172/2161-0703.1000170.
- Doughari H.J., Ndakidemi P.A., Human I.S. and Benade S. The ecology, biology and pathogenesis of Acinetobacterspp.: an overview. Microbes Environ., 2011; 26: 101–112.
- Basri R., Zueter A.R., Mohamed Z., Alam M.K., Norsa’adah B., Hasan S.A.,HasanH and Ahmad F. Burden of bacterial meningitis: a retrospective review of laboratory parameters and factors associated with death in meningitis, Kelantan Malaysia. Nagoya J. MED, SCI. 2015; 77: 5933.
- Momta Z.H., Safety S. and Tavakol M. Determining the Prevalence and Detection of the Most Prevalent Virulence Genes in Acinetobacter baumannii Isolated from Hospital Infections. International Journal of Medical Laboratory, 2015; 2(2): 87-97.
- Siroy A., mole V., Lemaitre-Guillier C., Vallenet D., Pestel-Caron M., Cozzone A.J., Jouenne T. and De E. Channel formation by Cairo, the carbapenem resistance-associated outer membrane protein of Acinetobacter baumannii. Antimicrob. Agents Chemother, 2005; 49: 4876–4883.
- Mussi M., Gaddy J.A., Arivett B. and Actis L.A. The opportunistic human pathogen Acinetobacter baumannii senses and responds to light. Journal of Bacteriology, 2010; 192(24): 6336–6345.
- Gentile V., Frangipani E., Bonchi C.,Minandri F. and Runci F. Iron and Acinetobacter baumannii biofilm formation. Pathogens, 2014; 3:704-719.
- McQuearyC.M., Kirkup B.C., and Zurawski D.V. Extracellular stress and lipopolysaccharide modulate Acinetobacter baumannii surface-associated motility. Journal of Microbiology, 2012; 50(3): 434–443.
- Ali M.R., & Khudhair A.M. Detection of Colony Adhesion Factors and Genetic Background of Adhesion Genes Among Multidrug-Resistant Uropathogenic Escherichia coli Isolated in Iraq.J. Pure Appl. Microbiol., 2018; 12(4): 2017-2025.
- Queenan A.M. and Bush, K. Carbapene-mases: the versatile beta-lactamases. Clin. Microbiol. Rev., 2007; 20: 440-58.
- Khanda A.A. and Omer F. Sh. Detection of metals b-lactamase enzyme in some gram negative bacteria isolate from burn patients in Sulamani city IRAQ. European Scientific Journal, January edition,. 2014; 10(3) : 1857 – 7881.
- Rodrigo E., Katia A., Monteiro J., Castanheira, M., Soraya S. Ana C., Antonio C. and Sergio T. Rapid Detection and Identification of Metallo-ג-lactamase Encoding Genes by Multiplex Real-Time PCR Assay and Melt Curve Analysis. J. Clin. Microbiol., 2007; 54(2): 544–547.
- Young D., Toleman M.A. and Giske C.G. Characterization of a new metallo-beta-lactamase gene, bla (NDM-1), and a novel Erythromycin esterase gene carried on a unique genetic structure in Klebsiellapneumoniae sequence type 14 from India. Antimicrob Agents Chemother, 2009; 53: 546–5054.
- Statistical Analysis System SAS, S. User’s Guide: Statistics, Version 9.1. SAS Institute, Cary, NC, USA.2003.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.