ISSN: 0973-7510
E-ISSN: 2581-690X
Escherichia coli (E. coli) is a significant species and a common dweller in the guts of humans and animals causing urinary tract infections, wound infection, otitis media, bloodstream infections, and other complications in humans. Worldwide resistance to antimicrobials is a serious public health problem. β-lactamases production by E. coli is responsible for resistance to frequently used beta-lactam antibiotics. 1) To determine antibiotic susceptibility pattern of Escherichia coli isolated from pus and urine samples 2) To determine ESBL producing Escherichia coli and its antibiotic susceptibility pattern. Retrospective study of E. coli isolates from urine and pus samples was performed by collecting data from bacteriology registers. Gender, age details of patients, number of E. coli isolates, and their antimicrobial susceptibility profiles were collected from the records. Out of 747 samples 102 E. coli were isolated and among them 60 (59%) were ESBL producers. Male preponderance was seen i.e. 59 (57.84%) and majority 68 (66.66%) were isolated from the age group of <18 years. E. coli showed maximum sensitivity to imipenem 88 (86.27%), followed by piperacillin-tazobactam 84 (82.35%), aminoglycoside group 76 (74.5%) and maximum resistance was seen to penicillin groups 92 (90.19%), followed by cephalosporins 71 (69.6%). Urinary isolates showed maximum sensitivity to nitrofurantoin (93.67%). ESBL producers showed resistance to ciprofloxacin 47 (78%). This study helps in the periodic formulation of regional antimicrobial policies and also giving continuous information to the clinicians with respect to the sensitivity pattern along with ESBL production which can help to avert further drug resistance.
Antimicrobial susceptibility, Escherichia coli, ESBL, Combination disk test
In the genus Escherichia, Escherichia coli (E. coli) is the most important species and a common dweller in the guts of humans and animals, and also dwells in vegetation, water, and soil. It is the predominant pathogen that causes urinary tract infections, and the most common isolated bacteria that cause wound infection, infections of the bloodstream, otitis media, and other human problems.1
Based on the severity of illness, pathogens isolated, the local pattern of antibiotic sensitivity, standard antibiotics are used to empirically treat these infections. In recent years, the therapeutic options have drastically decreased due to increased and high recurrence rates of antibiotic resistance. Acquired plasmids which code for extended-spectrum β-lactamases (ESBLs) harboured by E. coli are of concern as they are increasing globally.2 Unfortunately, a very narrow range of effective antibiotics is left as these plasmids rapidly spread resistance to all other important antibiotic groups including aminoglycosides and the fluoroquinolones.3
A key example of antibiotic resistance that can cause fatal infections is multidrug-resistant (MDR) and ESBL-producing Escherichia coli and any deferral in providing appropriate therapy can cause severe complications. Significant variations have been observed in the antibiotic resistance pattern between various geographic areas in India. As empirical treatment is given to most of the infections, the choice of the antibiotics should be based on the pathogen most likely to be isolated and its anticipated antimicrobial susceptibility.3 The present research was therefore carried out in order to understand the local antimicrobial susceptibility trends of the E. coli along with its ESBL pattern for providing a prudent empirical treatment.
Aims and Objectives
- To determine the antibiotic susceptibility pattern of Escherichia coli isolated from pus and urine samples
- To determine ESBL producing Escherichia coli and its antibiotic susceptibility pattern.
Method of Data Collection
A retrospective analysis was conducted in the Department of Microbiology at the tertiary care hospital in the Chamarajanagar district of Karnataka. Ethical Clearance for the study was obtained from Institutional ethical clearance committee. For the analysis, secondary data of Escherichia coli isolated from urine and pus samples, maintained in the laboratory registers in the department of Microbiology, for a duration of 1 year from January 2019 to December 2019 were collected. The following information has been noted – name, age, gender details of patients, number of E. coli isolates, and their antimicrobial susceptibility profiles.
Bacterial isolation and identification
After receiving the urine and pus samples in the laboratory, they were processed according to the standard procedure for direct microscopy, aerobic culture, and sensitivity. Samples were streaked on to the respective medias and isolates were identified on the basis of colonial morphology, Gram stain, and biochemical reactions. In urine samples, significant bacteriuria was considered if culture yielded≥105 CFU/ml.4
Antimicrobial susceptibility testing
Antibiotic sensitivity testing of all isolates was performed using antibiotics according to CLSI guidelines by the adapted Kirby-Bauer disc diffusion procedure on Mueller Hinton agar.5 The following drugs were tested- Ampicillin (10μg), Amoxy-clavulanic acid (30μg), Gentamicin (10μg), Amikacin (30μg), Ciprofloxacin (5μg), Cotrimoxazole (25μg), Piperacillin-tazobactam (100/10μg), Imipenem (10μg), Cephotaxime (30μg), Ceftriaxone (30μg), Cefepime (30μg), Ceftazidime (30μg), Ceftazidime-clavulanic acid (30/10μg) and Nitrofurantoin (300μg) [only for urine].
Criteria for selection of ESBL strains
Isolates showing zone of inhibition of ≤ 22mm for Ceftazidime, ≤ 27mm for cefotaxime, and ≤ 25mm for Ceftriaxone by disc diffusion method were suspected of ESBL production.
ESBL detection
This was carried out as per the recommendations of the CLSI. Antibiotics used were Ceftazidime (30μg) disc alone and Ceftazidime in combination with clavulanic acid (ceftazidime + clavulanic acid, 30/10 μg disc). An increase of ≥ 5mm in the inhibition zone of the combination disc compared to the ceftazidime disc alone was considered to be the ESBL producer.
Statistical analysis
MS Office Excel 2010 was used for statistical analysis.
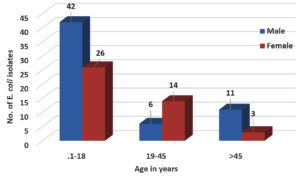
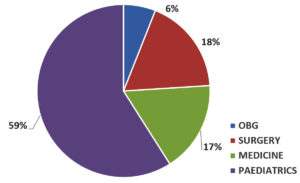
Out of 747 samples, 102 Escherichia coli were isolated. The prevalence of E. coli was 13.65% as depicted in Table 1. Male preponderance was seen 59 (57.84%) and the most i.e. 68 (66.66%) of the E. coli were isolated from the age group of <18 years followed by 20 (19.6%) in 19-45 years and 14 (13.72%) in >45 years as shown in Figure 1 & Table 2. 59% of the isolates were from the pediatric department as seen in Fig. 2.
Table (1):
Prevalence of Escherichia coli isolates.
Samples |
Urine |
Pus |
Total |
|---|---|---|---|
No. of samples tested |
523 |
224 |
747 |
No. of E. coli isolates |
79 |
23 |
102 |
Prevalence |
15.1% |
10.26 % |
13.65 % |
Table (2):
Distribution of Escherichia coli isolates according to age and gender (n= 102).
| Variables | No. | % |
|---|---|---|
| Gender | ||
| Male | 59 | 57.84 |
| Female | 43 | 42.15 |
| Age (years) | ||
| <18 | 68 | 66.66 |
| 19-45 | 20 | 19.6 |
| >45 | 14 | 13.72 |
Table 3 shows the antibiotic susceptibility pattern of E. coli. Isolates showed maximum sensitivity to imipenem 88 (86.27%), followed by piperacillin-tazobactam 84 (82.35%), aminoglycoside group 76 (74.5%). Maximum resistance was seen to penicillin group 92 (90.19%), followed by cephalosporin group 71 (69.6%). Urinary isolates showed maximum sensitivity to nitrofurantoin (93.67%).
Table (3):
Antibiotic susceptibility pattern of Escherichia coli (n= 102).
| Antibiotics | No. of isolates (n-102) | |||
|---|---|---|---|---|
| Sensitive | Resistant | |||
| No. | (%) | No. | (%) | |
| Penicillin group | 10 | (9.8) | 92 | (90.19) |
| Cephalosporins group | 31 | (30.39) | 71 | (69.6) |
| Aminoglycoside group | 76 | (74.5) | 26 | (25.49) |
| Ciprofloxacin | 41 | (40.19) | 61 | (59.8) |
| Cotrimoxazole | 47 | (46.07) | 55 | (53.92) |
| Piperacillin-Tazobactam | 84 | (82.35) | 18 | (17.64) |
| Imipenem | 88 | (86.27) | 14 | (13.72) |
| Nitrofurantoin | 74/79 | (93.67) | 05/79 | (6.32) |
**Nitrofurantoin tested only for urinary isolates (n=79).
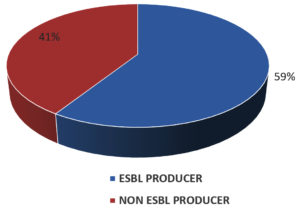
Out of 102 E. coli isolates, 60 (59%) were ESBL producers as seen in Fig. 3. Table 4 depicts the susceptibility pattern of ESBL producing E. coli. The most effective antimicrobial agents were imipenem 50 (83%) followed by piperacillin-tazobactam 43 (72%), aminoglycoside group 39 (65%), and maximum resistance to ciprofloxacin 47 (78%), cotrimoxazole 43 (72%).
Table (4):
Antibiotic susceptibility pattern of ESBL producing Escherichia coli (n= 60).
| Antibiotics | ESBL producing E. coli No. of isolates (n=60) |
|||
|---|---|---|---|---|
| Sensitive | Resistant | |||
| No. | (%) | No. | (%) | |
| Aminoglycoside group | 39 | (65) | 21 | (35) |
| Ciprofloxacin | 13 | (22) | 47 | (78) |
| Cotrimoxazole | 17 | (28) | 43 | (72) |
| Piperacillin-Tazobactam | 43 | (72) | 17 | (28) |
| Imipenem | 50 | (83) | 10 | (17) |
E. coli occurrence and susceptibility profiles show significant variations in various populations, environments, and geographical areas. The prevalence of E. coli in our study was 13.65%. These results are similar to the findings from other research.1,6,7 Many researchers have reported a higher prevalence rate ranging from 60%-80%.8-12
Resistance to antibiotics in E. coli has been reported globally and its rising rates in both developed and emerging nations are a serious concern as it complicates the treatment of infections. This may be attributed to increasing unreasonable intake or non-compliance with medication, self-medication of antibiotics available without a prescription, sales of drugs of inferior quality, eating food from animals that have obtained antibiotics, and spreading resistant isolates among individuals.1 By and large, E. coli resistance to antibiotics in this analysis was high.
Among the several antibiotics tested, E. coli isolates showed maximum sensitivity to Imipenem (86.27%), piperacillin-tazobactam (82.35%), and aminoglycoside group (74.5%), and maximum resistance to penicillin group (90.19%), cephalosporin group (69.6%) and ciprofloxacin (59.8%). These findings are consistent with other studies.1,3,8,9,11,13-18 Maximum sensitivity (93.67%) to nitrofurantoin was observed in this study, which is similar to the previous studies.6,7,15 As this drug is sparsely used in the outpatients’ department for the treatment of uncomplicated lower urinary tract infections as it does not achieve bloodstream therapeutic concentrations and is used for complicated cystitis only.
ESBL producing E. coli has been considered as the main risk to public health and is associated with a high death rate, increased duration of hospital stays, and high cost.13 ESBL producing E. coli was higher (59%) in our study which is in concordance with other studies2,3,12,14,15,19 and this may be attributed to the increasing use of antibiotics both in outpatient and hospitalized patients and definitely needs to be used with caution as an empirical choice.
Concurrent administration of β- lactamase inhibitors such as clavulanate or sulbactam, along with β- lactam antibiotics increases the spectrum of activity. ESBL producing E. coli. showed maximum sensitivity to imipenem (83%) followed by piperacillin-tazobactam (72%), aminoglycoside group (65%) and resistance to ciprofloxacin (78%), cotrimoxazole (72%). Other researchers have reported similar results.20
Unfortunately, the mechanism of drug resistance by ESBL-producing organisms comprises the plasmids that carry the genes of beta-lactamase that transmit the resistance to other classes of an antibiotic such as the aminoglycosides and the fluoroquinolones, allowing limited therapeutic choice and increased threat of treatment failure in patients infected with such strains.16 In our study fluoroquinolones resistance was seen by ESBL producing E. coli.
Carbapenems are the last choice for the treatment of many bacterial infections and lately, warnings have been raised about the spread of resistance to carbapenem groups due to New Delhi metallo-β-lactamase (NDM-1) production. Presently, no new antibiotics are in the pipeline to battle carbapenem resistance, and the spread of this gene globally. Resistance gene spread is considered to be a potentially frightening situation.8 Our study revealed 86.27% sensitivity to imipenem and this drug could be a reserved therapeutic option for infections that have not responded to other antibiotics.
E. coli is one of the most common pathogens associated with UTI and pus-forming lesions, worldwide. The increased rates of antibiotic resistance have left clinicians with fewer therapeutic options. Since drug resistance is an evolving process, its recognition by clinical microbiologists becomes the need of the hour. To detect ESBL producers test like the combination disk method can be regularly done. The hospitals should have its own antibiotic policy, based on the local susceptibility profile of area-specific pathogens and the clinicians should be constantly informed about the periodically revised antibiotic policy.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
TBN drafted the manuscript, gathered information from the literature, compiled data. VM collected data and designed tables. AB supervised and reviewed the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript.
- Gautam R, Chapagain ML, Acharya A, et.al. Antimicrobial susceptibility patterns of Escherichia coli from various clinical sources. Journal of Chitwan Medical College. 2013;3(1):14-17.
Crossref - Pandit R, Awal B, Shrestha SS, Joshi G, Rijal BP, Parajuli NP. Extended-Spectrum β-Lactamase (ESBL) genotypes among Multidrug-Resistant uropathogenic Escherichia coli clinical isolates from a teaching hospital of Nepal. Interdiscip Perspect Infect Dis. 2020;2020:6525826.
Crossref - Mukherjee M, Basu S, Mukherjee SK, Majumder M. Multidrug-resistance and Extended-spectrum beta-lactamase production in uropathogenic E. Coli which were isolated from hospitalized patients in Kolkata, India. J Clin Diagn Res. 2013;7(3):449-453.
Crossref - Collee JG, Barrie P, Marmion AG, Fraser A. Simmons. Mackie and McCartney Practical Medical Microbiology, 14th ed. Edinburgh: Churchill Livingstone. 2007.114.
- Clinical Laboratory Standards Institutes (CLSI). Performance Standards for antimicrobial susceptibility testing, XXI International Supplement (M100-S27). Wayne, Pennsylvania, USA: National Committee for Clinical Laboratory Standards. 2019;104.
- Acharya A, Gautam R, Subedee L. Uropathogens and their antimicrobial susceptibility pattern in Bharatpur, Nepal. Nepal Med Coll J. 2011;13(1):30-33. PMID: 21991698
- Rijal A, Ghimire G, Gautam K, Barakoti A. Antibiotic susceptibility of organisms causing urinary tract infection in patients presenting to a teaching hospital. J Nepal Health Res Counc. 2012;10(20):24-27. PMID: 22929632
- Sabir S, Anjum AA, Ijaz T, Ali MA, Rehman MR, Nawaz M. Isolation and antibiotic susceptibility of E. coli from urinary tract infections in a tertiary care hospital. Pak J Med Sci. 2014;30(2):389-392.
Crossref - Odoki M, Aliero AA, Tibyangye J, Maniga JN, Wampande E, Kato CD. Prevalence of bacterial urinary tract infections and associated factors among patients attending hospitals in Bushenyi district, Uganda. Int J Microbiol. 2019:1-8.
Crossref - Preethishree, P, Rai R, Vimalkumar K, Ashapai KB, Bhat PU. Uropathogens and their antibiotic susceptibility pattern at a tertiary care teaching hospital in Coastal Karnataka, India. Int J Curr Microbiol App Sci. 2016;5(1):23-31.
Crossref - Kalpana S, Hegadi SS, Ramesh K. Characterization and Antimicrobial susceptibility testing of Uropathogens from urinary tract infections. Int J Curr Microbiol App Sci. 2015:4(2):1010-1016.
- Wadekar MD, Jagdish L, Swaroopa RNB, Gupta RK. Extended-spectrum β lactamase- producing Escherichia coli in urinary tract infections tip-off to evaluate treatment practice. Indian J Microbiol Res. 2016;3(2):175-179.
Crossref - Patel HB, Soni ST, Bhagyalaxmi A, Patel NM. Causative agents of urinary tract infections and their antimicrobial susceptibility patterns at a referral center in Western India: An audit to help clinicians prevent antibiotic misuse. J Family Med Prim Care. 2019;8:154-159.
Crossref - Alharbi NS, Khaled JM, Kadaikunnan S, Alobaidi AS, Sharafaddin AH, Alyahya SA et.al. Prevalence of Escherichia coli strains resistant to antibiotics in wound infections and raw milk. Saudi J Biol Sci. 2019:26;1557-1562.
Crossref - Prasada S, Bhat A, Bhat S, Mulki SS, Tulasidas S. Changing antibiotic susceptibility pattern in uropathogenic Escherichia coli over a period of 5 years in a tertiary care center. Infect Drug Resist.2019:12:1439-1443.
Crossref - Mandal DK, Sah SK, Mishra SK, et al. Carriage of Extended-Spectrum-β-Lactamase- and AmpC- β-Lactamase-producing Enterobacteriaceae (ESBL-PE) in a healthy community and outpatient department (OPD) patients in Nepal. Can J Infect Dis Med Microbiol. 2020:1-9.
Crossref - Kalaiselvi G, Shanmugavadivoo N, Padmavathi BK, Usha B. Changing prevalence and antimicrobial susceptibility profiles by uropathogens – A study in India. Journal of Evolution of Medical and Dental Sciences. 2015:4(65);11342-11351.
Crossref - Wadekar MD, Sathish, JV. Changing Antimicrobial Susceptibility Pattern of E. coli Isolated from Urine and Pus Samples. Int J Curr Microbiol App Sci. 2017;6(7):2504-2511.
Crossref - Erb A, Sturmer T, Marre R, Brenner H. Prevalence of antibiotic resistance in Escherichia coli: an overview of geographical, temporal, and methodological variations. Eur J Clin Microbial Infect Dis. 2007;26:83-90.
Crossref - Wadekar MD, Swarooparani NB. ESBL producing Escherichia coli– its prevalence and antibiogram. Indian J Microbiol Res. 2017;4(3):287-290.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.