ISSN: 0973-7510
E-ISSN: 2581-690X
Candida tropicalis is the fourth main infective agent of Candida species in several developing nations and leads to the greatest fatality rate among the non-albicans Candida (NAC) species that cause candidemia. Seventy clinically known Candida isolates are isolated in positively flagged blood samples from BacT/ALERT 3D from various wards. Virulence factors like hemolysin production, coagulase activity, phospholipase activity, and biofilm formation were studied and antifungal susceptibility testing was and Minimum Inhibitory Concentration (MIC) values were interpreted. Of the total 70 clinical Candida isolates, the most predominant organism isolated was found to be C. tropicalis 27 (38.57%) which is succeeded by 19 (27.14%) C. albicans, 13 (18.57%) C. parapsilosis, 6 (8.57%) C. glabrata, and 5 (7.14%) C. krusei respectively. Among the70 Candida isolates, 49 (70%) showed hemolysin production, 43 (61.42%) isolates demonstrated phospholipase activity, 34 (48.57%) showed coagulase activity and 55 (78.57%) isolates showed biofilm production by crystal violet assay. A high level of Fluconazole resistance has been observed in 23 (32.85%) Candida isolates in comparison with other antimicrobials utilized in this study. The higher MIC value of: ≥ 64 µg/mL Fluconazole was shown by 4 (57.14%) isolates of C. tropicalis by broth microdilution method. The interpretation of various virulence factors and antifungal drug resistance were seen mostly among NAC species, thus hence signifying its pivotal role in immunocompromised individual treatment.
Candida tropicalis, Candidemia, Virulence Factors, Antifungal Drug Resistance
Candida species causing bloodstream infections (BSIs) are also called candidemia which exists mostly as a pervasive type of invasive candidiasis.1 Candidemia is one of the most prevalent reasons for BSIs among hospitalized patients in the United States, and it frequently leads to prolonged hospitalization as well as mortality.2 C. tropicalis is the fourth main infective agent of Candida species in several developing nations and contributes the greatest fatality rate among the NAC species that cause candidemia. The extensive utilization of antifungal agents increases C.tropicalis antifungal resistance, especially to azoles, which would ultimately result in the treatment failure of candidemia.3,4
C. tropicalis is considered the first and most predominant species causing candidemia in growing nations, wherein an enormous number of cases are treated by Fluconazole as a consequence of the outrageous price of echinocandins.5 In comparison to other continents, tropical Asia and Latin America have the relatively greatest proportion of candidemia caused by C. tropicalis.6 As stated by the Centers for Disease Control and Prevention (CDC), around 25,000 candidemia incidences were reported nationwide each year. C. tropicalis (38%) is the commonest organism in India among candidemia cases.7 On the other hand, isolates of C. tropicalis have spread globally, and since 2010, this situation has particularly become debatable in the Asia-Pacific area. However, a dramatic rise in pan azole-resistance and Amphotericin B resistance of C. tropicalis was shown in the majority of studies conducted on candidemia studies.8-11
Several virulence factors of Candida species cause BSIs including hemolytic activity, coagulase, proteinase, phospholipase, esterase, lipase, and biofilm formation. All of these multiple virulence factors could raise BSIs of Candida species by escape mechanism which is part of the defence system in order to impair the host tissue.12 The manifestation of virulence factors of Candida species can be differentiated based on species, type of infection, geographical region, host reaction and stage, and the site of infection. Understanding these virulence factors is crucial to know the pathophysiology, and also assists researchers to discover novel antifungal targets for more effective therapeutic regimens.13
As per the CDC monitoring statistics, the percentage of Fluconazole-resistant Candida isolates remained relatively stable over the last 20 years.14 In India, nationwide candidemia research has been held revealing that the origination of multidrug-resistant (MDR) C. tropicalis was in an equivalent percentage to what was perceived for MDR C. auris.15 The progression in drug tolerance enables the organism to procure persistent modifications of genes that result in the development of antifungal resistance, therapeutic failure, and death.16 Indeed, various research suggests C. tropicalis are tolerant to azoles as the efficiency of Fluconazole on these isolates from blood samples was diminished, which would ultimately result in Fluconazole therapeutic failure.17 Despite the progression of resistance, one or more antimicrobial agents should represent a severe concern among candidemia cases caused by C. tropicalis since there are so few antifungals available to treat, especially in underdeveloped nations, where Fluconazole is one of the most often antifungals used.18
Despite the fact that we comprehend how azole resistance developed in C. tropicalis, molecular researchers suggest that azole resistance in C. tropicalis may mostly be raised due to mutations in Cdr1 and Mdr1 efflux pumps, as well as in ergosterol production gene (ERG11).19 Alternative azole resistance pathways have also been linked to biofilm formation mitochondrial abnormalities and other virulence factors.20 Hence, this present work is done for analyzing the complete mycological profile of Candida species and emphasizing virulence factors as well as the antimicrobial sensitivity pattern of C. tropicalis isolates in order to determine effective ways of management to curb candidemia based on the new regimen.
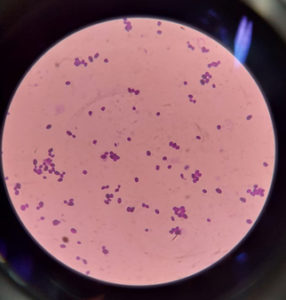
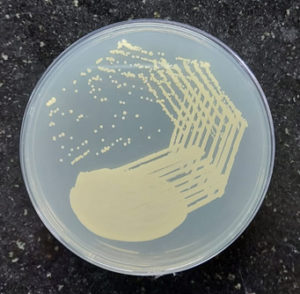
A cross-sectional study has been held in a tertiary care centre and ethical approval was obtained from the Institute Ethical Committee (Human Studies) between August 2021 to March 2022. Seventy clinically known Candida isolates were isolated from positively flagged blood samples incubated in BacT/ALERT 3D collected from various wards and gram stained directly to determine the existence of gram-positive budding yeast-like cells (Figure 1). The standard culture techniques were carried out on Sabouraud Dextrose Agar (SDA) with antibiotics, MacConkey agar, and Blood agar and were incubated at 25°C and 37°C for 24 to 48 hours. After that, every plate was studied macroscopically and the colony characteristics were examined. Colonies that appeared on SDA were smooth, creamy, convex, and pasty colonies (Figure 2). Then, the colonies were Gram stained and examined under the microscope.
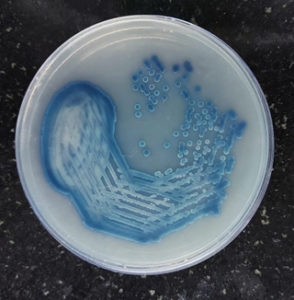
Following the culture characterization, species identification was performed by inoculating the colonies onto Candida CHROM agar plates which were incubated at 37°C for 48 hours and they were identified and speciated based on the color produced by the organisms on the chromogenic medium, i.e. C. albicans – Apple green, C. tropicalis–Metallic Blue,
C. krusei – Pink, C. parapsilosis – Cream to pink, and C. glabrata – purple. Further, germ tube formation test and chlamydospore formation by Cornmeal agar was also performed to identify the Candida species that could not be distinguished by color on Candida CHROM agar medium26 (Figure 3).
Virulence factors like hemolysin production, coagulase activity, phospholipase activity, and biofilm formation were studied and the outcomes were compared with the antifungal susceptibility testing along with the detection of Minimum Inhibitory Concentration (MIC) values which may be linked with the azole resistance pathway.
Germ tube formation method
A colony of yeast was suspended with 0.5 ml of human, sheep, or fetal bovine serum in a sterile Eppendorf tube and incubated at 37°C for 2-4 hours. Subsequently, a single drop of serum was placed onto a glass slide and a cover slip was kept over it. Then, it was examined microscopically under a low-power objective. The appearance of long tube-like projections attached to the yeast cells was called germ tube formation which is typically present in C.albicans or C. dubliniensis.
Haemolysin Production
Haemolysin production was demonstrated on SDA containing sheep blood along with gentamicin as described by Manns et al.21 The medium was aseptically injected with 10 µL of inoculum prepared with the isolates. Petri plates containing the above-described medium were cultured for 48 hours at 37°C and ATCC 90028 C.albicans is used as a control. Hemolysin activity was demonstrated by the existence of a halo zone of hemolysis surrounding a colony. The colony diameter to the transparent hemolysis zone (in mm) ratio is calculated as hemolytic activity (Hz).
Coagulase Activity
Using the method developed by Yigit et al.,22 coagulase production was identified. A 500 50µL of rabbit plasma was added to a tube and it was aseptically inoculated with 0.1mL of an inoculum that has been kept overnight aseptically. After incubation of 2, 4, 6, and 24 hours at 35°C, the tubes were checked. A positive coagulase test shows clot formation which can not be revived by mild shaking. Positive control: ATCC 25923 Staphylococcus aureus and negative control: ATCC 14990 S. epidermidis, respectively.
Phospholipase Production
The method developed by Samaranayake et al.,23 was used to identify phospholipase production. A standard test strain inoculum containing a volume of 5µL was inoculated aseptically onto agar containing egg yolk. Petri plates are dehydrated at ambient temperature, they were incubated for 48 hours at 37°C. The formation of a zone of precipitation circling the colony was checked and the existence of a precipitation zone suggested that the phospholipase enzyme was exhibited. The positive control was used as ATCC 10231 C.albicans. The colony diameter to the combined and precipitation zone ratio is calculated as phospholipase index (Pz). When Pz was 1, it indicates no phospholipase production.; Pz< 1 denotes positive activity. The test was carried out three times in triplicate for each isolate to reduce experimental error.
Biofilm formation
A growth from SDA has suspended in a sterile brain-heart infusion (BHI) broth, and cultured overnight. The next step was a 1:100 dilution in BHI. Then, a commercially available polystyrene, pre-sterilized, round-bottomed, 96-well microtiter plate (HiMedia) was used to incubate 100µL of diluted broth at 37°C overnight for the development of biofilm, and with distilled water, the microtiter plates were cleaned.
Crystal violet assay
Crystal violet assay was performed to demonstrate the biofilm production as described by Kuhn et al.24 The microtiter plate was cleaned with distilled water before being stained at ambient temperature for 15 minutes with 0.1% aqueous solution of crystal violet(120 µL). The wells were cleaned with sterile distilled water for 4 times. Then, 125 µL of 95% methanol was used to de-stain, and the wells were incubated at room temperature for 15 minutes. An enzyme-Linked Immunosorbent Assay (ELISA) reader was used to read the de-stained wells spectrophotometrically at 570 nm. 12 Duplicate runs of each sample were done.
The biofilm formation was read as follows:
OD – Optical Density
ODC – Optical Density cutoff
SD – Standard Deviation
Avg NC – Average OD value of negative control
Avg NC=0.194, SD=0.013
ODC =Avg NC+3׳SD, ODC =0.194+0.039, ODC=0.233
OD≤ODC= non-biofilm producers
ODC<OD≤2 ODC=weak biofilm producers
2ODC<OD≤4 ODC=moderate biofilm producers
4ODC<OD=strong biofilm producers
Antibiogram
The antibiogram pattern for antifungals like Fluconazole (25µg), Amphotericin B (100 U), Itraconazole (10µg), and Voriconazole (1µg) (Hi-Media Laboratories) was performed by the Kirby-Bauer disk diffusion method. To improve the determination of growth margins, it was carried out on Mueller-Hinton agar which was added to 0.5 g of methylene blue per milliliter and 2% glucose. The zone sizes were interpreted according to the Clinical and Laboratory Standards Institute (CLSI) guidelines. Then, Fluconazole resistant isolates were subjected to broth microdilution M27-A3 in order to determine MIC values, and results were interpreted according to the CLSI guidelines.25
In the course of the research period (16 months), blood samples were received from various wards, and those flagged with positive signals were subjected to the examination by microscope and culture findings (SDA, Blood agar, and MacConkey agar) for Candida species. A total of 70 consecutive and non-duplicate Candida species were comprised in our study. Organisms other than Candida isolates were excluded.
Among the 70 Candida isolates, 13 (18.57 %) isolates indicated a positive result for germ tube formation test, and 57 (81.42%) isolates of Candida species did not show germ tube formation. Of the total 70 clinical Candida isolates, the most predominant organism isolated was C. tropicalis 27 (38.57%)succeeded by 19 (27.14%) C. albicans, 13 (18.57%) C. parapsilosis, 6 (8.57%) C. glabrata, and 5 (7.14%) C. krusei respectively (Table 1). However, all the isolates indicated negative findings on the urease test.
In the interpretation of virulence factors of the 70 Candida isolates, phospholipase activity was demonstrated by 43 (61.43%) isolates, out of which 25 (58.13%) were C. tropicalis followed by 11 (25.58%) were C. albicans, 4 (9.30%) were C. glabrata, 2 (4.65%) isolates of C. parapsilosis and 1 (2.32%) isolate of C.krusei (Table 1).
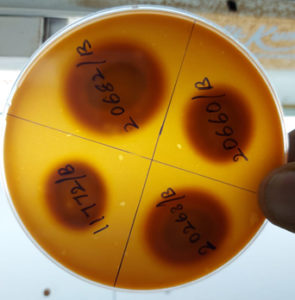
With respect to hemolysin, 49 (70% out of the 70 Candida isolates) showed hemolysin production (Table 1) (Figure 4), from which 27 (55.10%) were C. tropicalis, 12 (24.48%), C. albicans, C. parapsilosis 5 (10.20%), C. glabrata 3 (6.12%), and C. krusei 2 (4.08%). Coagulase activity was demonstrated by 34 (48.57%) Candida isolates, of them, 13 (38.24%) C. albicans produced the maximum coagulase activity followed by C. tropicalis 11 (32.35%), 6 (17.64%) C. parapsilosis, 3 (8.82%) C. glabrata and 1 (2.94%) C. krusei respectively (Table 1) (Figure 5).
Biofilm production was conducted by crystal violet assay for all the 70 isolates of Candida species and was interpreted as follows, 55 (78.57%) isolates showed biofilm production and among them, 25 (45.46%) C. tropicalis isolates showed biofilm production, subsequently followed by 13 (23.63%) C. albicans, 5 (9.09%) C. krusei, 9 (16.36%) C. parapsilosis, and 3 (5.45%) C. glabrata. C. tropicalis stood out among them as the most frequent cause of biofilm development. Of them, 18 isolates were found to produce strong biofilm production (3+), 10 isolates were observed to show moderate biofilm production (2+). Also, 8Candida species showed weak biofilm production (1+)(Table 1).
Table (1):
Mycological profile of Candida isolates isolated from Candidemia cases (n=70).
| Candida species | Species Distribution (%) |
Phospholipase activity (%) |
Haemolysin production (%) |
Coagulase activity (%) |
Biofilm production (%) |
|---|---|---|---|---|---|
| C. tropicalis | 27 (38.57%) | 25 (58.14%) | 27 (55.10%) | 11 (32.35%) | 25 (45.45%) |
| C. albicans | 19 (27.14%) | 11 (25.58%) | 12 (24.48%) | 13 (38.24%) | 13 (23.64%) |
| C. parapsilosis | 13 (18.57%) | 2 (4.65%) | 5 (10.20%) | 6 (17.64%) | 9 (16.36%) |
| C. glabrata | 6 (8.57%) | 4 (9.30%) | 3 (6.12%) | 3 (8.82%) | 3 (5.45%) |
| C. krusei | 5 (7.14%) | 1 (2.32%) | 2 (4.08%) | 1 (2.94%) | 5 (9.09%) |
| Total (n=70) | 43 (61.43%) | 49 (70%) | 34 (48.57%) | 55 (78.57%) | |
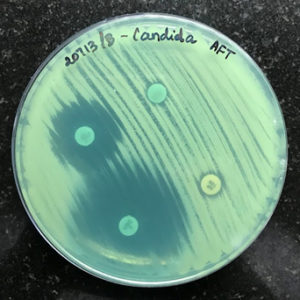
The antibiogram profile of the 70 isolates of Candida was demonstrated for Voriconazole, Amphotericin B, Fluconazole, and Itraconazole. A high level of resistance among Fluconazole has been observed in 23 (32.86%) Candida species in comparison with other antimicrobials utilized in this study. Among 23 (32.86%) Fluconazole resistant Candida species, C. tropicalis (7 out of 27 isolates) were more predominant and all C. krusei (5 out of 5 isolates) were resistant to Fluconazole. With respect to Itraconazole, 14 (20%) out of 70 Candida isolates showed resistance and 56 (80%) were sensitive. For Voriconazole, 6 (8.57%) isolates showed resistance, and 64 (91.43%) were sensitive. While the Amphotericin B resistance was interpreted in only 1 (1.43%) isolate of C.tropicalis and the rest 98.57% of Candida species were sensitive (Table 2) (Figure 6).
Table (2):
Antifungal susceptibility pattern of Candida isolates by Kirby Bauer disc diffusion method (n=70).
| Candida species | FLC | ITR | VRC | AMP B | ||||
|---|---|---|---|---|---|---|---|---|
| S | R | S | R | S | R | S | R | |
| C. tropicalis (n=27) | 20 | 7 | 23 | 4 | 24 | 3 | 26 | 1 |
| C. albicans (n=19) | 17 | 2 | 16 | 3 | 18 | 1 | 19 | 0 |
| C. parapsilosis (n=13) | 9 | 4 | 11 | 2 | 13 | 0 | 13 | 0 |
| C. glabrata (n=6) | 1 | 5 | 3 | 3 | 5 | 1 | 6 | 0 |
| C. krusei (n=5) | 0 | 5 | 3 | 2 | 4 | 1 | 5 | 0 |
| Total (n=70) | 47 | 23 | 56 | 14 | 64 | 6 | 69 | 1 |
| Percentage (%) | 67.14% | 32.86% | 80% | 20% | 91.43% | 8.57% | 98.57% | 1.43% |
*S – Sensitive, *R – Resistant, *FLC – Fluconazole, *ITR – Itraconazole, *VRC – Voriconazole, *AMP B – Amphotericin B
Fluconazole MICs for all C. tropicalis isolates were determined using microdilution method and the results were interpreted based on CLSI guidelines in which 1 (14.28%) C. tropicalis isolate showed MIC of ≤ 2µg/mL and 2 (28.57 %) isolates showed intermediate resistance to Fluconazole exhibiting MIC from 4-32µg/mL. The higher Fluconazole MIC value of ≥ 64µg/mL was shown by 4 (57.14%) isolates of C.tropicalis (Table 3).
Table (3):
MIC of Fluconazole-resistance C. tropicalis isolates (n=7).
| Name of the Isolate | MIC of Fluconazole (0.25- ≥ 64 μg/mL) | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| ≥ 64 µg/mL | 32 µg/mL | 16 µg/mL | 8 µg/mL | 4 µg/mL | 2 µg/mL | 1 µg/mL | 0.5 µg/mL | ≤ 0.25 µg/mL | |
| C. tropicalis (n=7) | 4 | – | 1 | – | 1 | 1 | – | – | – |
Candidemia is a major concern, especially among immunocompromised patients. The most prevalent causative agent of candidemia has shifted in recent years over C. albicans to NAC species. In Southern India, the most prevalent isolates of candidemia were C. tropicalis and C. parapsilosis.26 However, when compared to other Candida species, candidemia brought on by C. tropicalis isolates is a potentially fatal infection that increases patient mortality. There are risk factors that might enhance a person’s vulnerability to candidemia including: increase usage of antibiotics and corticosteroids, repeated hospitalisation, neutropenia, malignancy AIDS, intravascular catheterization, chemotherapy, and other illnesses that impair immunity.27,28
Over the last several years, a steady rise in the isolation NAC species from candidemia patients has been noticed.28-30 Numerous investigations have isolated and noted similar patterns of NAC species, but other research indicates that C. glabrata and C. albicans predominate.31-34 In this study, 70 consecutive, non-repetitive Candida isolates were speciated by the conventional method. In this study, NAC species account for about 72.85% while C. albicans (27.14%) and similar results were demonstrated by Sachin C et al.35 Among them, 27 (38.57%) C. tropicalis were the most frequent isolates of Candida species and similar results were also reported in the study findings of Tak V et al. and Chakrabarti et al.36,37
Since C. albicans was reported as the predominant harmful organism of candidemia, there have been much research on these virulence factors. However, a lot of research publications discuss the development of virulence among NAC species.38-40 The demonstration of virulence factors shows that 43 (61.43%) isolates showed phospholipase activity and 25 isolates of C.tropicalis showed maximum phospholipase activity. These results were correlated with the study findings of Sachin C et al. which showed that 60.9% of isolates demonstrated phospholipase activity and 76% and phospholipase activity was predominantly produced by 19 isolates of C. tropicalis.35 Conversely, Khater et al. resulted that phospholipase activity was more predominant among Candida albicans (45.62%) which was higher than that of the NAC species.41 However, on the contrary, Figueiredo-Carvalho et al. reported no phospholipase activity among NAC species.42
Among 70 Candida isolates, 49 (70%) showed hemolysin production and the present study results were correlated with the study findings of Luo et al. and M. A. Galen-Ladero et al. which showed 85.7% and 77.2%.43,44 Although studies have revealed that C. tropicalis produces hemolysin, it is crucial to know if the hemolytic activity shown is actually occurring or whether it is the consequence of phospholipase synthesis.45 Therefore, it appears necessary for conducting more sophisticated research based on the molecular study in order to elucidate the hemolysin production which contributes to the pathogenesis of C. tropicalis.22 Coagulase activity was demonstrated by 34 (48.57%) Candida isolates, of them, 13 C. albicans produced the maximum coagulase activity and these results were similar to the findings reported by Yigit N et al. which showed that 50.6% Candida isolates had produced coagulase activity. Among them 14 C. albicans produced coagulase activity.22
The development of a mature, highly organized biofilm happens when the yeast cell attaches to a surface and starts to multiply. Biofilm development is believed to be a powerful infectious feature causing therapeutic flops and infections that are recurring.46,47 Out of 70 Candida isolates, 55 (78.57%) strains showed biofilm production, and these findings correlated with the study results of Sanyuktha Tulasidas et al. which showed biofilm activity in 74% of Candida isolates.48 It was interpreted that 25 (45.46%) of Candida tropicalis isolates showed maximum biofilm production. This finding is in concordance with the results obtained in a study conducted by Sasani E et al. and similarly reported C. tropicalis (47%) as the prevalent isolate that produced biofilm activity.49 On many medical devices, especially intravascular ones, Candida isolates more easily colonize to create in vitro biofilms that might result in catheter-associated bloodstream infections. The development of biofilms may have a major impact on how patients with candidemia respond to therapy.50 The invasiveness and spread potential of C. tropicalis are influenced by biofilm formation, an essential virulence component. However, our study findings also demonstrate that most of the biofilm-producing C. tropicalis isolates were showing a high degree of Fluconazole resistance than non-biofilm producers as it is considered an essential virulence factor endowing pathogenicity among candidemia patients.51
With respect to an antibiogram pattern, maximum of isolates were sensitive for all drugs used: Amphotericin B (98.57%), Voriconazole (91.43%), Itraconazole (80%), and Fluconazole (67.14%). These findings were similar to other studies.52-54 A high level of resistance among Fluconazole has been observed in 23 (32.86%) Candida isolates in comparison with other antimicrobials in the current study. Of them, C. tropicalis was found to show maximum resistance to Fluconazole and only 1 (1.43%) isolate of C. tropicalis was identified as resistant. This was similar to the study findings of Giri et al.55 Notably, all strains of C. krusei were Fluconazole-resistant as these species are intrinsically resistant to Fluconazole. A few more isolates also showed resistance to Itraconazole (20%), Voriconazole (8.57%), and lower rate of resistance was reported in Amphotericin B (1.43%) which were in concordance with various study reports.56-58
The maximum resistance pattern was demonstrated by C. tropicalis as it is emerging as one of the most significant and alarming Candida isolates causing antifungal resistance.59 So, MIC of all Fluconazole-resistant C. tropicalis isolates were demonstrated and interpreted by the Micro dilution method as per CLSI guidelines.60 The current study results were similar to the study findings of Gandham et al. and it showed that Fluconazole showed higher MIC of ≥ 2 µg/ml in 4 isolates of C. tropicalis and 1 of C. parapsilosis, and 2 of C. albicans were resistant.61 The most likely cause of this change appears to be the overuse of Fluconazole, which allowed for the survival of resistant species like NAC species.62 The changing epidemiology and rising antifungal resistance highlight the importance of constant monitoring of these candidemia cases. Therefore, the management of the infections and reducing death rates may be aided by a proper diagnosis of Candida species and a strict antifungal stewardship program.63,64
In this study, the presence of candidemia among the patients from various wards occurs with the predominance C. tropicalis. Amphotericin B has the highest antifungal action, whereas Fluconazole has the least activity expressed in the highest antifungal resistance seen. The interpretation of various virulence factors and antifungal drug resistance were seen mostly among NAC species, thus hence signifying its pivotal role in the immunocompromised individual treatment. To enhance outcomes, particularly for critically sick newborns and immunocompromised people, timely identification of Candida BSI and knowledge of its resistance profile are very essential. Thus, this information on antifungal susceptibility and virulence variables are required to minimise the overall impact of Candida infections that are rising, therapeutic failures, and financial burden severely.
ACKNOWLEDGMENTS
None.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the Institutional Ethical Committee of SRM Medical College Hospital and Research Centre, India, with reference number 2896/IEC/2021.
- Morgan J, Meltzer MI, Plikaytis BD, et al. Excess mortality, hospital stay, and cost due to candidemia: a case-control study using data from population-based candidemia surveillance. Infect Control HospEpidemiol. 2005;26(6):540-547.
Crossref - Magill SS, O’Leary E, Janelle SJ, et al. Changes in Prevalence of Health Care-Associated Infections in U.S. Hospitals. N Engl J Med. 2018;379(18):1732-1744.
Crossref - Selvan P, Vajravelu LK, Mohanraj H, Ramakrishna MS. Monitoring the Spectrum of Candidemia and its Anti-fungal Resistance in A Tertiary Care Centre – An Emerging Global Alarm. J Pure Appl Microbiol. 2022;16(4):2704-2711.
Crossref - Adhikary R, Joshi S. Species distribution and anti-fungal susceptibility of Candidemia at a multi super-specialty center in southern India. Indian J Med Microbiol. 2011;29(3):309-311.
Crossref - Negri M, Silva S, Henriques M, Oliveira R. Insights into Candida tropicalis nosocomial infections and virulence factors. Eur J ClinMicrobiol Infect Dis. 2012;31(7):1399-1412
Crossref - Pfaller MA, Messer SA, Moet GJ, Jones RN, Castanheira M. Candida bloodstream infections: comparison of species distribution and resistance to echinocandin and azole antifungal agents in Intensive Care Unit (ICU) and non-ICU settings in the SENTRY Antimicrobial Surveillance Program (2008-2009). Int J Antimicrob Agents. 2011;38(1):65-69.
Crossref - Statistics. Centers for Disease Control and Prevention. https://www.cdc.gov/fungal/diseases/candidiasis/invasive/statistics.html. Published January 13, 2023. Accessed April 18, 2023.
- Chen PY, Chuang YC, Wu UI, et al. Mechanisms of Azole Resistance and Trailing in Candida tropicalis Bloodstream Isolates. J Fungi. 2021;7(8):612.
Crossref - Arastehfar A, Daneshnia F, Hafez A, et al. Antifungal susceptibility, genotyping, resistance mechanism, and clinical profile of Candida tropicalis blood isolates. Med Mycol. 2019;58(6):766-773.
Crossref - Tulyaprawat O, Pharkjaksu S, Chongtrakool P, Ngamskulrungroj P. An Association of an eBURST Group With Triazole Resistance of Candida tropicalis Blood Isolates. Front Microbiol. 2020;11:934.
Crossref - Czechowicz P, Nowicka J, Go ciniak G. Virulence Factors of Candida spp. and Host Immune Response Important in the Pathogenesis of Vulvovaginal Candidiasis. Int J Mol Sci. 2022;23(11):5895.
Crossref - Robbins N, Caplan T, Cowen LE. Molecular Evolution of Antifungal Drug Resistance. Ann Rev Microbiol. 2017;71(1):753-775.
Crossref - Whiteway M, Bachewich C. Morphogenesis in Candida albicans. Annu Rev Microbiol. 2007;61:529-553.
Crossref - Candidiasis. Centers for Disease Control and Prevention. https://www.cdc.gov/fungal/diseases/candidiasis/. Published June 28, 2022. Accessed April 18, 2023.
- Tortorano AM, Prigitano A, Morroni G, Brescini L, Barchiesi F. Candidemia: Evolution of Drug Resistance and Novel Therapeutic Approaches. Infect Drug Resist. 2021;14:5543-5553.
Crossref - Lee Y, Puumala E, Robbins N, Cowen LE. Antifungal Drug Resistance: Molecular Mechanisms in Candida albicans and Beyond. Chem Rev. 2021;121(6):3390-3411.
Crossref - Forastiero A, Mesa-Arango AC, Alastruey-Izquierdo A, et al. Candida tropicalis antifungal cross-resistance is related to different azole target (Erg11p) modifications. Antimicrob Agents Chemother. 2013;57(10):4769-4781.
Crossref - Myoken Y, Kyo T, Fujihara M, Sugata T, Mikami Y. Clinical significance of breakthrough fungemia caused by azole-resistant Candida tropicalis in patients with hematologic malignancies. Haematologica. 2004;89(3):378-380.
- Vandeputte P, Larcher G, Berges T, Renier G, Chabasse D, Bouchara JP. Mechanisms of azole resistance in a clinical isolate of Candida tropicalis. Antimicrob Agents Chemother. 2005;49(11):4608-4615.
Crossref - Song J, Zhou J, Zhang L, Li R. Mitochondria-Mediated Azole Drug Resistance and Fungal Pathogenicity: Opportunities for Therapeutic Development. Microorganisms. 2020;8(10):1574.
Crossref - Manns JM, Mosser DM, Buckley HR. Production of a hemolytic factor by Candida albicans. Infect Immun. 1994;62(11):5154-5156.
Crossref - Yigit N, Aktas E, Dagistan S, Ayyildiz A. Investigating Biofilm Production, Coagulase and Hemolytic Activity in Candida Species Isolated From Denture Stomatitis Patients. Eurasian J Med. 2011;43(1):27-32.
Crossref - Samaranayake LP, Raeside JM, Macfarlane TW. Factors affecting the phospholipase activity of Candidaspecies in vitro. Med Mycol. 1984;22(3):201-207.
Crossref - Kuhn DM, Chandra J, Mukherjee PK, Ghannoum MA. Comparison of biofilms formed by Candida albicans and Candida parapsilosis on bioprosthetic surfaces. Infect Immun. 2002;70(2):878-888.
Crossref - Clinical and Laboratory Standards Institute. Reference method for broth dilution antifungal susceptibility testing of yeasts, 4th ed Approved standard M27 Clinical and Laboratory Standards Institute, Wayne, PA. 2017.
- Ahmad S, Kumar S, Rajpal K, et al. Candidemia Among ICU Patients: Species Characterisation, Resistance Pattern and Association With Candida Score: A Prospective Study. Cureus. 2022;14(4):e24612.
Crossref - Bhattacharjee P. Epidemiology and antifungal susceptibility of Candida species in a tertiary care hospital, Kolkata, India. Curr Med Mycol. 2016;2(2):20-27.
Crossref - Delaloye J, Calandra T. Invasive candidiasis as a cause of sepsis in the critically ill patient. Virulence. 2014;5(1):161-169.
Crossref - Magalhדes YC, Bomfim MR, Melפnio LC, et al. Clinical significance of the isolation of Candida species from hospitalized patients. Braz J Microbiol. 2015;46(1):117-123.
Crossref - Kaur H, Singh S, Rudramurthy SM, et al. Candidemia in a tertiary care centre of developing country: Monitoring possible change in spectrum of agents and antifungal susceptibility. Indian J Med Microbiol. 2020;38(1):110-116.
Crossref - Sanches MD, Mimura LAN, Oliveira LRC, et al. Differential Behavior of Non-albicans Candida Species in the Central Nervous System of Immunocompetent and Immunosuppressed Mice. Front Microbiol. 2019;9:2968.
Crossref - Seyoum E, Bitew A, Mihret A. Distribution of Candida albicans and non-albicans Candida species isolated in different clinical samples and their in vitro antifungal suscetibity profile in Ethiopia. BMC Infect Dis. 2020;20(1):231.
Crossref - Berkow EL, Lockhart SR. Fluconazole resistance in Candida species: a current perspective. Infect Drug Resist. 2017;10:237-245.
Crossref - Zuo XS, Liu Y, Cai X, Zhan L, Hu K. Association of different Candida species with catheter-related candidemia, and the potential antifungal treatments against their adhesion properties and biofilm-forming capabilities. J Clin Lab Anal. 2021;35(4):e23738.
Crossref - Sachin CD, Ruchi K, Santosh S. In vitro evaluation of proteinase, phospholipase and haemolysin activities of Candida species isolated from clinical specimens. International Journal of Medicine and Biomedical Research. 2012;1(2):153-157.
Crossref - Tak V, Mathur P, Varghese P, Gunjiyal J, Xess I, Misra MC. The Epidemiological Profile of Candidemia at an Indian Trauma Care Center. J Lab Physicians. 2014;6(02):96-101.
Crossref - Chakrabarti A, Chatterjee SS, Rao KL, et al. Recent experience with fungaemia: change in species distribution and azole resistance. Scand J Infect Dis. 2009;41(4):275-284.
Crossref - Singh DP, Kumar Verma R, Sarswat S, Saraswat S. Non-Candida albicans Candida species: virulence factors and species identification in India. Curr Med Mycol. 2021;7(2):8-13.
Crossref - Nouraei H, Pakshir K, Zare Shahrabadi Z, Zomorodian K. High detection of virulence factors by Candida species isolated from bloodstream of patients with candidemia. Microb Pathog. 2020;149:104574.
Crossref - Sriphannam C, Nuanmuang N, Saengsawang K, Amornthipayawong D, Kummasook A. Anti-fungal susceptibility and virulence factors of Candida spp. isolated from blood cultures. J Mycol Med. 2019;29(4):325-330.
Crossref - Khater ES, Al-Nory MH. Exoenzymes Activity and Biofilm Production in Candida Species Isolated from Various Clinical Specimens in Benha University Hospital, Egypt. Br Microbiol Res J. 2014;4(6):654-667.
Crossref - Figueiredo-Carvalho MHG, Ramos LS, Barbedo LS, et al. Relationship between the Antifungal Susceptibility Profile and the Production of Virulence-Related Hydrolytic Enzymes in Brazilian Clinical Strains of Candida glabrata. Mediators Inflamm. 2017;8952878.
Crossref - Luo G, Samaranayake LP, Yau JY. Candida species exhibit differential in vitro hemolytic activities. J ClinMicrobiol. 2001;39(8):2971-2974.
Crossref - Galan-Ladero MA, Blanco MT, Sacristan B, Fernandez-Calderon MC, Perez-Giraldo C, Gomez-Garcia AC. Enzymatic activities of Candida tropicalis isolated from hospitalized patients. Med Mycol. 2010;48(1):207-210.
Crossref - Malcok HK, Aktas E, Ayyildiz A, Yigit N, Yazgi H. Hemolytic activities of the Candida species in liquid medium. Eurasian J Med. 2009;41(2):95-98.
- Zuza-Alves DL, Silva-Rocha WP, Chaves GM. An Update on Candida tropicalis Based on Basic and Clinical Approaches. Front Microbiol. 2017;8:1927.
Crossref - Rodriguez-Cerdeira C, Martinez-Herrera E, Carnero-Gregorio M, et al. Pathogenesis and Clinical Relevance of Candida Biofilms in Vulvovaginal Candidiasis. Front Microbiol. 2020;11:544480.
Crossref - Tulasidas S, Rao P, Bhat S, Manipura R. A study on biofilm production and antifungal drug resistance among Candida species from vulvovaginal and bloodstream infections. Infect Drug Resist. 2018;11:2443-2448.
Crossref - Sasani E, Khodavaisy S, Rezaie S, Salehi M, Yadegari MH. The relationship between biofilm formation and mortality in patients with Candida tropicalis candidemia. Microb Pathog. 2021;155:104889.
Crossref - Pereira R, Dos Santos Fontenelle RO, de Brito EHS, de Morais SM. Biofilm of Candida albicans: formation, regulation and resistance. J Appl Microbiol. 2021;131(1):11-22.
Crossref - Ponde NO, Lortal L, Ramage G, Naglik JR, Richardson JP. Candida albicans biofilms and polymicrobial interactions. Crit Rev Microbiol. 2021;47(1):91-111.
Crossref - Ghannoum MA, Rice LB. Antifungal Agents: Mode of Action, Mechanisms of Resistance, and Correlation of These Mechanisms with Bacterial Resistance. Clin Microbiol Rev. 1999;12(4):501-517.
Crossref - Arastehfar A, Gabaldon T, Garcia-Rubio R, et al. Drug-Resistant Fungi: An Emerging Challenge Threatening Our Limited Antifungal Armamentarium. Antibiotics. 2020;9(12):877.
Crossref - Pfaller MA, Diekema DJ. Progress in antifungal susceptibility testing of Candida spp. by use of Clinical and Laboratory Standards Institute broth microdilution methods, 2010 to 2012. J Clin Microbiol. 2012;50(9):2846-2856.
Crossref - Giri S, Kindo AJ, Kalyani J. Candidemia in intensive care unit patients: a one year study from a tertiary care center in South India. J Postgrad Med. 2013;59(3):190-195.
Crossref - Silva LN, Ramos LS, Oliveira SSC, et al. Insights into the Multi-Azole Resistance Profile in Candida haemulonii Species Complex. J Fungi. 2020;6(4):215.
Crossref - Fesharaki SH, Aghili SR, Shokohi T, Boroumand MA. Catheter-related candidemia and identification of causative Candida species in patients with cardiovascular disorder. Curr Med Mycol. 2018;4(2):7-13.
Crossref - Ksiezopolska E, Gabaldon T. Evolutionary Emergence of Drug Resistance in Candida Opportunistic Pathogens. Genes. 2018;9(9):461.
Crossref - Awasthi AK, Jain A, Awasthi S, Ambast A, Singh K, Mishra V. Epidemiology and microbiology of nosocomial pediatric candidemia at a northern Indian tertiary care hospital. Mycopathologia. 2011;172(4):269-277.
Crossref - Clinical and Laboratory Standards Institute (CLSI), Reference Method For Broth Dilution Antifungal Susceptibility Testing of Yeastsed, Approved standard M27-A2, Clinical Laboratory Standard Institute,Wayne, Ind, USA, 2nd edition, 2002.
- Gandham N, Vyawahare C, Jadhav S, Misra R. Candidemia: Speciation and Antifungal susceptibility testing from a Tertiary Care Hospital in Maharashtra, India. Medical Journal of Dr DY Patil University. 2016;9(5):569-599.
Crossref - Pappas PG, Kauffman CA, Andes DR, et al. Clinical Practice Guideline for the Management of Candidiasis: 2016 Update by the Infectious Diseases Society of America. Clin Infect Dis. 2016;62(4):e1-e50.
Crossref - Johnson MD, Lewis RE, Ashley ESD, et al. Core Recommendations for Antifungal Stewardship: A Statement of the Mycoses Study Group Education and Research Consortium. J Infect Dis. 2020;222(Suppl 3):S175-S198.
Crossref - Parslow BY, Thornton CR. Continuing Shifts in Epidemiology and Antifungal Susceptibility Highlight the Need for Improved Disease Management of Invasive Candidiasis. Microorganisms. 2022;10(6):1208.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.