ISSN: 0973-7510
E-ISSN: 2581-690X
The growing concern over environmental pollution caused by petroleum and oil spills, as well as the accumulation of hydrocarbons in soil and aquatic bodies, has intensified sustainable bioremediation strategies. Petroleum hydrocarbons pose a serious ecological and public health hazard due to their toxic and bioaccumulative nature. In contrast to conventional physical and chemical methods, bioremediation with hydrocarbon-degrading bacteria is eco-friendly and cost-effective in addressing environmental pollution. In the current research, we isolated and characterized four bacterial species from soil samples collected from salt lakes in Gujarat, evaluating their ability to degrade different hydrocarbons under varying salt concentrations. All four isolates (T1, T2, T3, T4) were found to grow in the presence of petrol-supplemented medium, and two of the isolates (T2 and T3) grew well in the presence of other hydrocarbons such as benzene, toluene, naphthalene, phenol, and diesel. They could tolerate up to 15% salt in the media and could degrade petrol and diesel into salt-supplemented media. All four isolates demonstrated biofilm-forming ability in the presence of various hydrocarbons in the media. The petrol degradation efficiency of the isolates, both in the presence and absence of salt in the media, demonstrated high petrol-degrading efficiencies. These isolates showed motility and endospore formation and were gram-positive rods belonging to the Bacillus and Paenibacillus genera.
Bioremediation, Hydrocarbon-degrading Bacteria, Petroleum, Biofilm
Petroleum oil is a vital global resource needed to meet energy demands; however, its extensive use has resulted in a tremendous increase in environmental pollution.1 Some of the main reasons for this are spills and discharges that occur during the storage, transportation, and processing of petroleum, blowout accidents, leakage accidents, and during overhaul of production equipment.2 Although large spills are eliminated, the inability to retrieve the contaminants increases the risk of toxicity to the surrounding ecosystem and biological communities.3 The studies carried out,4,5 suggest that petroleum contamination and oily sludge in industrial settings are intensively studied and treated using microbial consortia. Moreover, the study by Ghorbannezhad et al.6 demonstrates persistence of HMW hydrocarbons (pyrene/tetracosane) and the challenges of degrading them under saline conditions.
The accumulation of hydrocarbons in soil and water is a major environmental problem, as they pose severe health hazards, including carcinogenicity, to human life, marine life, plants and other organisms. Several physical and chemical methods, such as photo-oxidation, volatilization, bioaccumulation, and chemical oxidation, have been explored to combat this issue, but none of these methods is economic, effective, or safe.7 However, a modern comparative perspective of biosurfactant-assisted microbial remediation as a cost-effective approach was suggested by Chen
et al.8
Exposure to petroleum and related hydrocarbons has facilitated the evolution of hydrocarbon-degrading bacteria (HDB), as have evolved specific pathways (genomic evidence) and biosurfactant genes.9 Many of these bacterial species are isolated, identified, and used in different sectors, such as agricultural, chemical production, and food processing, for the degradation of waste products. Their use in the degradation of environmental pollutants is slowly gaining prominence due to their eco-friendly design and low cost.10,11 These bacteria form the foundation of microbial remediation technology, that are extensively employed to degrade petroleum hydrocarbon pollutants.12
Two of the main approaches of bioremediation include bio-stimulation and bioaugmentation. During bio-stimulation, the soil is nutritionally and physico-chemically enriched through controlled nutrition, aeration, temperature, and pH levels, and biosurfactants enhance the proliferation of hydrocarbon-degrading microbes, thereby improving biodegradation efficiency in contaminated sites. In contrast, bioaugmentation refers to adding specialized microbes in soil that can degrade hydrocarbons, thereby reinforcing native degradation communities and enhancing the biodegradation process of hydrocarbon pollutants.13
For both above processes, it is extremely important to have a working knowledge of hydrocarbon-degrading bacterial species and the conditions under which they thrive and degrade hydrocarbons. Understanding environmental factors, metabolic pathways, and key enzymes is vital for the hydrocarbon degradation process14 and bacterial consortia have already been applied effectively for the remediation of petroleum-contaminated soils.15
A consortium-based approach utilizing multiple bacterial species has been introduced in petroleum hydrocarbon biodegradation. With this newer method, enhanced degradation efficiency by complementary enzymatic systems of participating strains shows practical success in the remediation of petroleum-hydrocarbon-polluted soils.14,15
A variety of hydrocarbon-degrading bacteria have been isolated from soil samples.16 Among them, Staphylococcus,17 along with other Gram-positive bacteria such as Rhodococci18 play a significant role. Additionally,19 reported the hydrocarbon-degrading abilities of Acinetobacter baumannii, Pseudomonas aeruginosa, and Klebsiella pneumoniae recovered from soils in Delhi-NCR. Despite these findings, the diversity of bacterial genera capable of degrading hydrocarbons remains limited, highlighting the need for further studies to isolate and characterize such microorganisms.
The present study aimed to isolate and characterize bacterial species capable of degrading different hydrocarbons in a saline environment isolated from soil samples from the Wild Ass Sanctuary wetland and the salt lakes of the Little Rann of Kachchh, Gujarat, India. This study also investigated the biofilm formation capabilities of these bacterial species in a saline environment.
Study site and sample collection
Site
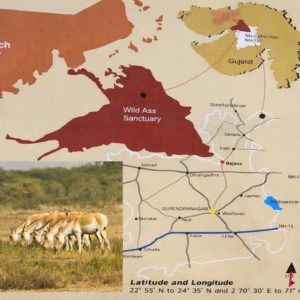
The Little Rann of Kachchh, constituting a Biosphere Reserve, extends over about 6,500 km² of low-saline desert with discontinuous patches of uplands. Of this area, 4,841 km2 has been declared as the Indian Wild Ass Sanctuary, which is the habitat of the critically endangered Indian Wild Ass (Equus hemionus khur)20 (Figure 1).
Figure 1. Sampling site from Indian Wild Ass Sanctuary of Little Rann of Kachchh (Source: https://en.wikipedia.org/wiki/Indian_Wild_Ass_Sanctuary)
Soil samples were obtained from two different locations (site 1 and 2) in the Wild Ass Sanctuary of Little Rann of Kachchh, Gujarat. Samples were obtained during April, the pre-monsoon period to reduce the impact of rainfall and surface runoff on microbial communities. Soil was obtained from a depth of around 0-15 cm at each location by avoiding contamination with sterile spatulas and then transferred to non-reactive sterile plastic bottles immediately. Bottles were filled up to two-thirds of capacity to provide sufficient aeration during transportation. Bottles were tightly capped and marked with all the site-specific details, date, location, sample code, etc. Sealed bottles were brought to the laboratory aseptically.
Culture media and chemicals
The reagents used in this study were of analytical grade. Nutrient agar media and nutrient broth were used to grow bacteria. BHM is a suitable medium for testing and investigating the hydrocarbon degradation potential of microorganisms. The compositions of the different media prepared for the experiments were as follows:
Bushnell Haas Agar (BHM Agar, g/L)
MgSO4·7H2O (0.2), CaCl2·2H2O (0.02), KH2PO4 (1.0), K2HPO4 (1.0), NH4NO3 (1.0), FeCl3 (0.05), and agar (20.0).21
Basal salt medium (BSM per 1000 ml distilled water)
NaCl (15.0 g), KH2PO4 (0.8 g), K2HPO4 (1.2 g), NH4NO3 (1.0 g), MgSO4·7H2O (0.2 g), FeCl3 (50 mg), CaCl‚ (20 mg), MnSO4 (1.0 mg), Na2MoO4 (0.2 mg); pH adjusted to 7.2. For a solid medium, 1.5% agar was incorporated.
Luria Bertani (LB) broth, per 1000 ml distilled water
NaCl (10.0 g), peptone (10.0 g), yeast extract (5.0 g); pH adjusted to 7.2.
Phosphate Buffered Saline (PBS, per 1000 ml distilled water)
NaCl (8.0 g), KCl (0.2 g), Na2HPO4 (1.44 g), KH2PO4 (0.24 g); pH adjusted to 7.2.22
Quantification of heterotrophic bacteria in a soil
The viable heterotrophic bacterial population present in the collected soil samples was assessed by the serial dilution technique. Ten grams of the soil sample were added to 90 ml of nutrient broth, and serial dilutions were made in 9 ml saline taken in sterile and labeled tubes. From each dilution, 0.1 ml was inoculated onto individual BHM agar plates and incubated at 37 °C for 24-48 hrs. After incubation, the bacterial colonies formed were enumerated, and the result is expressed as CFU/ml.23
Screening of Hydrocarbon-Utilizing Bacteria (HUB)
Primary screening of hydrocarbon-utilizing isolates from soil
To obtain pure hydrocarbon-degrading colonies, soil samples were processed by enrichment, which was carried out in 50 mL of minimal salt medium (MSM) adjusted at pH 6.5 and containing 1% (v/v) of sterile petrol as a carbon source and incubated at 30 °C, with shaking at 120 rpm for 7 days. The final enrichment culture broth was plated on Bushnell Haas (BH) agar plates supplemented with (1% v/v) petrol and incubated at 30 °C for 24 h. BH agar is a selective medium for monitoring microbial hydrocarbon degradation. Distinct colonies were picked and streaked on nutrient agar and maintained through sub-culturing for subsequent experiments.
Secondary screening of hydrocarbon-utilizing isolates
Pure culture isolates from primary screening were transferred into BHM broth and incubated overnight at 37 °C and centrifuged at 5000 RPM. The supernatant was discarded, and the pellet was washed with PBS twice. The washed pellets were resuspended in BHM broth and incubated at 37 °C for 24 hours. 0.1 ml of these fresh cultures is added to 9.9ml of BHM broth supplemented with 0.5%, 1%, 2%, 5% and 10% petrol in separate sterile test tubes. Tubes were incubated at 37 °C for 48 hrs, and bacterial growth was monitored by taking OD at 600 nm to evaluate the utilization of petroleum as a carbon source by isolates. The culture with >0.4 OD was chosen for further characterization.
Screening salt tolerance in hydrocarbon-utilizing isolates
The isolates selected from secondary screening were tested for their salt tolerance. These were also grown on nutrient agar plates supplemented with various concentrations of NaCl (0.5%-30%). The isolates that were growing on the highest concentration of salt-supplemented plates were selected for further screening.
Screening of hydrocarbon-utilizing ability in the presence of salt
Salt-tolerant isolates were selected and grown on BHM agar plates containing 5% salt and 15% petrol. The plates were placed in an incubator for 48 hours at 37 °C. The growth of cultures on these plates was recorded.
Determination of the multiple hydrocarbon-utilizing ability of isolates
100 µl of selected microbes from 2.4.2 and 2.4.4 were added to BHM agar plates that were supplemented with 1% of various hydrocarbons (diesel, benzene, toluene, naphthalene, and phenol) as carbon sources. Plates were placed at 37 °C for 48 hours and subsequently monitored for growth. Growth curve response of these isolates for utilizing these hydrocarbons was also monitored for seven days. Fresh overnight culture of the selected isolates was added to labelled flasks containing BHM supplemented with 1% of petrol, diesel, benzene, toluene, naphthalene, and phenol each as carbon sources. Flasks were kept at 37 °C at 160 rpm. The aliquots were taken, and OD was recorded spectrophotometrically (Shimadzu).
Biofilm formation
The glass tube assay24 was used to assess biofilm formation. The overnight bacterial cultures in LB broth (1 ml) were added to glass tubes and incubated at 37 °C for 48 hrs. After the incubation medium was removed, the tubes were gently washed with PBS to eliminate non-adherent cells. Biofilms were identified by staining with 0.2% crystal violet and later destained with 95% ethanol. The presence of a crystal violet ring representing cell mass was considered positive for biofilm formation. Biofilm growth was quantified by microtiter plate assay. Briefly, 200 µl of bacterial culture was inoculated in triplicate into a 96-well plate in triplicate and incubated at 37 °C for 48 hours. After incubation, the culture was discarded, and the wells were washed twice with PBS, air dried, and stained with 200 µl of 0.2% crystal violet for 5 min. The excess stain was removed by washing with distilled water, and then the plate was air-dried. 200 µl of 95% ethanol was added to solubilize the bound dye for 30 min, and absorbance was measured at 595 nm using an ELISA plate reader (Shimadzu). The extent of Biofilm formation was quantified in terms of the absorbance of ethanol solution containing solubilized crystal violet.
Quantitative estimation of petrol degradation
The petrol-degrading ability of the isolates is determined by the DCPIP method.25,26 Isolates are grown to mid-exponential phase, adjusted to an OD600 of 0.01, and inoculated into sterile LB media in a 96-well microtiter plate. 50 µl of 1% (v/v) aqueous solution of 2,6-dichlorophenolindophenol (DCPIP), a redox indicator, was dispensed to each well. After inoculation plates were maintained at 30 °C for 72 hrs. Cultures from the microtiter plate were removed, and wells were rinsed thoroughly with phosphate-buffered saline. The adherent cells were stained for 15 min with crystal violet (1% w/v) and then washed with PBS three times. The stain was solubilised with an acetone-alcohol mixture (1:4). The absorbance of the solubilized dye was determined at 595 nm. Negative controls were set in parallel wells, one with no culture, one with no hydrocarbon, and one with no DCPIP. The assay is considered positive for hydrocarbon degradation when the colour of the medium becomes colourless and negative when no change in colour, i.e., remains blue, is observed.
Identification of selected HUB
The chosen bacterial isolates were determined morphologically by staining, and by their biochemical properties according to the standard procedures outlined in Bergey’s Manual of Determinative Bacteriology.27 Molecular identification was done by 16S rRNA gene sequencing.28 The 16S rRNA gene was amplified from the bacteria using universal primers: forward primer 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) and reverse primer 1492R (5′-GTATTACCGCGGCTGCTGG-3′). PCR parameters consisted of an initial denaturation for 5 min at 95 °C, after which there were 30 cycles of denaturation for 30 s at 95 °C, annealing for 30 s at 55 °C, and extension for 90 s at 72 °C, followed by a final extension for 10 min at 72 °C. Amplicons were sequenced on an ABI 3500 Genetic Analyzer. These resulting sequences were compared using the Basic Local Alignment Search Tool (BLAST; http://www.ncbi.nlm.nih.gov/blast) at the National Center for Biotechnology Information (NCBI) to establish sequence homology. Consensus sequences were aligned against stored 16S rDNA sequences of GenBank using BLASTN. Phylogenetic analysis was performed in MEGA X24 using the neighbor-joining DNA distance algorithm, and trees were built. Partial 16S rRNA gene sequences of the four isolates were deposited in GenBank (NCBI), and accession numbers were acquired.
Statistical analysis
Statistical analysis of data on petrol degradation by the isolates was conducted using one-way ANOVA with post hoc multiple comparisons carried out by Tukey’s HSD test in SPSS v.18.0 (SPSS Inc., IBM Corp., Chicago, IL, USA). Results were statistically significant at p < 0.05 and are given as mean ± standard deviation of triplicates (n = 3).
Soil Sample enrichment, quantitative enumeration of heterotrophic bacteria in soil samples, and primary screening of soil samples
The total culturable bacteria count in the soil was determined by plating the serial dilutions of the Soil enriched in the nutrient agar. Qualitatively count is represented as colony-forming units (CFU/ml). Site 1 and site 2 showed 4.5 x 106 CFU/ml and 5 x 106 CFU/ml, respectively (Table 1).
Table (1):
CFU/ml from sampling site 1 and site 2
Sampling Site |
Total heterotrophic bacteria (CFU/ml) |
|---|---|
1 |
4.5 x 106 |
2 |
5 x 106 |
Secondary screening of selected isolates
Soil-derived bacterial isolates were screened using the selective enrichment with Bushnell-Haas medium containing 1% petrol. The aliquots of culture from this BHM supplemented with 1% v/v petrol were spread on plates containing 0.5%, 1%, 2% and 5% and 10% petrol. Morphologically distinct colonies growing on the plates with the highest petrol concentration (5%) were selected, inoculated, and grown overnight at 37 °C in BHM broth.
The growth patterns of each of the 4 isolates on BHM supplemented with different concentrations of petrol are represented in Table 2. All 4 isolates showed growth on all concentrations of petrol. Therefore, all these isolates were selected for further characterization. These isolates were named as T1, T2, T3, and T4.
Table (2):
Growth patterns of isolates on BHM plates supplemented with varying concentrations of petrol (v/v)
Isolate |
0.5% Petrol |
1% Petrol |
2% Petrol |
5% Petrol |
|---|---|---|---|---|
T1 |
+++ |
++ |
++ |
++ |
T2 |
+++ |
++ |
++ |
++ |
T3 |
++ |
++ |
++ |
++ |
T4 |
++ |
++ |
++ |
++ |
Screening of growth on salt-supplemented plates
All isolates obtained from the secondary screening process were assessed for their growth in nutrient broth supplemented with varying concentrations of salt. The growth patterns of the 4 isolates on 5%, 10%, 15%, and 20% salt concentrations in the media are shown in Table 3. Out of 12 isolates, only four isolates had considerable growth in the presence of 5% and 10% salt in the media whereas moderate growth was observed in media supplemented with 15% salt. However, none of the isolate’s growth was observed in the presence of 20% salt in the media (Table 3).
Table (3):
Growth patterns of isolates on BHM supplemented with 5%, 10%, 15%, and 20% salt concentrations
Isolate |
5% NaCl |
10% NaCl |
15% NaCl |
20% NaCl |
|---|---|---|---|---|
T1 |
++++ |
++++ |
++ |
– |
T2 |
++++ |
++++ |
++ |
– |
T3 |
++++ |
++++ |
++ |
– |
T4 |
++++ |
++++ |
++ |
– |
Screening of growth on salt and hydrocarbon-supplemented media
The four isolates, named T1, T2, T3, and T4, were found to grow on all salt concentrations except 20%. These isolates exhibiting different colony morphologies were selected and further screened for hydrocarbon-utilizing ability under salt stress by inoculating them on plates enriched with 1% of petrol and diesel and 5% salt concentration. It was seen that all 4 isolates showed considerable growth in the presence of 5% salt + 1% petrol, as well as 5% salt + 1% diesel, indicating that these isolates were capable of metabolizing hydrocarbons in a saline environment (Table 4). Therefore, these 4 isolates were selected for further characterization.
Table (4):
Growth patterns of isolates on media supplemented with salt and hydrocarbon
Isolate |
5% Salt + 1% Petrol |
5% Salt + 1% Diesel |
|---|---|---|
T1 |
+++ |
+++ |
T2 |
+++ |
+++ |
T3 |
+++ |
+++ |
T4 |
+++ |
+++ |
Multiple hydrocarbons utilization potential
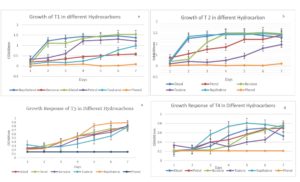
The overnight cultures (100 µl) of T1, T2, T3, and T4 were added to BHM agar plates that were supplemented with different hydrocarbons (1%), namely diesel, benzene, toluene, naphthalene, and phenol as sole carbon sources to assess their utilization capacity over a period of 24 and 48 hrs. The growth results of each of the 4 isolates selected in primary screening on BHM supplemented with 1% of different hydrocarbons are shown in Table 5. From this table, isolates T1, T2, T3, and T4 exhibited growth in the presence of petrol, diesel, benzene, and naphthalene as carbon sources in the medium. They showed mild growth in the presence of toluene at 24 hrs and showed no growth in the presence of phenol in the media plates (Figure 2(a-d)).
Table (5):
Growth of isolates on BHM supplemented with 1% hydrocarbons (petrol, diesel, benzene, toluene, naphthalene and phenol) recorded at 24 h and 48 h
| Isolate | 1% Petrol | 1% Diesel | 1% Benzene | 1% Toluene | 1% Naphthalene | 1% Phenol | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 24 h | 48 h | 24 h | 48 h | 24 h | 48 h | 24 h | 48 h | 24 h | 48 h | 24 h | 48 h | |
| T1 | ++ | ++ | ++ | ++ | ++ | ++ | + | + | ++ | ++ | – | – |
| T2 | ++ | ++ | ++ | ++ | ++ | ++ | + | + | ++ | ++ | – | – |
| T3 | ++ | ++ | ++ | ++ | ++ | ++ | + | + | ++ | ++ | – | – |
| T4 | ++ | ++ | ++ | ++ | ++ | ++ | + | + | ++ | ++ | – | – |
Figure 2. Growth characteristics of isolated strains (T1, T2, T3, T4) on different hydrocarbon (diesel, petrol, benzene, toluene, naphthalene, phenol) supplemented media growing on salt-supplemented media. (a) Growth pattern of T1; (b) Growth pattern of T2; (c): Growth pattern of T3; (d) Growth pattern of T4
Biofilm formation ability
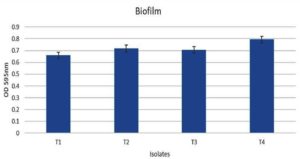
The four selected hydrocarbon-utilizing isolates were evaluated for their biofilm-forming ability using the crystal violet staining assay. Each isolate was cultured in nutrient agar enriched with petrol, diesel, naphthalene, benzene, and toluene in separate tubes for 7 days, after which the culture media were removed and the cells and exopolysaccharide films on the test tube walls were stained with crystal violet. All the isolates exhibited growth with OD595 greater than 0.2 and demonstrated biofilm formation in the presence of petrol in the media. The biofilm formation, as seen by OD595 greater than 0.65 for all isolates, represents good biofilm-forming ability of the isolates. Out of the 4 isolates, the maximum biofilm formation ability was seen in T4. The OD595 values of the biofilms formed by the 4 isolates are shown in Figure 3.
Petrol degradation assay
Petrol degradation efficiencies by DCPIP method
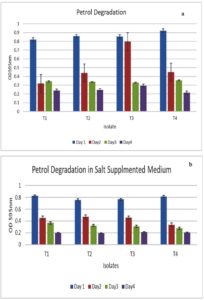
The hydrocarbon degradation ability of the selected isolates was quantified using the DCPIP assay. DCPIP, a redox indicator, is blue in the oxidized state and colourless when reduced, and the colour change of this indicator can be quantified by measuring OD600. To perform this assay, the selected isolates were cultured on BHM supplemented with 1% petrol and incubated at 37 °C on a shaker incubator set at 180 rpm for 4 days. The time taken for decolourization was monitored at regular intervals during incubation at 24 hrs, 48 hours, and 72 hours, and OD600 was measured at each time point. The colour changes of DCPIP, demonstrated by the disintegration of the oil layer, were observed within 24 hrs. In case of T3 and T4, the time for decolourization, represented by the reduction of DCPIP due to petrol-degrading activity of bacteria, was 48 hrs. The results of this assay are represented in Figure 4a, with OD600 values taken at 24 hr intervals. T1 and T2 also showed considerable degradation in 48 hours (Figure 4a).
Petrol degradation in salt-supplemented media
The hydrocarbon-degrading potential of the selected isolates under salt stress was evaluated by inoculating and incubating them in BHM supplemented with 1% petrol and 5% salt for 3 days at 37 °C. T1 and T4 show better levels of petrol-degrading ability, and the presence of a saline environment did not alter the hydrocarbon-degrading ability of the isolates. T3 and T4 show a similar level of hydrocarbon degradation. DCPIP was added to this medium also, and the time taken for decolourization was monitored by measuring OD600 at regular time points. The hydrocarbon-degrading capability of the isolates in the presence of salt, quantified by the DCPIP method, is given in Figure 4b. All four isolates were able to degrade 50% petrol in 48 hours (Figure 4b).
Figure 4. (a) Quantification of hydrocarbon (Petrol) degrading ability of the strains using the DCPIP assay method; (b): Quantification of hydrocarbon degrading ability of the isolates in the presence of salt using the DCPIP assay method
Identification and Characterization
Morphological characterization
The various morphological characteristics of the four isolates, namely T1, T2, T3, and T4, were studied as per standard methods. All the isolates showed motility and could form endospores. The colony characteristics and other morphological features of the isolates are given in Table 6.
Table (6):
Morphological profiling of the isolates
| Isolate | Colony Characteristics | Gram Staining | Motility | Endospore Formation | ||
|---|---|---|---|---|---|---|
| Colour | Surface | Shape and size | ||||
| T1 | Whitish | Smooth | – | Gram-positive rods | Motile | + |
| T2 | White/Translucent | Smooth | Dendron Tree-shaped | Gram-positive | Motile | + |
| T3 | Whitish | Smooth | Irregular | Gram-positive rods | Motile | + |
| T4 | Grey, White | Smooth | Granular | Gram-positive rods | Motile | + |
Biochemical characterization
The four isolates were biochemically characterized by performing various biochemical tests, as shown in Table 7. All isolates tested positive for catalase and positive and were able to utilize glucose as a carbon source in the medium. Isolates T2 and T3 were found to be indole and methyl red positive, and isolates T1 and T4 were found to be Voges-Proskauer and citrate positive. Only T2 was capable of starch hydrolysis, whereas T1 and T3 were capable of sucrose and mannitol utilization. Except for T2, all isolates showed positive for nitrate reduction. However, none of the isolates showed the capability for sulphur reduction or urease production (Table 7).
Table (7):
Biochemical profiling of the isolates
Biochemical Test |
T1 |
T2 |
T3 |
T4 |
|---|---|---|---|---|
Indole Test |
– |
+ |
+ |
– |
Methyl Red Test |
– |
+ |
+ |
– |
Voges Proskauer Test |
+ |
– |
– |
+ |
Citrate Test |
+ |
– |
– |
+ |
Catalase Test |
+ |
+ |
+ |
+ |
Starch Hydrolysis |
– |
+ |
– |
– |
Glucose Utilization |
+ |
+ |
+ |
+ |
Sucrose Utilization |
+ |
– |
+ |
+/- |
Mannitol Utilization |
+ |
– |
+ |
– |
Nitrate Reduction |
+ |
– |
+ |
+ |
Sulphur Reduction |
– |
– |
– |
– |
Urease Production |
– |
– |
– |
– |
Oxidase Test |
+ |
+ |
+ |
+ |
16S rRNA sequencing
To characterize the bacterial isolates, 16S rRNA gene sequencing was performed for the four strains, and the findings are tabulated in Table 8. Based on partial 16S rRNA sequences and BLAST analysis, isolate T1 was found to be Bacillus sp., T2 to be Paenibacillus dendritiformis strain ANSK05, T3 to be Paenibacillus sp., and T4 to be Bacillus cereus strain IARI-BC-1. Sequences have been deposited at the NCBI GenBank under accession numbers MH684905.1, MH660379, MH660380, and MH684904.1 (Table 7). The phylogenetic tree was created with MEGA X and is presented in Figure 5.
Table (8):
Sequencing results of 16S rRNA of the isolates and their identification
Isolate |
Accession Number |
Number of Nucleotides |
Alignment |
Identity |
Nearest Phylogenetic Neighbour |
|---|---|---|---|---|---|
T1 |
MH684905.1 |
1467 |
100% |
97% |
Bacillus sp. |
T2 |
MH660379 |
1459 |
100% |
99% |
Paenibacillus dendritiformis strain ANSK05 |
T3 |
MH660380 |
1409 |
100% |
99% |
Paenibacillus sp. |
T4 |
MH684904.1 |
1492 |
100% |
100% |
Bacillus cereus strain IARI-BC-1 |
Figure 5. Phylogenetic tree showing the relationship of isolates T1, T2, T3, T4 with other bacteria as obtained by MEGAX: Phylogenetic tree of four isolates is based on 16S rRNA gene sequence analysis. Numbers at nodes are bootstrap values (%) based on neighbor-joining analysis. The scale bar indicates the number of nucleotide substitutions per site
The present investigation was undertaken to evaluate hydrocarbon-degrading and biofilm-forming bacteria in soil samples from the Wild Ass Sanctuary in Gujarat. We were able to isolate four distinct bacterial species that demonstrated degradation of different hydrocarbons such as petrol, diesel, benzene, toluene, and naphthalene, to varying extents, and could also tolerate salt up to 15% in the medium. Furthermore, these four isolates demonstrated hydrocarbon degradation even in the salt-supplemented growth media. Salinity-tolerant Hydrocarbon degradation is a critical ecological attribute of soil bacteria, as it enables the soil bacteria to survive and bioremediate hydrocarbon-contaminated saline habitats. Petrol and diesel degrading bacteria are gaining importance in both environmental microbiology as well as agricultural microbiology due to their ability to clear the environment of toxic pollutants and to maintain soil health.
Research has documented that soil often harbors bacteria with promising potential of breaking down hydrocarbons.29 isolated Bacillus cereus, Bacillus subtilis, and Pseudomonas aeruginosa, from soil collected from Mansehra, Pakistan, that could degrade diesel. Similarly, Rehman et al.30 reported that biosurfactants producing B. cereus exhibited significant oil degradation properties in hydrocarbon-polluted soils. In early studies,31 isolated the DNA of Pseudomonas sp. from contaminated soil and identified the enzymes present which were able to break down naphthalene, suggesting the potential use in remediation of naphthalene-contaminated environments. Later, Shen et al.32 identified that Paenibacillus ehimensis could efficiently degrade oils with different viscosities.
The applications of Paenibacillus sp. have also been described by Shibulal et al.,33 in the degradation of naphthalene and phenanthrene. Biosurfactants released by Paenibacillus dendritiformis have been reported to be capable of motor oil sludge degradation.34 The present investigation highlights the presence of B. cereus and P. dendritiformis in soil samples from high salinity environments in Gujarat, along with other Bacillus and Paenibacillus sp., and demonstrates their capability to degrade different hydrocarbons. However, the production of biosurfactants from these strains, as reported in other studies, is yet to be investigated.
The abilities of hydrocarbon-degrading bacteria to thrive in a high-salinity environment are important plant growth-promoting attributes, as excess salt has adverse effects on plants. In such high salinity conditions, microbes like algae and cyanobacteria demonstrate high biological productivity, which leads to the accumulation of hydrocarbons in the soil.35 Additionally, an increase in industrial production has led to an overabundance of oil, brine, and petroleum in the soil, affecting both soil fertility as well as plant growth.36 As a result, it has become increasingly important to identify bacterial species that are capable of degrading different hydrocarbons in a saline environment, so that these can be used to maintain soil health and promote plant growth.
Biofilm formation is an additional important attribute of hydrocarbon-degrading bacteria in an environment exposed to saline stress, as it protects and facilitates adaptation in stressed habitats.37 In vitro tests demonstrated that biofilm formation enhances the effectiveness of these microbes in oil degradation by forming biofilms on surfaces like glass slides and gravel particles.38 Al-awadhi et al.,39 have reported enhanced biofilm-mediated bioremediation of hydrocarbon mixtures, monoaromatic compounds, and cyclic compounds by 3 different Pseudomonas sp. Most studies focus on biofilm formation and hydrocarbon degradation in bacteria collected from water samples, and there are limited studies that report hydrocarbon degradation efficiency by biofilm-forming soil bacteria.40,41 have suggested haloarchaea mechanisms and biosurfactant production under saline conditions. This research assessed the biofilm-forming potential of soil isolates when exposed to diverse hydrocarbons, including petrol, and exhibited pronounced biofilm development. This implies that the efficiency of isolates in developing biofilms may play an important role in enhancing hydrocarbon-degradation efficiency in saline soils.
Efficient petroleum degradation by bacteria, especially in saline soils, is an essential ecological mechanism for combating environmental pollution associated with petroleum and its derivatives.42 The salinity and nutrient effects for the biofilm formation under saline conditions and improved degradation rates by the halophiles.42,43 Findings from Fatima et al.44 reported alleviation of the adverse effects of salinity and enhancement of petroleum degradation after inoculation of saline soils with biosurfactant-producing Pseudomonas sp. It was observed that there was a considerable increase in the activity of the dehydrogenase enzyme and a reduction in soil toxicity by up to 30%. Research following this type of approach are important for enhancing the current perspective of microbial interventions for the remediation of hydrocarbon-contaminated saline soils. A study conducted by Ebadi et al.45 reported the effects of salinity on hydrocarbon-degrading abilities of soil bacteria, stating that moderate salinity enhanced hydrocarbon degradation of bacteria, while higher salt content led to a decrease in microbial load, thereby affecting biodegradation. Therefore, it is important to identify hydrocarbon-degrading bacteria that demonstrate resilience and metabolic functionality in salt-stressed soil.
In the present investigation, we used the DCPIP method to evaluate the petrol degradation efficiency of the isolates in an environment with or without salinity stress. Our results demonstrated petrol degradation that could be observed in the culture tubes, and it was quantitatively determined by measuring absorbance readings at regular time-points, which decreased with an increase in incubation period, signifying a decrease in DCPIP in the medium. Further, when comparing petrol degradation efficiencies of bacteria in media with and without salt supplementation, the results show similar trends, suggesting that even in the presence of a saline environment, the isolates were capable of degrading petrol to a similar extent as in the absence of salinity. This result is significant for the identification of suitable microbes for the process of in situ bioremediation at contaminated sites such as oil refineries for the microbial-mediated degradation of heterogeneous hydrocarbon compounds in the soil.46
An advantage of our study is that it tested the biodegradation abilities of the isolates on not only petrol, but also its constituents such as benzene, toluene, and naphthalene. The ability of bacteria to metabolize multiple hydrocarbons enhances their suitability for petrol remediation, as they are superior to bacteria with restricted ability to degrade petrol alone. Additionally, bacteria capable of degrading multiple hydrocarbons also have the advantage of not needing the application of a bacterial consortium or exploiting the phenomenon of catabolism in the soil.
Globally, petroleum and petroleum products are considered dangerous pollutants in the environment, with several adverse consequences for living organisms. Use of microorganisms for bioremediation of hydrocarbons is an eco-sustainable and cost-efficient approach that is currently being explored by the scientific community. Multiple bacterial species involved in degrading hydrocarbons have already been described in scientific studies; however, the search is still on for finding other bacterial species with enhanced abilities for hydrocarbon degradation. In this work, we aimed to investigate bacterial isolates with multiple hydrocarbon degradation abilities from soil samples of salt fields in Gujarat. Two of these isolates were identified as Paenibacillus sp. and B. cereus by 16S rRNA sequencing. Preliminary results showed that both isolates are promising candidates for hydrocarbon degradation and biofilm formation in a hypersaline environment. The presence of dual properties suggests that these isolates are good candidates for hydrocarbon remediation in contaminated sites and merit further detailed characterization. Also, soils from other regions can be sampled to discover other bacterial species capable of hydrocarbon degradation that can be used for the process of bioremediation.
Research shows that salt-tolerant and biofilm-forming bacteria, which can degrade petroleum and its constituents such as benzene, toluene, and naphthalene, can survive in saline soils in the Wild Ass Sanctuary, Gujarat. Hydrocarbon degradation abilities of isolates in both salty (saline) and non-salty environments, indicating high adaptability to salt stress. Identified isolates, Bacillus cereus and Paenibacillus sp. are promising in their possible application in the bioremediation of hydrocarbons.
The synergistic characteristics of multi-hydrocarbon degradation, salinity tolerance, and biofilm formation indicate that the bacteria would be good candidates for the in situ remediation of the hydrocarbon-contaminated saline soils. Future research must be aimed at further explaining the degradation pathways, evaluating the production of biosurfactants, and authenticating field-level to boost the sustainable bioremediation approach.
ACKNOWLEDGMENTS
The authors thank the authorities of Amity University Uttar Pradesh for providing the facilities to conduct this research.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
NB and TA conceptualized the study. MY, RD, RK, TA and NB applied methodology. NB and TA performed data curation. NB supervised the study and performed formal analysis. TA and NB wrote the original draft. SA and SS reviewed the manuscript. SS wrote the manuscript. NB, SA and SS revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies on human participants or animals performed by any of the authors.
- Xue J, Yu Y, Bai Y, Wang L, Wu Y. Marine Oil-Degrading Microorganisms and Biodegradation Process of petroleum hydrocarbon in marine Environments: A review. Curr Microbiol. 2015;71(2):220-228.
Crossref - Wang C, Liu X, Guo J, Lv Y, Li Y. Biodegradation of marine oil spill residues using aboriginal bacterial consortium based on Penglai 19-3 oil spill accident, China. Ecotoxicol Environ Saf. 2018;159:20-27.
Crossref - Xu X, Liu W, Tian S, et al. Petroleum Hydrocarbon-Degrading bacteria for the remediation of oil pollution under aerobic conditions: A Perspective analysis. Front Microbiol. 2018;9.
Crossref - Hentati D, Ramadan AR, Abed RMM, Abotalib N, Nayal AME, Ismail W. Biotreatment of oily sludge by a bacterial consortium: effect of bioprocess conditions on biodegradation efficiency and bacterial community structure. Front Microbiol. 2022; 13:998076.
Crossref - Hentati D, Ramadan AR, Abed RMM, Abotalib N, Nayal AME, Ismail W. Functional and structural responses of a halophilic consortium to oily sludge during biodegradation. Appl Microbiol Biotechnol. 2024; 108:116.
Crossref - Ghorbannezhad H, Moghimi H, Dastgheib SMM. Biodegradation of high-molecular-weight hydrocarbons under saline condition by halotolerant Bacillus subtilis and its mixed cultures with Pseudomonas species. Sci Rep. 2022;12:13227.
Crossref - Dasgupta D, Ghosh R, Sengupta TK. Biofilm-Mediated enhanced crude oil degradation by newly isolated Pseudomonas species. ISRN Biotechnol. 2013;1-13.
Crossref - Chen W, Sun J, Ji R, et al. Crude oil biodegradation by a biosurfactant-producing bacterial consortium in high-salinity soil. J Mar Sci Eng. 2024;12(11):2033.
Crossref - Nayarisseri A, Singh SK. Genome analysis of biosurfactant producing bacterium, Bacillus tequilensis. PLoS One. 2023;18(6):e0285994.
Crossref - Guerra AB, Oliveira JS, Silva-Portela RCB, et al. Metagenome enrichment approach used for selection of oil-degrading bacteria consortia for drill cutting residue bioremediation. Environ Pollut. 2018; 235:869-880.
Crossref - Pal S, Hait A, Mandal S, Roy A, Sar P, Kazy SK. Crude oil degrading efficiency of formulated consortium of bacterial strains isolated from petroleum-contaminated sludge. 3 Biotech. 2024;14(10):220.
Crossref - Dvorak P, Nikel PI, Damborsky J, de Lorenzo V. Bioremediation 3.0: Engineering pollutant-removing bacteria in the times of systemic biology. Biotechnol Adv. 2017;35(7):845-866.
Crossref - Abed RMM, Al-Sabahi J, Al-Maqrashi F, Al-Habsi A, Al-Hinai M. Characterization of hydrocarbon-degrading bacteria isolated from oil-contaminated sediments in the Sultanate of Oman and evaluation of bioaugmentation and biostimulation approaches in microcosm experiments. Int Biodeterior Biodegradation.2014;89:58-66.
Crossref - Hassanshahian M, Zeynalipour MS, Musa FH. Isolation and characterization of crude oil degrading bacteria from the Persian Gulf (Khorramshahr provenance). Mar Pollut Bull. 2014;82(1-2):39-44.
Crossref - Wu M, Li W, Dick WA, et al. Bioremediation of hydrocarbon degradation in a petroleum-contaminated soil and microbial population and activity determination. Chemosphere. 2016; 169:124-130.
Crossref - Varjani SJ, Upasani VN. Biodegradation of petroleum hydrocarbons by oleophilic strain of Pseudomonas aeruginosa NCIM 5514. Bioresour Technol. 2016; 222:195-201.
Crossref - DeRito CM, Pumphrey GM, Madsen EL. Use of Field-Based Stable Isotope Probing to Identify Adapted Populations and Track Carbon Flow through a Phenol-Degrading Soil Microbial Community. Appl Environ Microbiol. 2005;71(12):7858-7865.
Crossref - Kaplan CW, Kitts CL. Bacterial succession in a petroleum land treatment unit. Appl Environ Microbiol. 2004;70(3):1777-1786.
Crossref - Bhagat NN, Roy NP, Singh NS, Allen NT. Isolation and characterization of bacteria with multiple traits: Hydrocarbon degradation, antibiotic-resistant and metal tolerant. Int J Res Pharm Sci. 2019;10(4):3789-3795.
Crossref - UNESCO World Heritage Centre. (n.d.). Wild Ass Sanctuary, Little Rann of Kutch. UNESCO World Heritage Centre. Retrieved December 17, 2017, from https://whc.unesco.org/en/tentativelists/2105/
- Sawadogo A, Harmonie OC, Sawadogo JB, Kabore A, Traore AS, Dianou D. Isolation and Characterization of Hydrocarbon-Degrading Bacteria from Wastewaters in Ouagadougou, Burkina Faso. J Environ Prot (Irvine, Calif). 2014;05(12):1183-1196.
Crossref - Mangwani N, Kumari S, Das S. Taxonomy and Characterization of Biofilm Forming Polycyclic Aromatic Hydrocarbon Degrading Bacteria from Marine Environments. Polycycl Aromat Compd. 2019;41(6):1249-1262.
Crossref - Cappuccino JG. and Sherman N.. Microbiology – a Laboratory Manual, 4th edn. Addison Wesley Longman, Inc., Harlow. 1996.
- O’Toole GA, Kolter R. Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol Microbiol. 1998;30(2):295-304.
Crossref - Marchand C, St-Arnaud M, Hogland W, Bell TH, Hijri M. Petroleum biodegradation capacity of bacteria and fungi isolated from petroleum-contaminated soil. Int Biodeterior Biodegradation. 2016 ;16:48–57.
Crossref - Hanson KG, Desai JD, Desai AJ. A rapid and simple screening technique for potential crude oil degrading microorganisms. Biotechnol Tech. 1993;7(10):745–748.
Crossref - Buchanan RE, Gibbons NE. Bergey’s Manual of Determinative Bacteriology. 1975.
- Jain K, Parida S, Mangwani N, Dash HR, Das S. Isolation and characterization of biofilm-forming bacteria and associated extracellular polymeric substances from oral cavity. Ann Microbiol. 2013;63(4):1553-1562.
Crossref - Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol 2018;35(6):1547-1549.
Crossref - Rehman ZU, Khan K, Shah Faisal, et al. Isolation and Identification of Diesel-Degrading Bacteria from Oil Contaminated Soil in Mansehra, Pakistan. Int J Eng Sci Res 2015;6(10):1-1
- Janaki S, Thenmozhi S, Muthumari R. A study on Hydrocarbon Degradation by Biosurfactant Producing Bacillus cereus in Oil Contaminated Soil Samples. SSR Inst Int J Life Sci. 2016;2(4).
Crossref - Shen Y, Stehmeier LG, Voordouw G. Identification of Hydrocarbon-Degrading Bacteria in Soil by Reverse Sample Genome Probing. Appl Environ Microbio. 1998 64(2):637-645
Crossref - Shibulal B, Al-Bahry SN, Al-Wahaibi YM, Elshafie AE, Al-Bemani AS, Joshi SJ. The potential of indigenous Paenibacillus ehimensis BS1 for recovering heavy crude oil by biotransformation to light fractions. PLoS ONE. 2017;12(2): e0171432.
Crossref - Daane LL, Harjono I, Barns SM, Launen LA, Palleron NJ, Haggblom MM. PAH-degradation by Paenibacillus spp. and description of Paenibacillus naphthalenovorans sp. nov., a naphthalene-degrading bacterium from the rhizosphere of salt marsh plants. Int J Syst Evol Microbiol. 2002;52(1):131-139.
Crossref - Bezza FA, Chirwa EMN. Biosurfactant from Paenibacillus dendritiformis and its application in assisting polycyclic aromatic hydrocarbon (PAH) and motor oil sludge removal from contaminated soil and sand media. Process Saf Environ Prot. 2015; 98:354-364.
Crossref - Oren A. Saltern evaporation ponds as model systems for the study of primary production processes under hypersaline conditions. Aquat Microb Ecol. 2009;56:193-204
Crossref - Le Borgne S, Paniagua D, Vazquez-Duhalt R. Biodegradation of organic pollutants by halophilic bacteria and archaea. J Mol Microbiol Biotechnol. 2008;15(2-3):74-92.
Crossref - Wimpenny J, Manz W, Szewzyk U. Heterogeneity in biofilms. FEMS Microbiol Rev. 2000;24(5):661-671.
Crossref - Al-Awadhi H, Al-Hasan RH, Sorkhoh NA, Salamah S, Radwan SS. Establishing oil-degrading biofilms on gravel particles and glass plates. Int Biodeterior Biodegradation. 2003;51(3):181-185.
Crossref - Martinez-Espinosa RM. Halophilic archaea as tools for bioremediation technologies. Appl Microbiol Biotechnol. 2024;108(1):401.
Crossref - de Fatima FRM, Junior PSS, Leite MS, et al. Integrated process of biosurfactant production by Bacillus atrophaeus ATCC-9372 using an air-lift bioreactor coupled to a foam fraction column. Fermentation. 2023;9(11):959.
Crossref - Meliani A, Densoltane A. Enhancement of hydrocarbons degradation by use of pseudomonas biosurfactants and biofilms. J Pet Environ Biotechnol. 2014;05(01).
Crossref - Zhang S, Zhang M, Han F, et al. Enhanced degradation of petroleum in saline soil by nitrogen stimulation and halophilic emulsifying bacteria Bacillus sp. Z-13. J Hazard Mater. 2023; 459:132102.
Crossref - Fatima K, Imran A, Amin I, Khan QM, Afzal M. Plant species affect colonization patterns and metabolic activity of associated endophytes during phytoremediation of crude oil-contaminated soil. Environ Sci Pollut Res Int. 2016;23(7):6188-6196.
Crossref - Ebadi A, Sima NAK, Olamaee M, Hashemi M, Nasrabadi RG. Effective bioremediation of a petroleum-polluted saline soil by a surfactant-producing Pseudomonas aeruginosaconsortium. J Adv Res. 2017;8(6):627-633.
Crossref - Cai P, Ning Z, Liu Y, He Z, Shi J, Niu M. Diagnosing bioremediation of crude oil-contaminated soil and related geochemical processes at the field scale through microbial community and functional genes. Ann Microbiol. 2020;70(1).
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.