ISSN: 0973-7510
E-ISSN: 2581-690X
Twister disease is a major disease of onion (Allium ascalonicum L.). The causal organisms of the anthracnose/twister disease of onion were characterized and identified using molecular markers to assess the diversity of the pathogens and the identity of the pathogen was confirmed by sequencing the amplified ITS rDNA region, and analysing in NCBI BLAST program and identified as C. gloeosporioides, C. acutatum, G. acutata , F. oxysporum and G. moniliformis. The phylogenetic tree showed that all five isolates of Colletotrichum spp. Framed into 3 clusters. Since, there is no gene sequences of C. gloeosporioides C. acutatum and F. oxysporum causing onion twister disease in the Genbank we have deposited our sequences Genbank. Specific DNA amplification of region CgInt was fairly consistent band of 450 bp for C. gloeosporioides and for C. acutatum it was 490 bp which was positive for all isolates of C. acutatum. For F. oxysporum amplification was at 550-570 bp. The products of the PCR ITS rDNA digestion with DraI revealed that no restriction sites were present. The digestion with HaeIII resulted in a characteristic pattern of three fragments in 19 C. gloeosporioides isolates. In 24 isolates of C. acutatum isolates and 25 isolates F. oxysporum showed four clusters.
Onion Twister disease, Molecular variability, Colletotrichum, Fusarium
Onion (Allium cepa L.) rightly called as queen of kitchen is one of the oldest important bulbous vegetable crops grown in India. It belongs to the family Alliaceae. However, in spite of the increase in areas planted, maximum production has not been attained. This is due to the proliferation of insect pests and diseases that continue to affect onion in almost all production areas in the country. Several factors have been identified for the low productivity of onion in India. The most important factors responsible are the diseases like purple blotch, downy mildew, Stemphylium blight, basal rot, storage rots and now recently twister disease.
In the recent years, twister disease has become epidemic on onion crop in coastal tract and other major onion growing districts in Karnataka. This disease vernacularly in Sri Lanka called as disco, in Indonesia as seven whorl and in Karnataka as haavu suruli roga, tiruguni roga, gone roga, octopus disease and creeping disease of onion. This disease causes heavy yield loss, leads to shortage in supply to the market resulting in higher prices to a common man. Presently, onion is extensively cultivated in all the parts of Karnataka. Both seed and bulb crops are infected with disease severity of 20-30 per cent and 50-70 per cent respectively1.
Several Colletotrichum species within which having diverse morphological, pathogenic and genetic diversities. Information on pathogen diversity and geographic distribution of the pathogen populations is therefore a prerequisite for designing effective disease management practices including accurate assessment of suitable resistant germplasm in breeding programs2,3,4. Hence, knowledge on the presence of different pathogenic isolates of Colletotrichum and Fusarium spp., the causal organisms study was to determine the morphological and molecular variations among isolates of Colletotrichum and Fusarium isolated from infected onions collected from different agroclimatic zones.
Due to environmental influences on the stability of morphological traits and the existence of intermediate forms, morphological criteria are not always adequate for reliable differentiation of Colletotrichum species. Similar limitations have been reported the identification of Fusarium species as well. The overlap of morphological characters and phenotypes among species makes the classification difficult. Molecular techniques provide alternative methods for taxonomic studies and are important tools in solving the problems of species delineation. The ribosomal RNA genes (rDNA) possess characteristics that are suitable for the detection of pathogens at specific level. These rDNA are highly stable and exhibit a mosaic of conserved and diverse regions within the genome. The objective of our investigation was to determine the morphological and molecular diversity of the causal agents of onion leaf twister disease.
Isolation DNA and Polymerase chain reaction
The genomic DNA was isolated from Coletotrichum spp. and Fusarium sp. isolates by following standard CTAB method with certain modifications5. The rDNA have been employed to analyze evolutionary events because it is highly conserved, where as ITS rDNA is more variable. Hence, it has been used for investigation of these species level relationships. Primer sequences used are given below.
Sequencing of ITS region
The ITS region was sequenced in five isolates belonging to five different geographical regions to confirm organism and to know the variability present in them.
Sequencing and in silico analysis
The PCR product was sequenced using forward and reverse primers at Chromos Biotech Ltd., Bangalore. Homology search was done using BLAST algorithm available at http://www.ncbi.nlm.nih.gov. Multiple alignments for homology search were performed using the Clustral W algorithm software and phylogenetic tree was constructed5.
Analysis of the genetic variability among the Fusarium sp, C. gloeosporioides and C. acutatum by by using PCR-RFLP
Aliquots of four ìl of the amplicons were subjected to enzymatic digestion with the restriction enzymes DraI, HaeIII and MspI (Invitrogen), according to the instructions. Agarose gel electrophoresis was performed to resolve the amplified product using 3.0 per cent agarose in 1X TBE (Tris Borate EDTA) buffer, 0.5 ??g ml-1 of ethidium bromide and loading buffer (0.25% Bromophenol Blue in 40% sucrose). Electrophoresis was carried at 65 V for 1.5 h. The gel was observed under UV light and documented using gel documentation unit. DNA fragments were visualised under UV light. The fragment sizes were estimated by a comparison with a 100 bp marker.
Amplification of ITS1 and ITS4 region
The full length ITS rDNA region was amplified primers for isolates of Fusarium sp. and Colletotrichum spp. DNA amplicon was observed at the region 560 bp with a concentration of around 437 ng/ µl. The amplified products were checked on 1.4 per cent Agarose gel electrophoresis. This is to be the first report of amplification of ITS rDNA region of Fusarium sp. and Colletotrichum spp. isolated from onion which is casual agent of twister disease of onion in India.
| Organism | Sequence | |
|---|---|---|
| Universal fungal ITS | ITS-1 – f | 5’-AACGTTACCAAACTGTTG-3’ |
| ITS-4 – r | 5’-AAGTTCAGCGGGTATTCCT-3’ | |
| C. gloeosporioides | CgInt -f | 5-´GGCCTCCCGCCTCCGGGCGG-3’ |
| ITS-4 – r | 5´- TCCTCCGCTTATTGATATGC-3’ | |
| C. acutatum | CaInt-2 – f | 5´- GGGGAAGCCTCTCGCGG -3’ |
| ITS-4 – r | 5-´TCCTCCGCTTATTGATATGC-3’ | |
| F. oxysporum | Fu1-f | 5-’ACAACTCATAACCCTGTGAACAT-3’ |
| Fu1-r | 5’-CAGAAGTTGGGTGTTTTACGG-3’ | |
PCR reaction mixture
Particular |
Quantity(μl) |
|---|---|
Template DNA (40 ng) |
1.0 |
Primers (5pmol)-F and R |
1.0 |
dNTP’s (2 mM) |
2.0 |
Taq buffer A (10X) |
2.0 |
TaqDNA polymerase (3U/ìl) |
0.3 |
Sterile water |
13.7 |
Total |
20.0 |
PCR condition for ITS, Fusarium sp, C. gloeosporioides and C. acutatum
| Universal ITS | Fusarium spp. | C. gloeosporioides | C. acutatum | |||||
|---|---|---|---|---|---|---|---|---|
| Step | Temp (°C) | Duration (min) | Temp (°C) | Duration (min) | Temp (°C) | Duration (min) | Temp (°C) | Duration (min) |
| Initial denaturation | 94 | 5 | 95 | 5 | 95 | 5 | 95 | 5 |
| Denaturation | 94 | 1 | 58 | 1 | 94 | 1 | 94 | 1 |
| Annealing | 54 | 1 | 94 | 1 | 62 | 1 | 62 | 1 |
| Extension | 72 | 2 | 72 | 2 | 7 | 1 | 72 | 1 |
| Final extension | 72 | 10 | 72 | 8 | 72 | 7 | 72 | 7 |
| Hold | 4 | 20 | 4 | 20 | 4 | 20 | 4 | 20 |
| No. of cycles | 30 | 40 | 25 | 25 | ||||
DNA sequencing
The DNA sequences were obtained for ITS rDNA. DNA sequences of selected isolates were compared using bioinformatics tool like NCBI (National Centre for Bioinformatics) blast programme. Based on sequence comparison, the identification of Fusarium sp and Colletotrichum spp. isolates were confirmed All the ITS rDNA sequences of isolates were confirmed as Colletotrichum gloeosporioides, Colletotrichum acutatum and Fusarium oxysporum. The list of isolates, accession number, per cent homology and name identified are given in Table 1.
Table (1):
Comparison and identity of isolates with gene bank in NCBI BLAST program
Sl. No |
Accession number |
NCBI BLAST Hit results |
Max. Ident. |
|---|---|---|---|
1 |
KC895535.1 |
Colletotrichum gloeosporioides isolate CJBB21-25 18S ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and 28S ribosomal RNA gene, partial sequence |
99% |
2 |
KC010540.1 |
Colletotrichum gloeosporioides isolate C8 18S ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and 28S ribosomal RNA gene, partial sequence |
100% |
3 |
KC493156.1 |
Colletotrichum gloeosporioides strain CR-5 18S ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and 28S ribosomal RNA gene, partial sequence |
93% |
4 |
GU183326.1 |
Glomerella acutata culture-collection BRIP:27048 internal transcribed spacer 1, partial sequence; 5.8S ribosomal RNA gene, complete sequence; and internal transcribed spacer 2, partial sequence |
100% |
5 |
AJ749694.1 |
Colletotrichum acutatum ITS1, 5.8S rRNA gene and ITS2, isolate PT227 |
94% |
6 |
EF158026.1 |
Gibberella moniliformis strain F3 internal transcribed spacer 1, partial sequence; 5.8S ribosomal RNA gene, complete sequence; and internal transcribed spacer 2, partial sequence |
99% |
7 |
JN232187.1 |
Fusarium oxysporum isolate 847 18S ribosomal RNA gene, partial sequence; internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence; and 28S ribosomal RNA gene, partial sequenc |
94% |
8 |
KC351189.1 |
Fusarium oxysporum 18S ribosomal RNA gene, partial sequence |
97% |
9 |
KC577176.1 |
Fusarium oxysporum isolate FoxySIN1 internal transcribed spacer 1, partial sequence; 5.8S ribosomal RNA gene and internal transcribed spacer 2, complete sequence; and 28Sribosomal RNA gene, partial sequence |
100% |
10 |
JQ929907.1 |
Uncultured Fusarium clone f103 internal transcribed spacer 1, partial sequence; 5.8S ribosomal RNA and internal transcribed spacer 2, complete sequence; and 28S ribosomal RNA, partial sequence |
100% |
Phylogenetic Analysis
The ITS rDNA region sequence was used in these analysis because it has been shown to be more informative and closest phylogenetic relative in Colletotrichum gloeosporioides, Colletotrichum acutatum and Fusarium oxysporum. In order to evaluate whether the grouping pattern obtained on the basis of the ITS -sequences of the ex-type strains would be useful frame to identify and align, these isolates were identified at the species level by morphological characters using the existing taxonomic criteria analysis and analysis of their ITS rDNA region gene sequences.
In these the isolates identified as C. gloeosporioides, C. acutatum and F. oxysporum are markedly similar to the reference strain of NCBI BLAST Genbank. This information provided by these studies gives more useful tips for assessing the taxonomy. In present case, isolate of C. gloeosporioides, C. acutatum and Fusarium oxysporum, was used as an out group vice versa to interpret the clustering of isolates as distinct or related out group of genus. Phylogenetic analysis of ITS sequences showed three clusters in Colletotrichum spp. In cluster-I (Dharwad, Chitradurga), cluster-II (Uttara Kannada) and Cluster-III (Chikamagaluru, Haveri). In Fusarium spp. there are three clusters in cluster-I (Chikamagaluru and Dharwad,), cluster-II (Uttara Kannada) and Cluster-III (Chitradurga, Haveri) (Fig.1).
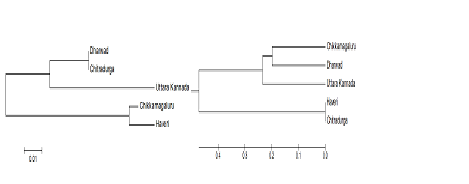
Fig. 1. Phylogram of selected isolates of Colletotrichum spp. and Fusarium spp. of ITS region sequence analysis
Since, there is less number of gene sequences of C. gloeosporioides C. acutatum and F. oxysporum infecting the onion for twister disease in the Genbank, deposited our sequences and the accession numbers are KF264963.1and HQ700357.1. It is first information in sequencing ITS rDNA region and molecular variability of pathogens causing the onion twister disease in India.
Molecular characterization of Fusarium isolates from wilted onion by analysis of the ITS regions of ribosomal DNA. The 32 isolates were grouped into eight types (four types for F. oxysporum, one for F. verticillioides and three for F. solani). The ITS region of non-pathogenic Fusarium strain of tomato were amplified. By using the NCBI BLAST program, they confirmed these isolates as Fusarium species. Some of the closely related strains are endophytic in nature (7). The pathogenic isolated showed 100% similarity to the standard isolates F. oxysporum. f. sp. lycoparsici . Based on the comparison of DNA sequence to GenBank (an open access sequence database produced by NCBI Information), the unidentified fungal species showed high similarity with G. moniliformis (synonym G. fujikoroi) the causal organisms of bakanae disease of rice causing abnormally long and weak internodes and ear and stalk rot to maize. Thus, it was inferred that the unidentified fungal species is G. moniliformis it causes the elongation and twisting symptoms of onions. In the present investigation, emphasis on ITS for amplification and obtained partial sequences to the same but still there is need to go for terminal elongation factor (TEF).
Specific gene amplification and restriction analysis
After ITS rDNA region identification of pathogens C. gloeosporioides and C. acutatum further studies were carried out for specific DNA amplification of region CgInt and it was fairly consistent band of 450 bp for C. gloeosporioides which was positive for isolates. Whereas C. acutatum at 490 bp positive for all isolates. ITS region amplified from a selection of different Fusarium species were at 550-570 bp. Diagnosis of pathogens by species specific primers helped us for quick detection and rapid identification.
The products of the PCR ITS rDNA digestion with DraI revealed that no restriction sites were present. The digestion with HaeIII resulted in a characteristic pattern of three fragments. C. gloeosporides isolates shown four clusters, in cluster-I (1,8,17,18), cluster – II (2,4,5,6,16,14,12,13,7), cluster – III (3,9,1510,11) and cluster – IV (19) . C. acutatum isolates shown six clusters, in cluster-I (1,5,7,24,23,20,19), cluster – II (2,6,11,12,14,18,17), cluster – III (13), cluster – IV (4,10), cluster – V (3,8, 16,9) and cluster – VI (15,21,22). F. oxysporum isolates shown five clusters, in cluster-I (1,4,6,24,23,20,19), cluster – II (2,5,9,11,22,12,18,17,13,24), cluster – III (3,7) cluster – IV (8,20) and cluster – V (15,25,21,16)(Fig.2).
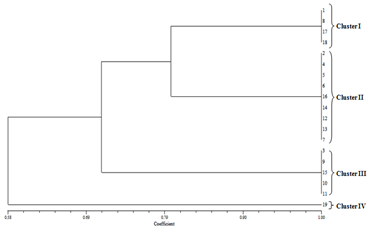
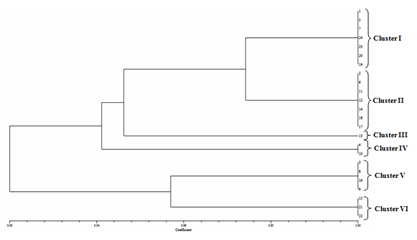
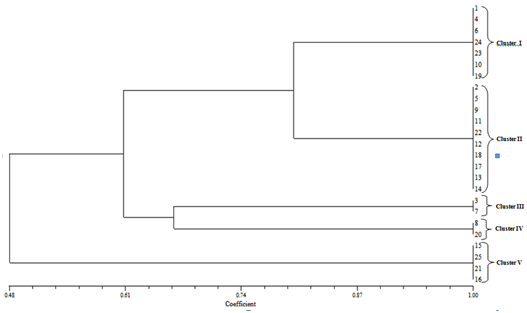
Fig. 2. Relationship between isolates using different molecular PCR-RFLP markers through UPGMA analysis
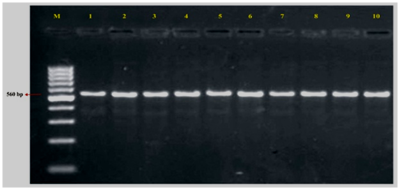
Plate 1. Amplification of ITS1 and 4 region of representative Colletotrichum spp. and Fusarium spp.
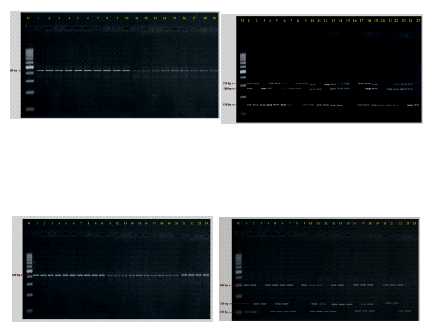
Plate 2. Specific amplification and PCR-RFLP pattern C. gloeosporioides (CgInt), C. acutatum (CgInt) and F. oxysporum (Fu-ITS)
- www. govya. lk/agri_learning/red_onionrearch_red/rog_p/I. pdf accesed on 12thMay 2011
- Guerber, J.C., Liu, B., Correll, J.C. and Johnston, P.R.. Characterization of diversity in Colletotrichum acutatum sensu lato by sequence analysis of two gene introns, mtDNA and intron RFLPs, and mating compatibility. Mycologia, 2003; 95: 872-895.
- Rao, V.P., Thakur, R.P. and Mathur, K. Morphological and pathogenic diversity among grain sorghum isolates of C. gramanicola in India. Indian Phytopathol, 1998; 51(2):164-174.
- Abang, M.M.. Genetic diversity of Colletotrichum gloeosporioides Penz. causing anthracnose disease of yam (Dioscorea spp.) in Nigeria. Bibliotheca Mycologia, 2003;197:20–33.
- Patil, S.,. Effect of non-pathogenic Fusarium spp. in bio- control of Fusarium wilt of tomato. M. Sc. (Agari.) Thesis, University of Agricultural Sciences, Bangalore.2009
- Dissanayake, M. L. M. C., Kashima, R., Tanaka, S. and Ito, S., Pathogenic variation and molecular characterization of Fusarium species isolated from wilted Welsh onion [Allium fistulosum] in Japan. J. Gen. Pl. Path. (Japan). 2009 ; 75(1) 37-45.
© The Author(s) 2016. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


