ISSN: 0973-7510
E-ISSN: 2581-690X
The aim of the present study was to identify and recognize the genotype for cystic Echinococcosis that infects goats (n=19) in Iraq. The hydatid cyst was collected from different parts of the body, i.e. lungs, liver, heart, spleen and peritoneal cavity. The current study was conducted in five different regions of Iraq (Anbar, Baghdad, Saladdin, Karkuk, Babylon during October 2018 to July 2019. The mitochondria DNA was extracted and screened for the presence of (CO1) cytochrome C oxidase subunit 1 gene using polymer chain reactions (PCR). Amplification size was around 450bp. The amplicon was isolated and purified. The isolated CO1 amplicon was sequenced. The CO1 putative sequence was BLAST with available sequence from NCBI. The genetic tree was deduced. The genotype G1 is the most frequently spread strains and considered as a key source for infection in goats of Iraq.
Mitochondria DNA, Cytochrome C oxidase, Genotype, Goats
E. granulosus causes Hydatid Cyst disease in goats.1 The prevalence of disease spreading is significantly high worldwide. Hence, it is also considered as one of the significant zoonotic diseases.1,2 The goats can be infected by consumption of infected water and food.3 The contamination can be spread through infected dogs litter containing eggs of worms.3
The mutations in the E. granulosus strains can affect the characteristics, life circle, evolution rate of disease, drug sensitivity, drugs evolution against this disease in Iraq.4 Cystic Echinococcosis is considered as one of the most significant pandemic diseases that have an important hazard for human beings and finally to animal health.5-9 Recently, about 10 genotypes (range from G1 to G10) have been diagnosed internationally depending on nucleotide sequence using genes for ND1 and CO1. This disease was found to be infect various animals such as camels, pigs, sheep, cattle, horses, cervids, and goats.10-15
The genotype G1 reported to infect human beings as well as sheep and cattle.16 For genotype recognisation, number of marker genes and various advanced techniques are commonly used in many countries.17 The strain causing hydatid cyst disease in Iraqi Goats has been identified based on the ND1 gene sequences. Similar gene can be used to detect the G1 strain in goat.19 The aim of the current study was to identify and recognize the cystic echinococcosis genotype that infects goats in Iraq. The PCR technique was used to recognize the minor strains which depending on CO1 gene amplification.
Sample Collection
The current study was conducted in five different regions of Iraq. Anbar, Baghdad, Saladdin, Karkuk, Babylon during October 2018 to July 2019. The samples (n=19) were obtained through custom massacres and from veterinarians, who contributed to goats infected diagnosis with Hydatid cyst where the samples were collected by me in the field.
Sample Preparation and Protoscoleces Isolation
The isolated cyst was washed twice with normal saline to remove contamination. Further, the cyst was washed with 70% ethanol as per the protocol described by Mc-Manus and Smyth.20 Each Hydatid Cyst was divided into two regions, viz. Outer cover and internal liquid containing protoscoleces. The protoscoleces were extracted using 10 ml sterile syringes into new sterile containers. The Hydatid Cyst was opened vertically and the liquids were separated. The protoscoleces were collected in a new sterilized container. The liquids was centrifuged at speed (3000) revolutions per minute (rpm), room temperature, for ten minutes. The whole protoscoleces pellet was collected. For isolation of germinal membrane, the procedure described by the Rishi and McManus21 was followed. The germinal membrane was washed twice by using (pH 2) Hanks Saline which contains 0.2% Pepsin(W/V). Followed by centrifugation at speed (3000) revolutions per minute rpm RT for ten minutes. The remaining protoscoleces were collected as a pallet. The pallet was washed 3 times with normal saline and centrifuged to decant the liquid. The protoscoleces pallet was stored by using (70%) ethanol at 4°C for further processing.
DNA Extraction from Protoscoleces
The stored protoscoleces was washed with phosphate buffer saline to remove ethanol. The samples were subjected for the DNA extraction21 using a Wizard purification DNA kit (USA).22,23 Briefly, around 20 ng sample were used for the DNA extraction. The extracted DNA were evaluated for the presence of mitochondrial cytochrome C oxidase subunit 1 (CO1) gene. Amplified CO1 gene by using CO1 specific primers according to the procedure of PCR24 (Table 1).
Table (1):
The primers of (CO1) gene.
Sequence of primer |
Sense |
Size |
|---|---|---|
5’ –TTT_TTT_GGG_CAT_CCT_GAG _GTT_TAT- 3’ |
Forwards |
450 bp |
5’-TAA_AGA_AAG_AAC_ATA_ATG_AAA_ATG-3’ |
Reverses |
450 bp |
The PCR condition was as follows denaturation temperature at 94°C for 5 minutes then followed by 35 cycles of three steps. These three steps were denaturation at a temperature 94°C for 45 seconds, followed annealing step at temperature 58°C for 45 seconds and finally extension step at temperature 72°C for 45 seconds. These steps were followed by a final step for extension at temperature 72°C as extra time for 7 minutes.
In silico analysis of CO1 gene
After CO1 gene amplification, the applicant was obtained and proceeded for extraction from the gel. The extracted amplicon was sequenced and BLAST with the reference CO1 sequences available in NCBI (www.ncbinlm.n.h.gov)
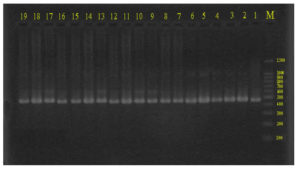
The extracted DNA from 19 Hydatid Cyst samples were screened for the amplification of the CO1 gene using PCR and CO1 specific primer. Fig. 1 showed CO1 gene amplification on agarose gel. Around 450bp amplicon was observed on the gel.
Fig. 1. PCR amplification of Co1 gene. Electrophoresed on 2 % agarose gel (80 V, 70 Amp) (M, 100 base pair, DNA ladder, lanes 19_1, E. granulosus isolates).
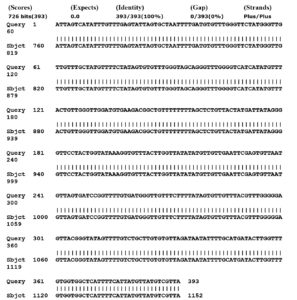
The pieces of CO1 gene were amplified. The amplified amplicon was extracted and sequenced. The putative CO1 sequence was compared with the available CO1 sequences on NCBI. This putative sequence was corresponding with the Genotype G1 the main causative E. granulosus (Fig. 2).
Fig. 2. Mitochondrial cytochrome C oxidase subunit 1 gene alignment of Echinococcus granulosus, by utilizing Gene bank.
In the present study, the registered sequences in Gene Bank have been accomplished by using Bioedit program which is considered as one of the dependent programs in analyzing the DNA. The isolated CO1 gene sequence showed 100% matching with the sheep strain G1 genotype (Accession no MN787561).25
In the present study, mitochondrial cytochrome C oxidase subunit 1 gene was amplified using available primers. These primers were found to be specific for the goat CO1 gene as all 19 samples showed positive results in PCR. The amplicon size was 450bp. Similar CO1 amplicon size was reported by various authors.16,23,24 The genetic type G1 and particular sheep strain were affected by E. granulosus. This infection was proving to be a local host. It is worth to mention that the examined places are close and everlasting availability in the pastoral with these animals. This closeness and food sharing lead to increase the infection possibility of Hydatid Cyst disease.15 This can be one of the major reasons behind the prevalence of infection.
It is worth referring that the remarkable spread of sheep strain G1 was common in the extreme inhabitance regions in Iraq. These results are in accordance with previous studies.15, 18, 24, 26,27 It is the main reason of being such strain as more frequent and spread in the intermediate hosts for these reasons.16,28 E. granolosus strain detection in domestic animals as well as in wild animal in this region will be epidemiologically significant. The genotype G1 is also considered as a disease that infects the other intermediate hosts such as goats.29,30 From the aforementioned, the genotype G1 is considered as the most frequent type and the most effective strain in the Iraqi goats. In the present study, all 19 samples showed presence of genotype G1 i.e. CO1 gene, is a particular for sheep strain.
ACKNOWLEDGMENTS
I would like to express my heartfelt thanks to Dr. Ayad Hammood for English editing of the manuscript and my heartfelt thanks to Veterinarian (Ass. Professor Dr. Mohammed A, Hammed) for helping me diagnose the samples.
FUNDING
None.
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript
- Thompson RC. The taxonomy, phylogeny and transmission of Echinococcus. Exp Parasitol. 2008 ;119(4): 439-446.
Crossref - Altintas N, M Oztatlici, Unver A, Sakarya A. Molecular analysis of cattle isolates of Echinococcus granulosus in manisa province of Turkey. Kafkas Univ Vet Fak Derg. 2013;19(3): 455-459.
- Fasihi HM, Hobbs RP, Adams PJ, Mobedi I., Morgan-Ryan UM, Thompson RCA. Molecular and morphological characterization of Echinococcus granulosus of human and animal origin in Iran. Parasitolo. 2002; 125(4): 367- 373.
Crossref - McManus DP, Thompson RCA. Molecular epidemiology of cystic echinococcosis. Parasitology. 2003;127(Suppl):S37-S51.
Crossref - Casulli, A. Recognising the substantial burden of neglected pandemics cystic and alveolar echinococcosis. Lancet Glob. Health. 2020;8: e470–e471.
Crossref - Piseddu T, Brundu D, Stegel G, et al. The disease burden of human cystic echinococcosis based on HDRs from 2001 to 2014 in Italy. PLoS Negl. Trop. Dis. 2017;11: e0005771.
Crossref - Khan A, Naz K, Ahmed H, et al. Knowledge, attitudes and practices related to cystic echinococcosis endemicity in Pakistan. Infect Dis Poverty. 2018;7:4.
Crossref - Ehsan M, Akhter N, Bhutto B, Arijo A, Gadahi JA. Prevalence and genotypic characterization of bovine Echinococcus granulosus isolates by using cytochrome oxidase 1 (Co1) gene in Hyderabad, Pakistan. Vet Parasitol. 2017;239:80-5.
Crossref - Khan A, Ahmed H, Simsek S. Molecular epidemiology of Echinococcus species in Pakistan. Asian Pacific J Trop Med. 2018;13:36.
Crossref - Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol and Evol. 2018;35:1547-1549.
Crossref - Romig T, A. Dinkel. 2006. Mackenstedt U. The present situation of echinococcosis in Europe. Parasitol. Int. 55: 187-191.
Crossref - Varcasia A, Canu S, Kogkos A, Pipia AP, Scala A, Garippa G, Seimenis A. Molecular characterization of cystic echinococcosis in small ruminants in Peloponnesus, Greece. Parasitol. Res. 2007; 101(4):1135-1139.
Crossref - Busi M, Snabel V, Vercasia A, Garippa G, Perrone V, De-Liberato C, D’Amelio S. Genetic variation within and between G1 and G3 genotypes of Echinococcus granulosus in Italy revealed by multilocus DNA sequencing. Vet. Parasitol. 2007; 150(1-2):75-83.
Crossref - Nakao M, Manus DPMc, PM Schantz, PS Craig, Ito A. A molecular phylogeny of the genus Echinococcus inferred from complete mitochondrial genomes. Parasitol. 2006; 134(5): 713-722.
Crossref - Sanchez E, Caceres O, Naguira C, et al. Molecular characterization of Echinococcus granulosus from Peru by sequencing of the mitochondrial cytochrome C oxidase subunit 1 gene. Oswaldo Cruz, Rio de Janeiro. 2010;105(6): 221-232.
Crossref - Sánchez E, Cáceres O, Náquir AC, Miranda E, Samudio F, Fernandes O. Echinococcus granulosus genotypes circulating in alpacas (Lama pacos) and pigs (Sus scrofa) from an endemic region in Peru. Mem Inst Oswaldo Cruz, Rio de Janeiro. 2012 ;107(2):275-278.
Crossref - Eryıldız C , N Şakru. Molecular Characterization of Human and Animal Isolates of Echinococcus granulosus in the Thrace Region, Turkey. Balkan Med J. 2012;29: 261-267.
Crossref - Muhaidi MJ, Omar H, Ahmed MN, Abdul-Rahman GM. Genetic Polymorphism in ND1 Gene of Hydatid Cyst in Iraqi goats. Research Jour. of Biotech. 2019;14 (Special Issue I).
- McManus DP, SD Smyth. Differences in the chemical composition and carbondrate metabolisim of Echinococcus granulosus (horse and sheep strains) and E. multilocularis. Parasitol. 1978 ;77(1):103-109.
Crossref - Rishi AK, D McManus. Genomic cloning of human Echinococcus granulosus DNA: isolation of recombinant plasmids and their use as genetic markers in strain characterization. Parasitol. 1987;94(2):369-383.
Crossref - Al-Azawiy AK. Immunization of Experimental Mice by DNA of Protoscoleces of Hydatid Cyst: Immunological and Histopathological Studies. Ph.D. Dissertation. College of Veterinary Medicine, University of Baghdad. Iraq. PP:5-15.
- Miller SA, DD Dykes, HF Polesky. A simple salting out procedure for extracting DNA from human nucleated cells. Nucl. Acids Res. 1988; 16(3): 1215.
Crossref - Ergin S, S Saribas, P Yuksel. Genotypic characterisation of Echinococcus granulosus isolated from human in Turkey. Afr. J. Microbiol. Res. 2010;4(7):551-555.
- Baraak MJ. Molecular Study on Cystic Echinococcosis in Some Iraqi Patients. Ph.D. Dissertation, College of Science, University of Baghdad. Iraq. 2014;10-19.
- Laurimae T, L Kinkar, V Andresiuk, et al. Genetic diversity and phylogeography of highly zoonotic Echinococcus granulosus genotype G1 in the Americas(Argentina, Brazil, Chile and Mexico) based on 8279bp of mtDNA. Infect. Genet. Evol. 2016;45: 290-296.
Crossref - Craig PS, MT Rogan. Campos-Ponce M. Echinococcosis: disease, detection and transmission. Parasitol.2003 ; 127: 5-20.
Crossref - Muhaidi MJ. Determination Of The Infective Strain Of Hydatid Cyst In Iraqi Cattle By Using Co1 Gene. The Iraqi Jour. of Agricul.l Scien. 2017; 466-466: 48(2).
Crossref - Rinaldi L, Maurelli MP, Capuano F, Perugini AG, Veneziano V, Cringoli S. “Molecular update and cystic echinococcosis in cattle and water buffaloes of southern Italy”, J. Blackwell verlag. Zoonosis, Pub. Heal. 2008;55(2):119-123.
Crossref - Eckert J, RC Thompson. Intraspesific variation of E.granulosus and related species with emphasis on their infectivity to humans. Acta. Trop. 1997; 64(1-2): 19-34.
Crossref - Mwambete KD, Ponce-Gordo F, Cuesta-Bandera C. Genetic identification and host range of the Spanish strains of Echinococcus granulosus. Acta. Trop. 2004 ; 91(2): 87- 93.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.