ISSN: 0973-7510
E-ISSN: 2581-690X
Nine lactic acid bacteria (LAB) were isolated from food, fruits and vegetables (beet root, dosa batter, mango, banana, pine apple, radish, tomato, milk and watermelon) on de Man Rogosa and Sharpe (MRS) agar medium and characterized as LAB based on their morphological and biochemical characters. These bacteria were further identified into species by 16S rRNA gene sequence as L. fermentum (3 isolates), L. buchneri (4 isolates), L. brevis, (1 isolate) and Weisella ciberia (1 isolate). The genetic diversity of Lactobacillus species were analyzed using Random Amplified Polymorphic DNA (RAPD). Based on RAPD analysis, the nine Lactobacillus species were grouped into two clusters. The LAB-1 (L. buchneri), LAB-5 (L. brevis), LAB-6 (L. buchneri), LAB-7 (L. buchneri) LAB-8 (L. buchneri), LAB-9 (Weisella ciberia) formed cluster-I and LAB-2 (L. fermentum), LAB-3 (L. fermentum), LAB-4 (L. fermentum), formed the cluster-II. Further, milk coagulation study showed that L. brevis, two strains of L. buchneri (LAB-6 and -7) and Weisella ciberia coagulated milk at 12 h of incubation.
Lactobacillus species, Bacteria, LAB
Lactic Acid Bacteria (LAB) comprises a wide range of genera including a considerable number of species. They are gram positive, micro aerophillic and facultative anaerobes, usually do not form endospores and produce lactic acid as a major end product of carbohydrate fermentation. The genera of these bacteria are Lactobacillus, Pediococcus, Lactococcus, Leuconostoc, Streptococcus etc., The genus Lactobacillus is the largest group among the Lactobacteriaceae and contains over 110 species (Dellaglio et al., 2005; Satokari et al., 2003). These bacteria can be found in/on plants, animals, food stuff niches, and also in gastrointestinal tract of human beings. LAB’s are ‘Generally Recognized as Safe’ (GRAS) and have been used as starter culture in many food industries due to their ability to produce desirable taste and flavour (Fathabad et al., 2011). In addition, LAB are known to produce antimicrobial compound known as ‘bacteriocins’ which are considered as natural bio preservatives to control food borne pathogens (Paul et al., 2004). Several Lactobacillus strains have been isolated from different sources and used as probiotics in commercial food products (Patil et al., 2009; Ashraf et al., 2009; Mallesha et al., 2010; Fathabad et al., 2011). Lactobacillus species can be selectively isolated using de Man Rogosa Sharpe (MRS) medium (De Man et al., 1960). Identification of Lactobacillus species can be done by morphological and biochemical characters. However, the molecular technique (16S rRNA gene sequence) has been found more useful and supportive for identification of microorganisms. Further, molecular markers are the powerful tools to assess genetic diversity between the species. Among the different markers used to assess the diversity, the random amplified polymorphic DNA (RAPD) method is widely used to ascertain the dominance of strains. In this study, we isolated nine Lactobacillus species from food, fruits and vegetable sources and identified by using 16S rRNA gene sequence. Genetic diversity of the species were analysed by using RAPD markers.
Sample collection and isolation
Fruits and vegetables were collected from the local market, milk from Karnataka Milk Federation (KMF), Bengaluru and dosa batter from the food canteen of University of Agricultural Sciences, Bengaluru. Ten grams of each sample was dissolved in sterile water and LAB were isolated by dilution plate method using MRS agar medium which is a selective medium for LAB. The plates were incubated at 37°C for 7 days. The colonies grown were subcultured on MRS agar medium and purified by repeated streaking on the same agar medium dispensed plates and preserved under refrigerator.
Milk fermentation test
Raw milk obtained from the KMF, Bengaluru was dispensed in 5 ml test tubes and sterilized by autoclaving at 121°C, 15 lb/sq.inch for 15 minutes. To the sterilized milk, individual isolates were inoculated and incubated at 37°C for 38 hours. Observation for coagulation of milk was recorded at 12, 24 and 38 h intervals and the change in pH was recorded at 38 h of incubation.
Molecular identification
Total genomic DNA from the bacterial cells was extracted by following the protocol established by Sambrook et al. (1989). The bacteria were grown in 5 ml MRS broth at 37°C for 48 h and the cells were pelleted by centrifugation at 8,000 rpm for 3 minutes. The pellet was resuspended in extraction buffer (1M Tris HCL pH 8.0, 0.5M EDTA and 5M NaCl) and incubated at 60°C for 30 minutes. To the extract, 100µl 3M Potassium acetate solution was added and placed on ice for 10 minutes. Clear supernatant was collected by centrifugation at 10,000 rpm for 10 min and the DNA was precipitated using 0.6 volumes of ice cold Isopropanol. The DNA pellet was collected by centrifugation at 6,000 rpm for 15 min and washed with 70% ethanol, dried, dissolved in 10mM Tris EDTA buffer and stored in aliquots at -20°C.
PCR amplification of genomic DNA by 16SrRNA primers
A 22 bp of FGPS6 forward primer 5’-GGA GAG TTA GAT CTT GGC TCA G-3’and 20 bp of FGPS1509 reverse primer 5’-AAG GAG GGG ATC CAG CCG CA-3’was used for the present study. The annealing temperature for primer pair was standardized and PCR was performed in a 40µl reaction volume containing 27.4µl of sterile distilled water, 4µl of 10X buffer with MgCl2 (1.5mM), 4µl of dNTPs (10mM), 2µl of forward and reverse primers (0.5mM each), 0.6µl of Taq DNA polymerase (3U/µl Genei, Bengaluru, India) and 2µl of template DNA (50ng). Amplification was carried out with an initial denaturation at 96°C for 3 min followed by 35 cycles consisting of 94°C for 1 min, 50°C for 30 sec, 72°C for 1 min and a final extension step at 72°C for 10 minutes. The PCR products were electrophoresed on agarose gel (1.0%, Sigma, USA) and documented using gel documentation unit (HeroLab, Germany).
The PCR product was eluted from the gel using Gene JETTM Gel Extraction Kit [MBI, Fermentas Life Sciences, USA (#K0691)] and the eluted product was cloned into pTZ57R/T cloning vector using InsT/A clone PCR product cloning kit [MBI, Fermentas Life Sciences, USA (#K1214)] after determining the appropriate vector: insert ratios (Sambrook et al.,1989). The ligation reaction was performed in a 10µl reaction volume at 16°C overnight. The ligated product was used to transform competent E.coli (DH5±) cells using heat shock method (Sambrook et al., 1989) and plated on Luria Berton (LB) agar medium containing antibiotic (ampicillin, 100 µg/ml). The recombinant clones were initially screened by blue white selection, followed by colony PCR using M13 forward and reverse primers (Sambrook et al., 1989). The transformed colony was inoculated in 3ml LB broth containing ampicillin and incubated overnight at 37°C. The cells were harvested by centrifugation and the recombinant plasmid was isolated using GenEluteTMHP Plasmid MiniPrep Kit (Sigma, USA) following the manufactures protocol. The isolated plasmid was sequenced at Sci Genome Labs Private Ltd. Kerala, INDIA using M13 forward and reverses primers.
Sequence analysis and homology search
Sequence results were analysed with Vec Screen online software from NCBI (http://www.ncbi.nlm.nih.gov/BLAST) (Altschul et al., 1990) for removing the vector contamination. Forward and reverse primer sequences were checked against each other by generating the reverse complement of the “reverse” sequence using Fast PCR Professional (Experimental test version 5. 0. 83) and aligning it with the “forward” sequence with the help of CLUSTAL W Multiple Sequence Alignment Programme using the online software SDSC Biology Workbench (San Diego Supercomputer Center). The full length gene homology search was performed with blast programme of National Centre for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/BLAST).
Analysis of genetic diversity by RAPD
The preliminary screening was carried out with 20 decamer primers. Each RAPD-PCR was performed in a total volume of 25µl reaction mixture composed of 2.5µl 10X PCR buffer with MgCl2 (1.5mM), 2µl dNTPs (10mM), 3µl random primer (10mM), 0.5µl Taq DNA polymerase (3U/µl Genei, Bengaluru, India) and 2µl (40ng) template DNA and 15µl sterile distilled water. The PCR conditions were standardised with initial denaturation at 96°C for 3 min, followed by 35 cycles consisting of 94°C for 1 min, 37°C for 1 min, 72°C for 1 min and a final extension at 72°C for 10 minutes. The PCR mixture was separated on agarose gel (1.0%, Sigma, USA) stained with ethidium bromide (EtBr) and photographs were obtained with gel documentation unit (HeroLab, Germany).
Data analysis
Amplified products were scored as 1 and 0 for the presence and absence of bands. Pair wise comparisons based on both unique and shared amplification products were employed to calculate genetic diversity. The data was subsequently analysed for polymorphism using STATISTICA version 7.0 Software (Stat Soft, Tulsa, OK, USA).
Isolation and identification of LABs from different sources
A total of 9 LABs were isolated from banana, beet root, dosa batter, mango, milk, pine apple, radish, tomato and watermelon by serial dilution method. One isolate per sample was obtained and all were primarily identified on the basis of colony/cell morphology, staining and milk coagulation test. All the isolates formed characteristic whitish-creamy round colonies on MRS agar medium and were Gram-positive. These strains were able to grow under anaerobic conditions suggested the relatedness of these species with lactic acid bacteria and hence named as lactic acid bacteria 1 to 9 (LAB-1 to LAB-9). The cell morphology of all isolates was observed under microscope and they are rod shaped with round and blunt ends. The LAB-2, LAB-3, LAB-4 and LAB-9 were round ends and LAB-1, LAB-5, LAB-6 and LAB-8 were shown to be blunt ends (Table 1). LABs are bacteria that have features of rod or coccus shapes, with negative catalase, homo fermentative or hetero fermentative and growing in low acid conditions (Holzapfel et al., 2001). These belong to a group of gram positive bacteria that produce lactic acid as their main fermentation product and generally recognized as safe (Konings et al., 2000). These are one of the microorganisms that dominate fermented foods (Guasch et al., 2005; Robert, 2008).
Table (1):
Morphological characters of Lactic acid bacteria isolated from different edible sources
SL. No. |
Source |
Isolate |
Colony colour |
Colony shape |
Gram’s reaction |
Cell shape |
|---|---|---|---|---|---|---|
1 |
Banana |
LAB-1 |
Creamy white |
Round |
Gram Positive |
Rods with round ends |
2 |
Beetroot |
LAB-2 |
Creamy white |
Round |
Gram Positive |
Rods |
3 |
Dosa batter |
LAB-3 |
Creamy white |
Round |
Gram Positive |
Rods |
4 |
Mango |
LAB-4 |
Creamy white |
Round |
Gram Positive |
Rods |
5 |
Milk |
LAB-5 |
Creamy white |
Round |
Gram Positive |
Rods with round ends |
6 |
Pine |
LAB-6 |
Creamy white |
Round |
Gram Positive |
Rods with round ends |
7 |
Radish |
LAB-7 |
Creamy white |
Round |
Gram Positive |
Rods with round ends |
8 |
Tomato |
LAB-8 |
Creamy white |
Round |
Gram Positive |
Rods with round ends |
9 |
Watermelon |
LAB-9 |
Creamy white |
Round |
Gram Positive |
Rods |
Note: LAB = Lactic acid bacteria
Milk coagulation test
The milk coagulation test showed all the isolates have the ability to coagulate milk at 36 h of incubation at a temperature of 37°C. There was significant difference observed in the milk coagulation capacity of different isolates. Four of the 9 isolates LAB-4, LAB-5, LAB-6, and LAB-8 took only 12 h to coagulate the milk (Fig. 1) whereas; LAB-2, LAB-3 and LAB-7 took 24 h to coagulate the milk. Of the 9 isolates, LAB-1 and LAB-9 took the longest time 36 h to coagulate the milk that indicating their slow growth. The pH is an important factor which indicates the fermentation of milk and producing lactic acid (Hoque et al., 2010). In the present study, we observed reduction of pH from 3.6 to 5.66 (Table 2). A lower pH of 3.94, 3.6, 3.98 and 3.94 were recorded for LAB-4, LAB-5, LAB-6 and LAB-8 isolates, respectively that coagulated milk at 12 h of incubation (Table 2). A comparatively higher pH was observed for LAB-1, LAB-2, LAB-3, LAB-7 and LAB-9 isolates that indicated the lactic acid production is less compared to LAB-4, LAB-5, LAB-6 and LAB-8 isolates. Among the 9 Lactobacillus isolates, the L. brevis isolated from milk with a pH of 3.60 was found to be efficient in milk fermentation.

Fig. 1. Coagulated milk inoculated with LAB-4, LAB-5, LAB-6 and LAB-8 at 12 h of incubation
Table (2):
Coagulation of milk by Lactobacillus species at different time intervals
| Sl.No | Source | Isolate | Time taken for coagulation | pH after coagulation | ||
|---|---|---|---|---|---|---|
| 12 hours | 24 hours | 36 hours | ||||
| 1 | Blank | – | – | – | – | 6.48 |
| 2 | Banana | LAB-1 | – | – | + | 5.24 |
| 3 | Beetroot | LAB-2 | – | + | + | 4.8 |
| 4 | Dosa batter | LAB-3 | – | + | + | 4.68 |
| 5 | Mango | LAB-4 | + | + | + | 3.94 |
| 6 | Milk | LAB-5 | + | + | + | 3.60 |
| 7 | Pine apple | LAB-6 | + | + | + | 3.98 |
| 8 | Radish | LAB-7 | – | + | + | 4.20 |
| 9 | Tomato | LAB-8 | + | + | + | 3.94 |
| 10 | Watermelon | LAB-9 | – | – | + | 5.65 |
NOTE: + = Positive for coagulation – = Negative for coagulation
Molecular characterization using 16S rRNA gene sequence
The comparison of gene encoding the 16S ribosomal RNA (16S rRNA) are excellent phylogenetic markers for genus and species level identification and their sequences can generate vast information about the composition of complex bacterial system (Naeem et al., 2012). A considerable part of the 16S rRNA gene is conserved in all bacterial genera, whereas a smaller part is variable and this enable us to estimate genealogical distances, from which phylogenies are derived (Sabin et al., 2012). In the present study, 9 isolates were identified up to species level using a 1500 bp PCR amplified product of 16S rRNA gene sequence. All the 9 LAB isolates identified to the species level in this study showed 99% sequence homology with the existing Lactobacillus species sequences reported in the NCBI, expect the LAB-9 that showed 99% sequence homology with Wiesella ciberia species (Table 3). One of the representative picture showing the PCR amplification and Blastn analysis is shown in Figure 2. Ali et al. (2012) isolated lactic acid bacteria from a fresh plant polygonum minus (kesum) and identified as Lactococcus lactis, Pediococcus pentosaceus and Lactobacillus curvatus using 16S rRNA amplification with 99% homology. Fathima et al. (2012) characterized 7 Xanthomonas strains comprising 4 species using the 16S rRNA gene amplification to confirm the identity of strains. Similarly, 16S rRNA gene sequence comparison was employed by Naeem et al. (2012) to identify 15 bacterial strains isolated from different fruit juice samples to the species level.
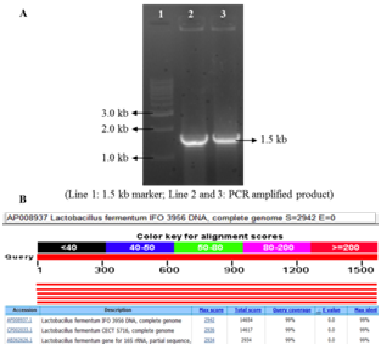

Fig. 2. A. PCR amplified product of L. fermentum
Table (3):
List of Lactobacillus species identified using 16S rRNA gene sequencing
Sl. No. |
Source |
Strain name |
Organism |
Accession number |
Amplified product (bp) |
|---|---|---|---|---|---|
1. |
Banana |
LAB-1 |
L. buchneri CD034 |
CP003043.1 |
1560 |
2. |
Beet root |
LAB-2 |
L. fermentum IF03956 DNA |
AP008937.1 |
1570 |
3. |
Dosa batter |
LAB-3 |
L. fermentum CECT 5716 |
CP002033.1 |
1567 |
4. |
Mango |
LAB-4 |
L. fermentum CECT 5716 |
CP002033.1 |
1567 |
5. |
Milk |
LAB-5 |
L. brevis ATCC367 |
CP000416.1 |
1570 |
6. |
Pine apple |
LAB-6 |
L. buchneri NRRL B-30929 |
CP002652.1 |
1571 |
7. |
Radish |
LAB-7 |
L. buchneri NRRL B-30929 |
CP002652.1 |
1569 |
8. |
Tomato |
LAB-8 |
L. buchneri CD034 |
CP003043.1 |
1472 |
9. |
Watermelon |
LAB-9 |
Wiesella ciberia |
NRIC0136 |
1571 |
bp= base pair
Among the 9 isolates, 4 were L. buchneri species that were isolated from banana, pineapple, radish and tomato. Three strains were L. fermentum species that were isolated from beet root, dosa batter and mango. One species each of L. brevis and W. ciberia was isolated from milk and water melon, respectively (Table 3). These results revealed the occurrence of different species of Lactobacilli with different fruits and vegetable sources. However, Kam et al. (2012) isolated Lactobacilli from fermented liquid sourdough and identified by 16S rRNA gene partial sequence analysis as L. plantarum and L. fermentum which were predominant bacteria in liquid sour dough. Similarly, Hamid and Abulfazl (2012) isolated and identified potentially probiotic lactobacilli in traditional white cheese of Tabriz in Iran using 16S rDNA gene. In this study, each sample yielded a single organism.
Table (4):
Polymorphism generated using random primers for the 9 Lactobacillus species
| Primers | Primer sequence | No. of amplified fragments | No. of polymorphic bands | No. of Mono morphic bands | ||
|---|---|---|---|---|---|---|
| Shared | Unique | |||||
| 1 | OPAH5 | 5’ TTGCAGGCAG 3’ | 6 | 5 | 1 | 0 |
| 2 | OPP3 | 5’ CAAGCTGGGT 3’ | 10 | 8 | 1 | 1 |
| 3 | OPP5 | 5’ CAGTGGGGAG 3’ | 10 | 8 | 2 | 0 |
| 4 | OPAH16 | 5’ ACTGAACGAC 3’ | 5 | 3 | 1 | 1 |
| 5 | OPAH17 | 5’ CTGATACGCC 3’ | 7 | 6 | 1 | 0 |
| 6 | OPN6 | 5’ CCCCGGTAAC 3’ | 8 | 6 | 1 | 1 |
| 7 | OPS1 | 5’ GTTTCGCTCC 3’ | 7 | 6 | 1 | 0 |
| 8 | OPS10 | 5’ CTGCTGGGAC 3’ | 7 | 5 | 1 | 1 |
| 9 | OPS11 | 5’ GTAGACCCGT 3’ | 7 | 6 | 0 | 1 |
| 10 | OPY5 | 5’GGCTGCGACA3’ | 3 | 2 | 0 | 1 |
| Total | 70 | 55 | 9 | 6 | ||
| Percentage | 100 | 78.57 | 12.85 | 8.57 | ||
| Mean | 7 | 5.5 | 0.9 | 0.6 | ||
Genetic diversity of Lactobacillus species
The RAPD technology is very useful, fast and informative in differentiating Lactobacillus species (Venturi et al., 2012) and analysis of genetic diversity and it is necessary to obtain information on variations between species within particular group. Therefore, to characterize genetic diversity of the species isolated, a total of 20 primers were screened for PCR amplification. Finally, 10 primers were selected for fingerprinting and diversity analysis based on the reproducibility and the banding pattern (Table 4). The 10 selected random primers showed considerable degree of polymorphism among the isolated species (Fig. 3). A total of 70 amplified bands were obtained with 10 primers, of which 64 (91.42%) were found to be polymorphic and 6 (8.57%) were monomorphic. Among the 64 polymorphic bands, 55 (78.57%) were polymorphic shared and 9 (12.85%) were polymorphic unique. A maximum of 10 bands were obtained in primer OPP5 and a minimum of 3 bands in primer OPY5 with an overall average of 7 bands. The high level of polymorphism (91.42%) obtained in this study indicates greater genetic diversity between the isolated strains. Weiss et al. (2005) used RAPD PCR using 14 arbitrary primers to differentiate 8 strains. Kumar et al. (2013) characterized 19 Bacillus thuringiensis strains using 14 random primers and reported 93 amplified fragments. Out of 93 fragments, 57 (61.20%) were found to be polymorphic shared, 29 (31.28%) were polymorphic unique and 7 (7.52%) were monomorphic.
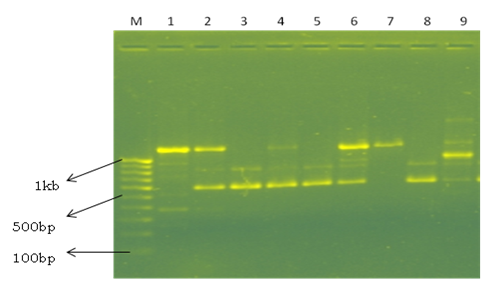
Fig. 3. Representative picture showing the RAPD profile in different Lactobacillus species by primer OPAH 5. Line M – 100 bp marker; Line 1- L. buchneri; Line 2- L. fermentum; Line 3- L. fermentum; Line 4- L. fermentum; Line 5- L. brevis; Line 6- L. buchneri; Line 7-L.buchneri; Line 8- L. buchneri; Line 9- Wiessella ciberiai.
The cluster analysis of the 9 lactobacillus isolates identified formed two major clusters. Six of the 9 isolates, LAB-1 (L. buchneri-1), LAB-5 (L. brevis), LAB-6 (L. buchneri-2), LAB-7 (L. buchneri-3), LAB-8 (L. buchneri-4) and LAB-9 (Weisella ciberia) were grouped in cluster 1 and 3 of them LAB-2 (L. fermentum-1), LAB-3 (L. fermentum-2) and LAB-4 (L. fermentum-3) grouped into clusters II. Further, the clusters I formed two sub clades and the sub clade 1 consists of LAB-1 (L. buchneri-1), LAB-5 (L. brevis), LAB-6 (L. buchneri-2), LAB-7 (L. buchneri-3) and the sub clade II contains LAB-8 (L. buchneri-4) and LAB-9 (W. ciberia) (Fig. 4). The dissimilarity matrix of 9 isolates of Lactobacillus revealed the highest dissimilarity (5.74%) was found between LAB-4 (L. fermentum-3) and LAB-9 (Weisella ciberia), whereas the lowest dissimilarity (3.74%) was found between LAB-2 (L. fermentum-1) and LAB-4 (L. fermentum-3) (Table 5). The study concluded that the L. fermentum was associated with beet root, dosa batter and mango, L. buchneri was present in banana, pine apple, radish, tomato and milk contains L. brevis and the W. ciberia was unique to watermelon. The diversity analysis indicated variation between the species isolated from different sources.
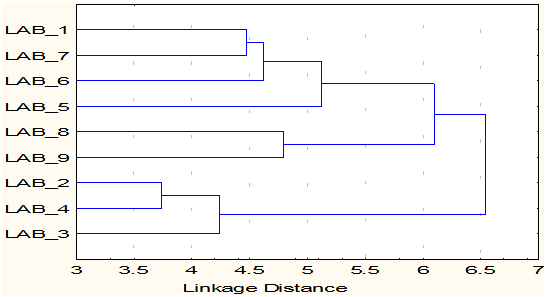
Fig. 4. Cluster diagram of 9 Lactobacillus species
Table (5):
Dissimilarity matrix of Lactobacillus isolates generated by RAPD analysis
LAB-1 |
LAB-2 |
LAB-3 |
LAB-4 |
LAB-5 |
LAB-6 |
LAB-7 |
LAB-8 |
LAB-9 |
|
|---|---|---|---|---|---|---|---|---|---|
LAB-1 |
0.0 |
||||||||
LAB-2 |
4.58 |
0.0 |
|||||||
LAB-3 |
5.10 |
3.87 |
0.0 |
||||||
LAB-4 |
5.20 |
3.74 |
4.36 |
0.0 |
|||||
LAB-5 |
5.10 |
4.80 |
5.29 |
4.58 |
0.0 |
||||
LAB-6 |
4.58 |
5.10 |
5.20 |
5.10 |
4.80 |
0.0 |
|||
LAB-7 |
4.47 |
4.58 |
5.10 |
5.00 |
4.90 |
4.58 |
0.0 |
||
LAB-8 |
5.39 |
5.66 |
4.80 |
4.90 |
5.20 |
4.90 |
5.00 |
0.0 |
|
LAB-9 |
5.29 |
5.39 |
5.29 |
5.74 |
5.29 |
5.39 |
5.29 |
4.80 |
0.0 |
- Ali, B., Ling, H.F., Sieo, C.C. and Rahim, R.A., Isolation, Identification and characterization of lactic acid bacteria from Polygoum minua. Romanian Biotechnol. Lett., 2012; 17(3): 7245-7252.
- Altschul, S. F., Gish, W. and Lipman, W. W., Basic Local Alignment Search Tool. J. Mol. Biol., 1990; (215)3: 403-410.
- Ashraf, M. Arshad, M., Siddique. and Muhammad, G., In vitro screening of locally isolated Lactobacillus species for probiotic properties. Pakistan Vet. J., 2009; 29(4): 186-190.
- De Man, J. D., Rogosa, M. and Sharpe, M. E., A medium for the cultivation of Lactobacilli. J. Appl. Bacterial., 1960; 23: 130-135.
- Dellaglio, Giovanna. and Efelis., Taxonomy of Lactobacillus and Bifidobacteria. Curr. Issues. Int. Microbiol., 2005; 8: 44-61.
- Fahathabad, E. H. And Eslamifer, M., Isolation and application of one strain of Lactobacillus paraplantarum from tea leaves (Camellia sinensis). Amer. J. Food Technol., 2011; 6(5): 429-434.
- Fathima, S., Anjum, T., Bajwa, R. and Ali, A., Identification of indigenous Xanthomonas isolates through 16S rRNA gene amplification and SDS page analyses. Afr. J. Microbiol. Res., 2012; 6(15): 3651-3655.
- Guasch, J. M., Andrés, L. C., Jáuregui, O. and Lamuela, R. R., First evidence of white wine in ancient Egypt from Tutankhamun’s tomb. J. Archaeol. Sci., 2005; 33: 1075-1080.
- Hamid, M. and Abulfazl, B., Isolation and molecular study of potentially probiotic Lactobacilli in traditional white cheese of Tabriz in Iran. Annals. Biol. Res., 2012; 3(4): 2019-2022.
- Holzapfel, E. H., Haberer, P., Geisen, R., Björkroth, J. and Schillinger, U., Taxonomy and important features of probiotic microorganisms in food and nutrition. Am. J. Clin. Nutr., 2001; 73(2): 365S-373S.
- Hoque, M. Z., Akter, F., Hossain, K. M., Rahman, M. S. M., Billah M. M. and Islam K. M. D., Isolation, Identification and Analysis of Probiotic Properties of Lactobacillus Spp. From Selective Regional Yoghurts. World J. DairyFood Sci., 2010; 5(1): 39-46.
- Kam, W. Y., Wanaida, W. M. and Sahilah, A. M., Identification of redominant Lactobacillus species in liquid sourdough fermentation, Int. Food Res. J., 2012; 19(4): 1739-1743.
- Konings, W. N., Kok, J., Kuipers, O. P. and Poolman, B., Lactic acid bacteria: the bugs of the new millennium. Curr. Opin Microbiol., 2000; 3(3): 276-282.
- Kumar, K., Yashaswini, K. S. and Earanna, N., Molecular Characterization of Lepidopteran specific Bacillus thuringiensis strains isolated from Hilly Zones of Karnataka, India. Afr. J. Biotech., 2013; 12(20): 2924-2931.
- Mallesha, R., Shylaja, D., Selvakumar. and Jagannath, J. H., Isolation and Identification of Lactic acid bacteria from raw and fermented products and their antibacterial activity. Recent Res. Sci. Technol., 2010; 2(6): 42-46.
- Naeem, M., Ilyas, M., Haider, S., Baig, S. and Saleem M., Isolation characterization and identification of lactic acid bacteria from fruit juices and their efficacy against antibiotics. Pak. J. Bot., 2012; 44: 323-328.
- Patil, M. M., Pal, A., Anand, T. and Ramanna, K.V., Isolation and characterisation of lactic acid bacteria from curd and cucumber. Indian J. Biotechnol., 2010; 19: 166-172.
- Paul, N., Smidt, I., Jelenas., Brilene, T., Annuk, H. and Mikelsaar, M., Inhibition of Clostridium difficile strains by intestinal Lactobacillus species., J. Med. Microbiol., 2004; 53: 551-554.
- Robert, S., Ecology of fermented foods. Human Ecology Rev., 2008; 15: 25-31.
- Sabin, Fatima, Tehmina, A., Rukhsana, B. and Ahmad, A.S., Identification of indigenous Xanthomonas isolated through 16S gene amplification and SDS PAGE analysis. Afri. J. Microbial. Res., 2012; 6(15): 3651-3655.
- Sambrook, J., Fritsch, F. F. and Maniatis, T., Molecular cloning: A laboratory manual, Cold Spring Harbour Laboratory Press, Cold Spring Harbor, NY., 1989; 931-957.
- Satokari, R. M., Vanguan, E. E. and Favier, C.F., Diversity of Bifidobacterium and Lactobacillus species in the breast fed and formula fed infants as assessed by 16S rDNA sequence differences. Microbial ecology, 2002; 14(2): 139-145.
- Venturi, M., Guerrini, S., Granchi, L. and Vincenzin, M., Typing of Lactobacillus sanfranciscensis isolates from traditional Sourdoughs by combining conventional and multiplex RAPD-PCR profiles. Int. J. Fd. Microbiol., 2012; 156: 122-126.
- Weiss, A., Hanspeter, P. L., Walter, K., Helmut, K. M. and Wolfgang, K., Identification of probiotic Lactobacillus reuteri strains. Food. Technol. Biotech., 2005; 43(3): 295-300.
© The Author(s) 2016. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


