ISSN: 0973-7510
E-ISSN: 2581-690X
The bacterium Staphylococcus can cause various health problems, particularly in hospitalized patients. Therefore, the current study aimed to isolate methicillin-resistant Staphylococcus aureus (MRSA) strains, test their capability to form a biofilm, and detect genes related to virulence and biofilm formation. Bacterial isolates were collected from the King Faisal Specialist Hospital and Children’s hospital in Taif Governorate, Saudi Arabia, and identified using primers for mecA and nuc1. They were tested for resistance against twelve widely distributed antibiotics and biofilm formation capability. The MRSA isolates were tested for fnbA, fnbB, and SCCmec. Among 100 isolates, 24 were identified as Staphylococcus aureus, and most of them were MRSA. Most isolates were resistant to cefrizine and cefepime (96%). The isolates showed higher resistance to amoxicillin and ampicillin (92%), followed by aztreonam (83%). Two isolates, S15 and S17, were high-grade positive for biofilm formation, 62.5% were medium-grade, and 20.8% were low-grade positive. Two of the isolates, S11 and S16, tested negative for biofilm formation. Furthermore, mecAI. ncu1 was found in all of the isolates, except S11. Most isolates had SCCmecIII and SCCmecV. All isolates were habituated to fnbB, while fnbA was not found in S3 and S11. These results indicated that PCR techniques offer rapid, simple, and accurate determination of the genetic profile and biofilm production capability of MRSA, and can be used in clinical diagnosis as well as to monitor the spread of antibiotic-resistant S. aureus strains.
Staphylococcus aureus, Biofilm Formation, Virulence genes, Antibiotic Resistance
Methicillin-resistant Staphylococcus aureus (MRSA) is one of the most pathogenic microbes that cause diseases in humans and animals. This gram-positive bacterium is found on human skin and nasal mucous membranes as a part of normal bacterial flora.1-3 Indiscriminate use of antibiotics has resulted in an increase in antibiotic-resistant bacteria, owing to which patients fail to respond to antibiotic treatment. The effectiveness of antibiotics used for treating humans and animals is threatened by resistant species and strains of microorganisms.4-6 S. aureus is a prevalent and endemic pathogen found in hospitals, and biofilm-forming and MRSA strains have become a serious clinical problem.7-9
S. aureus is cluster-shaped and has the ability to form a three-dimensional biofilm that surrounds the cluster of cells, allowing them to resist unfavorable conditions. MRSA can develop antibiotic resistance owing to its biofilm formation capability and hence poses a great threat to hospitalized patients.2,9 The biofilm is an organized structure and a major virulence factor that is important in protecting Staphylococcus cells from exposure to antibiotics.10 Biofilm-forming bacteria infect biofilms in most human infections, with Staphylococcus aureus being the most harmful biofilm-producing species.7,8 MRSA biofilms can spread to the hippocampus, and are not affected by antibiotics as the biofilm acts as a protective shield that increases resistance to antibiotics and other immune factors.9,11
MRSA causes chronic infections owing to its ability to resist various antibiotics by forming a biofilm on artificial heart valves, catheters, and medically implanted prostheses.12,13 The spread of MRSA along with other staphylococcal diseases has led to a significant increase in the use of antibiotics at an estimated annual cost of $450 million, with increased disease rates associated with biofilm-mediated infection.14 Therefore, an understanding of the evolution of staphylococcal biofilms at the molecular level is necessary to generate new treatment strategies for biofilm-associated infections and reduce the burden caused by these pathogens. Therefore, this study aimed to isolate antibiotic-resistant staphylococci and study their genetic details in 50 relation to the capability of biofilm formation.
Collection of Staphylococcus isolates
The study protocol was approved by Taif University Medical Ethics Review Board (Project No. 1-437-5371) in accordance with the guidelines for human protection. Samples were collected from patients at King Faisal Hospital and Children’s hospital in Taif City, Saudi Arabia, from October 2020 to November 2021, with documented patient consent. Approximately 100 bacterial isolates were collected and identified using a fully automated VITEK-2 COMPACT microbiology system (Bio Mtrieux, Inc., Durham, NC, USA).
Antibiotics susceptibility
The antibiotic susceptibility of S. aureus was determined according to the National Committee for Clinical Laboratory Standards (National Committee for Clinical Laboratory Standards, 2018)15 by disc diffusion method using Mueller-Hinton (MH) agar. The inocula were streaked onto MH agar plates using a sterile swab, and discs were incubated. The antibiotics tested were cefotaxime (30 µg), amoxicillin (25 µg), cefepime (30 µg), ampicillin (10 µg), cefixime (10 µg), cefatrizine (10 µg), gentamicin (10 µg), chloramphenicol (30 µg), lincomycin (15 µg), aztreonam (30 µg), oxacillin (5 µg), and norfloxacine (5 µg).
Biofilm forming capability
Biofilm formation by S. aureus isolates was determined using crystal violet assay on microtiter plates as described previously16. The optical density of each well was measured at 570 nm (OD570) using an automated Multiskan reader (Bio-Rad, Germany). Biofilm formation was interpreted as highly positive (OD570 ≥ 1), moderate-grade positive (0.4 ≤ OD570 < 0.9), low- 72 grade positive (0.1 ≤ OD570 < 0.4), or negative (OD570 < 0.1). Each isolate was tested three times.
DNA isolation
DNA was extracted from the isolates using a DNeasy Bacterial Mini Kit (QIAGEN, USA) according to the manufacturer’s instructions.
Detection of the mecA, SCCmec, and fibronectin-binding protein genes
PCR was performed using Go Taq® Green Master Mix (Promega, USA), according to the manufacturer’s instructions. The primers used and conditions for each gene are listed in Table 1.2,17 Amplicons were observed after electrophoresis on 1.5% agarose gel using a 100 bp DNA ladder (Fermentas, Lithuania, USA).
Table (1):
Primer sequences and amplicon sizes of tested genes fibronectin-binding protein mecA and SCCmec
Primers |
Sequence |
Size (bp) |
Annealing temp. |
|---|---|---|---|
mecAI-F |
(F) TCC AGA TTA CAA CTT CAC CAG G |
162 |
56 |
(R) CAA TTC ATA TCT TGT AAC G |
|||
ncu1 |
(F) TCC AGA TTA CAA CTT CAC CAG G |
301 |
52 |
SCCmec-III |
(R) CAA TTC ATA TCT TGT AAC G |
243 |
58 |
(F) CATTTGTGAAACACAGTACG |
|||
(R) GTTATTGAGACTCCTAAAGC |
|||
SCCmec-V |
(F) GAACATTGTTACTTAAATGAGCG |
325 |
58 |
(R) TGAAAGTTGTACCCTTGACACC |
|||
fnbA |
(F) CAT AAA TTG GGA GCA GCA TCA |
127 |
54 |
(R) ATC AGC AGC TGA ATT CCC ATT |
|||
fnbB |
(F) GTA ACA GCT AAT GGT CGA ATT GAT ACT |
||
(R) CAA GTT CGA TAG GAG TAC TAT GTT C |
524 |
54 |
Data analyses
Pearson’s simple linear correlation coefficient (r) and their significance (P) were assessed using SPSS 20.
Isolation of antibiotic-resistant bacteria
Approximately 100 clinical samples including urine and stool swabs were collected and analyzed for antibiotic-resistant bacteria. Of the 100 cultures tested, 24 bacterial isolates resistant to multiple antibiotics were identified as Staphylococcus and tested for biofilm formation, antibiotic resistance, and detection of virulence and biofilm genes. These isolates were assigned codes S1–S24.
Antibiotic susceptibility testing
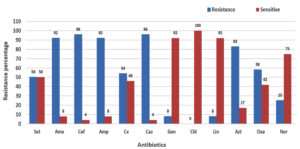
In total, 12 antibiotics were tested, and the isolates showed high variability of resistance. Most isolates showed highest resistance to cefrizine and cefepime (96%). The isolates also showed high resistance to amoxicillin and ampicillin (92%), followed by aztreonam (83%). All the isolates were sensitive to chloramphenicol. The isolates showed low resistance to lincomycin and gentamicin (8%) (Figure 1 and Table 2). The most sensitive isolates were S11, S16, and S18, which were resistant to amoxicillin, cefepime, cefrizine, and aztreonam. Isolates S15 and S17 showed the highest resistance to most of the antibiotics tested. The other isolates showed moderate resistance.
Figure 1. Antibiotic resistance pattern among S. aureus isolates, Whereas, Sxt = Trimethoprim / sulfamethoxazole, Amx = Amoxicillin, Cef = Cefepime, Amp = Ampicillin, Cx = Cefixime, Caz = Cefatrizine, Gen = Gentamicin, Chl = Chloramphenicol, Lin = Lincomycin, Azt = Aztreonam, Oxa = Oxacillin and Nor = Norfloxacine
Table (2):
The number of isolates and antibiotic resistance profile of Staphylococcus aureus isolates
Isolates |
Antibiotic Profile |
|---|---|
S -1 |
Sxt, Amx, Cef, Amp, Cx, Caz, Lin, |
S -2 |
Amx, Cef, Amp, Cx, Caz, Azt |
S -3 |
Amx, Cef, Amp, Cx, Caz, Azt, Oxa |
S -4 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa |
S -5 |
Amx, Cef, Amp, Cx, Caz, Azt |
S -6 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa, Nor |
S -7 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa, Nor |
S -8 |
Cef, Amp, Cx, Caz, Lin, Azt, Oxa |
S -9 |
Amx, Cef, Amp, Cx, Caz, Azt |
S -10 |
Sxt, Cef, Amp, Cx, Caz, Azt |
S -11 |
Amx, Cef, Caz, Azt |
S -12 |
Amx, Cef, Amp, Cx, Caz |
S -13 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa |
S -14 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa |
S -15 |
Sxt, Amx, Cef, Amp, Cx, Caz, Azt, Gen, Oxa, Nor |
S -16 |
Amx, Cef, Amp, Caz, Azt |
S -17 |
Sxt, Amx, Cef, Amp, Cx, Caz, Azt, Gen, Oxa, Nor |
S -18 |
Amx, Cef, Cx, Azt |
S -19 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa, Nor |
S -20 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa, Nor |
S -21 |
Amx, Cef, Amp, Cx, Caz, Azt, Oxa |
S -22 |
Amx, Cef, Amp, Cx, Caz |
S -23 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa |
S -24 |
Sxt, Amx, Cef, Amp, Caz, Azt, Oxa |
Whereas, Sxt = Trimethoprim / sulfamethoxazole (1.25/23.75µg), Amx = Amoxicillin (25µg), Cef = Cefepime (30 µg), Amp = Ampicillin (10µg), Cx = Cefixime (10µg), Caz = Cefatrizine (10µg), Gen =Gentamicin (10µg), Chl = Chloramphenicol (30 µg), Lin = Lincomycin (15 µg), Azt = Aztreonam (30µg), Oxa = Oxacillin (5 µg) and Nor = Norfloxacine (5 µg)
Determination of slime production
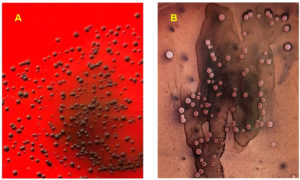
Phenotypic slime production was assessed by culturing isolates on CRA plates. Among the isolates, S1, S2, S13, S14, S15, and S17 were slime-producers, developing almost black colonies. The remaining isolates were considered as non-producers because they produced white colonies on CRA plates (Figure 2 and Table 3).
Figure 2. Colorimetric scale for colony analysis of slime production by Staphylococcus aureus 288 S15 and S11 using Congo Red agar assay. A: slime-producing strain (almost black); B: non-289 producing strain (white)
Quantitative biofilm formation
All 24 S. aureus isolates were screened for adherence to polystyrene microplates. Most isolates were able to form biofilms, and S15 and S17 were considered high-grade positive (OD570 values of 1.004 ± 0.007 and 1.011 ± 0.017, respectively). Isolates S1, S3, S4, S6, S7, S8, S13, S14, S19, S20, S21, S22, S23, and S24 were considered moderate-grade positive, with OD570 ranging from 0.670 ± 0.056 to 0.939 ± 0.025. Moreover, the isolates S2, S9, S10, S12, and S18 were considered low-grade positive, with OD570 ranging from 0.129 ± 0.003 to 0.390 ± 0.072. while S11 and S16 were negative for biofilm formation (Table 3).
Table (3):
Biofilm formation, grade of biofilm and production of the slime of Staphylococcus aureus isolates
Isolates |
Biofilm |
Grade of biofilm |
Production of slime |
|---|---|---|---|
S -1 |
0.670± 0.056 |
Mediate-grade positive |
Negative |
S -2 |
0.258±0.003 |
Low-grade positive |
Negative |
S -3 |
0.768±0.037 |
Mediate-grade positive |
Negative |
S -4 |
0.800±0.138 |
Mediate-grade positive |
Negative |
S -5 |
0.728±0.011 |
Mediate-grade positive |
Negative |
S -6 |
0.939±0.025 |
Mediate-grade positive |
Positive |
S -7 |
0.928±0.109 |
Mediate-grade positive |
Positive |
S -8 |
0.762±0.132 |
Mediate-grade positive |
Positive |
S -9 |
0.390±0.072 |
Low-grade positive |
Negative |
S -10 |
0.136±0.036 |
Low-grade positive |
Negative |
S -11 |
0.093±0.002 |
Negative |
Negative |
S -12 |
0.129±0.003 |
Low-grade positive |
Negative |
S -13 |
0.815±0.164 |
Mediate-grade positive |
Positive |
S -14 |
0.899±0.037 |
Mediate-grade positive |
Positive |
S -15 |
1.004±0.007 |
High-grade positive |
Positive |
S -16 |
0.070±0.010 |
Negative |
Negative |
S -17 |
1.011±0.017 |
High-grade positive |
Positive |
S -18 |
0.197±0.071 |
Low-grade positive |
Negative |
S -19 |
0.916±0.043 |
Mediate-grade positive |
Negative |
S -20 |
0.919±0.028 |
Mediate-grade positive |
Negative |
S -21 |
0.837±0.007 |
Mediate-grade positive |
Negative |
S -22 |
0.737±0.117 |
Mediate-grade positive |
Negative |
S -23 |
0.819±0.059 |
Mediate-grade positive |
Negative |
S -24 |
0.827±0.041 |
Mediate-grade positive |
Negative |
PCR analysis for mecAI, ncu1, and SCCmec genes
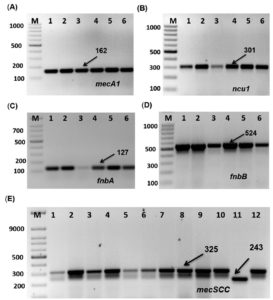
The PCR amplification products of mecA and SCCmec in S. aureus isolates are shown in Figure 3 and Table 4. All isolates carried mecA I with a size of approximately 162 bp. The ncu1 amplicon, approximately 301 bp in size, was found in most isolates except S11. This suggested that all isolates were MRSA, except S11, which is a weak isolate incapable of biofilm formation. Moreover, PCR amplicons of SCCmec were produced in all isolates, as shown in Figure 3. Most isolates had SCCmec III with a size of approximately 243 bp; however, isolates S16, S18, S19, S20, S21, S22, S23, and S24 did not contain SCCmec III. The SCCmec V amplicon, with a size of approximately 325 bp, was found in most tested isolates except in S11, S16, and from S18 to S24.
Figure 3. Amplification of Biofilm formation and MRSA genes of Staphylococcus aureus isolates by single PCR. (a) Amplification of mecA1 gene (162 bp). (b) Amplification of cnu1 gene (301 bp). (c) Amplification of fnbA gene (127 bp). (d) Amplification of fnbB gene (524 bp). (e) Amplification of mecSccIII and mecSccV genes (243 and 325bp). M: 100-bp DNA ladder
Detection of fnbA and fnbB
The presence of adhesive genes was confirmed by 127 bp and 524 bp bands of fnbA and fnbB, respectively. Almost all isolates were found to possess genes for both the homologous fibronectin-binding proteins fnbA and fnbB, except S3 and S11, which contained only the fnbA (Table 4 and Figure 2).
Table (4):
The number of Staphylococcus aureus isolates that habitat the Biofilm and SCCmec genes
| Isolates | Detection genes | |||||
|---|---|---|---|---|---|---|
| mecA1 | ncu1 | fnbA | fnbB | SCCmec III | SCCmec V | |
| S -1 | + | + | + | + | + | + |
| S -2 | + | + | + | + | + | + |
| S -3 | + | + | ــ | + | + | + |
| S -4 | + | + | + | + | + | + |
| S -5 | + | + | + | + | + | + |
| S -6 | + | + | + | + | + | + |
| S -7 | + | + | + | + | + | + |
| S -8 | + | + | + | + | + | + |
| S -9 | + | + | + | + | + | + |
| S -10 | + | + | + | + | + | + |
| S -11 | + | ــ | ــ | + | + | ــ |
| S -12 | + | + | + | + | + | + |
| S -13 | + | + | + | + | + | + |
| S -14 | + | + | + | + | + | + |
| S -15 | + | + | + | + | + | + |
| S -16 | + | + | + | + | ــ | ــ |
| S -17 | + | + | + | + | + | ــ |
| S -18 | + | + | + | + | ــ | ــ |
| S -19 | + | + | + | + | ــ | ــ |
| S -20 | + | + | + | + | ــ | ــ |
| S -21 | + | + | + | + | ــ | ــ |
| S -22 | + | + | + | ــ | ــ | ــ |
| S -23 | + | + | + | + | ــ | ــ |
| S -24 | + | + | + | + | ــ | ــ |
Biofilm formation is characteristic of several bacterial species and is related to their virulence. Chronic bacterial infections are closely related to biofilm formation.18,19 The current study showed that two isolates, S15 and S17, had high capability to form biofilms. Moreover, 55% of the isolates were medium-grade positive for biofilm formation. The biofilm-forming strains cause chronic infection associated with polymeric implants.20-22 MRSA is characterized by its adhesiveness, an important trait for infecting humans. The characterization of genes related to biofilm formation may improve the understanding of the phenotypic and genetic characteristics of infection caused by biofilms.7-9 Several genes responsible for biofilm formation have been characterized, example, genes encoding adhesion molecules in S. aureus.23,24 In this study, a polystyrene microtiter plate was used to detect biofilm formation. Molecular methods to detect the presence of biofilm-forming genes require the development of biofilm and polysaccharide adhesion between bacterial cells, which is brought about by genes encoding intracellular adhesion enzymes.8,9 The expression of genes related to biofilm formation is regulated by multiple genes, such as fnbA and fnbB, which may interact with each other and regulate biofilm formation. fnbA and fnbB contribute to the invasion and adhesion of this bacterial species, and therefore, may be related to its ability to form biofilms.25 The prevalence of Staphylococcus carrying these genes have been previously observed,26,27 and the differences in availability of the genes might be due to varied primer sequences or the location of these genes in the bacterial chromosome. In the current study, 96% of the Staphylococcus isolates carried fnbB and 92% had fnbA, which is in accordance with previous studies.25 The presence of fnbB may be related to the ability to form biofilms. The antibiotic resistance gene, mecA, has recently become prevalent; however, it is not indigenous to S. aureus and has been acquired recently from unknown sources.17 The mecA gene product is a penicillin-binding protein (PBP), specifically BP2a. S. aureus produces four PBPs,10 which are cytoplasmic membrane-fixing enzymes involved in cell wall formation.4,28,29 Eleven species containing SCCmec have been assigned to the Staphylococcus species.4,30 However, the prevalence of the fourth type is universal, while the rest of the species differ according to the location of bacterial isolates.30-32 All the isolates carried mecA and 96% of them had ncu1. Moreover, 79% of the isolates carried SCCmecIII, and 71% had SCCmecV. Several subtypes of SCCmec, including IIA to E, IVa to IVg, and VT have been reported.2 Two possible explanations for the ability of Staphylococcus species to colonize synthetic materials, such as catheters or hospital plastics, are the production of polysaccharide slime by Staphylococcus isolates and the presence of adhesives on biomaterial surfaces to host matrix proteins that are absorbed in vivo.6,9
PCR is an easy, fast, and cheap way to characterize pathogenic S. aureus isolates capable of forming biofilms and carrying related genes such as fnbA and fnbB. In addition, MRSA isolates are resistant to many antibiotics and can cause health issues. The genes mecA and SCCmec are virulence markers in Staphylococcus isolates.
ACKNOWLEDGMENTS
The author extends his appreciation to King Faisal Hospital, Health Affairs and Children’s Hospital at Taif, Saudi Arabia, for facilitating the collection of bacterial samples and antibiotic resistance assessment.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by Taif University Medical Ethics Review Board (Project No. 1-437-5371).
INFORMED CONSENT
Written informed consent was obtained from the participants before enrolling in the study.
- World Health Organization. Antimicrobial resistance: global report on surveillance. World Health Organization. 2014. https://apps.who.int/iris/handle/10665/112642
- Alharthi AA, Gaber A, Hassan MM. Molecular characterization of mecA and SCCmec genes in pathogenic Staphylococcus spp. collected from hospitals in Taif Region, KSA. Biotechnology. 2016;15(1-2):26-34.
Crossref - Kot B, Wierzchowska K, Piechota M, Gruzewska A. Antimicrobial resistance patterns in methicillin-resistant Staphylococcus aureus from patients hospitalized during 2015-2017 inhospitals in Poland. Med Princ Pract. 2020;29(1):61-68.
Crossref - Hafez EE, Al-Sohaimy SA, El-Saadani MA. The effect of the mecA gene and its mutant form on the response of S. aureus to the most common antibiotics. Int J Immunol Stud. 2009;29(1):106-122.
Crossref - Hassan MM , Soliman MM , Alotaibi SS, Sayed S, El-Shehawi AM, Ben-Abdallah F. Ameliorative impacts of rough cocklebur leaf extracts against methicillin-resistant Staphylococcus aureus. Fres Env Bull. 2022; 31: 6553-6560
- Saeidi S, Ravan H, Sanadgol N, Khaleghi M, Bazi S, Shojaei P. Methicillin-resistance Staphylococcus aureus in Southeast Iran: Herbal control and detection methods comparison. J Med Sci. 2014;14(3):123-129.
Crossref - Vasudevan P, Nair MKM, Annamalai T, Venkitanarayanan KS. Phenotypic and genotypic characterization of bovine mastitis isolates of Staphylococcus aureus for biofilm formation. Vet Microbiol. 2003;92(1-2):179-185.
Crossref - Gunther F, Wabnitz G, Stroh P, et al. Host defence against Staphylococcus aureus biofilms infection: phagocytosis of biofilms by polymorphonuclear neutrophils (PMN). Mol Immunol. 2009;46(8-9):1805-1813.
Crossref - Ben Abdallah F, Lagha R, Gaber A. Biofilm inhibition and eradication properties of medicinal plant essential oils against methicillin-resistant Staphylococcus aureus clinical isolates. Pharmaceuticals. 2020;13(11):369.
Crossref - Sajith-Khan AK, Shetty PJ, Lakshmi SY. Detection of mecA genes of methicillin-resistant Staphylococcus aureus by polymerase chain reaction. IJHRS. 2012;1(2):64-68.
Crossref - Simoes M, Bennett RN, Rosa EAS. Understanding antimicrobial activities of phytochemicals against multidrug-resistant bacteria and biofilms. Nat Prod Rep. 2009;26(6):746-757.
Crossref - Ribeiro M, Monteiro FJ, Ferraz MP. Infection of orthopedic implants with emphasis on bacterial adhesion process and techniques used in studying bacterial-material interactions. Biomatter. 2012;2(4):176-194.
Crossref - Parvizi J, Pawasarat IM, Azzam KA, Joshi A, Hansen EN, Bozic KJ. Periprosthetic joint infection: the economic impact of methicillin-resistant infections. J Arthroplasty. 2010;25(Suppl 6):103-107.
Crossref - Song X, Perencevich E, Campos J, Short BL, Singh N. Clinical and economic impact of methicillin-resistant Staphylococcus aureus colonization or infection on neonates inintensive care units. Infect Control Hosp Epidemiol. 2010;31(2):177-182.
Crossref - Clinical and Laboratory Standards Institute. Performance standards for antimicrobial disk susceptibility tests, 13th ed CLSI standard M02 Clinical and LaboratoryStandards Institute, Wayne, PA.2018.
- Ben-Abdallah F, Chaieb K, Zmantar T, Kallel H, Bakhrouf A. Adherence assays and Slime production of Vibrio alginolyticus and Vibrio parahaemolyticus. Braz J Microbiol. 2009;40(2):394-398.
Crossref - Murai M, Moriyama H, Hata E, Takeuchi F, Amemura-Maekawa J. Variation and association of fibronectin-binding protein genes fnbA and fnbB in Staphylococcus aureus Japanese isolates. Microbiol Immunol. 2016;60(5):312-325.
Crossref - Mohamed JA, Huang DB. Biofilm formation by enterococci. J Med Microbiol. 2007;56(Pt 12):1581-1588.
Crossref - Abou-Elhassan Y, Zhang Y, Jin S, Huigens RW. Transcript profiling of MRSA biofilms treated with a halogenated phenazine eradicating agent: a platform for defining cellular targets and pathways critical to biofilm survival. Angew Chem Int Ed Engl. 2018;57(47):15523-15528.
Crossref - Costerton JW, Stewart PS, Greenberg EP. Bacterial biofilms: a common cause of persistent infections. Science. 1999;284(5418):1318-1322.
Crossref - Gotz F. Staphylococci in colonization and disease prospective targets for drugs and vaccines. Curr Opin Microbiol. 2004;7(5):477-487.
Crossref - Gotz F. Staphylococcus and biofilms. Mol Microbiol. 2002;43(6):1367-1378.
Crossref - Singh R, Kumar R, Yadav B. Distribution of pathogenic factors in Staphylococcus aureus strains isolated from intra -mammary infections in cattle and buffaloes. Indian J Biotechnol. 2011;10:410-416.
- Wood TK. Strategies for combating persister cell and biofilm infections. Microb Biotechnol. 2017;10(5):1054-1056.
Crossref - Wong HC, Chung YC, Yu JA. Attachment and inactivation of Vibrio parahaemolyticus on stainless steel and glass surface. Food Microbiol. 2002;19(4):341-350.
Crossref - Boles BR, Thoendel M, Roth AJ, Horswill AR. Identification of genes involved in polysaccharide-independent Staphylococcus aureus biofilm formation. PLoS One. 2010;5(4):e10146.
Crossref - Gurusamy KS, Koti R, Toon CD, Wilson P, Davidson BR. Antibiotic therapy for the treatment of methicillin-resistant Staphylococcus aureus (MRSA) in non-surgical wounds. Cochrane Database Syst Rev. 2013;18(11):CD010427.
Crossref - Ito T, Katayama Y, Asada K, et al. Structural comparison of three types of Staphylococcal Cassette Chromosome mec integrated in the Chromosome in methicillin resistant Staphylococcus aureus. Antimicrob Agents Chemother. 2001;45(5):1323-1336.
Crossref - Laupland KB, Lyytikainen O, Sogaard M, et al. The changing epidemiology of Staphylococcus aureus bloodstream infection: A multinational population-based surveillance study. Clin Microbiol Infect. 2013;19(5):465-471.
Crossref - Ghaznavi-Rad E, Shamsudin MN, Sekawi Z, van Belkum A, Neela V. A simplified multiplex PCR assay for fast and easy discrimination of globally distributed staphylococcalcassette chromosome mec types in methicillin-resistant Staphylococcus aureus. J Med Microbiol. 2010;59(10):1135-1139.
Crossref - Sechi LA, Deriu A, Falchi MP, Fadda G, Zanetti S. Distribution of virulence genes in Aeromonas spp. Isolated from Sardinian waters and from patients with diarrhoea. J Appl Microbiol. 2002;92(2):221-227.
Crossref - Lakhundi S, Zhang K. Methicillin-resistant Staphylococcus aureus: Molecular characterization, evolution, and epidemiology. Clin Microbiol Rev. 2018;31(4):e00020-18.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.