ISSN: 0973-7510
E-ISSN: 2581-690X
Microorganism’s cycle exposure to reactive oxidants from internal metabolism and abiotic stress conditions, e.g. oxidative stress, salinity, drought, low temperature, high temperature, and high light. Synechocystis PCC6803 genomes typically encode different types of antioxidant scavenging enzymes including Peroxiredoxins (Prxs). There are five genes similar to Prxs were found inthe Synechocystis PCC 6803 genome. Based on sequence homology analysis of Synechocystis PCC 6803sll1621 gene, it is categorized into type II Prx (PrxII). The presumed amino acid sequence of Synechocystis PCC6803 PrxII protein exhibited identity about 44%-99% to other PrxII proteins from human, plants, algae, and other different cyanobacterial cells. In the last decade, the genetically controllable model organism E. coli has been introduced as a viable biotechnological model for genetically modified microorganisms. The Synechocystis PCC6803 sll1621 gene was overexpressed in pTYB21 expression vector and the resulting was named as pTYB21/sll1621. The pTYB21/sll1621 was overexpressed in Escherichia coli BL21 (DE3) host cell. The overexpressed protein of PrxII gave the recombinant E. coli cells the ability to survive under high concentrations of salinity stress, whereas the viability of wild type cells was completely inhibited at the same high concentrations of salinity stress. In conclusion, the present research documented the expressing of the sll1621 gene into E. coli cells. This result approved the absence of species barrier in relations to the function of Synechocystis PCC 6803 PrxII protein in different microorganism.
Synechocystis PCC6803, sll1621,PrxII, salinity stress, E. coli
Peroxiredoxins (Prxs) are a universal group of antioxidant proteins that promote the scavenging of several hydroperoxides1. In the past decades, Prxs have gotten extensive consideration as a group of thiol-specific antioxidant proteins. Additionally, they named as thioredoxin peroxidases and/or alkylhydroperoxide reductase proteins2. It started back to 1989 when an alkyl hydroperoxide reductase activity was isolated from Salmonellatyphimurium and E. coli. The enzyme was able to decrease cumene hydroperoxide as an electron acceptor with NADH or NADPH as an electron donor3. Yet, in a way autonomous of selenium, glutathione and heme can replace NADH and NADPH4. Thus, in such situation, two enzymes named AhpF (52 kDa) and AhpC (22 kDa) correspondingly made the activity4. According to their genetic background and catalytic activities, there are four different groups of enzymes that belong to Prxs which are 1-Cys Prx, 2-Cys Prx, PrxII, and PrxQ5.
Synechocystis PCC 6803 contains five Prxs genes6. These genes are a part of each reported groups as sll1621 (PrxII), sll0755 (2-Cys Prx), slr1198 (1-Cys Prx), and slr0242, sll0221 (Prx Q)6. Investigations of cyanobacterial Prxs mutant strains propose that the physiological roles of these proteins are focused on the adjustment of the growth of the cyanobacterial cells at high light intensities, however the catalytic activity may not generally include peroxide detoxification1,7. Interestingly, Synechocystis PCC 6803 PrxII disrupting mutant strain showed a seriously diminished development and growth rate comparative to the growth rate of wild type cells under even normal light intensities1,7. Interestingly, a mutant strain of Anabaena PCC 7120 was failed to express one of its four PrxQ, displayed a slow growth rate at moderate light intensities and was hypersensitive to methyl viologen8. Perez-Perez et al.9 stated that all five enzymes of Synechocystis PCC 6803 Prxs could utilize thioredoxins as an electron donor. The most noteworthy catalytic ability was acquired for the two enzymes of PrxII and TrxQ with thioredoxin as an electron donor9.Kobayashi et al.7indicated that the expression level of Synechocystis PCC6803 sll1621 gene was significantly induced under oxidative stress through the treatment of Synechocystis PCC6803 wild-type cells to methyl viologen for 15 min under high light intensities. Additionally, numerous defensive mechanism genes are liable for adaptation under a biotic stress conditions in Synechocystis PCC6803 cells. For example, 1-Cys Prx (slr1198) and PrxII (sll1621) genes are overexpressed under highlight treatment10. Also,another antioxidant enzyme in Synechocystis PCC6803, NADPH-dependent glutathione peroxidase-like protein (slr1992), was found to be overexpressed and enhance the transgenic Arabidopsis plants under several abiotic stress including salinity, drought, chilling and highlight11,12. Additionally, it was reported that the mRNA levels of sll1621 under conditions of paraqout-induced oxidative stress increased seven times more than the untreated cells13. All these data clearly show the importance of the sll1621 gene to scavenge the free radicals of reactive oxygen species (ROS) particularly under different abiotic stress conditions such as the exposure to high light and methyl viologen.
The aim of the present researchis to further analyze the effect of the overexpression of the Synechocystis PCC 6803 PrxII protein on the growth of E. coli BL21 cells under salt stress condition that induced oxidative stress. The present data will help to more emphasizing of the physiological role of Synechocystis PCC 6803 PrxII under salt stress and its use for the scavenging of ROS that generated under salt stress.
Chemicals and bacterial strains
Ampicillin, X-gal and IPTG purchased from UFC Biotechnology (USA). The restriction enzyme EcoRI and E. coli BL21 strain were acquired from NEB (USA). Other different synthetic compounds were obtained with the highest caliber financially accessible. Cloning vector (pGEM-T easy) was bought from Promega (USA); whereas, the expression vector pTVB21 was gotten from NEB (USA).
Growth conditions
Synechocystis sp. PCC 6803 cells was grown in Allen’s medium 14 under dim light condition with shaking for five to seven days.Cloning E. coli host cell (DH5a) and the expression E. coli host cells (BL21 DE3) were cultured in LB broth media at 36°C. Cells harboring recombinant plasmid was cultured and kept on LB media accompanied with ampicillin (100 mg/mL).
Gene expression of Synechocystis PCC6803 sll1621 in E. coli
Total DNA of Synechocystis PCC6803 cells was isolated as described previously15. The specific sll1621 gene was isolated using PCR with the following primer: 5’-CATATGACCCCC-GAACGAGTTCC-3’ (forward primer) and 5’-CTCGAGTTAGCCG-ACAAAAGCTTTAACG-3’ (reverse primer). Both primers were designed to introduce restriction enzymes sites of NdeI in forward primer and XhoI in reverse primer. The purified PCR product was successfully cloned into E. coli DH5a host cells (Promega, USA) using pGEM-T easy vector according to the instruction of the supplier. The positive transformed colony was confirmed by PCR and DNA sequencer. Next, the confirmed sequenced gene was over expressed in E. coli BL21 (DE3) (NEB, USA) host cell using pTYB21 vector (NEB, USA). The resulting construct was designated as pTYB21/sll1621.
Production of the recombinant protein
The recombinant pTYB21/sll1621vector was cultured in five mL LB broth accompanied with ampicillin (100 mg/mL)at two different temperatures (16°C or 36°C), then left overnight. Next, the overnight cultures were inoculated to a new 100 mL of LB broth. When the OD600 = 0.4, IPTG (400 mmol/L) was added and the culture cells were grown for additional three hours to induce the recombinant protein. In parallel, wild type strain harboring empty pTYB21 vector was cultured. Both cells were harvested, and SDS-PAGE was achieved in 12% (w/v) as descripted before14.
Evaluate of salt stress tolerance of recombinant E. coli cells
The recombinant (pTYB21/sll1621) and wild type (pTYB21/empty) cells were cultured in 50 mL LB broth accompanied with ampicillin (100 mg/mL) at 36°C with shaking. After the induction of the recombinant protein, both cells were collected by centrifugation at 5000 rpm/10 min. Next, the collected cells were re-inoculated in five ml of LB broth and distributed by equal concentration on LB agar medium including several concentrations of NaCl (0, 400, 800, and 1000mM) and were grown overnight at 36°C.
Protein sequence evaluation
The amino acids sequence of Synechocystis PCC 6803 PrxIIwas alignment with other PrxIIproteins from different organisms using ClustalW program16. The molecular information of PrxII protein (amino acids composition, molecular weight and pI value)wereestimatedusing the ProtParam resourcesfrom ExPASy website as described previously15.
Characterization of the sll1621 gene
To defend the cells and organ against oxidative stress and ROS, aerobic organisms have developed a profoundly classy and complex antioxidant prevention system17. It includes a diversity of components, both endogenous and exogenous in source. Both components work collaboratively to remove free radicals17. Type II Prxs are one of the most important peroxiredoxins proteins that can catalyze the reduction of various hydroperoxides1,2. Investigations of cyanobacterial Prxs mutant cells propose that these proteins work in adjustment to growth and development of cyanobacteria at high-light intensity, however the catalytic activity may not generally contain peroxide detoxification1. It was found that the interruption of the sll1621 gene had a reduction consequence on the survival growth rate of Synechocystis PCC 6803 cells under normal or dim light circumstances, suggesting that PrxII enzyme is important for the viability of this cells7. The purpose of the present research is to further explore the effect of the overexpression of Synechocystis PCC6803 sll1621 genein E. coli cells especially the survive rate under salt stress condition.
Table (1):
Identity and similarity percent of type II Peroxiredoxin proteins of Synechocystis PCC 6803 with other type II peroxiredoxin proteins.
| Type II Prx protein from Synechocystis PCC 6803 | ||||||
|---|---|---|---|---|---|---|
| S. PCC 6714 | S. PCC 7003 | C. reinhardtii | Jatropha curcas | Oryza sativa | Homo sapiens | |
| Identity | 99% | 85% | 48% | 48% | 43.4% | 44.2% |
| Similarity | 99% | 91% | 66% | 66% | 60.7% | 62.8% |
The sll1621 gene was shown to contain of 570 base pair that encoded a protein of 189 amino acids. The molecular mass of the PrxII protein was assessed to be 21.167 kDa with a pI of 4.94. The identity percentage of PrxII protein with other PrxIIs was ranged between 44.2% to 99% (Table 1). Total amino acid sequences of PrxII from different organisms including prokaryotic algae [Synechocystis PCC 6714 (189 amino acid) and Synechococcus PCC 7003 (187 amino acid)], eukaryotic algae [Chlamydomonas reinhardtii (194 amino acid)], higher plants [Jatropha curcas (227 amino acid) and Oryza sativa sub sp. japonica (225 amino acid)], and human (214 amino acid) were compared with the sequences of Synechocystis PCC 6803 PrxII protein (Fig. 1). All PrxIIs proteins share high degree of the critical catalytic cysteine residue inside the active site consensus sequence of PGAFTP(T/G)CS18. Moreover, all PrxIIs proteins have two cysteine residues. The second cysteine residue separated from the first one by the same number of amino acid residues (Fig. 1)18.
Fig. 1. Comparison of the deduced amino acid sequences of Synechocystis PCC 6803 type II Prx (189 aa) with those of the type II Prx proteins from Synechocystis PCC 6714 (189 aa), Synechococcus PCC 7003 (187 aa), Chlamydomonas reinhardtii (194 aa), Jatropha curcas (227), Oryza sativa subsp. japonica (225 aa), and Homo sapiens (214 aa). Amino acid sequences were aligned by CLUSTALW method using MAFFT v7.429. The active site is pointed out by black frame, and the two conserved cysteine residues are marked by red asterisk.
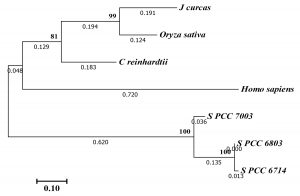
According to the phylogenetic tree analysis of PrxII proteins (Fig. 2), the Synechocystis PCC 6803 PrxII was clustered in same group with the similar PrxII proteins from other prokaryotic algae, including Synechocystis PCC 6714 and Synechococcus PCC 7003. The other PrxII proteins from eukaryotic algae, plants and human were clustered together in second group. Among all of these proteins, Synechocystis PCC 6803 PrxII was most nearly to PrxII from Synechocystis PCC 6714 (Fig. 2). The PrxII proteins of plants, Oryza sativa and Jatropha curcas, have an N-terminal extension about 38 amino acids than Synechocystis PCC 6803 PrxII (Fig. 1). Also, the human PrxII protein have an N-terminal about 25 amino acids than Synechocystis PCC 6803 PrxII. These extensions in plants or human PrxII proteins might be coded for a transit peptide anticipated to address the protein to chloroplast or mitochondria, respectively.
Fig. 2. Molecular phylogenetic tree by Maximum Likelihood method of the Synechocystis PCC 6803 type II Prx protein (S. PCC 6803) with those of the type II Prx proteins from other organisms including Synechocystis PCC 6714 (S. PCC 6714), Synechococcus PCC 7003 (S. PCC 7003), Chlamydomonas reinhardtii, Jatropha curcas, Oryza sativa, and Homo sapiens. The bootstrap consensus tree was from 1000 replicates. Numbers by nodes indicate Maximum Likelihood bootstrap. Evolutionary analyses were conducted in MEGA728.
Protein expression of PrxII in E. coli cells
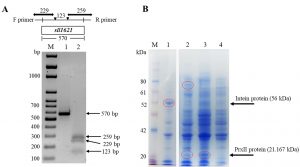
We amplify the sll1621 gene using PCR and the resulted specific product was used for cloning in E. coli DH5a cells (Fig. 3). Thesequence of the PCR product (570 base pair, Fig. 3A lane 1) was checkedusing an automated DNA sequencer apparatus. The results of DNA alignments of the gene productwith other genes in NCBI website indicated that the gene was 100% homologous to Synechocystis PCC 6803 sll1621 gene (data not shown). To further ensure about the full sequence of sll1621 gene, we performed a restriction gene map using EcoRI restriction enzyme. As expected, the restriction map of the sll1621 after cutting with EcoRI was given three fragments with lengths of 259 bp, 229 bp and 123 bp(Fig. 3A, lane 2).
Fig. 3. Isolation and expression of sll1621 gene in E. coli BL21 (DE3) cells. (A) PCR amplification of sll1621 gene that isolated from the genomic Synechocystis PCC 6803 (lane 1),while, lane 2 represented the restriction map of the sll1621 gene using EcoRI. (B) SDS-PAGE analysis of the recombinant PrxII protein after addition of IPTG. Lane 1, E. coli cells transformed with pTYB21/empty vector that cultured at 15°C for 16 hours; lane 2,E. coli cells transformed with pTYB21/sll1621vector that cultured at 15°C for 16 hours; lane 3, E. coli cells transformed with pTYB21/sll1621 vector that cultured at 36°C for 3 hours; lane 4, E. coli cells transformed with pTYB21/empty vector that cultured at 36°C for 3 hours; lane M, PiNK plus prestained protein ladder (GeneDireX).
The restriction sites of NdeI and XhoI that involved in the gene product were used to overexpress sll1621 gene into pTYB21 vector. The resulting was namedaspTYB21/sll1621, which was transformed into E. coli BL21 (DE3) cells to obtain there combinant protein. Subsequently, we examined the best conditions for the growth of BL21 (DE3) cells to produce PrxII protein with high expression (Fig. 3B). In this case two types of temperatures were used (15°C and 36°C). The reason for choosing 15°C, as it is recommended for the overexpression of the Intein protein, of the Saccharomyces cerevisiae, that fused at the N-terminus of pTYB21 vector (56 kDa). The recombinant protein was successfully expressed with a sufficient level in BL21 (DE3) cells at 36°C (Fig. 3B). The predicted molecular weight of the recombinant PrxII protein (21.167 kDa) was detected (Fig. 3B, lane 3). Whereas, we could not detect any protein band related to PrxII molecular weight in E. coli control cells (Fig.3B, lane 4). Interestingly, at 15°C, we detect a protein with a molecular weight of 77 kDa in recombinant BL21 (DE3) cells that harboring pTYB21/sll1621 vector (Fig. 3B, lane 2). This molecular weight is considered for the total molecular weight of the Intein protein (56 KDa) that expressed by pTYB21 at 15°C plus PrxII protein (21.16 KDa).
Synechocystis PCC 6803 PrxII protein is necessary for improve growth of E. coli under salt stress
Peroxiredoxins are proteins that have molecular weight around 20 to 30 kDa and widely spread in living organisms19. Initially, Prxs were identified according to its ability to protect cells especially proteins from abiotic stress particularly oxidative damage that leads to an increase of ROS20. Previously, Prxs was termed as the “protector protein” or “thiol-specific antioxidant” before being name again as Prxs21-24. Five different Prxs genes including sll1621 were found in the genome of Synechocystis PCC 68036. Also, Kobayashi et al.7 stated that the mRNA level of the sll1621 gene is remarkably up-regulated when Synechocystis PCC 6803 are exposed to methylviologen with light conditions. Also, Hosoya-Matsuda et al.1 concluded that PrxII is significantly important as scavenging ROS enzyme. They found that the growth of the knockout mutant cells lacking sll1621 gene was remarkably weak against oxidative stress1.
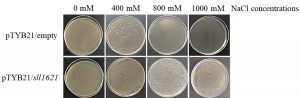
Oxidative injury to microorganism cells frequently happens under different abiotic stress conditions during bioprocess11,15. Thus, it has been addressed either the peroxiredoxin activity of PrxII protein assumes a significant useful role in recombinant E. coli, and additionally whether such activity might be useful under salinity stress. In the present research, we used the model microorganism, E. coli, to examine the ability of Synechocystis PCC 6803 sll1621 gene to increase the viability of growth of E. coli recombinant cells under salt stress. Salt stress was generated in-vitro by culturing the recombinant and wild type E. coli cells on LB agar provided with various concentrations of NaCl (Fig. 4). The pTYB21/empty cells were sensitive to 400mM NaCl, while, the growth was completely deleted in the presence of 800mM and 1000mM of NaCl (Fig. 4). On the other hand, the recombinant pTYB21/sll1621 cells increased the growth efficiency under the high concentration of NaCl (1000mM), demonstrating that the growth was, for some reason, improved compare to those of the pTYB21/emptycells (Fig. 4). Accordingly, this survival test under different concentration of NaCl showed that the sll1621 gene from Synechocystis PCC 6803 could function as an antioxidant protector enzyme to defend cells from oxidative damage under salt stress (Fig. 4). Similar results documented that the expression of PrxIIgene are typically affected by oxidative stress and abiotic stress that caused by hyperoxia, peroxides, UV, and ionizing radiation25-27.
In the present work, the Synechocystis PCC 6803 sll1621 gene was successfully over expressed in the cytoplasm of E. coli BL21 (DE3) cells. The over expressed protein was verified by SDS-PAGE and observ ableviability test. The present data specify that Synechocystis PCC 6803 sll1621 gene confers tolerance of recombinant E. coli cells to high concentration of salt stress. Consequently, these results are strong evidence for the success of gene expression and the absence of species barrier among different microorganisms.
ACKNOWLEDGMENTS
The authors would like to thank The Deanship of Scientific Research, Taif University.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
AG, MMF, AKI and MHH involved in the design of the research. AG and MHH designed the experimental procedure. AG performed the molecular cloning and gene expression experiments. MMF and AKI involved in SDS-PAGE experiment. AG wrote the original draft. MMF, AKI, MHH, SA and WFA involved in analysis the data. MMF, AKI, MHH, SA and WFA reviewed and edited the final version. All authors agreed with the final version of the manuscript.
FUNDING
The authors would like to thank The Deanship of Scientific Research, Taif University for funding this research [grant number1-438-5947].
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
AVAILABILITY OF DATA
All datasets analyzed in the study are involved in the manuscript and existing as tables and figures.
- Hosoya-Matsuda N, Motohashi K, Yoshimura H, et al. Anti-oxidative stress system in cyanobacteria: Significance of type II peroxiredoxin and the role of 1-Cys peroxiredoxin in Synechocystis sp. strain PCC 6803. J Biol Chem. 2005;280(1):840-846.
Crossref - Dietz KJ. Plant Peroxiredoxins. Annu Rev Plant Physiol Plant Mol Biol. 2003;54:93-107.
Crossref - Storz G, Jacobson FS, Tartaglia LA, Morgan RW, Silveira LA, Ames BN. An alkyl hydroperoxide reductase induced by oxidative stress in Salmonella typhimurium and Escherichia coli: genetic characterization and cloning of ahp. J Bacteriol. 1989;171(4):2049-2055.
Crossref - Jacobson, FS, Morgan RW, Christman MF, Ames BN. An alkyl hydroperoxide reductase from Salmonella typhimurium involved in the defense of DNA against oxidative damage. Purification and properties. J Biol Chem. 1989;264:1488-1496.
- Perelman A, Uzan A, Hacohen D, Schwarz R. Oxidative stress in Synechococcus sp. strain PCC 7942: various mechanisms for H2O2 detoxification with different physiological roles. J Bacteriol. 2003;185:3654-3660.
Crossref - Kaneko T, Sato S, Kotani H, et al. Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. strain PCC6803. II Sequence determination of the entire genome and assignment of potential protein-coding regions. DNA Res. 1996;3:109-136.
Crossref - Kobayashi M, Ishizuka T, Katayama M, et al. Response to oxidative stress involves a novel peroxiredoxin gene in the unicellular cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol. 2004;45:290-299.
Crossref - Latifi A, Ruiz M, Jeanjean R, Zhang CC. PrxQ-A, a member of the peroxiredoxin Q family, plays a major role in defense against oxidative stress in the cyanobacterium Anabaena sp. strain PCC7120. Free Radic Biol Med. 2007;42:424-431.
Crossref - Perez-Perez ME, Mata-Cabana A, Sanchez-Riego AM, Lindahl M, Florencio FJ. A Comprehensive Analysis of the Peroxiredoxin Reduction System in the Cyanobacterium Synechocystis sp. Strain PCC 6803 Reveals that All Five Peroxiredoxins Are Thioredoxin Dependent. J Bacteriol. 2009;191(24):7477-7489.
Crossref - Singh AK, Elvitigala T, Bhattacharyya-Pakrasi M, Aurora R, Ghosh B, Pakrasi HB. Integration of carbon and nitrogen metabolism with energy production is crucial to light acclimation in the cyanobacterium Synechocystis. Plant Physiol. 2008;148:467-478.
Crossref - Gaber A, Yoshimura K, Tamoi M, Takeda T, Nakano Y, Shigeoka S. Induction and Functional Analysis of Two Reduced Nicotinamide Adenine Dinucleotide Phosphate-Dependent Glutathione Peroxidase-Like Proteins in Synechocystis PCC 6803 during the Progression of Oxidative Stress. Plant Physiol. 2004;136(1):2855-2861.
Crossref - Gaber A, Yoshimura K, Yamamoto T, et al. Glutathione peroxidase-like protein of Synechocystis PCC 6803 confers tolerance to oxidative and environmental stresses in transgenic Arabidopsis. Physiol Plant. 2006;128:251-262.
Crossref - Gao H, Xu X. The Cyanobacterial NAD Kinase Gene sll1415 Is Required for Photoheterotrophic Growth and Cellular Redox Homeostasis in Synechocystis sp. Strain PCC 6803. J Bacteriol. 2012;194(12):218-224.
Crossref - Allen MM. Simple conditions for the growth of unicellular blue-green algae on plates. J Phycol. 1968;4:1-4.
Crossref - Gaber A, Hassan MM, El- Awady MA. The overproduction of Synechocystis sp. PCC 6803 heat-shock protein (Sll0170) protects Escherichia coli against high-temperature stress. Biotechnol Equip. 2015;29(6):1201-1207.
Crossref - Thompson JD, Higgins DG, Gibson TJ. CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673-4680.
Crossref - Lushchak, VI. Free radicals, reactive oxygen species, oxidative stress and its classification. Chem Biol Interact. 2014;224:164-175.
Crossref - Brehelin C, Meyer EH, de Souris JP, Bonnard G, Meyer Y. Resemblance and dissemblance of Arabidopsis type II peroxiredoxins: similar sequences for divergent gene expression, protein localization, and activity. Plant Physiol. 2003;132(4):2045-2057.
Crossref - Sharapov MG, Ravin VK, Novoselov VI. Peroxiredoxins as Multifunctional Enzymes. Mol Biol. 2014;48(4):520-545.
Crossref - Rhee SG, Woo HA, Kil IS, Bae SH. Peroxiredoxin Functions as a Peroxidase and a Regulator and Sensor of Local Peroxides. J Biol Chem. 2012;287(7):4403-4410.
Crossref - Kim K, Kim IH, Lee KY, Rhee SG, Stadtman, ER. The isolation and purification of a specific “protector” protein which inhibits enzyme inactivation by a thiol/Fe(III)/O2 mixed-function oxidation system. J Biol Chem. 1988;263:4704-4711.
- Chae HZ, Kim IH, Kim K, Rhee SG. Cloning, sequencing, and mutation of thiol-specific antioxidant gene of S. cerevisiae. J Biol Chem. 1993;268:16815-16821.
- Chae HZ, Robison K, Poole LB, Church G, Storz G, Rhee SG. Cloning and sequencing of thiol-specific antioxidant from mammalian brain: alkyl hydroperoxide reductase and thiol-specific antioxidant define a large family of antioxidant enzymes. Proc Natl Acad Sci USA. 1994;91:7017-7021.
Crossref - Rhee SG, Woo HA. Multiple functions of peroxiredoxins: peroxidases, sensors and regulators of the intracellular messenger H2O2, and protein chaperones. Antioxid Redox Signal. 2011;15:781–794.
Crossref - Kim HS, Kang SW, Rhee SG, Clerch LB. Rat lung peroxiredoxins I and II are differentially regulated during development and by hyperoxia. Am J Physiol Lung Cell Physiol. 2001;280:1212-1217.
Crossref - Ito T, Kimura S, Seto K, et al. Peroxiredoxin I plays a protective role against UVA irradiation through reduction of oxidative stress. J Dermatol Sci. 2013;12:S0923-S1811.
- Lee K, Park JS, Kim YJ, et al. Differential expression of Prx I and II in mouse testis and their up-regulation by radiation. Biochem Biophys Res Commun. 2002;296:337-342.
Crossref - Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016; 33:1870-1874.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.