ISSN: 0973-7510
E-ISSN: 2581-690X
Sheep and goat pox (SGPX) are systemic transmissible viral diseases characterized mainly by skin and internal lesions. This work aimed to provide the first molecular detection of sheep pox virus (SPV) in Iraq. Sheep pox virus (SPV) is a widely spread skin viral disease in Iraqi sheep and is associated with the formation of benign and malignant lesions in adult and young animals, resulting in notable economic losses in sheep and goat. In the current study, 24 sheep pox samples were collected from sheep from central Iraq. These samples were analyzed by polymerase chain reaction, and sequence analysis. Sheep pox virus DNA was detected in 18 of the samples (75%) in Iraqi sheep as the main causative agent for the disease. A partial sequence for the SGP P32 gene was deposited in Gene Bank. Phylogenetic analysis confirmed the presence of SPV-Iraqi isolate and showed that the origin of infection may be imported from India, china and USA sheep. This study presents the first report of SPV-infection in the Iraqi sheep and gives to spreading the knowledge of the origin of this disease. The results of this study will assist in the development of appropriate prophylaxis procedures and therapeutic measures.
Sheep pox, Molecular, Phylogenetic analyses.
Sheep and goat pox (SGPX) are systemic contagious viral diseases characterized mainly by skin and internal lesions which were observed as different diseases in the past1. Sheep and goat pox are endemic in greatest portions of the world such as the Middle East, Turkey, Iran, Iraq, Afghanistan, Pakistan, India, Nepal, and Africa north of the Equator and other parts of the world2-4. In Iraq, the virus was isolated for the first time from the northern part of Iraq and designated as Sersenik strain5,6.
The SGPX virus is a Capri pox virus belong to the family Poxviridae, one of the largest (170–260 nm by 300–450 nm) enveloped, double stranded DNA viruses, lengths of around 150 kbp. Strains from goatpox and sheep pox share at least 147 genes7. The SGPX seasonality could be described moreover by the ability of the viruses to survive for several months in wet and cold weather, in the lambing period, or by the deprived physiological disorder of flocks in the autumn1,8,9, while sheep and goat shows a main role in national economy , livestock diseases are the essential technical restrictions by reducing production and impeding skill in animal and animal products. In young animals the mortality rate may reach 100% while the overall mortality may be 50% of the flock10,11.
Sheep and goat are an important source of beef products in Iraq, and SGPX is responsible for significant economic losses for Iraqi sheep breeders, the importance and breadth of the disease in Iraq makes it necessary to study the epidemiology and main types causing this disease, which can help control the disease.
Ethical approval
The animals used in this study were defined as farm animals and no experimental work was performed on them, therefore, approval was not required from IACUC or anethics committee. The owner of the animals provided permission for these studies. The ethics committee at the Wahj Al –DNA and Iraqi Biotechnology Company Lab. Baghdad, Iraq reviewed the work and approved the waiver.
Animal and sample collection
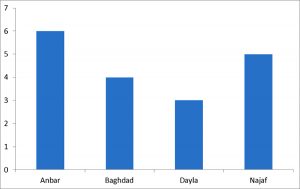
In this study, recorded veterinarians from the cities of Anbar, Dayla, Baghdad, and Najaf collected Sheep pox samples from 24 animals. In those different regions of Iraq, the study samples included sheep suffering from sheep pox virus (SPV) disease that were brought to private clinics by their owners between October 2017 and June 2018, sheep pox tissue sample were collected and blood samples were collected in EDTA tubes from this animals.
Six (2 tissue + 4 blood) samples were collected from Al-Anbar province, 5 blood sample were collected from Dayla city, 7 blood samples were collected from Baghdad city and 6 blood samples were collected from al- Najaf province. The blood samples transferred for laboratory test by EDTA tube and tissue samples were removed and transferred into sterile tubes separately and stored at -20°C for further use. These samples were submitted for examination directly by PCR.
The clinical significance of these animals showed benign lesion occur, particularly under the tail and head Figure 1.
Extraction and DNA amplification
A Magnesia® Genomic DNA Tissue Kit (Anatolia Gene works, Istanbul, Turkey) kit was used to extract viral DNA from frozen tissue and blood samples on a Magnesia® 16 magnetic bead extraction system (Anatolia Gene works) according to the manufacturer’s instructions. The primers were designed manually using A Plasmid Editor (ApE) software (v2.0.49, 2015) to align the complete genome sequences and to choose the conserved regions between sequences Table 1.
Table (1):
The primers were used in detection of virus.
| Primer Sequence | Tm (ᵒC) | GC (%) | Product size | |
|---|---|---|---|---|
| Forward | 5′- GCGGATCCTTTCTACAGGCT -3′ | 56.9 | 55 % | 551 base pair |
| Reverse | 5′ GCGGATCCACTATATATACGT-3′ | 51.1 | 42.9 % |
Ready Mix (Kapa Biosystems, Cape Town, South Africa). The amplification protocol included an initial denaturation at 94°C for 3 min, followed by 35 cycles of denaturation at 94°C for 20s, annealing at 44°C for 15s, an extension at 72°C for 30s, and a final extension at 72°C for 10min/kb.
The PCR products of the viral DNA were detected by electrophoresis on a 1.5% agarose gel containing ethidium bromide, at 5 volt/cm2. 1X TBE buffer for approximately 1:30 hours. The DNA was visualized under gel recommendation system.
The KAPA Universal Ladder Kit (10Kbp) is designed for determining the approximate size and quantity of double-stranded DNA on agarose gel.
DNA sequencing
A total of 18 sheep pox virus (SPV) positive specimens (as determined by PCR), which were selected as characteristic of the SPVs geographical distribution were sequenced to confirm viral genome type. For sequencing analysis, a PCR product which fragments genes were purified using a Strata Prep DNA Gel Extraction Kit (Agilent Technologies, USA); sequencing was performed at the National Instrumentation Center for Environmental Management (NICEM), College of Agriculture and Life Sciences, Seoul National University (South Korea). Sequencing reactions were performed using both forward and reverse primers in 10mL total reaction volume (ABI Big DYE V3.1 Ready-Reaction Kit; Applied Bio systems, USA) according to the manufacturer’s recommendations. The samples were analyzed on a 3730XL DNA Analyzer (Applied Biosystems, USA). Forward and reverse complementary sequences were aligned using ApE (A plasmid Editor) software (v2.0.49, 2015), and the obtained results were submitted to GenBank. ApE software was used to detect corresponding amino acid sequences.
Phylogenetic analysis
The sequences obtained in this study were submitted to phylogenetic analysis. As described above, the sequences were generated using a SGP specific primer set, and the products were deposited in GenBank under the primary accession number.
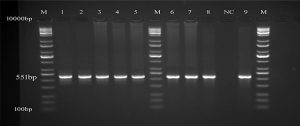
Samples were made to run out and assessed for sequencing analysis by using PCR reaction. The bands of Figure 2 represented the differentiation of strains of genotypes for the virus. These SGP positive samples produced DNA fragments of 551bp fragments of attachment gene, which were amplified using SGP P32 gene primers (Table 1), (Figure 3). PCR analysis showed that 75% of the analyzed sheep pox virus disease was induced by SGP in the measured Iraqi sheep populations, which included cross breeds in central Iraq (Table 2), (Figure 2).
Table (2):
Detection of SPV by PCR technique.
Total no. of cases |
SPV No. |
SPV % |
|---|---|---|
24 |
18 |
75 |
SPV: Sheep pox virus
PCR product the band size 551bp. The product was electrophoresis on 1.5% agarose at 5 volt/cm 2. 1x TBE buffer for 1:30 hours. M: DNA ladder (10Kbp), visualized under gel documentation system, Lane (1, 2, 3, 4, 5) SGP from tissue samples, Lane (6, 7, 8, 9) SGP from blood samples, NC negative control.
Sequencing
Sequencing analyses were conducted on the PCR products for SGP genes amplified from the collected samples that showed identical sequence. The results confirmed the presence of SPV with 99% identity for the majority of the NCBI BLAST-searched SPV sequences of sheep pox of India, China and USA strain which bear the accession No. in the Table 3. As we used SGP–specific primers to amplify the extracted DNA samples, SPV was the predominant genotype detected in all 18 SPV-positive (75%) scab and blood samples. This finding is the first genotype confirmation of the presence of SPV as a primary causative agent for sheep pox virus disease in Iraqi sheep.
Table (3):
NCBI reported virion protein gene in Sheep pox virus strain from India, china and USA.
Sr.N. |
Accession |
Country |
Host |
Compatibility |
|---|---|---|---|---|
1. |
ID: MG000157.1 |
India |
Sheep |
99% |
2. |
ID: MG000156.1 |
India |
Sheep |
99% |
3. |
ID: KT438551.1 |
China |
Ovis aries |
99% |
4. |
ID: KT438550.1 |
China |
Ovis aries |
99% |
5. |
ID: KY500996.1 |
India |
Sheep |
99% |
6. |
ID: KY500995.1 |
India |
Sheep |
99% |
7. |
ID: AY077834.1 |
USA |
Sheep |
99% |
8. |
ID: AY077833.1 |
USA |
Sheep |
99% |
9. |
ID: AY077832.1 |
USA |
Sheep |
99% |
Phylogenetic analysis
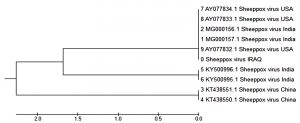
A phylogenetic tree was generated using retrieved genome sequences that were deposited in NCBI under the primary accession number (Figure 4). Sheep pox virus was the main sheep pox virus genotype recognized in (18 samples). The aligned sample sequences were classified as SPV The aligned sequences exposed high similarity to the nucleotide sequence of SPV (accession number MG000157.1, MG000156.1, KY500996, India strain) and to a sequence of a SPV (accession No. KT438551, KT438550,) isolate from china and (accession number. AY077834, AY077833, AY077832) isolate from USA.
In the current work, clinically and Molecular diagnosed cases of sheep pox disease were studied to identify and characterize the sheep pox virus genotype which was the most prevalent in the central Iraq region, which is a major location for large sheep breeding farm. The identification and characterization of the SPV genotype present in this region are important for effective disease control.
Polymerase chain reaction play a cut off tool in the identification of medically important sequences of SPV, PCR amplifies a specific region of a DNA target. Most PCR methods amplify DNA fragments of between 0.1 and 10 kilo base pairs (kbp), although some techniques allow for amplification of fragments up to 40 kbp in size12, The quantity of amplified product is determined by the available substrates in the reaction, which become limiting as the reaction progresses13.
In the presented study genotype specific primers contain P32 gene have been used14,15 to identify and characterize SPV, amplified by PCR and these results confirmed by sequencing and phylogenetic analysis, which are considered to be the best methods for these purposes according to the works16 and interestingly, based on distance analysis, this sequence was found to exhibit high similarity to sequences isolated from an India, China and USA. These results suggest the possibility of disease transmission via the importation of livestock from those countries17.
Sheep pox virus and LSDV generally have genome identities similar by at least 96%. Thus SPV vaccine can be used for prophylaxis against LSDV. In Iraq Allawe and Mohsen 2017confirmed a similarity between the lumpy skin disease virus and sheep pox vaccine up to 97-98%18-20. The current study is the first aimed at the recognition and description of SPV genotype in central Iraq. The results of this work are essential for assisting in the development of prophylactic and therapeutic procedures to reduce the economic losses associated with sheep and goat pox virus disease Iraq.
Conflict of Interest
The author(s) declare that there is no conflict of interest.
- Bhanuprakash V, Moorthy ARS, Krishnappa G, Srinivasagowda RN, Indrani BK. An epidemiological study of sheep pox in Karnataka state, Révue Scientifique et Technique (Office International des Epizooties) 2005; 24:909–920.
- Hussein M, El-Sadi HI, Zangana IK, El-Attar MY, Al-Bana AS. Clinical, pathological and virological studies on goat pox in Mosul, Iraq. Veterinary Medical Journal Giza. 1989; 37: 65–76.
- Office International des Epizooties 2008. Sheep pox and goat pox. In: Manual of Diagnostic Tests and Vaccines for Terrestrial Animals, 5th edn, World Organization for Animal Health, Paris, France.
- Zewdie S. Sheep and goat pox: causes, prevention and treatment. Technical bulletin No. 29. Ethiopia Sheep and Goat Productivity Improvement Programme, 2009.
- Tantawi HH, Al-Falluji MM. Laboratory characteristics of four strains of goat pox virus. Acta Virologica 1979; 23:455-460.
- Al-Bana AS, Abass AH. Growth characteristics, electron microscopy and immunoflourescence studies on goat pox virus isolated in Iraq. Iraqi J Vet Sci 1983; 9:75-84.
- Matthews RE. Classification and nomenclature of viruses. Intervirology 1982; 17:1-99.
- Yeruham I, Yadin H, Van Ham M. Economic and epidemiological aspects of an outbreak of sheeppox in a dairy sheep flock. Vet Rec 2007; 160:236-237.
- Tsegaye D, Belay B, Haile A. Prevalence of Major Goat Diseases and Mortality of Goat in Daro-Labu District of West Hararghe, Eastern Ethiopia. J Sci Innov Res 2013; 2:665-672.
- Jilo K, Abdela N, Adem A. Insufficient veterinary service as a major constrants in pastoral area of ethiopia: a review. Journal of Biology, Agriculture and Healthcare 2016, 6: 94-101.
- Ausvetplan. Australian Veterinary Emergency Plan. Disease Strategy, Lumpy skin disease, 1996.
- Cheng S, Fockler C, Barnes WM, Higuchi R: Effective Amplification of Long Targets from Cloned Inserts and Human Genomic DNA. Proc Natl Acad Sci USA 1994; 91(12): 5695-5699.
- Carr AC, Moore SD, Lucia, Alejandro ED. Robust quantification of polymerase chain reactions using global fitting. PLoS ONE. 2012; 7(5):e37640.
- Heine HG, Stevens MP, Foord AJ, Boyle DB. A capripoxvirus detection PCR and antibody ELISA based on the major antigen P32, the homolog of the vaccine virus H3L gene. J Immunol Methods 1999; 227:187-196.
- Tian H, Chen, Y, Wu JY, Shang YJ, Liu XT. Serodiagnosis of sheep pox and goat pox using an indirect ELISA based on synthetic peptide targeting for the major antigen P32. Viro J 2012; 7: 245.
- Fulzele S, Singh RK, Hosamani M, Mondal B, Yadav MP. Evaluation of duplex PCR and PCR-RFLP for diagnosis of sheep pox and goat pox. Int J Trop Med 2006; 1(2):66-70.
- Tulman ER, Afonso CL, Lu Z, Zsak L, Sur JH, et al. The genomes of sheep pox and goat pox viruses. J Virol. 2002; 76(12):6054-6061
- Allawe AB, Mohsen HH. Molecular characterization of lumpy skin disease virus in comparison with sheep pox vaccine. Res J Pharm Biol Chem Sci 2017, 8(2):2313.
- Le GC, Lamien CE, Fakhfakh E, Chadeyras A, Aba-Adulugba E et al. Capripoxvirus Gprotein-coupled chemokine receptor: a host-range gene suitable for virus animal origin discrimination. J Gen Virol 2009; 90:67–77.
- Ireland DC, Binepal YS. Improved detection of capripoxvirus in biopsy samples by PCR. J Virol Methods 1998; 74:1-7.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.