ISSN: 0973-7510
E-ISSN: 2581-690X
Rhizoctonia bataticola (Taub.) Butl. a omnipresent soil-borne fungus having pycnidial state as Macrophomina phaseolina (Tassi.) Goid., causes dry root rot disease in a wide range of hosts starting from cereal crops to horticultural and medicinal crops. The present study was aimed to find out genetic and morphological variability among the twenty five indigenous isolates of R. bataticola collected from geographically distinct regions (10 states) infecting 19 different crop plants. Sclerotial morphology of the isolates was studied and they were confirmed to species using species-specific primers. All the isolates were grouped into four different types of sclerotia. Genetic diversity of all these isolates was analysed using random amplified polymorphic DNA (RAPD) markers and inter simple sequence repeat (ISSR) markers. Ten random primers for RAPD and six primers for ISSR were tested for amplification of genomic DNA of R. bataticola isolates. Best six primers for RAPD and five primers for ISSR were further analysed. Unweighted pair-group method using arithmetic means (UPGMA) clustering of data showed that isolates did clearly differentiate to the specific group according to the geographical origins. ISSR markers were found more efficient than RAPD markers to correlate the genetic diversity with the grouping of isolates according to the geographical regions. But RAPD markers were found efficient todifferentiate R. bataticola isolates into different clusters according to their sclerotial morphology. It was the first study that investigated genetic relationships with two PCR-markers among different isolates of R. bataticola belongs to different hosts from India.
Dry root rot, Genetic diversity, ISSR, Rhizoctonia bataticola, RAPD, Sclerotial morphology.
Dry root rot of different agriculturally important crops viz. Black gram, Chick Pea, Cotton, Cowpea, Jute, Mungbean, Onion, Pea, Rice, Soybean, Spinach and Wheat etc. is caused by Rhizoctonia bataticola (Taub.) Butler in different parts of world including India 36. The perfect stage of R. bataticola is Thanatephorus cucumeris (Frank.) Donk. It also produces pycnidia and this stage is known as Macrophomina phaseolina (Maubl.) Ashby. R. bataticoa is very diversified and cosmopolitan soil borne fungus, producing different types of symptoms viz., blight, rot, wilt and decay 15. The hyphae are septate and 1.5 – 2.5 µm thick, and produce black and irregular sclerotia up to 100 µm in diam. resembles charcoal powder.
A number of workers have carried out a series of comprehensive studies with reference to its incidence, etiology and genetic variability 41,42,2,27.They found that isolates of R. bataticola are very diverse in nature with high phenotypic, pathogenic and genetic variation and even differences may occur in the different isolates from a single host. A high level variation in the sclerotial morphology of R. bataticola was reported by no. of workers 32,39.
PCR based molecular marker techniques were used to determine and evaluate the genetic diversity among different isolates of M. phaseoliona21. Genetic variability within the population was varied with the type of the marker 28. Restriction Fragment Length Polymorphism (RFLP) of rDNA-ITSregions2. Random Amplified Polymorphic DNA 1,2,44, Amplified Fragment Length Polymorphism(AFLP) 9,35 Universal Rice Primer PCR (URP-PCR) 18 Inter simple sequence repeats (ISSR) ,17, 30, 21 Repetitive Sequence-Based Polymerase Chain Reaction (Rep-PCR) 30 and Simple Sequence Repeats (SSR)6 were explored extensively to determine the variation of different isolates of R. bataticola mostly from single host plant. Of these techniques, RAPD is simple, low cost and requires small amount of material. Inter simple sequence repeat (ISSR) markers are powerful tools to reveal the variation in genome microsatellite regions. Both RAPD and ISSR markers have been successfully used to differentiate and identify many fungi 44, 21.
The objective of present investigation was to study the morphological and genetic variability among the different isolates of R. bataticola obtained from geographically distinct regions (10 states) over a range of 19 host plant species using RAPD and ISSR markers.
Collection of R. bataticola isolates
Twenty five isolates taken under consideration from different places and sources were collected from Indian Type Culture Collection (ITCC), Division of Plant Pathology, IARI, New Delhi (Table 1, Fig. 1). All the 25 isolates were taken for the morphological characterization and for variability studies using RAPD and ISSR markers.
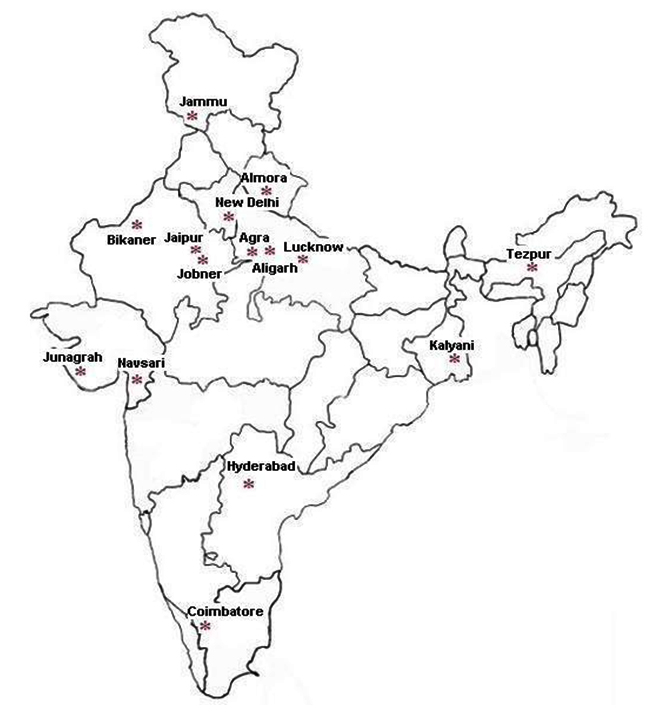 Fig. 1. Map* of India showing sampling regions of Rhizoctonia bataticola isolates
Fig. 1. Map* of India showing sampling regions of Rhizoctonia bataticola isolates(*Map was downloaded from www.cbse.aglasem.com)
Table (1):
Sources and location of different R. bataticola isolates.
S. No |
ITCC* Number |
Source |
Location |
|---|---|---|---|
1 |
1568 |
Jute |
Kalyani, West Bengal |
2 |
2474 |
Soil |
Hyderabad, Andhra Pradesh |
3 |
5196 |
Pea Seed |
Agra, Uttar Pradesh |
4 |
6037 |
Soybean |
Navasari, Gujarat |
5 |
6375 |
Chick Pea |
New Delhi, Delhi |
6 |
5519 |
Cotton root |
Junagadh, Gujarat |
7 |
249 |
Soybean |
New Delhi, Delhi |
8 |
5510 |
Balsam |
Aligarh, Uttar Pradesh |
9 |
5128 |
Onion |
Jaipur, Rajasthan |
10 |
250 |
black gram |
New Delhi |
11 |
4253 |
Cowpea Seed |
Jobner, Rajasthan |
12 |
1567 |
Soybean |
Kalyani, West Bengal |
13 |
6184 |
Spinach |
New Delhi, Delhi |
14 |
2204 |
Wheat |
New Delhi, Delhi |
15 |
652 |
Rice |
Coimbatore, Tamil Nadu |
16 |
5141 |
Soil |
New Delhi, Delhi |
17 |
481 |
Rice |
Jammu, Jammu &Kashmir |
18 |
5521 |
Castor root |
Junagadh, Gujarat |
19 |
2577 |
Balsam |
New Delhi, Delhi |
20 |
6267 |
Anola |
Bikaner, Rajasthan |
21 |
6256 |
Strawberry |
Jammu, Jammu &Kashmir |
22 |
3546 |
Mungbean |
Tezpur, Assam |
23 |
1569 |
Burning bush |
Kalyani, West Bengal |
24 |
5467 |
Rice |
Almora, Uttarakhand |
25 |
5499 |
Sarpgandha |
Lucknow, Uttar Pradesh |
*Indian Type Culture Collection
Morphological Identification of R. bataticola
The phenotypic characteristics of Rhizoctonia isolates were investigated based on sclerotial morphology.
Genomic DNA Extraction
Genomic DNA was extracted from frozen mycelium of R. bataticola based on Cetyltrimethyl ammonium bromide (CTAB) mini extraction method. 12
The DNA concentration and purity of the samples was determined with Nano Drop Spectrophotometer (Thermo Scientific).
Identification of R. bataticola isolates using species-specific primers:
The specific primers MpKFI (5’-CCGCCAGAGGACTATCAAAC- 3’) and MpKRI (5’-CGTCCGAAGCGAGGTGTATT-3’)4 were used during the course of this study for amplification of the isolates using PCR. Controls included DNA of isolates of Trichoderma harzianum, Colletotrichum gloeosporioides, Fusarium oxysporum and Chaetomium globosum .
PCR protocol was standardized varying the cycle number, annealing temperature and the concentration of target DNA. The optimized protocol is described here; PCR amplifications were performed in a total volume of 20 µl by mixing 2 µl of 10X Taq buffer, 0.5µldNTPs (100Mm dNTPs Mix), 5 pmol of each primer(0.5 µl), 1 U (0.3µl) of Taq DNA polymerase, 0.5 µl of 50 mMMgCl2and 35-50 ng of template DNA (2 µl). The PCR reaction was carried out for 25 cycles of denaturation at 95°C for 30s, followed by annealing at 56°C for 1 min, extension at 72°C for 2 min. and the final extension step of 72°C for 10 min.
Variability among the different isolates of R. bataticola using different RAPD Markers
DNA from 25 isolates was subjected to genetic diversity analysis by the RAPD method using 10 randomly chosen 10-base random primers. Among them six primers were chosen for further studies (Table 3). PCR reactions were carried out in 0.2 ml thin-walled PCR tubes with a total reaction volume of 25 µl containing 2.5¼l of 10X buffer, 2.5mM of each dNTPs( 0.5¼l) ,1¼l of primer, 0.5¼l MgCl2 (25mM) and 1U (0.2¼l) of Taq polymerase (Bangalore, Genei). The PCR amplification conditions were initial denaturation at 94°C for 3 min followed by 40 cycles of denaturation at 94°C for 1 min, primer annealing at 35°C for 1 min, primer extension at 72°C for 2 min, and a final primer extension at 72°C for 5 min. Six primers of RAPD were selected for final analysis based on informative banding patterns, clarity, and repeatability.
Variability among the different isolates of R. bataticola using different ISSR Markers
ISSR analysis was carried out amplifying the genomic DNA using six ISSR primers. Among them five primers were chosen for further studies (Table 3).The PCR amplification conditions were: Initial denaturation at 95°C for 5 min followed by 40 cycles of denaturation at 94°C for 1 min, primer annealing at 34°C for 1 min, primer extension at 72°C for 1 min, and a final primer extension at 72°C for 5 min. A total of six primers were screened initially. Five primers were selected for final analysis based on informative banding patterns, clarity, and repeatability.
Statistical analysis
RAPD and ISSR reproducible fragments were scored as present (1) or absent (0), and bands were entered in a computer file as a binary matrix, one for each molecular marker. The binary matrix was analyzed using the NTSYS-pc (Numerical Taxonomy and Multivariate Analysis System) package, version 2.0 to calculate the similarity values. After obtaining the similarity matrix, clustering was performed by sequential agglomerative hierarchical nested clustering (SAHN). The graphical representation of the cluster was obtained by using the unweighted pair group method of mathematical averages (UPGMA). Bootstrap analysis of the data was performed with 1000 replications using the WINBOOT program to estimate structural stability of clusters. Polymorphic information content (PIC) or average heterozygosity was calculated as per the formula of.34 Average heterozygosity (Hav) is obtained by taking the average of PIC values obtained for all the markers. Multiplex ratio (MR) for each assay was estimated by dividing the total number of bands (monomorphic and polymorphic) amplified by the total number of assays. Marker index (MI) was obtained by multiplying the average heterozygosity (Hav) with MR.28 A principal coordinate analysis (PCO) was performed EIGEN procedure in the NTSYSpc in order to highlight the resolving power of the ordination. To estimate the congruence among dendrograms, cophenetic matrices for each marker type were computed and compared using the Mantel matrix correspondence test. This test was performed using the MXCOMP procedure in the NTSYSpc.
The present research was undertaken to study the sclerotial and genetic variability among the different isolates of R. bataticola obtained from geographically distinct regions (10 states) infecting 19 different crop plants. All the isolates were divided into four different groups based on their high level variation in sclerotial shape, size and arrangement. RAPD and ISSR markers were used to estimate the genetic variability. ISSR markers were found more efficient than RAPD markers to correlate the genetic diversity with the grouping of isolates according to the geographical regions. But RAPD markers were found efficient to differentiate R. bataticola isolates into different clusters according to their sclerotial morphology.
Morphological Identification
Morphological features of 25 isolates of R. bataticola were observed by growing the isolates on PDA medium. Sclerotial morphology was considered to assess the possible variation among different isolates.
Variability in Sclerotial morphology
Among the isolates, the size of sclerotia ranged from 29.4-100 X 21.1-52.9 ¼m. Based on sclerotial shape, size and arrangements, the isolates were divided into four different groups (Table 2). The observations were made under the compound microscope and pictures were taken for further documentation (Fig. 2).
Table (2):
Variability in Sclerotial morphology in different R. bataticola isolates.
Groups |
Isolates |
Description of Sclerotia |
|---|---|---|
I |
2 and 15 |
Highly irregular and are connected by thick walled mycelium, 52.2 -73.6 X 42.1-52.6µm in size |
II |
14, 16, 17, 18, 19, 20, 21, 22, 23, 24 and 25 |
Round to irregular and free, 36.2-99.8 X 21.1- 47.4 µm in size |
III |
8 and 13 |
Round and free, 29.4-52.9 µm in size |
IV |
1, 3, 4, 5, 6, 7, 9, 10, 11 and 12 |
Round to irregular and are connected by thin mycelium, 42.1-100 X 31.6- 52.6 µm in size |
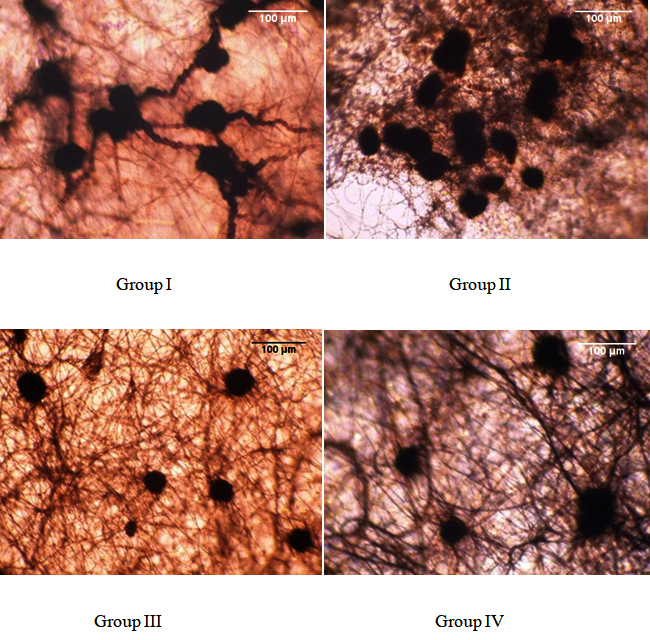 Fig. 2. Grouping of R. bataticola isolates based on their sclerotial morphology
Fig. 2. Grouping of R. bataticola isolates based on their sclerotial morphologyMolecular identification of R. bataticola based on species specific marker
Molecular characters of 25 isolates of R. bataticola were analyzed in order to resolve their identification based on species specific marker. The primers used yielded single amplified product of 350 bp with all the R. bataticola isolates tested (Fig. 3). The specificity of the primers was tested on representative species of common soil borne microbes like Trichoderma harzianum, Colletotrichum gloeosporioides, Fusarium oxysporum and Chaetomium globosum. The primer pair was found to be specific for R. bataticola as none of the other soil microbes tested could yield any amplification product under identical conditions of amplification.
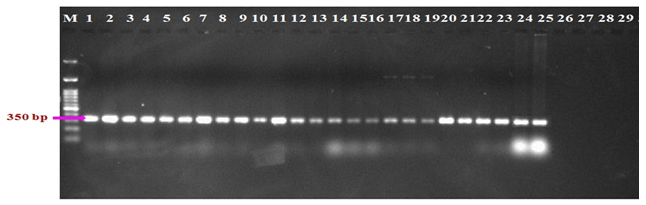 Fig. 3. Identification of R. bataticola with species specific primers. 1-25: R. bataticola isolates; 26: Trichoderma harzianum; 27:Colletotrichum gloeosporioides; 28: Fusarium oxysporum; 29: Chaetomium globosum
Fig. 3. Identification of R. bataticola with species specific primers. 1-25: R. bataticola isolates; 26: Trichoderma harzianum; 27:Colletotrichum gloeosporioides; 28: Fusarium oxysporum; 29: Chaetomium globosumRAPD Analysis of R. bataticola
A total of 238 bands were produced for Rhizoctonia isolates and all of them were found to be polymorphic. Maximum number (59) bands were obtained with OPA-11, followed by OPE-14 (45 bands), OPO-4 (41), OPN-4 (35), OPN-20 (30), OPS-18 (26) and all of them were polymorphic (Table 3). The size of the fragments varied from 0.4 to 2.5kb. The data obtained from RAPD analysis of 25 isolates of Rhizoctonia species with six primers was subjected to UPGMA analysis. A dendrogram was prepared using the similarity coefficient of RAPD marker. The dendrogram showed the presence of differences among R. bataticola isolates.
As per Fig. 4, one isolate (No. 1) of R. bataticola was not grouped. Rest all the isolates were grouped into four clusters.
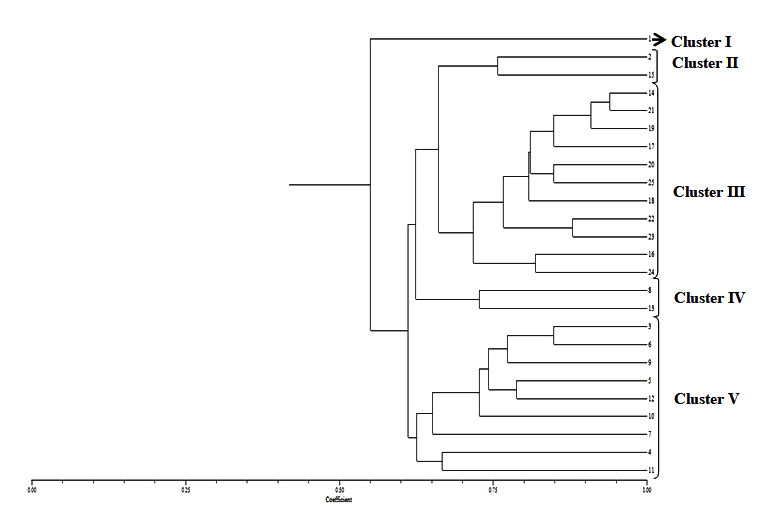 Fig. 4. Dendrogram constructed with UPGMA clustering method based on Random Amplified Polymorphic DNA (RAPD) -PCR for 25 isolates of R. bataticolafrom different geographical areas of India. Cluster I – Kalyani (1): Cluster II – Hyderabad(2), and Coimbatore (15); Cluster III – New Delhi (14, 16, 19) Jammu (17, 21), Bikaner (20), Lucknow (25), Junagadh (18), Tezpur (22), and Kalyani (23); Cluster IV – Aligarh (8) and New Delhi (13); Cluster V – Agra (3), Navasari (4), New Delhi (5, 7,10), Junagadh (6), Jaipur (9) and Kalyani (12)
Fig. 4. Dendrogram constructed with UPGMA clustering method based on Random Amplified Polymorphic DNA (RAPD) -PCR for 25 isolates of R. bataticolafrom different geographical areas of India. Cluster I – Kalyani (1): Cluster II – Hyderabad(2), and Coimbatore (15); Cluster III – New Delhi (14, 16, 19) Jammu (17, 21), Bikaner (20), Lucknow (25), Junagadh (18), Tezpur (22), and Kalyani (23); Cluster IV – Aligarh (8) and New Delhi (13); Cluster V – Agra (3), Navasari (4), New Delhi (5, 7,10), Junagadh (6), Jaipur (9) and Kalyani (12)The dendrogram obtained separated R. bataticola isolates (No. 2 and 15) completely from all other isolates and revealed more than 75 percent similarity among themselves. These isolates were characterized based on their sclerotial morphology which were large and highly connected. Isolate (Nos. 14,16,17,18,20,21,22,23,24,25) were having large, round to irregular and free sclerotia separated from all other isolates and show 72-94% similarity.
Similarly isolate 8 and 13 were characterized as the isolates having small, round and free sclerotia and also clustered together with 73% similarity. Isolates nos. 3,4,5,6,7,10,11 and 12 were clearly visualized in the same cluster with almost 63-84% similarity as these isolates having large, irregular and loosely connected sclerotia. Isolate 1 was not grouped with any other cluster based on RAPD analyses even though it has similar sclerotia later isolates.
This dendrogram revealed the presence of the variability among the different isolates by separating based on their sclerotial morphology.
ISSR Analysis of R. bataticola
A total of 250 bands were produced for Rhizoctonia isolates and all of them were found to be polymorphic. Maximum number (53) of bands were obtained with ISSR-1 and ISSR-2, followed by ISSR-3 (49 bands), ISSR-4 (46), ISSR-6 (49) and all of them were polymorphic (Table 3). The size of the fragments varied from 0.7 to 2.5kb. The data obtained from RAPD analysis of 25 isolates of R. bataticola with five primers was subjected to UPGMA analysis. A dendrogram was prepared using the similarity coefficient of ISSR marker. The dendrogram showed the presence of differences among R. bataticola isolates.
Table (3):
List of Random Amplified Polymorphic DNA (RAPD) and Inter-Simple Sequence Repeat (ISSR) primers and the resulted DNA polymorphisms applied to differentiate isolates of R. bataticola.
| Primers | Primer sequence (5’–3’) | Total Bands | Polymorphic Bands | Polymorphic Percentage | Monomorphic Bands |
|---|---|---|---|---|---|
| Selected RAPD primers | |||||
| OPA-11 | 5’–CAATCGCCGT–3’ | 59 | 59 | 100 | 0 |
| OPN-20 | 5’–GGTGCTCCGT–3’ | 32 | 32 | 100 | 0 |
| OPE- 14 | 5’–TGCGGCTGAG–3’ | 45 | 45 | 100 | 0 |
| OPO-4 | 5’–AAGTCCGCTC–3’ | 41 | 41 | 100 | 0 |
| OPN-4 | 5’–GACCGACCCA–3’ | 35 | 35 | 100 | 0 |
| OPS-18 | 5’–CTGGCGAACT–3’ | 26 | 26 | 100 | 0 |
| Total | 238 | 238 | —- | 0 | |
| Selected ISSR primers | |||||
| ISSR-1 | 5’–GAGAGAGAGAGA GAGAGAT–3’ | 53 | 53 | 100 | 0 |
| ISSR-2 | 5’–ACTGACTGACTGACTG–3’ | 53 | 53 | 100 | 0 |
| ISSR-3 | 5’–GAGAGAGAGAGA GAGAGAAT–3’ | 49 | 49 | 100 | 0 |
| ISSR-4 | 5’–ACCACCACCACCY–3’ | 46 | 46 | 100 | 0 |
| ISSR-6 | 5’–GATAGATAGATAGATA–3’ | 49 | 49 | 100 | 0 |
| Total | 250 | 250 | —- | 0 | |
OPA – Operon Series A primer, OPN – Operon Series N primer, OPE – Operon Series E primer, OPO – Operon Series O primer, OPS – Operon Series S primer
Isolates formed two main clusters with all the five primers tested (Fig. 5). ISSR primers were able to separate the R. bataticola isolates into 8 clusters (Table 4) on the basis from where they were collected. Cluster I consists the isolates of Kalyani (1, 12, 23) and Tezpur (22); Cluster II consists the isolates of Hyderabad (2), New Delhi (5) and Coimbatore (15); Cluster III consists the isolates of Agra (3), Aligarh (8) and New Delhi (7, 10, 16); Cluster IV consists the isolates of New Delhi (13, 14, 19); Cluster V consists the isolates of Navasari (4), Junagadh (6) and Jaipur (9); Cluster VI consists the isolates of Jobner (11), Junagadh (18) and Bikaner (20); Cluster VII consists the isolates of Jammu (17, 21); Cluster VIII consists the isolates of Almora (24) and Lucknow (25) and only Delhi Isolate (No. 5) was not cluster with other isolates of same origin. ISSR markers were found a much more efficient tool to differentiate variability among the different isolates of Rhizoctonia.
Table (4):
Different groups of R. bataticola isolates based on Inter-Simple Sequence Repeat (ISSR) -PCR analysis was found to coincided with their place of origin.
Group Numbers |
Source of Collection |
Isolate Numbers |
Similarity Coefficient |
|---|---|---|---|
I |
Kalyani and Tezpur |
1,12,22,23 |
0.5-0.6 |
II |
Hyderabad, New Delhi and Coimbatore |
2,5,15 |
0.55 |
III |
Agra, New Delhi, Aligarh, |
3,7,8,10,16 |
0.65 |
IV |
New Delhi |
13,19,14 |
0.55 |
V |
Navasari, Junagadh,Jaipur |
4,6,9 |
0.55 |
VI |
Jobner, Bikaner, Junagadh |
11,20,18 |
0.55 |
VII |
Jammu |
17,21 |
0.5-0.6 |
VIII |
Almora, Lucknow |
24,25 |
0.5-0.6 |
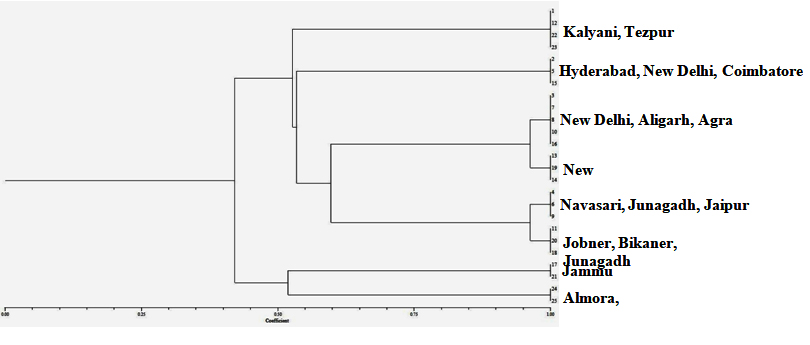 Fig. 5. Dendrogram constructed with UPGMA clustering method based on Inter-Simple Sequence Repeat (ISSR) markers for 25 isolates of R. bataticola from different geographical areas of India. Cluster I – Kalyani (1, 12, 23) and Tezpur (22); Cluster II – Hyderabad(2), New Delhi (5) and Coimbatore (15); Cluster III – Agra (3), Aligarh (8) and New Delhi (7, 10, 16); Cluster IV – New Delhi (13, 14, 19); Cluster V – Navasari (4), Junagadh (6) and Jaipur (9); Cluster VI – Jobner (11), Junagadh (18) and Bikaner (20); Cluster VII – Jammu (17, 21); Cluster VIII – Almora (24) and Lucknow (25)
Fig. 5. Dendrogram constructed with UPGMA clustering method based on Inter-Simple Sequence Repeat (ISSR) markers for 25 isolates of R. bataticola from different geographical areas of India. Cluster I – Kalyani (1, 12, 23) and Tezpur (22); Cluster II – Hyderabad(2), New Delhi (5) and Coimbatore (15); Cluster III – Agra (3), Aligarh (8) and New Delhi (7, 10, 16); Cluster IV – New Delhi (13, 14, 19); Cluster V – Navasari (4), Junagadh (6) and Jaipur (9); Cluster VI – Jobner (11), Junagadh (18) and Bikaner (20); Cluster VII – Jammu (17, 21); Cluster VIII – Almora (24) and Lucknow (25)Rhizoctonia bataticola is a non-host specific fungus causing dry root rot disease in a wide range of hosts and is responsible for causing losses in many number of cultivated and wild plant species.22,20 R. bataticola was reported on highly economic hosts viz., cotton, rice, maize, cucurbits, okra, and wheat 24,36 Physiological specialization of the fungus is not well demonstrated14 and reported a high level of variation in morphology, physiology and pathogenesis among different isolates obtained from different parts of the same plant.
Therefore, morphological and molecular variability were assessed in the present study. Twenty five isolates were collected from different host plants growing in different parts of the country. These isolates were characterized and grouped by their sclerotial morphology. All the isolates from Jute, Pea, Soybean, Chick Pea, Cotton, Balsam, Onion, black gram, Cowpea, Spinach, Wheat, Rice, Castor, Anola, Strawberry, Mungbean, burning bush and Sarpgandha were found variable in the shape and size of sclerotia. The size of the sclerotia was 29.4-100 X 21.1-52.9µm in the present study. The biggest sclerotia reported was 101.51µm in Gliricidia isolate of R. bataticola while cowpea isolate produced the smallest sclerotia (66.88 µm)10 where as 25 reported the size varied from 58.83 µm to 126.63 µm.29 also reported the morphological variability in R. bataticola collected from different pulse crops. Therefore, it is clear that variability exists in sclerotial morphology.
These morphologically characterized 25 isolates of R. bataticola were also assessed at their molecular level. PCR-RAPD analysis is one of the simplest tools available for the analysis of genetic differences at DNA level without demanding any prior information about the genomic sequences.
Molecular markers are useful tools for assessing variation rapidly within and among species11In this study, we have compared two different molecular marker systems, RAPD and ISSR, to define genetic relationships among isolates of R. bataticola, and to investigate which marker system can be more effectively used. In RAPD, a total of 238 bands were produced for R. bataticola isolates and all of them were found to be polymorphic. Maximum number (59) of bands were obtained with OPA-11 and all of them were polymorphic. The size of the fragments varied from 0.4 to 2.5kb. The higher level of polymorphism detected by RAPD markers revealed the selective capacity of this marker among R. bataticola isolates.40 In the present study, the variability based on PCR-RAPD was coinciding with the groups made based on sclerotial morphology.
The ISSR markers were successfully used to identify genetic differences in R. bataticola.17 The results of the present study clearly demonstrated that R. bataticola from different parts of the country were highly variable and ISSR markers are suitable to reflect the genetic diversity among the isolates. A high level of polymorphism in R. bataticola was found in the isolates collected from different hosts and regions using different molecular tools3,19,23,43 Many number of workers demonstrated the genetic diversity among M. phaseolina isolates .1,2,5,6,7,17,18,26,30,31,33,35 The high genetic diversity was observed not only in isolates from distinct hosts and geographical origins but also in isolates collected from a single host or single region.
The number of alleles at a locus and their frequency of distribution are responsible for Polymorphism thereby genetic variation in a population28. RAPD and ISSR markers target different portions of the genome resulted the difference in discrimination among the isolates.21 The genetic variation usually depends on the number of polymorphic bands obtained with each marker.8
It was the first study that investigated genetic relationships among different isolates of R. bataticola from different host plants from India and also that compared two PCR-markers to define genetic relationships among isolates of R. bataticola.
ACKNOWLEDGMENTS
The authors thank the Head, Division of Plant Pathology, Indian Agricultural Research Institute, New Delhi for the help in various aspects of this study. Financial support from Indian Council of Agricultural Research, Govt. of India, is gratefully acknowledged.
- Aboshosha, S.S., Attaalla, S.I., El-Korany, A.E., El-argawy E. Characterization of Macrophominaphaseolina isolates affecting sunflower growth in El-Behera governorate. International Journal of Agriculture & Biology, 2007; 9: 807″815.
- Aghakhani, M., Dubey, S.C. Determination of genetic diversity among Indian isolates of Rhizoctoniabataticolacausing dry root rot of chickpea. Antonie Leeuwenhoek, 2009; 96: 607-619.
- Almeida, A.M.R., Abdelnoor R.V., Arias, C.A.A., Carvalho, V.P., Martin S.R.R., Benato, L.C., Pinto, M.C., Carvalho, C.G.P. Genotypic diversity among brazilian isolates of Macrophominaphaseolina revealed by RAPD. FitopatologiaBrasileira, 2003; 28: 279″285.
- Babu, B.K., Saxena A.K., Srivastava, A.K., Arora, D.K. Identification and detection of Macrophominaphaseolina by using species-specific oligonucleotide primers and probe. Mycologia, 2007; 99: 797″803.
- Babu, B.K., Reddy, S.S., Yadav, M.K., Sukumar, M., Vijendra, M., Saxena, A.K., Arora, D.K. Genetic diversity of Macrophominaphaseolina isolates from certain agro-climatic regions of India by using RAPD markers. Indian Journal of Microbiology, 2010; 50: 199″204.
- Baird, R.E., Wadl, P.A., Allen T., McNeill D., Wang, X.W., Moulton J.K., Rinehart T.A., Abbas, H.K., Shier T., Trigiano, R.N. Variability of United States isolates of Macrophominaphaseolina based on simple sequence repeats and cross genus transferability to related genera within Botryosphaeriaceae. Mycopathologia, 2010; 170: 169″180.
- Beas-Fernandez, R., De Santiago-De Santiago, A., Hernandez- Delgado S., Mayek-Perez N. Characterization of Mexican and non-Mexican isolates of Macrophominaphaseolina based on morphological characteristics, pathogenicity on bean seeds and endoglucanase genes. Journal of Plant Pathology, 2006; 88: 53″ 60.
- Belaj, A., Satovic, Z., Cipriani G., Baldoni L., Testolin, R., Rallo L., Trujillo I. Comparative study of the discriminating capacity of RAPD, AFLP and SSR markers and of their effectiveness in establishing genetic relationships in olive. Theoretical and Applied Genetics, 2003; 107(4): 736-744.
- Brooker, N., Lord J.R., Long J., Jayawardhana, A. AFLP analysis of genetic diversity in charcoal rot fungal populations impacted by crop rotations. Communications in Agricultural and Applied Biological Sciences, 2008; 73: 7″19.
- Byadgi, A.S., Hedge R. Variation among the isolates of Rhizoctonia from different host plants. Indian Phytopathology, 1985; 38(2): 297–301.
- Chakravarthi, B.K., Naravaneni, R. SSR marker based DNA fingerprinting and diversity study in rice (Oryza sativa L.). African Journal of Biotechnology, 2006; 5: 684″688.
- Culling, K.W. Design testing of a plant specific PCR primer for ecological evolutionary studies. Molecular Ecology, 1992; 1: 233-240.
- Das, I.K., Fakrudin, B., Arora, D.K. RAPD cluster analysis and chlorate sensitivity of some Indian isolates of Macrophomina phaseolina from sorghum and their relationships with pathogenicity. Microbiological Research, 2008; 163: 215″224.
- Dhingra, O.D., Sinclair, J.B. Variation among isolates of Macrophomina phaseoli (Rhizoctonia bataticola) from the same soybean plant Phytopathology, 1972; 62: 1108″1115.
- Dhingra, O.D., Sinclair, J.B., Biology and pathology of Macrophomina phaseolina. ImprensaUniversitaria, Universidade Federal De Vicosa, Vicosa- 1978; 36.570, (pp. 166). Minas Gerais- Brazil
- Indera, K., Singh, T., Machado, C.C., Sinclair, J.B. Histopathology of soybean seed infection by Macrophomina phaseolina. Phytopathology, 1986; 76:532-535.
- Jana, T., Sharma, T.R., Singh, N.K. SSR-based detection of genetic variability in the charcoal root rot pathogen Macrophomina phaseolina. Mycological Research, 2005a; 109: 81″86.
- Jana, T., Singh, N.K., Koundal K., Sharma, T.R. Genetic differentiation of charcoal rot pathogen, Macrophomina phaseolina, into specific groups using URP-PCR. Canadian Journal Microbiology, 2005b; 51: 159″164.
- Jones R.W., Canada S., Wang H. Highly variable minichromosomes and highly conserved endoglucanase genes in the phytopathogenic fungus Macrophomina phaseolina. Canadian Journal Botany, 1998; 76: 694″698.
- Kunwar, IK, Singh T, Machado, CC, Sinclair JB. Histopathology of soybean seed and seedling infection by Macrophominaphaseolina. Phytopathology, 1986; 76:532–535
- Mahdizadeh, V., Safaie, N., Goltapeh, E.M. Diversity of Macrophomina phaseolina based on Morphological and Genotypic Characteristics in Iran. Plant Pathology Journal, 2011; 27(2): 128-137.
- Malaguti, G., Half a century of a plant pathologist in a tropical country Venezuela. Annual Review of Phytopathology, 1990; 28: 1–10.
- Mayek-Perez, N., Lpez-Castaeda, C., Gonzglez-Chavira M., Garcia- Espinosa, R., Acosta-Gallegos, J., de la Vega, O.M., Simpson J. Variability of Mexican isolates of Macrophomina phaseolina based on pathogenesis and AFLP genotype. Physiology and Molecular Plant Pathology, 2001; 59: 257″264.
- Mirza, J.H., Qureshi, M.S.A., Fungi of Pakistan, Department of Plant Pathology, University of Agriculture, 34 Faisalabad, Pakistan, (1978)
- Monga. D, Raj S. Cultural and pathogenic variations in the isolates of Rhizoctoniasp causing root rot of cotton. Indian Phytopathology 1994; 47(4):403–408.
- Omar, M.R., Abd-Elsalam, K.A., Aly, A.A., El-Samawaty, A.M.A., Verreet J.A. Diversity of Macrophominaphaseolina from cotton in Egypt: Analysis of pathogenicity, chlorate phenotypes, and molecular characterization. Journal of Plant Diseases and Plant Health Protection, 2007; 114: 196″204.
- Pande, S., Desai, S., Sharm M., Impact of Climate Change on Rainfed Crop Diseases: Current Status and Future Research Needs. In: National Symposium on Climate Change and Rainfed Agriculture, Indian Society of Dryland Agriculture, Central Research Institute for Dryland Agriculture, Hyderabad, India, February 2010; 18-20, 2010. pp. 55-59.
- Powell, W., Morgante, M., Andre, C., Hanafey, M., Vogel, J., Tingey, S., & Rafalski A. The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Molecular Breeding, 1996; 2(3): 225-238.
- Prameela, T., Singh, R.H. Cultural variation of Macrophomina phaseolina isolates collected from Vignamungo. Indian Phytopathology, 1998; 51(1): 292–293.
- Purkayastha, S., Kaur, B., Arora P., Bisyer, I., Dilbaghi, N., Chaudhury A. Molecular genotyping of Macrophomina phaseolina isolates: comparison of microsatellite primed PCR and repetitive element sequence-based PCR. Journal of Phytopathology, 2008; 156(6): 372-381.
- Rajkumar, F.B., Kuruvinashetti M.S. Genetic variability of sorghum charcoal rot pathogen (Macrophomina phaseolina) assessed by random DNA markers. Plant Pathology Journal, 2007; 23:45″50.
- Raut, J.C., Ingle, R.W. Variation in isolates of Rhizoctonia bataticola. Indian Phytopathology, 1989; 42: 506-508.
- Reyes-Franco, M.C., Hernandez-Delgado, S., Beas-Fernandez, R., Medina-Fernandez, M., Simpson J., Mayek-Perez, N. Pathogenic and genetic variability within Macrophomina phaseolina from Mexico and other countries. Journal of Phytopathology, 2006; 154: 447″453.
- Roldan-Ruiz, I., Dendauw, E., Van Bockstaele, A., Depicker., De Loose, M. AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Molecular Breeding, 2000; 6: 125-134.
- Saleh, A.A., Ahmed, H.U., Todd, T.C., Travers, S.E., Zeller, K.A., Leslie, J.F., Garrett, K.A. Relatedness of Macrophomina phaseolina isolates from tallgrass prairie, maize, soybean and sorghum. Molecular Ecology, 2010; 19: 79″91.
- Shehzad, S.A., Sattar, A., Ghaffar, A. Addition to the hosts of Macrophomina phaseolina. Pakistan Journal of Botany, 1988; 20: 151-152.
- Sinclair. J.B. Compendium of Soybean disease. 2nd Ed., American Phytopathology Society, St. Paul, Minnesota, USA. (1982)
- Su G., Suh S.O., Schneider R.W., Russin J.S. Host specialization in the charcoal rot fungus, Macrophomina phaseolina. Phytopathology, 2001; 91: 120″126.
- Sundravadana, S., Alice, D.,Thirumurugan, S. Exploration of variability in colony morphology and virulence of Rhizoctonia bataticola isolates causing dry root rot of pulses. Global Journal of Bio-Science and Biotechnology, 2012; 1(1): 91-97.
- Sundravadana, S., Thirumurugan, S., Devadason, A. Exploration of molecular variability in Rhizoctonia bataticola, the incitant of root rot disease of pulse crops. Journal of Plant Protection Research, 2011; 51(2): 184-189.
- Tripathi N.N., Sharma, B.K. Incidence of chickpea dry root rot (Rhizoctonia bataticola) in Southern Haryana. International chickpea newsletter, 1983; 8: 22-23.
- Trivedi, S., Gurha, S.N. Status of some soil borne pathogens infecting chickpea in Bundelkhand region of Uttar Pradesh. Indian Journal Pulses Research, 2006; 19: 88-90.
- Vandemark, G., Martinez, O., Pecina, V., Alvarado, M.D. Assessment of genetic relationships among isolates of Macrophomina phaseolina using a simplified AFLP technique and two different methods of analysis. Mycologia, 2000; 92: 656″ 664.
- Zade, M.S.A., Zadeh, N.A., Nejad, R.F., Rezaee S., Mahmoudi B. RAPD marker as a criterion to study differentiation of isolates of Rhizoctonia solani and Rhizoctonia bataticola (Macrophomina phaseolina). Phytopathology, 2009; 99: 113″113.
© The Author(s) 2016. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


