ISSN: 0973-7510
E-ISSN: 2581-690X
The geothermal springs are said to contain the greatest diversity of undiscovered microorganisms, making them the best source for enzymes with economic significance. The untapped microbial diversity living in the geothermal springs can be mined for novel genes, bioactive substances, and industrially significant biocatalysts using the metagenomics technique. Metagenome was extracted from soil samples of various geothermal springs of India. Metagenome was screened for various carbohydrate degrading enzymes (amylase, cellulase, xylanase, amylopullulanase) using degenerate primers-based Polymerase chain reaction amplifications. Further amplicons were cloned, sequenced and analysis of data was done using various bioinformatics tools, e.g., Blast analysis, Protparam and phre2 program. We have isolated numerous enzymes, including cellulase, amylase, amylopullulanase, and xylanase, from diverse geothermal spring in different parts of India using sequence and function-based metagenomics. In this study, we describe the metagenomics-based isolation of a thermostable amylase from the geothermal spring of Odisha. The amylase gene (1503 bp) was amplified using the metagenome as a template using degenerate primers and cloned into the linearized T vector. The putative gene was likely to encode a protein of 469 amino acids with a molecular weight of 53895.05 Da with pI-7.78. Sequence analysis showed its maximum identity of 98.95% with Bacillus licheniformis alpha-amylase gene. Homology modeling of the amylase protein was done using the phyre2 program, which shows it belongs to the (trans) glycosidase superfamily and contains the catalytic TIM alpha/beta-barrel fold. Hence, we can conclude that geothermal springs are hotspots for the mining of industrially robust biocatalysts.
Metagenomics, Alpha-Amylase, Geothermal Spring, Thermostable Enzyme, Industrial Biocatalyst
Geothermal springs are unique habitats of microflora mainly thermophilic microorganisms that can be searched for novel genes, bioactive compounds, and important hydrolytic enzymes for industry.1 The microbes living in thermal springs have adapted to this way of life to survive at high temperatures. They contain thermo-tolerant enzymes that are naturally able to stabilize and function at high temperatures and in a wide range of pH levels.2 Novel thermostable biocatalysts have been identified from thermal springs, including lipase, xylanase, amylase, cellulase, and protease.3-4 The most prevalent source of renewable carbon source is lignocelluloses which is present on earth as a major source of biomass waste and causes soil and water pollution and mostly made up of cellulose, hemicellulose, and lignin. With the help of Carbohydrate degrading enzymes these biopolymers were converted into simpler sugar forms that can be used to produce useful commercial products like bioethanol. Carbohydrate hydrolyzing enzymes like cellulase, pectinase, xylanase, and amylase are used in a variety of biotechnology procedures.5 Amylases are a class of economically significant catalysts that are widely used in many different industries, including the food business, fermentation, textiles, ethanol production, detergent industry, paper and pulp industry, and pharmaceutical industry.6 Amylolytic enzymes account for 25% of the global enzyme market and are mostly extracted from several strains of the Bacillus species for commercial applications (Bacillus licheniformis, Bacillus stearothermophilus, and Bacillus amyloliquefaciens). Due to their thermolabile nature, earlier amylases isolated from their mesophilic counterparts had numerous issues and were therefore ineffective for industrial usage. Researchers then began looking for amylolytic enzymes in thermophilic bacteria belonging to various taxonomic groupings since these microorganisms need these enzymes to break down natural polymeric substrates to survive. Amylases are divided into endoamylases (α-amylases), exoamylases (β-amylases), and other debranching enzymes based on the activity they perform. Alpha-amylase (EC 3.2.1.1), a member of the glycosyl hydrolase family GH13, is one of the most significant members of the amylase group. It functions as an endolytic enzyme that hydrolyzes the a-1, 4-glycosidic bond to generate low molecular weight dextrins, glucose, maltotriose, and maltose.7 These thermotolerant amylases have many benefits for many industries, including improving substrate solubility, cutting cooling costs, increasing diffusion rates, and remaining stable. Saxena et al. found that it was resistant to denaturing chemicals and reduced microbial contamination.8
Several thermostable/thermotolerant amylolytic catalysts have been examined for their hydrolytic activity at higher temperatures and their probable application in starch liquefaction and saccharification (100-110°C) to produce profitable items such as glucose, sugar syrup.9,10 Raw dry starch granules are heated at 105°C for 5 minutes, then at 95°C for 1 hour at pH 6; in the saccharification step, the solution is treated at 60°C at pH 4.5 for 3 hours.11 Most of the amylases are not functional at much higher temperatures, and for the saccharification step, pH adjustment and cooling become indispensable, so searching for a thermoacidophilic amylase becomes imperative. Amylases are majorly produced by microbes, animals and plants. Microorganisms, especially bacteria, among these organisms provide benefits for the production of enzymes due to their rapid growth and easy genetic manipulation.12 A thermostable amylase having an optimal temperature of 100°C from Pyrococcus furiosus was reported in 1990.13 Recently, an acid-stable amylolytic enzyme that operates at 80 °C was investigated and found to be a promising industrial catalyst14. Han et al. reported a thermo-active pullulanase (100 °C) from Thermococcus kodakarensis KOD115. However, the most thermoactive starch hydrolyzing enzyme (active at 120°C) is reported from Methanococcus jannaschii.16 Recently, a thermostable amylase was isolated from hot springs of Kashmir, having optimal activity at 70°C and at pH 7.0. Production of this enzyme was verified with response surface methodology and showed an increase in the enzyme activity with the addition of calcium and magnesium ions.17 The amylolytic enzymes regained their dominance as industrial catalysts owing to impetus in the bioenergy sector, where starch can be used as a source of starting material for bioethanol production. Even while thermophilic bacteria can produce enzymes of industrial use, only 0.1-1% of them can be grown using conventional techniques, making it challenging to isolate novel enzymes with superior features using culture methods. To get beyond the constraints of traditional culture techniques, metagenomic technologies have been devised. This technique is based on the study of whole community genome present in environmental samples and can be used to prospect novel enzymes that cultured techniques could never find. In this paper, we have isolated a gene coding for a thermophilic amylase from the geothermal spring of Odisha using a sequence-based metagenomics approach and cloning and sequence analysis were done to analyze the gene.
The reagents used in metagenome isolation, PCR, and cloning were all of the molecular grades. Bacterial strains like E. coli DH5α were purchased from Himedia Laboratories (HTBM017). GeNei Instant Cloning Linearized T-vector was used as a cloning vector. Himedia plasmid miniprep kit (MB508), as well as a manual plasmid isolation method,18 was used in plasmid preparation.
Sample collection
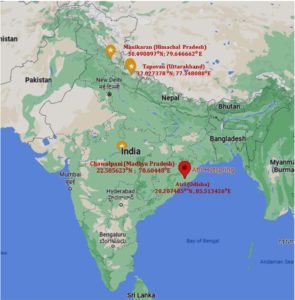
Samples of soil and sediment were taken from geothermal springs in Himachal Pradesh(30.027378°N and 77.348088°E), Uttarakhand (30.490897°N and 79.646662°E), Odisha (20.207485°N and 85.513426°E), and Madhya Pradesh (22.585623°N and 78.60448°E) (Figure 1) that range in temperature from 50°C to 95°C and have a pH of 6.8 to 8.0. The soil and sediment samples were collected in sterile sample collection bottles and stored at 4°C until further use.
Metagenome isolation
Using two published techniques by Zhou et al.19, Verma and Satyanarayan20, and the HiPurA Soil DNA Purification Kit (MB542), metagenomic DNA was recovered from different soil samples.
Synthesis of oligo-primers for the amylase gene and amplification using PCR
The degenerate primers for each enzyme gene were created using the gene’s nucleotide data from NCBI. However, in this paper, only amylase gene amplification, cloning, and sequence analysis are covered. The PCR oligo-primers designed for amylase genes based on the conserved amino and carboxyl terminals of already reported thermophilic amylases (FP-ATTGCTSACGCTGTTATTTGCGC and RP-CTTGGCTARAATTTKGCTYTTTTG *S=C/G, R=A/G, K=G/T, Y=C/T) were synthesized and PCR amplification was done by Himedia 2x PCR TaqMixture (MBT061) and PromegaGoTaq (Eppendorf MasterCycler EP Gradient Thermal Cycler 96 well). The PCR conditions were standardized by performing an annealing temperature gradient of 0.5°C ranging from 56°C to 62°C, and the optimum amplicon density was achieved at 58°C. Initial denaturation at 94°C for 5 minutes was followed by 35 cycles of denaturation at 94°C for 30 seconds, annealing at 58°C for 30 seconds, extension at 72°C for 2 minutes, and final extension at 72°C for 10 minutes.
Cloning of the PCR amplified amylase gene and bioinformatics analysis
The cloning of the PCR amplicon was performed by the GeNei Instant Cloning Kit (KT63A), which contains T4 DNA ligase and 2 x ligation buffers, which enhance the ligation process. After DNA ligation in the linearized T-vector, a highly competent E. coli DH5a (HTBM017) strain is used as a host for the cloning of the gene. E. coli DH5 cells were transformed with plasmids by the CaCl2 method.21 The clones were selected using the blue/white screening method.16 The white clones were sub-cultured for plasmid DNA isolation. After isolation, the plasmids were extracted from the electrophoresis gel by using the GeNei gel extraction kit (KT154L). These plasmids were sequenced by Apical Scientific Sdn Bhd Malaysia and, afterwards sequencing data were analyzed for the presence of amylase genes using the ORF finder tool, Protparam, nucleotide and protein BLAST analysis, and the Phyre 2 program for homology modeling.
In the current study, soil and sediment samples were collected from different geothermal springs in India to isolate carbohydrate-degrading enzymes such as cellulase, amylase, amylopullulanase, and xylanase (Table). The concoction of these carbohydrate metabolizing enzymes can be used to treat lignocellulosic biomass, which is present on earth as major waste biomass and converted into bioethanol and other consumable products.22 Through sequence and function-based metagenomics, we have isolated four different biocatalysts: amylase, xylanase, cellulase, amylopullulanase, and a transporter protein (auxin efflux carrier). High-quality metagenomic DNA was isolated from various soil samples of Chawalpani (Madhya Pradesh), Manikaran (Himachal Pradesh), Atri (Odisha), and Tapovan (Uttarakhand) using the HipurA soil DNA purification Kit (Himedia MB542) (Supplementary Figure 1). In this study, the authors focused on isolation and in silico studies on thermostable alpha-amylase genes.
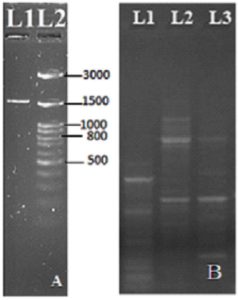
Figure 2. a) PCR amplicon of amylase gene ODS sample (Orissa sample-L1, Ladder L2), b) PCR amplicon of amylase gene present in different DNA samples (L1- TP1-1, L2- TP1-2, L3-TP2-2)
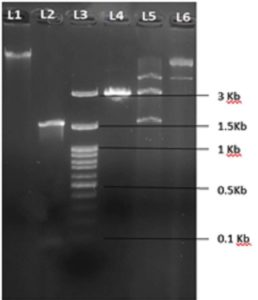
Figure 3. Recombinant plasmid of clone4, which was selected from the blue-white screening L1- metagenome-ODS DNA, L2- PCR amplicon of Amylase gene, L3-100 bp DNA ladder, L4-T vector without insert, L5- Recombinant plasmid digested with EcoRI restriction enzyme, L6- T vector along with insert
Out of all the isolated metagenome samples, only the Tapovan (TP) and Odisha (ODS) samples showed the presence of amylase genes (Figure 2a and 2b). TP1-1, TP1-2, and TP2-2 all had multiple bands, but the ODS sample only had one. The size of the ODS band was found to be around 1503 bp (Table). For further cloning in T-vector, a PCR amplicon from ODS from Atri geothermal spring in Odisha was chosen. After cloning, plasmids were isolated from recombinant strains using the Himedia plasmid miniprep kit (MB508) as well as from the manual plasmid isolation method.18 Recombinant plasmid of clone 4 of ODS sample was digested with EcoRI restriction enzyme to validate whether recombinant plasmid carry amylase gene or not (Figure 3). After confirmation, the recombinant plasmid of clone 4 was sent for sequencing. The cloned fragment was sequenced and contained 1503 bp (Genbank accession no. OP019318) (Supplementary Figure 2). Further instability index (II) was computed to be 23.98 and the protein was classified as stable. Homology modelling of cloned alpha-amylase was done using the phyre2 program that had 94% structural identity with template protein d1ob0a2 and contained a TIM barrel and three conserved carboxylic acid amino acids, i.e., Asp, Glu, and Asp as catalytic residues23 (Supplementary Figure 3). These catalytic residues play an important role in catalyzing starch hydrolysis.
Table:
Different enzymes and transporter proteins isolated through metagenomics approach.
No. |
Source |
Longitude |
Latitude |
Putative function |
pH |
Temperature ᵒC |
Size of gene |
% Identity |
|---|---|---|---|---|---|---|---|---|
1 |
Atri geothermal spring, Odisha |
20.15˚ |
85.30˚ |
Amylase |
– |
57 |
1.5 kb |
98.95% Bacillus lichniformis 584 alpha-amylase gene |
2 |
Atri geothermal spring, Odisha |
20.15˚ |
85.30˚ |
Xylanase |
– |
57 |
526 bp |
91.12% Bacillus halodurans endo-1,4- beta-xylanohydrolase (xyn10A) gene |
3 |
Tapta kund geothermal spring, Uttarakhand |
30.44˚ |
79.29˚ |
Auxin Efflux Carrier |
8.0 |
80 |
522 bp |
98.15% Multispecies of Bacillus |
4 |
Tapovan geothermal spring, Uttarakhand |
30.57˚ |
79.57˚ |
Amylopullulanase |
8.0 |
80 |
2.37 kb |
97.63% Cohnella sp. A01 amylopullulanase |
5 |
Tapovan geothermal spring, Uttarakhand |
30.57˚ |
79.57˚ |
Cellulase |
8.0 |
80 |
926 bp |
97.76% Geobacillus kaustophiluosGBlys |
Physicochemical studies revealed that the geothermal springs of different regions of India have a varying range of temperature and pH because they have different types of microorganisms. These microbes, being adapted to extreme conditions, are excellent sources of various novel biomolecules, antibiotics, and industrially important enzymes1. In present study cloned amylase gene isolated from Tapovan geothermal spring exhibited maximum identity of 98.95% with Bacillus licheniformis SRCM 103914, B. licheniformis strain CP6, and B. licheniformis 584, with 81% query coverage at the nucleotide level24 (Supplementary Figure 4). ORF predicted amino acid sequence exhibited 95.23 % identity with Bacillus licheniformis 1, 4-alpha-D-glucan glucanohydrolase25 (Supplementary Figure 5.). The complete nucleotide sequence of a Bacillus licheniformis gene coding for heat and pH-stable alpha-amylase: comparison of the amino acid sequences of three bacterial liquefying alpha-amylases deduced from DNA sequences,25 and 77.14% identity with B. amyloliquefaciens protein with 84% query coverage26. The only modification found was at the C-terminal of the protein, while the N-terminal was conserved. It has been shown that the alpha-amylase enzymes of Bacillus licheniformis and Bacillus amyloliquefaciens are conserved at their N-terminal.27 The thermophilic-amylase enzyme is an important enzyme for paper and pulp, as well as other starch processing industries (glucose, maltose, chocolate and corn syrup production, bakery, and food).28 Richardson et al. described the isolation of thermo-acid amylase genes using a metagenomics approach and demonstrated excellent activity at high temperatures and acidic pH ranges.29 A hyper thermostable amylase gene belong to GH57 family was mined from metagenomic fosmid library, prepared from a black smoker chimney sample 4143-1 from the hydrothermal vent of Mothra. It showed optimum activity at temperature 90°C and optimum pH of 7.5.30 Yun and his coworkers reported some alkalophilic amylases recovered from soil metagenomes.31 A thermophilic novel amylase (optimal activity at 60°C) was also mined from the soil metagenomic library of the Kerala Western Ghats.32 The thermostable alpha-amylase gene isolated in this study has a high potential for starch processing and can be used in the food industry. Motahar et al. isolated two thermostable enzymes which are α-amylase (PersiAmy2) and amylopullulanase (PersiPul1) from raw cow rumen metagenome and used them as a cocktail for fortification of bread using quinoa protein.33 Amylase enzymes along with other lignocellulosic waste degrading enzymes were isolated using functional and sequence-based metagenomics approach in our laboratory. Endo-1, 4-beta-xylanohydrolase (xyn10A) gene was isolated from Atri geothermal spring Odisha sample, exhibiting 91.12% similarity to Bacillus halodurans (Genbank accession no. OP019319). Amylopullulanase, and cellulase genes were isolated from Tapovan geothermal spring, Uttarakhand, using a metagenomic approach. Isolated amylopullulanase gene (Genbank accession no. OP080606) exhibited 97.63% similarity to the amylopullulanase gene of Cohnella sp. A01. A thermostable cellulase gene was also mined using a sequence-based metagenomics approach (Genbank accession no. OP080607) and exhibited 97.76% similarity to Geobacillus kaustophilous GBlys. Auxin efflux carrier (AEC) gene was isolated from Tapta Kund geothermal spring of Uttrakhand. Auxin efflux carrier (AEC) is a growth hormone transporter protein of bacteria, which is present in the rhizospheric microflora of plants growing near geothermal springs. The AEC gene exhibited 98.15% similarity to multispecies of Bacillus sp. A detailed mechanism of the functioning of this AEC transporter was published by our group in 2021.34 All these thermoactive enzymes have a wide range of applications in industries such as textiles, pulp and paper, food and beverages, baking, detergent, and biofuel.35 This study indicates that Atri and geothermal springs of Himalayan region can be a potential hot spot for exploring industrially robust enzymes.
Many sectors, like the food, beverage, detergent, textile, pulp and paper, and biofuel industries, will benefit from the metagenomics technique through the isolation of novel robust enzymes from natural geothermal springs. Most of the enzymes are unable to reach commercial application standards in the harsh conditions imposed by industrial bioprocesses and rapidly lose activity. The enzyme alpha-amylase, as well as other carbohydrate hydrolyzing enzymes such as cellulase, xylanase, and amylopullulanase, was retrieved using a metagenomics technique in this study, and the combination of these enzymes can be used to treat lignocellulosic biomass for bioethanol production. These enzymes’ potential can be increased through site-directed mutagenesis and efficiently exploited for a variety of purposes, such as enhancing the loaf’s quality and making it gluten-free, used in chicken feed processing and starch saccharification.
Additional file: Additional Figure S1-S5.
ACKNOWLEDGMENTS
The authors acknowledges UGC-Senior Research Fellowship, UGC-non-NET Ph.D. fellowship and the Project Assistant Fellowship from CST-UP.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
MS contributed to conceptualization of the problem and fund procurement. GC, VK. MA, AK, AS collected samples and carried out the experimental work. PK assisted in experimental work. GC and MS collected the data and wrote the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
This study was supported by Uttar Pradesh Council of Science & Technology with grant number CST/8275.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies with human participants or animals performed by any of the authors.
- Saxena A, Yadav AN, Rajawat M, et al. Microbial Diversity of Extreme Regions: An Unseen Heritage and Wealth. Indian J Genet Resour. 2016;29(3):256.
Crossref - Verma P, Yadav AN, Shukla L, Saxena AK, Suman A. Hydrolytic enzymes production by thermotolerant Bacillus altitudinis IARI-MB-9 and Gulbenkianiamobilis IARI-MB-18 isolated from Manikaran hot springs. Int J Adv Res. 2015;3:1241-1250.
- Panda MK, Sahu MK, Tayung K. Isolation and characterization of a thermophilic Bacillus sp. with protease activity isolated from hot spring of Tarabalo, Odisha, India. Iran J Microbiol. 2013;5(2):159-165.
- Parsai S, Choure K, Srivastava A, Rai P K, Agnihotri V, Singh Gour S. Lipase producing thermophilic bacteria isolation and characterization from hot springs of Central India. Natl J LIFE Sci. 2020;17(2):91-96.
Crossref - Gong G, Zhou S, Luo R, Gesang Z, Suolang S. Metagenomic insights into the diversity of carbohydrate-degrading enzymes in the yak faecal microbial community. BMC Microbiol. 2020;20:302-310.
Crossref - Sharma A, Satyanarayana T. Microbial acid-stable α-amylases: Characteristics, genetic engineering, and applications. Process Biochem. 2013;48(2):201-211.
Crossref - Sindhu R, Binod P, Madhavan A, et al. Molecular improvements in microbial Α-amylases for enhanced stability and catalytic efficiency. Bioresour Technol. 2017;245(Part B):1740-1748.
Crossref - Saxena RK, Dutt K, Agarwal L, Nayyar P. A highly thermostable and alkaline amylase from a Bacillus sp. PN5. Bioresour Technol. 2007;98(2):260-265.
Crossref - Elleuche S, Antranikian G. Starch-hydrolyzing enzymes from thermophiles. Thermophilic Microbes Environmental and Industrial Biotechnology. 2013:509-533.
Crossref - Elleuche S, Schroder C, Sahm K, Antranikian G. Extremozymes-biocatalysts with unique properties from extremophilic microorganisms. Curr Opin Biotechnol. 2014;29:116-123.
Crossref - Bertoldo C, Antranikian G. Starch-hydrolyzing enzymes from thermophilic archaea and bacteria. Curr Opin ChemBiol. 2002;6(2):151-160.
Crossref - Ardhi A, Sidauruk AN, Suraya N, Pratiwi NW, Pato U, Saryano. Molecular identification of amylase-producing thermophilic bacteria isolated from Bukit Gadang Hot Spring, West Sumatra, Indonesia. Biodiversitas. 2020;21(3):2085-4722.
Crossref - Koch R, Zablowski P, Spreinat A, Antranikian G. Extremely thermostable amylolytic enzyme from the archaebacterium Pyrococcus furiosus. FEMS Microbiol Lett. 1990;71(1-2):21-26.
Crossref - Sharma A, Satyanarayana T. Cloning and expression of acid stable, high maltose-forming, Ca2+ independent alpha-amylase from an acidophile Bacillus acidicola and its applicability in starch hydrolysis. Extremophiles. 2012;16(3):515-522.
Crossref - Han T, Zeng F, Li Z, et al. Biochemical characterization of a recombinant pullulanase from Thermococcus kodakarensis KOD1. Lett Appl Microbiol. 2013;57(4):336-343.
Crossref - Kim JW, Flowers LO, Whiteley M, Peeples TL. Biochemical confirmation and characterization of the family-57-like alpha-amylase of Methanococcus jannaschii. Folia Microbiol (Praha). 2001;46:467-473.
Crossref - Sharif S, Shah AS, Fariq A, Jannat S, Rasheed S, Yasmin A. Optimization of amylase production using response surface methodology from newly isolated thermophilic bacteria. Helion. 2023;9(1):e12901.
Crossref - Sambrook J, and Russell DW. Purification of Nucleic Acids by Extraction with Phenol: Chloroform. Cold Spring HarbProtoc1. 2006.
Crossref - Zhou J, Bruns MA, Tiedje JM. DNA recovery from soils of diverse composition. Appl Environ Microbiol. 1996;62(2):316-322.
Crossref - Verma D, Satyanarayana T. An improved protocol for DNA extraction from alkaline soil and sediment samples for constructing metagenomic libraries. Appl Biochem Biotechnol. 2011;165:454-464.
Crossref - Hanahan D. Studies on Transformation of Escherichia coli with Plasmids. J Mol Biol. 1983;166:557-580.
Crossref - Singh N, Singh V, Singh MP. Microbial degradation of lignocellulosic biomass for bioenergy production: A metagenomic-based approach. Biocatal Biotransformation. 2022; 41(1):15-25.
Crossref - Machius M, Declerck N, Huber R, Wiegand G. Kinetic stabilization of Bacillus licheniformis a-amylase through introduction of hydrophobic residues at the surface. J Biol Chem. 2003;278(13):11546-11553.
Crossref - Stephens MA, Ortlepp S A, Ollington J F, McConnell DJ. The nucleotide sequence of the 5’ region of the B. licheniformis alpha-amylase gene: comparison with the B. amyloliquefaciens gene. J Bacteriol. 1984;158:369-372.
Crossref - Yuukt T, Nomura T, Tezuka H, et al. Complete nucleotide sequence of a gene coding for heat- and pH-stable a-amylase of Bacillus licheniformis: Comparison of the amino acid sequences of three bacterial liquefying a-amylases deduced from the DNA sequences. J Biochem. 1985;98(5):1147-1156.
Crossref - Takkinen K, Pettersson RF, Kalkkinen N, Palva I, Soderlund H, Kaariainen L. The amino acid sequence of alpha-amylase from Bacillus amyloliquefaciens was deduced from the nucleotide sequence of the cloned gene. J Biol Chem. 1983;258(2):1007-1013.
Crossref - Kuhn H, Fietzek PP, Lampen JO. N-terminal amino acid sequence of Bacillus licheniformis a-amylase: Comparison with Bacillus amyloliquefaciens and Bacillus subtilis enzymes. J Bacteriol. 1982;149(1):372-373.
Crossref - Mojsov K. Microbial a-amylases and their industrial applications. Int J Manag. 2012;2:583-609.
- Richardson TH, Tan X, Frey G, et al. A novel, high performance enzyme for starch liquefaction discovery and optimization of a low pH, thermostable a-amylase. J BiolChem. 2002;277(29).
Crossref - Wang H, Gong Y, Xie W, et al. Identification and Characterization of a Novel Thermostable gh-57 Gene from Metagenomic Fosmid Library of the Juan De Fuca Ridge Hydrothemal Vent. Appl Biochem Biotechnol. 2011;164(8):1323-1338.
Crossref - Yun J, Kang S, Park S, et al. Characterization of a novel amylolytic enzyme encoded by a gene from a soil-derived metagenomic library. Appl Environ Microbiol. 2004;70(12):7229-7235.
Crossref - Vidya J, Swaroop S, Singh SK, Alex D, Sukumaran RK, Pandey A. Isolation and characterization of a novel a-amylase from a metagenomic library of Western Ghats of Kerala, India. Biologia (Bratisl). 2011;66(6):939-944.
Crossref - Motahar SFS, Salami M, Ariaeenejad S, et al. Synergistic Effect of Metagenome-Derived Starch-Degrading Enzymes on Quality of Functional Bread with Antioxidant Activity. Starch. 2021;74(1-2):2100098.
Crossref - Rai N, Kumar V, Sharma M, Akhter Y. Auxin transport mechanism of membrane transporter encoded by AEC gene of Bacillus licheniformis isolated from metagenome of Tapta Kund Hotspring of Uttrakhand, India. Int J Biol Macromol. 2021;185:277-286.
Crossref - Singh R, Kim SW, Kumari A and Mehta PK. An overview of Microbial a-amylase and recent Biotechnological applications. Curr. Biotechnol. 2022; 11(1): 11-12.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.