ISSN: 0973-7510
E-ISSN: 2581-690X
Haemophilus are fastidious Gram negative bacilli, which require factor X (hemin), factor V (NAD), or both for their growth. Haemophilus influenzae is the type species, and is considered to be the most pathogenic. They are associated with many invasive infections including meningitis, epiglottitis, pneumonia, and otitis media. Serotype b is most commonly associated with infections. Haemophilus species isolated from patients in a tertiary care centre in South India were studied. Identification, serotyping and biotyping were done and antibiotic susceptibility test was performed. The incidence of H. influenzae infections in our study was 65.3 cases/100,000 persons. Serotype b was the most common (66.67%), followed by non typeable H.influenzae (NTHi) (25%). Most isolates from adults were type b, while all isolates from pediatric population were non typeable. The most common biotype was type II, followed by type I and type III. Three of 24 isolates were β lactamase producers (12.5%). One isolate was β lactamase negative Ampicillin resistant (BLNAR). Resistance to ampicillin was 16.67%. Resistance to cephalosporins and fluoroquinolones was low (4-10%). Co-trimoxazole resistance was found to be very high (75%). All isolates were susceptible to azithromycin, tetracycline, chloramphenicol and meropenem. No isolates of H.influenzae type b were obtained from the paediatric population which may be due to the introduction of Hib vaccine. The increase in resistance to commonly used antibiotics is worrisome, especially penicillins and co-trimoxazole. Use of co-trimoxazole in empirical therapy of upper and lower respiratory tract infections has a high chance of failure in the current scenario.
Haemophilus, serotype b, Non typeable, biotype, BLNAR
Members of the genus Haemophilus are small, non-motile, fastidious, Gram negative pleomorphic bacilli which require X factor (hemin), and/or V factor (NAD), for their growth.
H. influenzae is the type species of which H. influenzae serotype b (Hib) is considered to be the most virulent1. It was estimated that it is responsible for more than 8 million cases of severe illness worldwide, and for 371,000 deaths in children under 5 years of age in the year 2000 worldwide2. It is estimated that annually, up to three million cases and more than 70,000 deaths in children under 5 years of age may be due to diseases caused by H. influenzae type b in India3. A report published in Lancet showed modelled estimates of deaths in children under 5 years due to Hib disease to have fallen to 15,600 (uncertainty range 9800-21500)4.
The findings from 6 teaching institutions affiliated to International Clinical Epidemiological Network in India showed that serotype b accounted for 97% of all infections due to Haemophilus influenzae5. In a prospective multicentric study in India, serotype b was seen as the main cause of pyogenic meningitis in children across many different places in India6. The incidence of Hib meningitis in infants was 32 per 100,000 and 19 per 100,000 in children under 2 years of age 7.
After the introduction of Hib vaccine, the incidence of invasive infection due to Hib has dropped significantly in developed countries, but in developing nations, it is still of major concern with rates ten times as high as developed nation in the pre-vaccination period2. A significant proportion of nasopharyngeal isolates of H. influenzae in a population of school children in India were identified as H. influenzae type b8.
The Indian government introduced Hib vaccination as a liquid pentavalent vaccine (LPV), in combination with DPT and Hepatitis B in the two South Indian states of Kerala and Tamil Nadu in 20119. There are lacunae in the data regarding the disease burden in these states, especially with regards to the impact of the vaccine.
The World Health Organization (WHO) has stressed the need for surveillance for Hib disease in developing countries10. This is necessary to estimate the morbidity and mortality due to Hib disease and to get an idea about the extent of benefit of introducing Hib vaccines in these regions. According to Peltola, the data on Hib disease for many poor or developing regions has not been very extensive11.
The emergence of and rise in resistance to commonly used antibiotics is also of grave concern. High resistance levels to many antibiotics used as first line drugs including ampicillin among isolates of H. influenzae from nasopharyngeal site of healthy, asymptomatic school going children was reported by Jain et al12. Similar findings were also reported by Saikia et al 8. Very high resistance was also reported in a multicentre study in India5, and in China13.
This study aims to shed some light on microbiological pattern among Haemophilus influenzae isolates from a tertiary care centre in a state in southern part of India in the period following introduction of the Hib vaccination.
The study was conducted on samples of sputum, ear and throat swabs, blood, CSF, and other samples including bronchoalveolar lavage (BAL) from patients coming to Amrita Institute of Medical Sciences and Research Centre, Kochi, which is a tertiary care referral centre, from May, 2014 to June, 2015.
The present study was cleared by the Dissertation Review Committee without any ethical, financial or scientific concerns being raised [Letter no. Amrita Vishwavidyapeetham Dissertation Review/MD/MS/2013/44].
The samples were cultured on chocolate agar and incubated at 36°C -37°C in a 5% CO2 atmosphere for 18 hours. Sputum samples were graded according to the quality by Bartlett scoring14, and the ones which did not have a Bartlett score of greater than 0 were rejected. Colonies with typical morphology were selected and identification was done with standard microbiological techniques like Gram staining, oxidase tests, satellitism and dependence on factor X (hemin) and factor V (NAD) using BD BBL™ Taxo™ discs (BD, Berkshire, U.K) for growth. H. influenzae (ATCC® 49247), Staphylococcus aureus (ATCC® 25923), E. coli (ATCC® 25922) and Pseudomonas aeruginosa (ATCC® 27853) (ATCC, Virginia, U.S) were used as controls.
Serotyping of H. influenzae was done by slide agglutination using polyvalent and type b specific antisera (BD, Berkshire, U.K). Due to financial considerations, type specific antisera for serotypes a, c, d, e, and f were not used.
Biotyping was done using indole, urease and ornithine decarboxylase tests, using peptone water, Christensen’s urease medium and Moeller’s ornithine decarboxylase broth supplemented with hemin and NAD, to a concentration of 10 µg/mL each15,16. Beta lactamase testing was done using cefinase discs (BD, Berkshire, U.K)
Antibiotic susceptibility testing by Kirby-Bauer disc diffusion method was done according to CLSI recommendations on Haemophilus Test Medium (HTM)17. Haemophilus test medium was prepared from Haemophilus test agar base [HiMedia, Mumbai, India] with added Haemophilus Growth Supplement [HiMedia, Mumbai, India]. Antibiotic discs [HiMedia, Mumbai, India] of the following strengths were used – Ampicillin (10 µg), Amoxycillin-clavulanate (20/10 µg), Ampicillin-sulbactam (10/10 µg), Cefuroxime (30 µg), Ceftriaxone (30 µg), Co-trimoxazole (1.25/23.75 µg), Azithromycin (15 µg), Ciprofloxacin (5 µg), Levofloxacin 5 µg, Tetracycline (30 µg), Chloramphenicol (30 µg), Meropenem (10 µg), and Rifampin (5 µg).
Statistical analysis of the findings was done by chi square test using IBM SPSS v.20.
A total of 36762 samples which included blood (26377 samples), CSF (681 samples), sputum (3141 samples), ear and nasal discharge (571 samples), and other samples including BAL (5992 samples) were processed from May 2014 till June 2015. No isolates of Haemophilus were recovered from blood, CSF, or pleural fluid during this period.
- Number of isolates from culture samples identified as Haemophilus- 65
- Number of sputum samples which did not meet the acceptance criteria – 9
- Number of isolates obtained from clinically significant samples- 56
Incidence of Haemophilus infection was 152.3 per 100,000 persons. Of the 56 Haemophilus isolates, 39 were from sputum samples, 8 from throat swabs, 5 from ear swabs, 3 from bronchoalveolar lavage samples, and one from nasal discharge.
For the purpose of statistical analysis, sputum and BAL were included as lower respiratory samples, while throat swabs, ear discharge and nasal discharge were included under upper respiratory tract samples.
H. influenzae, accounted for 24 out of clinically significant 56 Haemophilus isolates. Only 4 of those were from pediatric patients. The incidence of H. influenzae infections in the present study was 65.3 cases/100,000 persons.
Haemophilus influenzae serotype b accounted for two thirds of all isolates (16 of 24, 66.67%), followed by non typeable serotype (6 of 24, 25%) [Table 1].
Table (1):
Serotype distribution of H. influenzae by age.
| Serotype | Serotype b | Non type b | Non typeable |
|---|---|---|---|
| Age group (in years) | |||
| 0 – 18 | 0 | 0 | 4 |
| 18 – 60 | 7 | 2 | 1 |
| > 60 | 9 | 0 | 1 |
The incidence of capsulated H. influenzae and non-capsulated H. influenzae infections in the present study was 49 cases/100,000 persons and 16.3 cases/100,000 persons respectively. The incidence of H. influenzae type b infections in the current study was 43.5 cases/100,000 persons.
All sixteen of the isolates of serotype b were from adult patients [Table 1]. Also, all 4 isolates of Haemophilus influenzae from the pediatric population were non typeable. There was found to be an association between the age of the patient and serotype of H. influenzae isolated. Infection in adults was more likely to be with serotype b (p value= 0.007) than other serotypes of H. influenzae. In the paediatric age group, the isolates of H. influenzae were more likely to be non typeable (p value= 0.001).
The typeable strains of H. influenzae were almost equally distributed between the males (n=8) and the females (n=10). The majority of NTHI were isolated from females (5 out of 6), although the association was not found to be statistically significant (p=0.166).
All except one isolate of H. influenzae type b (15 of 16) were from lower respiratory tract samples. The only two isolates of capsulated non type b H. influenzae were both isolated from throat swabs [Table 2]. Serotype b of H. influenzae was more likely to be isolated from samples from the lower respiratory like sputum and BAL. (p value= 0.028), but the numbers are too few to conclusively establish this.
Table (2):
Distribution of serotypes of H. influenzae by sample.
| Serotype | Serotype b | Non type b | Non typeable |
|---|---|---|---|
| Sample | |||
| Sputum | 13 | 0 | 4 |
| Throat swab | 0 | 2 | 0 |
| BAL* | 2 | 0 | 0 |
| Ear swab | 1 | 0 | 1 |
| Nasal discharge | 0 | 0 | 1 |
*BAL- bronchoalveolar lavage
Of the biotypes of H. influenzae, biotype II was the most commonly isolated, accounting for 15 of the 24 (62.5%) isolates of H. influenzae, followed by biotype I (5 of 24, 20.83%) and biotype III (3 of 24, 12.5%) [Table 3].
Table (3):
Biotype distribution of H. influenzae by age.
| Biotypes | I | II | III | IV | V | VI | VII | VIII |
|---|---|---|---|---|---|---|---|---|
| Age group | ||||||||
| 0 – 18 | 3 | 1 | 0 | 0 | 0 | 0 | 0 | 0 |
| 18 – 60 | 1 | 7 | 1 | 1 | 0 | 0 | 0 | 0 |
| > 60 | 1 | 7 | 2 | 0 | 0 | 0 | 0 | 0 |
Biotype II was the most commonly isolated biotype (15 of 24) and was predominantly seen in the adult patients (14 of 15 isolates) as opposed to paediatric age group [Table 3]. Three out of four isolates in the paediatric age group belonged to biotype I. But there was no statistically significant association between age of the patient and the biotype of H. influenzae isolated (p=0.130).
Isolates of biotype II were almost equally distributed between males and females (7 and 8 respectively). 4 out 5 isolates of biotype I were recovered from males, but this association was not significant (p=0.423).There does not appear to be any association between the sample and the biotype of H. influenzae isolated (p=0.615).
Most of the isolates of serotype b belonged to biotype II (11 of 16) (68.75%) [Table 4]. Half of the six isolates of NTHI belonged to biotype I. There was no statistical significance between the serotype and biotype of H. influenzae.
Table (4):
Relation between biotypes and serotypes of H. influenzae.
| Biotypes | I | II | III | IV | V | VI | VII | VIII |
|---|---|---|---|---|---|---|---|---|
| Serotype | ||||||||
| Type b | 2 | 11 | 2 | 1 | 0 | 0 | 0 | 0 |
| Non type b | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 0 |
| Non typeable | 3 | 2 | 1 | 0 | 0 | 0 | 0 | 0 |
Susceptibility Pattern of H. influenzae
Out of 24 isolates, 20 were ampicillin sensitive (83.33%), 3 were β lactamase producers (12.5%), and one was β lactamase negative ampicillin resistant (BLNAR) (4.17%). Overall resistance to ampicillin was 16.67%.
Resistance to both amoxycillin-clavulanate and ampicillin-sulbactam was 12.5%. The β lactamase negative ampicillin resistant (BLNAR) isolate was also resistant to β lactam/ β lactamase inhibitor combinations. Two of three (66.7%) β-lactamase positive isolates showed resistance to both these drugs.
Resistance to cefuroxime was 8.33% (2 of 24 isolates) and ceftriaxone was 4.17% (one isolate).
Susceptibility to co-trimoxazole was very low. Three quarters (18 of 24) of all isolates of H. influenzae were resistant.
One of the 24 H. influenzae isolates was resistant to ciprofloxacin (4.17%). All the isolates, including the one above, were susceptible to levofloxacin.
One isolate was resistant to rifampin. This was β lactamase negative and showed susceptibility to all the other tested drugs except co-trimoxazole.
All the isolates showed susceptibility to azithromycin, tetracycline, chloramphenicol and meropenem.
In the present study, the incidence of infection due to Haemophilus species was 152.3 per 100,000 persons.
The incidence of H. influenzae infections in the current study was 65.3 cases/100,000 persons. These findings are much higher than seen in the United States (1.7/ 100000 persons)18.
The incidence of capsulated H. influenzae infections in the current study was 49 cases per 100,000 persons. The incidence of NTHi infections in the current study was 16.3 cases/100,000 persons. These findings are much higher than those found in a study in Spain (0.3 cases/100,000 persons and 0.7 cases/100,000 persons for capsulated and NTHi infections respectively)19.
In this study, the incidence of infection due to H. influenzae was found to be much higher than what is seen in the Western population. This maybe a direct consequence of the fact that rates of carriage of H. influenzae among healthy individuals in India was found to be high, which, in turn, may be due to poor socioeconomic status12.
Serotypes of H. influenzae
The incidence of H. influenzae type b infections in the current study was 43.5 cases/100,000 persons per year. It was the most common serotype of H. influenzae isolated, accounting for two thirds of all isolates (16 of 24) (66.67%), followed by non typeable serotype (25%).
This was similar to findings by Sharma et al. in North India20, by Weinberg et al in Pakistan21, and by Rahman et al in Bangladesh22. The proportion of infections due to serotype b was lesser among patients in 12 pediatric centres in Canada23.
The proportion of infection due to serotype b was found to be markedly lower in Spain, where nearly three quarters of H. influenzae infection was due to NTHi, and only 16.5% was due to serotype b19, and also in the UK (86% and 1% for NTHi and serotype b respectively)24, while a study in Ontario, Canada didn’t find any cases due to serotype b25.
A multicentre study in Canada on invasive disease due to H. influenzae in children found NTHi to be most common (35%), followed by serotypes a and f. Serotype b accounted for only 9% of the isolates26.
Even though the incidence of infection due to serotype b has decreased in European and North American populations, it still remains high in the Indian subcontinent. This study is in concordance with these findings.
In the present study, all four isolates of H. influenzae isolated from the paediatric population were non typeable [Table 1]. All sixteen isolates of H. influenzae type b were from the adult population. The number of NTHi isolated from the adult population was very few, accounting for 2 of the 20 H. influenzae isolates. We found an association between the age of the patient and serotype of H. influenzae isolated. Infection in adult population was more likely to be due to serotype b (p value= 0.007) than other serotypes of H. influenzae. In the paediatric age group, the isolates of H. influenzae were more likely to be non typeable (p value= 0.001). This may be due to the fact that Hib vaccine was introduced only recently in the population, and most of the adult population are not immunized. Conversely, it also lends support to the effectiveness of the recently introduced vaccination program.
These findings were similar to those seen in the annual Health Protection Report by the Public Health Department on H. influenzae24.
In the current study, all except one isolate of H. influenzae type b (15 of 16) were from lower respiratory tract samples [Table 2]. H. influenzae serotype b was more likely to be isolated from lower respiratory samples like sputum and BAL (p value= 0.028). This was similar to findings by Rahman et al22.
Biotypes of H. influenzae
Of the biotypes of H. influenzae, biotype II was the most commonly isolated, accounting for 15 of the 24 (62.5%) isolates of H. influenzae, followed by biotype I (5 of 24; 20.83%) and biotype III (3 of 24; 12.5%). Also, biotype II was predominantly seen in adult patients (14 of 15 isolates) (Table 3), similar to the findings of Oberhofer and Back27. Three out of four isolates in the paediatric age group belonged to biotype I.
Our study showed that 11 of the 16 isolates of serotype b belonged to biotype II (68.75%) (Table 4). This is similar to what was found in Pakistan21. Half of the six isolates of NTHI belonged to biotype I.
β-lactamase production
Among H. influenzae isolates, 3 of 24 isolates produced β lactamase enzyme (12.5%). This was similar to rates of β lactamase production by Jain et al in North India (19.6%)12, Lam et al in Germany (9.23%)28, and Hoban et al in many centres across the globe(14.3%)29.
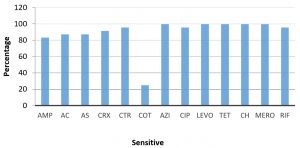
Susceptibility Pattern of H. influenzae (Fig. 1)
Out of 24 isolates, overall resistance to ampicillin was 16.67%. This was lower than that found in a study on invasive H. influenzae disease in children in Canada26. 20 were β lactamase negative ampicillin sensitive (83.33%), one was β lactamase negative ampicillin resistant (BLNAR) (4.17%), and 3 were β lactamase producers (12.5%). These findings are similar to those in other studies12,28-30. However, incidence of BLNAR as found by Hoban et al29 was very low (0.07%) compared to our findings. These findings were markedly different from those in a study on H. influenzae carriage in children, in which only 26% of isolates were susceptible to ampicillin31.
Fig. 1. H. influenzae Percentage Susceptibility
*AMP- Ampicillin, AC- Amoxicillin-Clavulanic acid, AS- Ampicillin-Sulbactam, CRX- Cefuroxime, CTR- Ceftriaxone, COT- Co-trimoxazole, AZI- Azithromycin, CIP- Ciprofloxacin, LEVO- Levofloxacin, TET- Tetracycline, MERO- Meropenem, RIF- Rifampin
Resistance in BLNAR strains is mediated by alteration in penicillin binding proteins, especially PBP3. In the β lactamase positive amoxycillin-clavulanic acid Resistant (BLPACR) strains, resistance is brought about by a combination of production of TEM-1 and ROB-1 β lactamases, and alteration in penicillin binding proteins32.
We found that resistance to both amoxicillin-clavulanic acid and ampicillin-sulbactam was 12.5%. The BLNAR isolate was also resistant to combinations. Two β lactamase positive isolates (8.33%) were resistant to both these drugs (β lactamase positive amoxicillin-clavulanate resistant) (BLPACR). These results are discordant with other studies, showing higher rates of prevalence than those found in North America by Karlowsky et al (BLPACR incidence 0.14%)33 and by Heilman et al (BLPACR incidence 0.4%)34, but similar to the findings of Kofteridis et al in Greece (BLPACR 7%)35.
Resistance to cefuroxime (8.33%) and ceftriaxone (4.17%) was higher than what was found in other studies- all isolates susceptible 5,19,22, cefuroxime resistance of 0.1%, no isolates resistant to ceftriaxone34, 0.5% resistant to ceftriaxone36.
Fluoroquinolone resistance is still relatively rare19,22,34,35, and our study also corroborates this, with one isolate (4.17%) being resistant to ciprofloxacin.
One isolate was rifampin resistant. All the isolates showed susceptibility to azithromycin, tetracycline, chloramphenicol and meropenem. Resistance to these tetracyclines, chloramphenicol and macrolides was similarly found to be very low in other studies19,29,34. However, the susceptibility patterns of tetracycline and chloramphenicol were markedly different from those observed in studies by IBIS group (60% isolates resistant to chloramphenicol)5, Saikia et al (56.25% and 38.75% resistance to tetracycline and chloramphenicol respectively)8, and Jain et al (chloramphenicol resistance 41.9%)12 from India, Rahman et al from Bangladesh (42% isolates resistant to chloramphenicol)22, and Shooraj et al from Iran (only 33% isolates susceptible to chloramphenicol and only 7% isolates susceptible to tetracycline)31.
In our study, co-trimoxazole resistance was very high. 75% of all isolates of H. influenzae were resistant. This may be because of high use and affordability of this drug leading to selective pressure. This is somewhat similar to the findings in other studies in the subcontinent by IBIS Group5 (55%), Jain et al12 (67.3%), and Rahman et al22 (44%), but much higher than the rates seen in Western population by Hoban et al29 (15.8% overall, as high as 70% in Brazil), and Heilman et al. 200534 (14.6%). Rates of co-trimoxazole resistance in the subcontinent are alarming, to say the least.
Haemophilus influenzae has been implicated in severe invasive illness with established mortality. Introduction of Hib vaccine targeted against serotype b, one of the most common and pathogenic serotypes, has been effective in lowering the rates of Hib infection worldwide, but incidence is still high in the Indian subcontinent. All isolates of serotype b were obtained from adult patients and none from the paediatric population, which may, in part, be due to introduction of Hib vaccine, which appears promising, but increase in infection due to NTHi and other serotypes remains a concern.
Also of grave concern is the increase in resistance to commonly used antibiotics including the penicillins, and appearance of BLNAR and BLPACR strains. An extremely high resistance against co-trimoxazole, especially in developing countries, is extremely worrying as it is one of the most commonly prescribed drugs because of availability and cheap cost, which makes it accessible to a majority of the population. But in view of the high resistance, use of co-trimoxazole in empirical therapy of upper and lower respiratory tract infections may have a high chance of failure in the current scenario.
The findings of this study support both the continued usage of the Hib vaccine, as well as the current recommendations regarding the use of amoxicillin-clavulanic acid plus macrolides for the empirical therapy of respiratory tract infections, and avoiding the use of co-trimoxazole. This also highlights the importance of microbiological confirmation of the etiology of respiratory tract infections, not just in tertiary care centres, but also at the primary level. This is because the healthcare needs of the vast majority of the Indian population are catered to at primary health centres and family health centres, and it would also reduce the burden on secondary and tertiary care centres. The authors recommend ramping up of microbiological diagnostic facilities in all primary healthcare facilities including those for performing culture and sensitivity and serological tests, along with appointing a medical officer specialized in microbiology, who can help to supervise and interpret the laboratory findings of the primary health centres in that area. This would help to achieve a more precise epidemiology of and a more targeted approach to the management of numerous infectious diseases of public importance.
Limitations
- Isolates were obtained from patients coming to a tertiary care referral centre. So, the incidence of infections due to different Haemophilus species may not fully represent the general population at large.
- Samples from many patients, especially those with upper respiratory tract infections, may not be sent for microbiological identification by culture and for susceptibility testing, and may be treated empirically by the clinician.
- Determination of factor requirement using X, V, and XV discs are known to have many inconsistencies, which can be overcome using δ-aminolevulinic acid (ALA) – porphyrin test. But in our study, this couldn’t be done due to non-availability of reagents.
- Serotyping of H. influenzae isolates using antisera specific for serotypes a, c, d, e, and f could not be done due to the prohibitive costs. Our study had to be limited to identifying the isolates as type b, non-type b, and non-typeable.
- Mechanisms of resistance of the different strains to each of the antimicrobial agents could not be elaborated upon as it was beyond the scope of this study.
ACKNOWLEDGMENTS
We would like to express our heartfelt thanks to Prof. Dr. KR Sundaram for providing assistance on bio statistical analysis.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors were involved in the design of the experiment. All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the Supplementary File.
- Peltola H. H.influenzae in the post-vaccination era. Lancet. 1993;341(8849):864-5.
Crossref - Watt JP, Wolfson LJ, O’Brien KL, et al. Burden of disease caused by Haemophilus influenzae type b in children younger than 5 years: global estimates. Lancet. 2009;374(9693):903-11. DOI: 10.1016/S0140-6736(09)61203-61204.
Crossref - NTAGI Subcommittee. NTAGI subcommittee recommendations on Haemophilus influenzae type b (Hib) vaccine introduction in India. Indian Pediatr, 2009;46:945-954.
- Wahl B, Sharan A, Knoll MD, et al. National, regional, and state-level burden of Streptococcus pneumoniae and Haemophilus influenzae type b disease in children in India: modelled estimates for 2000-15. Lancet. 2019;7(6):735-747.
Crossref - Invasive Bacterial Infections Surveillance (IBIS) Group of the International Clinical Epidemiology Network. Are Haemophilus influenzae infections a significant problem in India? A prospective study and review. Clin Infect Dis. 2002;34(7):949-957.
Crossref - Ramachandran P, Fitzwater SP, Aneja S, et al. Prospective multi-centre sentinel surveillance for Haemophilus influenzae type b & other bacterial meningitis in Indian children. Indian J Med Res. 2013;137(4):712-720.
- Minz S, Balraj V, Lalitha MK, et al. Incidence of Haemophilus influenzae type b meningitis in India. Indian J Med Res. 2008;128(1):57-64.
- Saikia KK, Das BK, Bewal RK, Kapil A, Arora NK, Sood S. Characterization of nasopharyngeal isolates of type b Haemophilus influenzae from Delhi. Indian J Med Res. 2012;136(5):855-861.
- Gupta SK, Sosler S, Lahariya C. Introduction of Haemophilus influenzae type b (Hib) as pentavalent (DPT-HepB-Hib) vaccine in two states of India. Indian Pediatr. 2012;49(9):707-709.
Crossref - World Health Organization (WHO). Global Programme for Vaccines and Immunization (GPV) – Vaccine Research and Development. Generic protocol for population-based surveillance of Haemophilus influenzae type b. 1996. WHO/VRG/GEN/95.05, WHO Geneva.
- Peltola H. Spectrum and burden of severe Haemophilus influenzae type b diseases in Asia. Bull World Health Organ. 1999;77(11):878-887.
- Jain A, Kumar P, Awasthi S. High ampicillin resistance in different biotypes and serotypes of Haemophilus influenzae colonizing the nasopharynx of healthy school-going Indian children. J Med Microbiol. 2006;55(Pt 2):133-137.
Crossref - WangH-J, Wang C-Q, Hua CZ, et al. Antibiotic resistance profiles of Haemophilus influenzae isolates from children in 2016: a multicenter study in China. Can J Infect Dis Med Microbiol. 2019;2019:6456321.
Crossref - Bartlett RC. A plea for clinical relevance in medical microbiology. Am J Clin Pathol. 1974;61(6):867-872. DOI:10.1093/ajcp/61.6.867.
Crossref - Kilian M. A taxonomic study of the genus Haemophilus with the proposal of a new species. J Gen Microbiol. 1976;93(1):9-62.
Crossref - Winn W, Allen S, Janda W, et al. Koneman’s Color Atlas and Textbook of Diagnostic Microbiology. 6th Edition, Lippincott Williams and Wilkins, New York. 2006.
- CLSI. Performance Standards for Antimicrobal Susceptibility Testing; Twenty-Fifth Informational Supplement. CLSI document M100-S25. Wayne PA. Clinical and Laboratory Standards Institute. 2015:76-79.
- Soeters HM, Blain A, Pondo T, et al. Current epidemiology and trends in invasive Haemophilus influenzae disease-United States, 2009-2015. Clin Infect Dis. 2018;67(6):881-889.
Crossref - Campos J, Hernando M, Roman F, et al. Analysis of invasive Haemophilus influenzae infections after extensive vaccination against H.influenzae type b. J Clin Microbiol. 2004;42(2):524.
Crossref - Sharma A, Kaur R, Ganguly NK, Singh PD, Chakraborti A. Subtype distribution of Haemophilus influenzae isolates from North India. J Med Microbiol. 2002;51(5):399-404.
Crossref - Weinberg GA, Ghafoor A, Ishaq Z, et al. Clonal analysis of Hemophilus influenzae isolated from children from Pakistan with lower respiratory tract infections. J Infect Dis. 1989;160(4):634-643. DOI:10.1093/infdis/160.4.634.
Crossref - Rahman M, Hossain S, Baqui AH, et al. Haemophilus influenzae type-b and non-b-type invasive diseases in urban children (<5years) of Bangladesh: implications for therapy and vaccination. J Infect. 2008;56(3):191-196.
Crossref - Mcconnell A, Tan B, Scheifele D, et al. Invasive infections caused by Haemophilus influenzae serotypes in twelve Canadian IMPACT centers, 1996-2001. Pediatr Infect Dis J. 2007;26(11):1025-1031.
Crossref - Public Health England. Laboratory reports of Haemophilus influenzae by age group and serotype (England and Wales): October to December 2014, and consolidated annual report for 2014. 2015. HPR 8:8. Available at https://www.gov.uk/government/uploads/system/uploads/attachment_data/file/408155/hpr0715_hib.pdf.
- Brown VM, Madden S, Kelly L, et al. Invasive Haemophilus influenzae disease caused by non-type b strains in Northwestern Ontario, Canada, 2002-2008. Clin Infect Dis. 2009;49(8):1240-1243.
Crossref - Frankel C, Alghounaim M, McDonald J, et al. Invasive Haemophilus influenzae disease in children: a Canadian Multicenter study on emerging serotypes. Open Forum Infect Dis. 2019;6(Suppl 2). Abstr.856 p.S19.
Crossref - Oberhofer TR, Back AE. Biotypes of Haemophilus encountered in clinical laboratories. J Clin Microbiol. 1979;10(2):168-174.
Crossref - Lam T-T, Claus H, Elias J, Matthias Frosch, Ulrich Vogel. Ampicillin resistance of invasive Haemophilus influenzae isolates in Germany 2009-2012. Int J Med Microbiol. 2015;305(7):748-755.
Crossref - Hoban D, Felmingham D. The PROTEKT surveillance study: antimicrobial susceptibility of Haemophilus influenzae and Moraxella catarrhalis from community-acquired respiratory tract infections. J Antimicrob Chemother. 2002;50(suppl 2):49-59.
Crossref - Omoding D, Bazira J. Isolation and antibiotic susceptibility testing of Haemophilus influenzae from nasopharynx of children under five years attending maternal and child health clinic in Mbarara Regional Referral Hospital. Can J Infect Dis Med Microbiol. 2019:6542919.
Crossref - Shooraj F, Mirzaei B, Fazlollah M, Hosseini F. Clonal diversity of Haemophilus influenzae carriage isolated from under the age of 6 years children. BMC Res Notes. 2019;12(1):565.
Crossref - Tristram SG, Jacobs MR, Appelbaum PC. Antimicrobial resistance in Haemophilus influenzae. Clin. Microbiol. Rev. 2007;20(2):368-389.
Crossref - Karlowsky JA, Critchley IA, Blosser-Middleton RS, et al. Antimicrobial surveillance of Haemophilus influenzae in the United States during 2000-2001 leads to detection of clonal dissemination of a b-lactamase-negative and ampicillin-resistant strain. J Clin Microbiol. 2002;40(3):1063-1066.
Crossref - Heilmann KP, Rice CL, Miller AL, et al. Decreasing prevalence of β-lactamase production among respiratory tract isolates of Haemophilus influenzae in the United States. Antimicrob Agents Chemother. 2005;49(6):2561-2564.
Crossref - Kofteridis D, Samonis G, Mantadakis E, et al. Lower respiratory tract infections caused by Haemophilus influenzae: clinical features and predictors of outcome. Med Sci Monit. 2009;15(4):CR135-9.
- Shenoy PA, Chawla K, Vishwanath S, Shaw D. Microbiological characterization of Haemophilus influenzae isolated from patients with lower respiratory tract infections in a tertiary care hospital, South India. J Clin Diagn Res. 2016;10(5): DC31-DC34.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.