ISSN: 0973-7510
E-ISSN: 2581-690X
The number of fatalities caused by multidrug-resistant (MDR) bacteria is over 700,000 annually due to widespread antibiotic usage. So, there is a need of new antibiotics, materials that work like antibiotics, or combinations of antibiotics with nanomaterials that could help in treating the infections which is caused by MDR bacteria. The present study describes the synthesis of ZnO nanoflakes using a co-precipitation method. The ZnO nanoflakes and ZnO nanoflakes combinations with carbapenem antibiotics were tested against carbapenem-resistant (CR) clinical isolates. The SEM analysis showed surface morphology of the synthesized nanoflakes-like structure of ZnO. All 67 CR isolates were tested and showed inhibitory action at varying concentrations of ZnO nanoflakes. ZnO nanoflakes were found to have an inhibitory effect against Escherichia coli and Klebsiella pneumoniae at lowest concentration of 1.25 mg.ml-1 of ZnO NPs with average zone size (mean ±SD) 1.91±2.94 mm and 2.00±4.14 mm and the average zone size of ZnO nanoflakes against Acinetobacter baumanni and Pseudomonas aeruginosa was 9.89±0.76 mm and 10.17±0.39 mm at 2.5 mg.ml-1 concentration. The combined action of ZnO nanoflakes with Meropenem 10 mcg demonstrated synergetic activity against CR pathogens, with an average zone of inhibition measuring 15.2 mm in diameter. ZnO nanoflakes illustrated considerable antibacterial activity against MBL-producing gram-negative clinical isolates at the lowest concentration. Chemically synthesized ZnO nanoflakes may offer a superior future expectation as a nano-antibiotic to treat the infection caused by CRE bacteria.
Antimicrobial, MBL, Nanoparticles, Synergetic Effect, Zinc Oxide
Today, we have many kinds of antibacterial compounds, such as penicillin, cephalosporins, carbapenems, and monobactams.1 As a result of the widespread use of antimicrobial agents, antibiotic resistance has become one of the largest threats to global health.2,3 The World Health Organization stated that antibiotic resistance is one of the top 10 public health threats to humanity due to the antibiotics used indiscriminately. The spread of drug-resistant pathogens is not the only concern; we continue to face threats to treating common infections as new antimicrobial resistance mechanisms emerge. The rapid spread of antibiotic-resistant bacteria (also called “superbugs”), which cause infections that cannot be treated with existing antibiotics, is of particular concern.4 Worldwide, the number of deaths caused by multidrug-resistant (MDR) microorganisms are estimated at approximately 700,000 every year due to the misuse of antibiotics on a large scale.5,6
Metallo β-lactamase (MBL) is a diverse group of metalloenzymes that can catalyze the hydrolysis of a wide range of β-lactam antibiotics.7 In the 1990s, Imipenemase Metallo beta-lactamase (IMP-type MBL) and Verona integron Metallo beta-lactamase (VIM-type MBL) were prevalent in Gram-negative pathogens. The recently discovered New Delhi Metallo β-lactamase (NDM1) provides an example of the ubiquitous potential of MBL.8,9 NDM1 was first identified in Klebsiella pneumoniae and E. coli patients returning from India to Sweden in 2008,10,11 NDM1 is frequently present in the Enterobacteriaceae family in India and worldwide.12
An increasing number of bacteria that belong to the Enterobacteriaceae family are becoming resistant to various groups of antibiotics, and clinicians use the broad-spectrum carbapenem as a last resort to effectively treat severe infections resulting from these pathogens. Consequently, the emergence and spread of carbapenem resistance is a primary medical and public health problem.11,13-15 No new classes of antibiotics have been reported in the last few decades so combination therapy can improve or expand the antimicrobial spectrum, reduce toxicity, prevent bacterial resistance during treatment, and achieve the synergistic activity.16 In this context, nanomaterials might resolve issues associated with multidrug resistance.
By integrating nanotechnology with biology, nanomaterials such as Cu, Ag, Au, and Zn have gained significant interest in developing new antimicrobial drugs with greater effectiveness. Various studies have reported that a few precious metals, such as gold and silver, have an interesting antibacterial effect. A higher concentration of these metal ions leads to toxicity, which remains a concern. In this context, antibiotics incorporated with relatively less toxic nanomaterials need to be explored.16
Zn and its oxide nanoparticles (ZnO-NPs) are active elements and reduce solid agents. The properties of ZnO-NPs suggest that they are anticancer, antibacterial, and antifungal agents.17,18 When used in conjunction with other therapeutic agents, ZnO-NPs synergize. ZnO is currently regarded as a relatively safe metal oxide approved by the United States Food and Drug Administration (USFDA) for cosmetic preparation.16 The data explores the antimicrobial action of ZnO-NPs and their synergetic effect on carbapenemase-producing MDR clinical isolates in India.
The present study described the synthesis of ZnO nanoflakes and their antimicrobial activity against carbapenem-resistant gram-negative clinical isolates and their synergetic effect with Meropenem to inhibit the MBL producing clinical isolates.
Chemicals
Zinc nitrate hexahydrate (Zn (NO3)2•6H2O) (≥98.0%), sodium hydroxides (NaOH) (≥97.0%) were procured from Sigma Aldrich India. The culture media were procured from Himedia Lab Pvt. Ltd., Mumbai, India. All the reagents used in this study were of analytical grade.
Synthesis of ZnO nanoflakes
ZnO nanoflakes were synthesized by a simple co-precipitation process using Zn (NO3)2•6H2O as a precursor and NaOH as a precipitating agent.19 Initially, approximately 9.12 gm of Zn (NO3)2•6H2O was suspended in 300 mL of deionized (DI) water and 2.4 gm of NaOH in 600 mL of DI water using a magnetic stirrer separately. Prepared solutions were labelled as solutions A and B, respectively. Solution B was added dropwise into solution A under continuous stirring at 65°C to produce a white precipitate solution. The mixture was subjected to constant stirring for 30 minutes and then switched off the hot plate. The solution was allowed to cool to produce ZnO nanoflakes.
The formed ZnO nanoflakes solution was washed twice with DI water using a centrifugation process (4000 rpm for 10 min). The supernatant was discarded, and the pellet containing ZnO nanoflakes was collected. The pellets were dried at 60°C for 24 hrs to produce ZnO nanoflakes powder for further analysis.
Material Characterization of ZnO nanoflakes
The prepared ZnO nanoflakes were characterized using a scanning electron microscope (SEM), energy-dispersive X-ray spectroscopy (EDX), X-ray diffraction (XRD), and Fourier transform Infrared spectroscopy (FT-IR).19,20 The surface morphology of ZnO nanoflakes was investigated using an SEM (JSM-7490LV, JEOL, Japan). EDX analysis was used to determine the elemental composition using JSM-7490LV, JEOL, Japan. The diffraction peaks were observed using XRD analysis (D8 Advance eco, Bruker, Germany). The ZnO nanoflakes’ crystal structure was characterized by Cu Kα radiation (l = 0.15418 nm). The prepared ZnO nanoflakes surface functional group was measured using FTIR (Nicole 6700, Thermo-Scientific, USA). The spectra were recorded with wavelengths ranging from 500–4000 cm-1.
Phenotypic and genotypic confirmation of carbapenem-resistant clinical isolates
A total of 67 carbapenem-resistant clinical isolates of Escherichia coli, Klebsiella pneumoniae, Acinetobacter baumanni, and Pseudomonas aeruginosa were isolated from various clinical samples (urine, pus, blood, sputum and body fluids etc.). The standard biochemical test (conventional test like, catalase test, oxidase test, indole, methyl red, Voges Proskauer, citrate utilization, triple sugar iron (TSI) test, urease test, various sugar fermentation test, nitrate reduction test, gelatin hydrolysis test etc.) was used for up to species level identification and further confirmed by the VITEK automated system. Antibiotic sensitivity tests were performed by the disc diffusion test following the Clinical and Laboratory Standards Institute CLSI 2018. All carbapenem-resistant isolates were then confirmed phenotypically using various standard methods (Combined disc synergy test (CDST), Modified Hodge Test (MHT), modified carbapenem inactivation method (mCIM), and EDTA-modified carbapenem inactivation method (eCIM).21 The genes responsible for carbapenemase production may initiating the resistance to carbapenem groups antibiotics were detected by genotypic method.
Genomic DNA was extracted from bacterial cells using HiPer® Bacterial Genomic DNA Extraction Teaching Kit (Column Based) Himedia Lab Pvt. Ltd. The procedure of DNA extraction was followed as per kit manufacturer.22
The qRT-PCR technique was used to amplify a targeted DNA sequence by use of hydrolysis probes that are short oligonucleotides that have a fluorescent reporter dye attached to the 5’ and a quencher dye to the 3’end. Hi-PCR Carbapenemase Gene (multiplex) test PCR pack (MBPCR132 Himedia Lab Pvt. Ltd) PCR kit is designed to detect the specific regions of the genes encoding the carbapenemase enzymes. Master mix 1 detect NDM, KPC, IMP, VIM in FAM, HEX, Texas Red and Cy5 channels and master mix 2 detect OXA-51, OXA-23, OXA-48, OXA-58 in FAM, HEX, Texas Red and Cy5 channels respectively.23
MM mix for PCR reaction
| Components | Volume to be added for 1 R (for 25 µL reaction) | |
|---|---|---|
| CRG1 tube | CRG 2 tube | |
| Hi-Quanti 2x Realtime PCR master mix | 12.5 µL | 12.5 µL |
| CRG 1 Primer-probe mix | 4 µL | – |
| CRG 2 Primer-probe mix | – | 4 µL |
| IC Primer probe mix | 1 µL | 1 µL |
| IC B DNA | 1 µL | 1 µL |
| Molecular grade water | 1.5 µL | 1.5 µL |
| Positive control/ Negative control/ template DNA | 5 µL | 5 µL |
| Total volume | 25 µL | 25 µL |
PCR program:
| Initial denaturation | 95ºC for 10 minutes | |
|---|---|---|
| Denaturation | 95ºC for 05 seconds | No. of cycle: 45 |
| Annealing and Extension | 60ºC for 1 minute (plate read) | |
| Hold | 4ºC for ∞ | |
Data Interpretation
The amplification data was interpreted as per RT-PCR kit (Hi-PCR Carbapenemase Gene) literature.
Ct value |
Result |
|---|---|
≤40 |
Detected (+) |
≥41 |
Not detected (-) |
| Target in Tube 1 (Primer probe mix) | Result Interpretation | ||||
|---|---|---|---|---|---|
| NDM (FAM) | KPC (HEX) | IMP(Texas Red) | VIM (Cy5) | IC (Cy5.5) | |
| Ct value ≤40 | Ct value ≤40 | Ct value ≤40 | Ct value ≤40 | Ct value ≤40 | Positive for NDM, KPC, IMP, VIM |
| Ct value ≥41 | Ct value ≥41 | Ct value ≥41 | Ct value ≥41 | Ct value ≤40 | Negative for NDM, KPC, IMP, VIM |
| No Ct | No Ct | No Ct | No Ct | No Ct | PCR inhibition or reagent failure repeat PCR or repeat extraction from original sample. |
| Target in Tube 2 (Primer probe mix) | Result Interpretation | ||||
| OXA-51 (FAM) | OXA-23 (HEX) | OXA-48 (Texas Red) | OXA-58 (Cy5) | IC (Cy5.5) | |
| Ct value ≤40 | Ct value ≤40 | Ct value ≤40 | Ct value ≤40 | Ct value ≤40 | Positive for OXA-51, OXA-23, OXA-48, OXA-58 |
| Ct value ≥41 | Ct value ≥41 | Ct value ≥41 | Ct value ≥41 | Ct value ≤40 | Negative for OXA-51, OXA-23, OXA-48, OXA-58 |
| No Ct | No Ct | No Ct | No Ct | No Ct | PCR inhibition or reagent failure repeat PCR or repeat extraction from original sample. |
Antibacterial activity
Preparation of ZnO nanoflakes suspension
A 40 mg/mL stock solution of synthesized ZnO nanoflakes was prepared in sterile deionized water and sonicated for 20 min at 35°C using a 24 Hz frequency and 400 rpm rotation through complete ultrasonication dispersion. The working solution was prepared by performing two-fold serial dilutions in sterile deionized water. All eight MCT tubes were filled with 1000 µL of sterile deionized water, except for the first tube. Add 1000 µL of stock solution in the second tube, mix well, and transfer the solution from the second to the third, followed by the last tube to get a final concentration of 40 mg.ml-1, 20 mg.ml-1, 10 mg.ml-1, 05 mg.ml-1, 2.5 mg.ml-1, 1.25 mg.ml-1, 0.625 mg.ml-1, 0.312 mg.ml-1, and 0.156 mg.ml-1 solutions.24
Well diffusion assay
An overnight culture of the test strain in trypticase soy broth (TSB) was used. The optical density matched McFarland turbidity standards (0.5), resembling a 1.5 × 108 CFU/mL bacterial count. Lawn culture was performed on the Mueller-Hinton agar (MHA) surface and allowed to dry. After drying the agar surface, a 6 mm well was punched with a well borer, and 50 µL of suspension was added in to the each well and incubate overnight at 37°C.24
Synergistic effect of ZnO nanoflakes with Meropenem
To determine the synergistic effect of ZnO nanoflakes against CR isolates, two Meropenem antibiotic discs (10 mcg) were placed on the inoculated agar surfaces with sterile forceps, and one antibiotic disc was impregnated with 50 µl of 5 mg.ml-1 ZnO nanoflakes suspension to test the synergistic effects of antibiotics with nanoflakes. Petri plates were meticulously labeled and incubated at 37°C for overnight incubation. After incubation, the inhibitory zones (mm) sizes around the discs were measured. The tests were performed in triplicate to minimizing the error, and the zone of inhibition of antibiotics alone and in combination with ZnO nanoflakes was recorded.
Statistical analysis
All outcomes were analyzed using Statistical R-Software ver. 4.1.2. Descriptive statistics were used for quantitative information like mean and standard deviation (SD), while ordered information was addressed as numbers and rates. Analysis of variance (ANOVA) and post hoc analysis was used to look at a method for differences between and within various groups. The Pearson correlation coefficient was utilized to check for a connection between two or more quantitative parametric variances. A two-tailed test was used for all evaluations, and a p-value of <0.05 was considered statistically significant.
The synthesized ZnO nanoflakes were characterized by using several physiochemical methods like scanning electron microscopy (SEM), Energy-dispersive X-ray spectroscopy (EDX), X-ray diffraction (XRD), and Fourier transform Infrared (FT-IR) spectroscopy.
SEM and EDX analysis
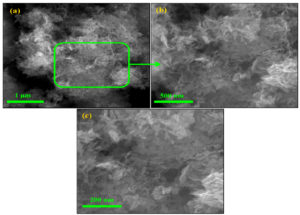
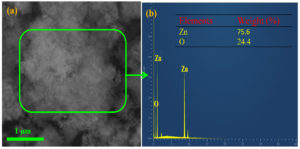
The SEM images showed that the precipitate comprises ZnO nanoflakes (Figure 1a). The ZnO nanoflakes contain a few nanometers, 20–50 nm in thickness. The higher magnification SEM images clearly show nanoflake structures. Moreover, interconnected ZnO nanoflakes produce pores (Figure 1 (b-c)). The porous structure of ZnO nanoflakes might increase the active sites, which might increase their antibacterial ability.
The presence of Zn and O confirms the synthesis of ZnO nanoflakes seen in the EDX analysis (Figure 2) and ensures the synthesis of ZnO nanoflakes.
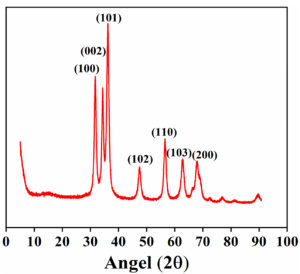
XRD analysis
The XRD spectra of synthesized ZnO nanoflakes (Figure 3) showed characteristic peaks at 2θ = 31.67°, 34.31°, 36.14°, 47.40°, 56.52°, 62.73°, and 66.28° were assigned to (100), (002), (101), (102), (110), (103), and (200) of the prepared ZnO nanoflakes material. It indicated that the samples had polycrystalline wurtzite structures. It was confirmed by comparing these data with known standard data published by the Joint Committee on Powder Diffraction Standard (Zincite, file no. JCPDS 5-0664). No different characteristic peaks of any impurities have been detected, suggesting that incredible ZnO nanoflakes have been synthesized. Scherrer’s formula calculated ZnO nanoflakes’ average crystallite size (d).
d= K l / β cosθ
Where k = 0.9 is the shape factor, l is the X-ray wavelength of Cu Ka radiation (1.54 Å), θ is the Bragg diffraction angle, and θ is the full width at half maximum of the respective diffraction peak. The average crystallite size of ZnO nanoflakes was 21.59 ± 4.89 nm.
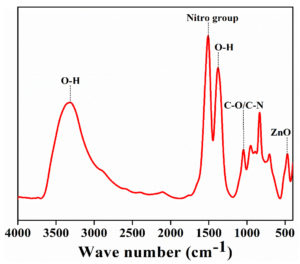
FTIR spectra
The characteristic peaks (Figure 4) were observed at 575, 1005, 1595, and 3330 cm-1. They were assigned to be ZnO stretching vibration, C-N bond or stretching of C-O bond, aromatic nitro compound, and hydroxyl group, respectively. The characteristic peaks at 1120, 1350, and 1385 cm-1 correspond to primary and secondary alcohol. The distinct peaks confirm the synthesis of the ZnO nanoflakes.
A total of 67 MBL producing clinical isolates consisting of Escherichia coli (32.8%), Acinetobacter baumannni (26.8%), Klebsiella pneumoniae (22.4%), and Pseudomonas aeruginosa (18%) were included in this study. The combined disc synergy test (CDST) showed 16.4% negative and 83.6% positive results in phenotypic methods. In contrast, the modified Hodge test (MHT), mCIM, and eCIM showed 100% positive for MBL producers.
In genotypic distribution (Table 1), the NDM gene was found most abundantly in 33 (49.2%), followed by VIM and OXA-48 (10, 14.9%), OXA-51 and OXA-23 (6, 8.9%). In this investigation, neither KPC nor OXA-58 was detected. The epidemiological findings of genotypic diversity concur with studies from India that have previously been published.25–27
Table (1):
Distribution of Carbapenem-resistant genes in Gram-Negative isolates N=67.
| Bacterial Isolates | Number of isolates | Carbapenemase-producing targeted gene | |||||||
|---|---|---|---|---|---|---|---|---|---|
| NDM | KPC | IMP | VIM | OXA-51 | OXA-23 | OXA-48 | OXA-58 | ||
| Member of Family Enterobacteriaceae n= 37 | |||||||||
| Escherichia coli | 22 | 12 | 0 | 3 | 4 | 0 | 0 | 3 | 0 |
| Klebseilla penumoniae | 15 | 6 | 0 | 0 | 3 | 0 | 0 | 7 | 0 |
| Acinetobacter baumannii | 18 | 6 | 0 | 0 | 0 | 6 | 6 | 0 | 0 |
| Pseudomonas aeruginosa | 12 | 9 | 0 | 0 | 3 | 0 | 0 | 0 | 0 |
| Total | 67 | 33 | 0 | 3 | 10 | 6 | 6 | 10 | 0 |
| Percentage | 49.2 | 0 | 4.4 | 14.9 | 8.9 | 8.9 | 14.9 | 0 | |
Antibacterial activity of ZnO nanoflakes
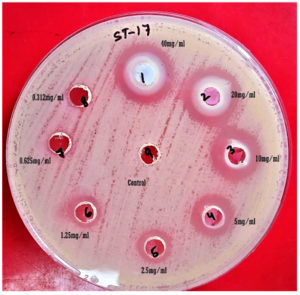
The ZnO nanoflakes were evaluated for antibacterial activity (Figure 5) against MBL-producing clinical isolates at various concentration and showed significant inhibitory effect than the control (DI water).
Figure 5. Antibacterial efficacy of ZnO nanoflakes against MDR E. coli by agar well diffusion methods
It was statistically significant when the concentration of ZnO NPs was reduced from 40 mg.ml-1 to 1.25 mg.ml-1. (Table 2) showed the descriptive analysis of various concentrations of ZnO nanoflake’s effectiveness against different clinical isolates. ZnO nanoflakes were found to have an inhibitory effect against Escherichia coli and Klebsiella pneumoniae at the lowest concentration of 1.25 mg.ml-1 of ZnO NPs with average zone size (mean ±SD) 1.91±2.94 mm and 2.00±4.14 mm and the average zone size of ZnO nanoflakes against Acinetobacter baumanni and Pseudomonas aeruginosa was 9.89±0.76 mm and 10.17±0.39 mm at 2.5 mg.ml-1 concentration.
Table (2):
Descriptive analysis of various concentrations of ZnO nanoflakes effectiveness against different clinical isolates N=67.
| ZnO nanoflakes Concentration | Bacterial Isolates | Mean ± S.D. | F- value | “p-value” |
|---|---|---|---|---|
| 40 mg.ml-1 | Acinetobacter baumanni. | 16.72 ±1.60 | 9.970 | 0.002 |
| Pseudomonas aeruginosa. | 13.08 ±0.90 | |||
| Escherichia coli | 14.91 ±2.07 | |||
| Klebsiella pneumoniae | 15.67 ±2.23 | |||
| 20 mg.ml-1 | Acinetobacter baumanni. | 15.50 ±1.95 | 10.193 | 0.007 |
| Pseudomonas aeruginosa. | 11.58 ±0.51 | |||
| Escherichia coli | 13.50 ±2.30 | |||
| Klebsiella pneumoniae | 14.20 ±2.04 | |||
| 10 mg.ml-1 | Acinetobacter baumanni. | 13.83 ±2.01 | 8.338 | 0.013 |
| Pseudomonas aeruginosa. | 10.42 ±0.51 | |||
| Escherichia coli | 12.14 ±2.12 | |||
| Klebsiella pneumoniae | 12.47 ±1.88 | |||
| 5 mg.ml-1 | Acinetobacter baumanni. | 11.67 ±1.57 | 3.622 | 0.018 |
| Pseudomonas aeruginosa. | 10.17 ±0.39 | |||
| Escherichia coli | 11.23 ±1.74 | |||
| Klebsiella pneumoniae | 10.60 ±0.74 | |||
| 2.5 mg.ml-1 | Acinetobacter baumanni. | 9.89 ±0.76 | 9.634 | 0.022 |
| Pseudomonas aeruginosa. | 0.00 ±0.00 | |||
| Escherichia coli | 9.23 ±2.14 | |||
| Klebsiella pneumoniae | 8.60 ±2.56 | |||
| 1.25 mg.ml-1 | Acinetobacter baumanni. | 0.56 ±2.36 | 1.221 | 0.031 |
| Pseudomonas aeruginosa. | 0.00 ±0.00 | |||
| Escherichia coli | 1.91 ±2.94 | |||
| Klebsiella pneumoniae | 2.00 ±4.14 | |||
| 0.75 mg.ml-1 | Acinetobacter baumanni. | 0.00* | – | – |
| Pseudomonas aeruginosa. | 0.00* | |||
| Escherichia coli | 0.00* | |||
| Klebsiella pneumoniae | 0.00* | |||
| 0.375 mg.ml-1 | Acinetobacter baumanni. | 0.00* | – | – |
| Pseudomonas aeruginosa. | 0.00* | |||
| Escherichia coli | 0.00* | |||
| Klebsiella pneumoniae | 0.00* |
*Mean & S.D. could not be computed due to zero value
Various researchers (Table 3) synthesize ZnO NPs using different methodologies and assess antibacterial activity against gram-positive and gram-negative bacterial isolates. Most researchers use bacterial (ATCC, NCTC, PTCC, RN, etc.) isolates with pre-defined properties like sensitivity and resistance to empirical antibiotics. In India, very few studies have reported the antimicrobial action of ZnO nanoparticles against CRE strains isolated from clinical samples.
Table (3):
Comparison of synthesis of ZnO NPs and antimicrobial actions by various methods.
Synthesis methods |
Particle size |
Isolates type |
Methods Use |
Antibacterial action at the lowest concentration |
Ref |
|---|---|---|---|---|---|
Not defined |
– |
E.coli, B. subtilis and S. aureus |
Disc Diffusion |
125 mg/ml |
[41] |
Sole gel |
20.20 nm |
E.coli, S. aureus |
MIC |
5mg/ml |
[29] |
Not defined |
08 nm |
S. aureus RN6390 |
MIC |
1.2 mg/ml |
[42] |
Sole gel |
21-38 nm |
S. aureus E.coli |
MIC & MBC |
Mic 78 µg/ml |
[43] |
Sole gel |
39 nm |
S. aureus |
MIC |
0.1041 mg.ml |
[44] |
Precipitation |
20-40 nm |
K. pneumoniae (ATCC70068) |
MIC |
0.75 mM Con. |
[45] |
Not defined |
03 nm |
S. auerus PTCC 1431 E. coli PTCC |
MIC & MBC. |
0.5 mg/ml & 8 mg/ml 1 mg/ml & 16 mg/ml |
[46] |
Procured (Sigma) |
70 nm |
ESBL producing E.coli Klebsiella spp. |
Inhibitory concentration methods |
1 mg/ml |
[47] |
Not defined |
Not defined |
E.coli ATCC 25922, ESBL & AmpC producing E.coli P. aeruginosa ATCC 27853, ESBL & AmpC producing P. aeruginosa |
MIC & MBC |
1000 µg/ml 500µg/ml 1000µg/ml |
[26]
|
Chemical & Green Synthesis method |
30 nm |
carbapenem-resistant Acinetobacter baumannii (RS-307, RS-6694, and ATCC-19606 strains) |
Disc diffusion Micro broth dilution |
Not define IC50 2mM |
[48] |
Precipitation |
MBL producing Clinical isolates E.coli, K. pneumoniae, P. aeruginosa, A. baumanni |
Well Diffusion |
Average lowest concentration 1.25 mg/ml Average highest concentration 5 mg/ml |
Present study |
ZnO-NPs with an average size of 30 nm were synthesized by Vishvanath Tiwari et al. (2018) and exhibited good antibacterial action against carbapenem-resistant Acinetobacter baumannii (RS-307, RS-6694, and ATCC-19606 strains).24
According to Asfia Sultan et al. (2015), ZnO NPs have antibacterial efficacy against ESBL-producing enterobacteria. She observed that the minimum and maximum MIC values were 1000 and 8,000 µg/ml, respectively, while the minimum and maximum MBC values were 2,000 and 16,000 µmg/ml, respectively. The minimal MIC and MBC values for standard strain E. coli were 1000 µg/ml and 2,000 µg/ml, respectively, while for P. aeruginosa, they were 8000 µg/ml and 16000 µg/ml.26
Elaheh Sadat Nazoori et al. (2018) revealed that the MIC of ZnO nanoparticles against E. coli was more significant at 2.5 mg/mL, while the MIC of other bacteria was 5.0 mg/mL. The minimum bactericidal concentration (MBC) of ZnO nanoparticles against P. aeruginosa, A. baumannii, K. pneumoniae, and S. aureus was 10 mg/mL, whereas other bacteria required 20 mg/mL.26
According to several researchers, bacteria’s cell walls are also responsible for their protective action against antimicrobial compounds. At the lowest concentration, gram-positive bacteria were readily inactivated compared to gram-negative bacteria. Research on the antibacterial activity of ZnO microspheres (MS-ZnO) against S. aureus and E. coli by Shinde et al showed that variations in their cell walls might explain the variation in susceptibility of MS-ZnO in the two bacteria. NPs of smaller sizes may readily penetrate through the peptidoglycan barrier and extremely capable of causing damage.28 The peptidoglycan layer is made up of repeated amino acid and carbohydrate units. As a result, ZnO NPs may bind with carboxylic acid and amino groups, inhibiting biological activities. However, since the peptidoglycan layer of Gram-negative bacteria is thinner than that of Gram-positive bacteria, cell membrane breakage is simpler.29
Tayel et al, reported ZnO NPs were able to kill Gram-positive bacteria more effectively than Gram-negative bacteria.30 Reddy et al found that the MIC of ZnO NPs against S. aureus (a Gram-positive bacterium) was 1 mg/mL. Still, the MIC against E. coli (a Gram-negative bacterium) was 3.4 mg/mL, indicating that larger doses of ZnO NPs are necessary to inhibit Gram-negative bacteria. Most likely, this is because the cell wall components of Gram-negative bacteria, such as lipopolysaccharides, can defend against ZnO attack within the cell. In contrast, the peptidoglycan layer surrounding Gram-positive bacteria may enhance ZnO attack inside the cell.30,31 Agua et al. evaluated the antibacterial activity of textiles containing ZnO NPs using the agar diffusion technique and observed comparable findings.32 The current investigation also revealed that gram-negative bacteria need more ZnO Nps to be inhibited than gram-positive bacteria.
Treatment of bacterial infections is a major concern in recent years due to the growing problem of resistance to traditional antibiotics. MBL-producing Gram-negative microorganisms have now been identified in a variety of geographical areas.33 The rise of GNB makes MBL a problem for microbiology labs because there are no standardized rules for how to find them. Plasmids easily spread MBLs, so they quickly spread through an institution and cause bad results when they infect someone.34-36
The synergistic effect of ZnO nanoflakes with Meropenem against CR clinical isolates were observed, but Meropenem (10 mcg/ml) alone showed no activity against the pathogens tested. However, at a 5 mg.ml-1, the combined action of antibiotics and ZnO nanoflakes demonstrated high synergistic activity against all pathogens, with an average zone of inhibition measuring 15.22 mm in diameter (Table 4).
Table (4):
Significance of synergism of ZnO NPs with and without Meropenem.
Meropenem (10mcg) |
Meropenem (10mcg) + NP (5mg/ml) 50µl |
|
|---|---|---|
N |
67 |
67 |
Mean ± S.D. |
9.89 ±2.96 |
15.22 ±2.69 |
r-value |
1 |
.780** |
p-value |
0.001 |
** Correlation is significant at the 0.01 level (2-tailed)
The statistical findings (Table 4) summarized the synergistic effects of ZnO nanoflakes with Meropenem against four types of CR clinical isolates. It was found that the relationship between Meropenem and ZnO nanoflakes with Meropenem had a statistically significant difference (p-value <0.005). The synergistic effect of ZnO nanoflakes with Meropenem on the test pathogens validated its efficacy as a combination therapy.37
Gram-negative bacteria are more prevalent than gram-positive bacteria, causing community and hospital-acquired illnesses, such as urinary tract infections, wounds, and lower respiratory tract infections14 Gram-negative bacteria also having more carbapenem resistance than Gram-positive bacteria.15 Infections induced by these resistant bacteria have a greater death rate than infections caused by carbapenem-susceptible bacteria. Most cases are multidrug-resistant (MDR) infections that need long-term antibiotic treatment, which is associated with high healthcare expenditures and contributes to a rise in antibiotic tolerance in bacterial cells that survive. Antimicrobial agents such as metal oxide nanoparticles (NPs) have emerged as promising weapons against MDR bacteria. Multiple mechanisms of action are responsible for their antimicrobial effectiveness.38,39 We utilized ZnO nanoparticles in this investigation with several reasons, including their inexpensive cost compared to other metal NPs, effective bactericidal action, and commercial antibacterial uses. ZnO is graded as a “GRAS” substance by the US FDA; Zn plays an essential role in the human body as one of the vital microelements.17,40 It is found in all the body tissues, such as muscle and bone (85% of the total body Zn content), skin (11%), and all other tissues; it is intracellular, primarily residing in the nucleus cytoplasm and cell membrane of the cells due to their unique ability to induce ROS and apoptosis, thereby promising biomedical potential. These properties make ZnO receive more attention in biomedical applications.27
The broad-spectrum class of carbapenems is the last resort to effectively treating severe infections caused by sensitive pathogens. Combination therapy is the last resort in treating patients infected with MDR pathogens. Various drug combinations are available on the market but are toxic for the patient in long-term treatment.
Our results showed that simple precipitation methods could achieve syntheses of ZnO nanoflakes. It effectively killed or inhibited confirmed MBL producing clinical isolates.
ZnO nanoflakes illustrated considerable antibacterial activity against MBL-producing gram-negative clinical isolates. Chemically synthesized ZnO nanoflakes may offer a superior future expectation as a nano-antibiotic against carbapenem-resistant bacteria. The cell-targeted delivery of ZnO nanoflakes in the animal model needs further evaluation for improving the mechanistic understanding of synthesized ZnO nanoflakes in vitro settings. The cytotoxicity of chemically produced ZnO-NP can be studied in cell lines and animal models to find the effective non-cytotoxic dose of ZnO.
ACKNOWLEDGMENTS
The authors would like to thank Members of the Microbiology Research Laboratory at Index Medical College Hospital & Research Centre, Indore, India. The authors also would like to thank Indian Institute of Technology, Kanpur, India; and the University of Sophisticated Instrumentation Centre (USIC) at Babasaheb Bhimrao Ambedkar University (Central University), Lucknow, for providing support.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
BNS designed and performed the experiments. AS performed statistical analysis. BNS and PSP wrote the manuscript. HS, MA and GCU revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the ethics committee of Index Medical College, Hospital & Research Centre affiliated with Malwanchal University, Indore (MP) (M.U./Research/E.C./Ph.D./2018/14).
- Hauser AR. The ABCs of Choosing the Right Antibacterial Agent Antibiotic Basics for Clinicians. 2nd ed. 2013:1-253.
- Davis M, Whittaker A, Lindgren M, Djerf-Pierre M, Manderson L, Flowers P. Understanding media publics and the antimicrobial resistance crisis. Glob Public Health. 2018;13(9):1158-1168.
Crossref - Baluja Z, Nabi N, Ray A. Challenges in Antimicrobial Resistance: An Update. 2018;6(8):65-77.
- Antimicrobial resistance fact sheet. 2021. https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance
- Betts JW, Hornsey M, la Ragione RM. Novel Antibacterials: Alternatives to Traditional Antibiotics. Adv Microb Physiol. 2018;73(1):123-169.
Crossref - Padiyara P, Inoue H, Sprenger M. Global Governance Mechanisms to Address Antimicrobial Resistance. Infect Dis. 2018;11(11):788-887.
Crossref - Nascimento GGF, Locatelli J, Freitas PC, Silva GL. Antibacterial activity of plant extracts Antibacterial Activity of Plant Extracts and Phytochemicals on Antibiotic-Resistant Bacteria. Braz J Microbiol. 2000;31:247-256.
Crossref - Laraki N, Galleni M, Thamm I, et al. Structure of In31, a bla IMP-Containing Pseudomonas aeruginosa Integron Phyletically Related to In5, Which Carries an Unusual Array of Gene Cassettes. Antimicrob Agents Chemother. 1999;43(4):890-901
Crossref - Lauretti L, Riccio ML, Mazzariol A, et al. Cloning and Characterization of bla VIM, a New Integron-Borne Metallo-Lactamase Gene from a Pseudomonas aeruginosa Clinical Isolate. Antimicrob Agents Chemother. 1999;43(7):1584-1590.
Crossref - Yong D, Toleman MA, Giske CG, et al. Characterization of a new metallo-β-lactamase gene, bla NDM-1, and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob Agents Chemother. 2009;53(50):5046-5054.
Crossref - Nordmann P, Naas T, Poirel L. Global spread of carbapenemase producing Enterobacteriaceae. Emerg Infect Dis. 2011;17(10):1791-1798.
Crossref - Mohapatra PR. Metallo-β-lactamase 1 – why blame New Delhi & India? Indian J Med Res. 2013;137(1):213-215. PMCID: PMC3657891
- Vital Signs: Carbapenem-Resistant Enterobacteriaceae. Morbidity and Mortality Weekly Report (MMWR). Center for Disease control and Prevention. 2013;62(09):165-170. https://www.cdc.gov/mmwr/preview/mmwrhtml/mm6209a3.htm
- U.S. Department of Health and Human Service C for DC and P. Antibiotic Resistance Threats in the United States. 2013. http://www.cdc.gov/drugresistance/threat-report-2013/
- World Health Organization. Antimicrobial Resistance Global Report on surveillance. 2014. http://www.who.int/drugresistance/documents/surveillancereport/en/.
- Raghupathi KR, Koodali RT, Manna AC. Size-Dependent Bacterial Growth Inhibition and Mechanism of Antibacterial Activity of Zinc Oxide Nanoparticles. Langmuir. 2011;27(40):4020-4028.
Crossref - Colon G, Ward BC, Webster TJ. Increased osteoblast and decreased Staphylococcus epidermidis functions on nanophase ZnO and TiO2. J Biomed Mater Res A. 2006;78A(1):595-604.
Crossref - Cheesman M, Ilanko A, Blonk B, Cock I. Developing new antimicrobial therapies: Are synergistic combinations of plant extracts/compounds with conventional antibiotics the solution? Pharmacogn Rev. 2017;11(22):57-72.
Crossref - Salahuddin NA, El-Kemary M, Ibrahim EM. Synthesis and Characterization of ZnO Nanoparticles via Precipitation Method: Effect of Annealing Temperature on Particle Size. Nanoscience and Nanotechnology. 2015;5(4):82-88.
Crossref - Ashfaq M, Verma N, Khan S. Copper/zinc bimetal nanoparticles-dispersed carbon nanofibers: A novel potential antibiotic material. Materials Science and Engineering: C. 2016;59(9):938-947.
Crossref - CLSI. M100-performance standards for antimicrobial susceptibility testing, 28th edition. Clinical and Laboratory. 2018.
- HTBM008 HiPer® Bacterial Genomic DNA Extraction Kit (Column Based). https://www.himedialabs.com/eu/htbm008-hiper-bacterial-genomic-dna-extraction-teaching-kit-column-based.html
- MBPCR132 Hi-PCR® Carbapenemase Gene (Multiplex) Probe PCR Kit. https://www.himedialabs.com/eu/mbpcr132-hi-pcr-carbapenemase-gene-multiplex-probe-pcr-kit.html
- Karthick S, Namasivayam R, Prasanna M, Subathra S. Synergistic antibacterial activity of zinc oxide nanoparticles with antibiotics against the human pathogenic bacteria. J Chem Pharm Res. 2015;7(13):3-8.
- Tiwari V, Mishra N, Gadani K, Solanki PS, Shah NA, Tiwari M. Mechanism of anti-bacterial activity of zinc oxide nanoparticle against Carbapenem-Resistant Acinetobacter baumannii. Front Microbiol. 2018;9:1218.
Crossref - Sultan A, Khan HM, Malik A, Ansari MA, Azam A, Perween N. Antibacterial Activity of ZnO Nanoparticles against ESBL and Amp-C Producing Gram Negative Isolates from Superficial Wound Infections. Int J Curr Microbiol App Sci. 2015.
- Nazoori ES, Kariminik A. In Vitro Evaluation of Antibacterial Properties of Zinc Oxide Nanoparticles on Pathogenic Prokaryotes. J Appl Biotechnol Rep. 2018;5(4):162-165.
Crossref - Sirelkhatim A, Mahmud S, Seeni A, et al. Review on Zinc Oxide Nanoparticles: Antibacterial Activity and Toxicity Mechanism. Nanomicro Lett. 2015;7(2):219-422.
Crossref - Shinde V, Dalavi DS, Mali SS, Hong CK, Kim JH, Patil PS. Surfactant free microwave assisted synthesis of ZnO microspheres: Study of their antibacterial activity. Appl Surf Sci. 2014;307:495-502.
Crossref - Tayel AA, El-Tras WF, Moussa S, et al. Antibacterial Action of Zinc Oxide Nanoparticles Against Foodborne Pathogens. J Food Saf. 2011;31(2):211-218.
Crossref - Reddy KM, Feris K, Bell J, Wingett DG, Hanley C, Punnoose A. Selective toxicity of zinc oxide nanoparticles to prokaryotic and eukaryotic systems. Appl Phys Lett. 2007;90(21):213021.
Crossref - Borda d’ Agua R, Branquinho R, Duarte MP, et al. Efficient coverage of ZnO nanoparticles on cotton fibres for antibacterial finishing using a rapid and low cost in situ synthesis. New Journal of Chemistry. 2018;42(1):1052-1060.
Crossref - Walsh TR, Toleman MA, Poirel L, Nordmann P. Metallo-β-Lactamases: The Quiet before the Storm? Clin Microbiol Rev. 2005;18(2):306-625.
Crossref - Hirakata Y, Yamaguchi T, Nakano M, et al. Clinical and Bacteriological Characteristics of IMP-Type Metallo-β-Lactamase-Producing Pseudomonas aeruginosa. Clin Infect Dis. 2003;37(1):26-32.
Crossref - Laupland KB, Parkins MD, Church DL, et al. Population-Based Epidemiological Study of Infections Caused by Carbapenem-Resistant Pseudomonas aeruginosa in the Calgary Health Region: Importance of Metallo-β-Lactamase (MBL)-Producing Strains. J Infect Dis. 2005;192(16):1606-1612.
Crossref - Peleg AY, Franklin C, Bell JM, Spelman DW. Dissemination of the Metallo-beta-Lactamase Gene blaIMP-4 among Gram-Negative Pathogens in a Clinical Setting in Australia. Clin Infect Dis. 2005;41(15):1549-1556.
Crossref - Ahmed MNM,; Al-Azaim NGA, Al-Sehrawey AA-S, Elgendy AelAlAeAA. Effect of Silver nanoParticles on Carbapenem Resistant Gram-negative Bacteria. The Egyptian Journal of Hospital Medicine. 2018;72(4):4389-4391.
Crossref - Rasmussen JW, Martinez E, Louka P, Wingett DG. Zinc oxide nanoparticles for selective destruction of tumor cells and potential for drug delivery applications. Expert Opin Drug Deliv 2010;7(10):1063-1077.
Crossref - Rai M, Ingle AP, Birla S, Yadav A, Santos CA dos. Strategic role of selected noble metal nanoparticles in medicine. Crit Rev Microbiol 2015;42(5):696-719.
Crossref - Zhang L, Jiang Y, Ding Y, Povey M, York D. Investigation into the antibacterial behaviour of suspensions of ZnO nanoparticles (ZnO nanofluids). J Nanopart Res. 2007;9(4):79-89.
Crossref - Baek Y-W, An Y-J. Microbial toxicity of metal oxide nanoparticles (CuO, NiO, ZnO, and Sb2O3) to Escherichia coli, Bacillus subtilis, and Streptococcus aureus. Sci Total Environ. 2011;409(8):1603-1608.
Crossref - Jones N, Ray B, Ranjit KT, Manna AC. Antibacterial activity of ZnO nanoparticle suspensions on a broad spectrum of microorganisms. FEMS Microbiol Lett. 2008;279(1):71-76.
Crossref - Lallo da Silva B, Caetano BL, Chiari-Andreo BG, Pietro RCLR, Chiavacci LA. Increased antibacterial activity of ZnO nanoparticles: Influence of size and surface modification. Colloids Surf B Biointerfaces. 2019;177(1):440-447.
Crossref - Arakha M, Saleem M, Mallick BC, Jha S. The effects of interfacial potential on antimicrobial propensity of ZnO nanoparticle. Sci Rep. 2015;5(1):9578.
Crossref - Reddy LS, Nisha MM, Joice M, Shilpa PN. Antimicrobial activity of zinc oxide (ZnO) nanoparticle against Klebsiella pneumoniae. Pharm Biol. 2014;52(13):88-97.
Crossref - Emami-Karvani Z, Chehrazi P. Antibacterial activity of ZnO nanoparticle on Gram-positive and Gram-negative bacteria. Afr J Microbiol Res. 2012;5(18):1368-1373.
Crossref - Mustafa S, Khan HM, Shukla I, et al. Effect of ZnO nanoparticles on ESBL producing Escherichia coli & Klebsiella spp. Eastern J Med. 2011; 16(4): 253-257.
- R. Jalal, Elaheh K., Goharshadi, Maryam Abareshi, Majid Moosavi, AbbasYousef, Paul Nancarrow. ZnO nanofluids: Green synthesis, characterization, and antibacterial activity. Materials Chemistry and Physics. 2010;121 (1–2): 198-201.
Crossref
© The Author(s) 2023. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.